Submitted:
18 April 2025
Posted:
22 April 2025
Read the latest preprint version here
Abstract
Keywords:
Introduction
Interface
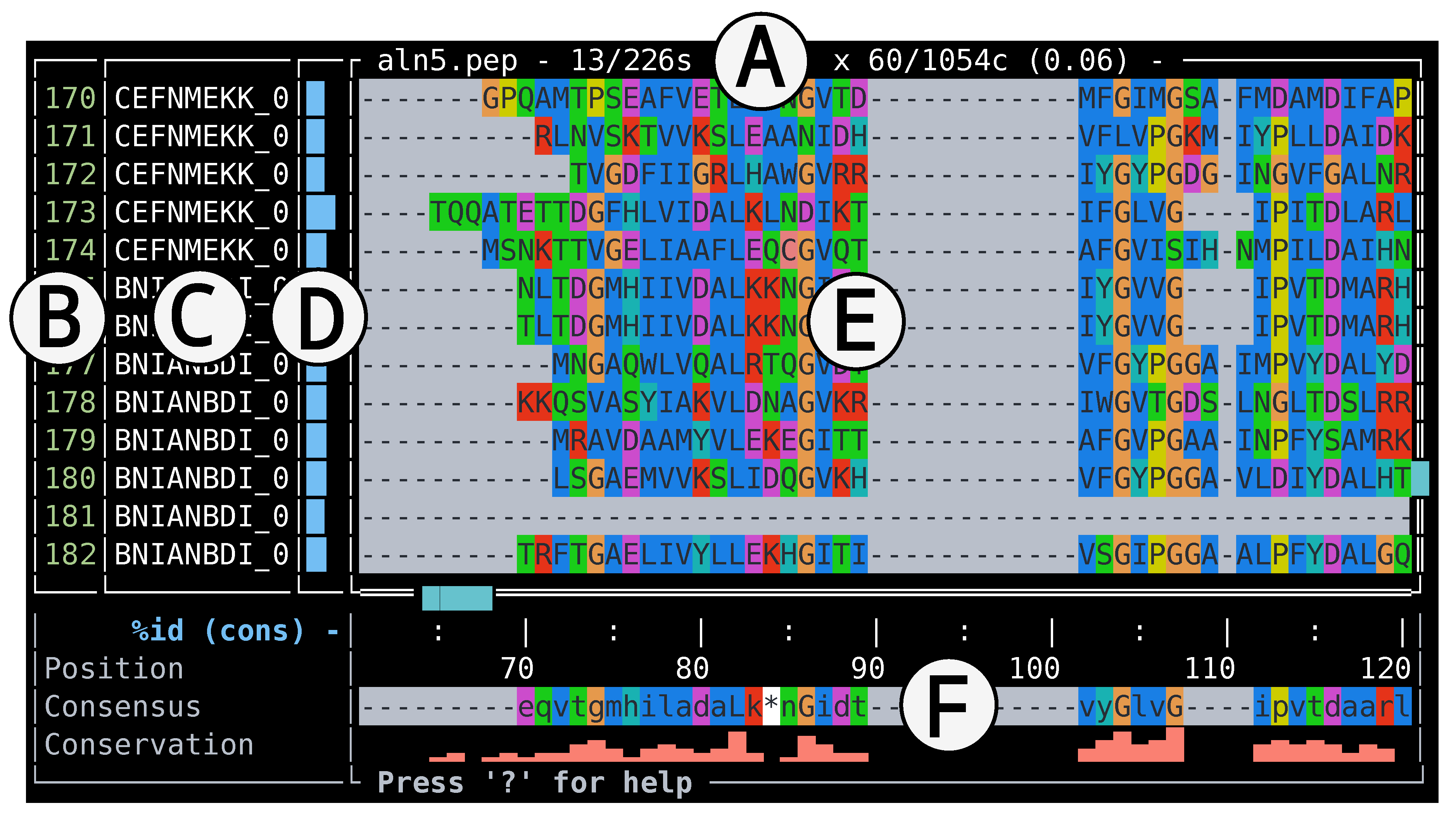
Performance and Limitations
Comparison with Existing Tools
Availability
Conclusion
Acknowledgments
References
- Dunne, M. Alan: A lightweight alignment viewer for the terminal. https://github.com/mpdunne/alan, 2020. Accessed 2025-04-13.
- Rice, P.; Longden, I.; Bleasby, A. EMBOSS: the European Molecular Biology Open Software Suite. Trends in Genetics 2000, 16, 276–277.
- Nissen, J. alen: Interactive terminal-based MSA viewer. https://github.com/jakobnissen/alen, 2023–2024. Accessed 2025-04-13.
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.; Clamp, M.; Barton, G.J. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191.
- Larkin, M.; Blackshields, G.; Brown, N.; Chenna, R.; McGettigan, P.; McWilliam, H.; Valentin, F.; Wallace, I.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948.
- Lesk, A.M. Introduction to bioinformatics; Oxford university press, 2019.
- Wilm, J.; contributors. Alacritty - A cross-platform, GPU-accelerated terminal emulator. https://github.com/alacritty/alacritty, 2017–2024. Accessed 2025-04-13.
- Goyal, K. Kitty - The fast, feature-rich, GPU-based terminal emulator. https://sw.kovidgoyal.net/kitty/, 2017–2024. Accessed 2025-04-13.
- Hashimoto, M. Ghostty - A modern, GPU-accelerated terminal emulator. https://github.com/mitchellh/ghostty, 2023–2024. Accessed 2025-04-13.
| Feature | termal | alen | alv | alan | showalign |
|---|---|---|---|---|---|
| Basics | |||||
| Language | Rust | Rust | Python | Shell | C |
| Interactivity | Yes (full TUI) | Yes (TUI) | No (pager output) | No (pager output) | No (static output) |
| Handles large alignments | Yes (tested >15k×1500) | Yes | Yes (slower) | Yes (but static) | Moderate |
| Display features | |||||
| Scrolling / navigation | Yes | Yes | No | No | No |
| Zooming | Yes | No | No | No | No |
| Label pane toggle | Yes | No | No | No | No |
| Consensus display | Yes | Yes (reference toggle) | Yes (static) | No | Optional |
| Conservation display | Yes | No | No | No | No |
| Similarity histogram | Yes (vertical bars) | No | No | No | No |
| Colour schemes | Multiple, configurable | Fixed | Fixed | Basic ANSI | Limited |
| Sequence numbering | Yes | No | No | No | Yes |
| Supported formats | FASTA | FASTA | FASTA, Clustal (via BioPython) | FASTA | FASTA, Clustal |
| Sorting / filtering | |||||
| Sort by consensus similarity | Yes | No | No | No | No |
| Sort by sequence length | Yes | No | No | No | No |
| Manual row reordering | No | Yes | No | No | No |
| Regex search | No | Yes | No | No | No |
| Integration / output | |||||
| Output style | TUI | TUI | STDOUT (to pager) | STDOUT (to pager) | Plain text output |
| HPC / CLI friendly | Yes (single binary) | Yes | Needs Python env | Yes | Yes (installed via EMBOSS) |
| License | MIT | MIT | MIT | Various | GPL |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




