Submitted:
03 April 2025
Posted:
07 April 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Deep Generative Models: Core Architectures
2.1. Variational Autoencoders (VAEs)
2.2. Generative Adversarial Networks (GANs)
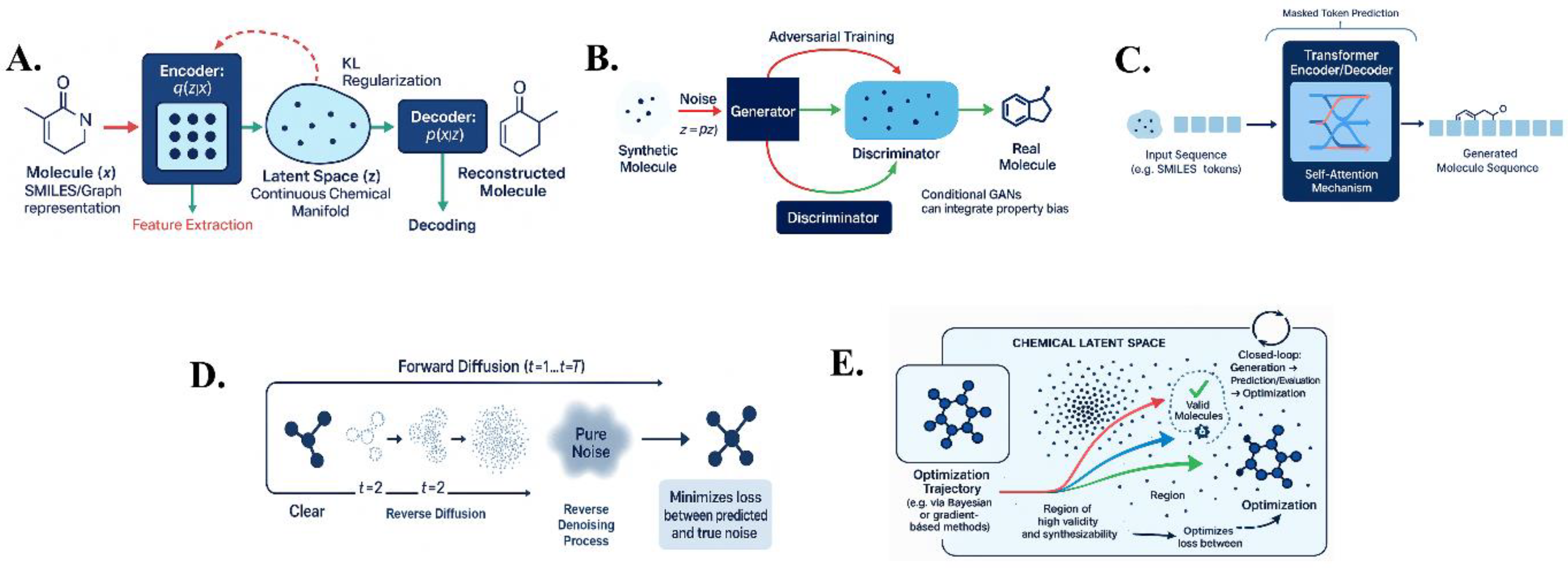
2.3. Transformer-Based Models
2.4. Denoising Diffusion Models (DDPMs)
2.5. Theoretical Considerations in Chemical Space Exploration
2.5.1. Latent Space Optimization and Chemical Manifolds
2.5.2. Validity and Synthesizability Constraints
2.5.3. High-Dimensional Chemical Space
3. Generative AI for Molecular Structure Prediction and Optimization
3.1. AI-Driven Small Molecule Design
3.1.1. Self-Supervised Learning for Molecular Representations
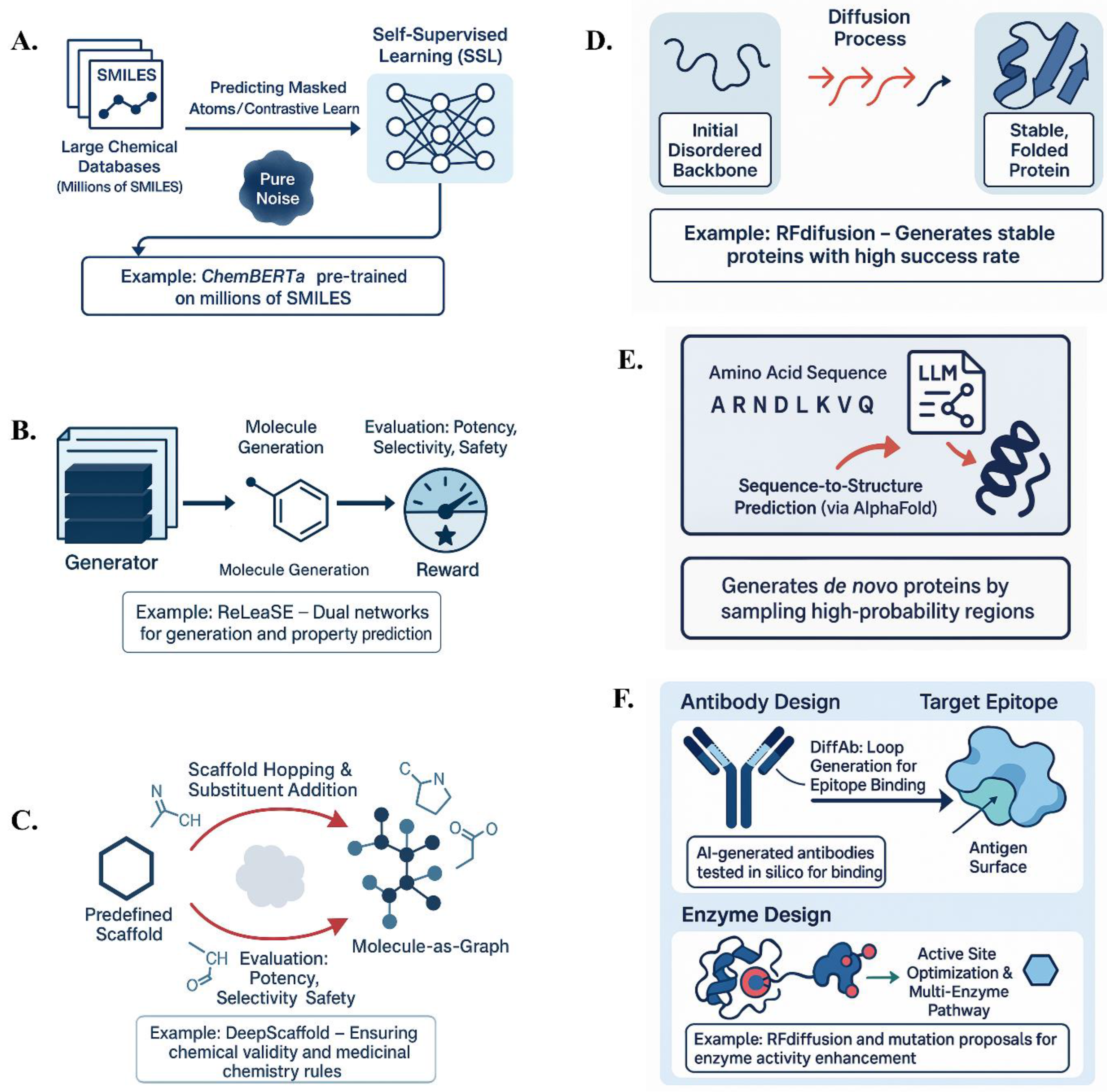
3.1.2. Reinforcement Learning (RL) for Molecular Optimization:
3.1.3. Graph-Based Generative Models:
3.2. AI-Driven Protein Design
3.2.1. Diffusion Models for Protein Folding & Stability Prediction:
3.2.2. LLMs for De Novo Protein Sequence Generation:
3.2.3. Antibody and Enzyme Design Using AI:
- Antibody Design: AI can design antibodies by generating complementarity-determining region (CDR) sequences likely to bind a target antigen or by generating 3D conformations of antibody loops that complement antigen surfaces (112). DiffAb, a diffusion model, generates antibody structures conditioned on the 3D structure of the target antigen’s epitope, effectively growing an antibody loop to fit into the epitope pocket (113). The success of AbSci’s model in creating functional antibodies in silico indicates that these methods can produce viable therapeutic candidates (114).
- Enzyme and Biocatalyst Design: Enzymes catalyze chemical reactions, and AI is transforming enzyme design by improving active site modeling and exploring backbone arrangements. RFdiffusion has been used to design enzyme active sites, with some designs showing promising activity. AI can also optimize existing enzymes by proposing mutations that stabilize them or alter their substrate scope. Generative models can propose multi-enzyme pathways for synthetic routes, offering a new approach to metabolic network design (115–117).
4. Computational Strategies for AI-Guided Drug–Target Interactions
4.1. DiffDock and Beyond: AI in Molecular Docking
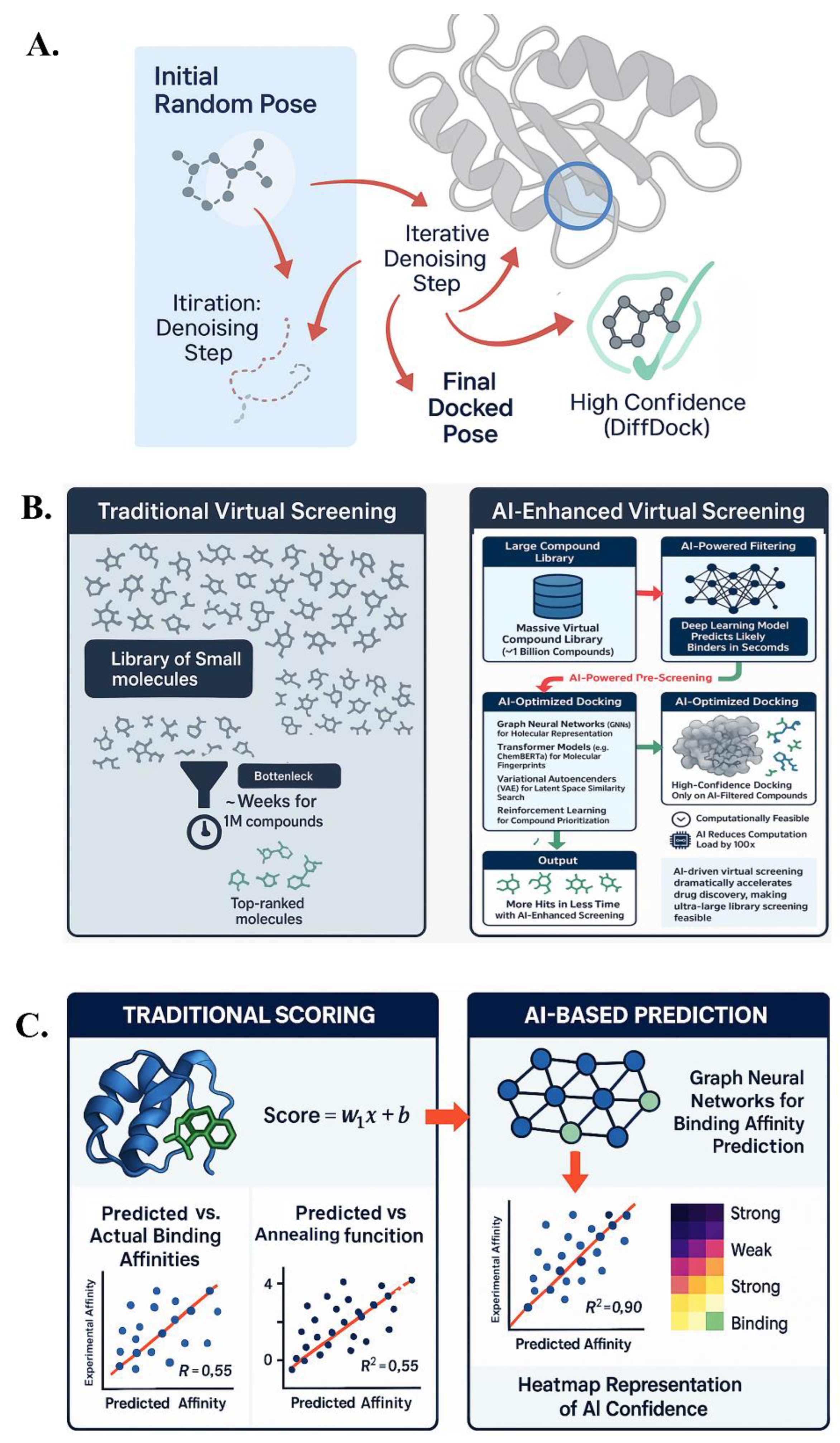
4.2. Protein–Ligand Binding Affinity Prediction
4.3. Large-Scale Virtual Screening
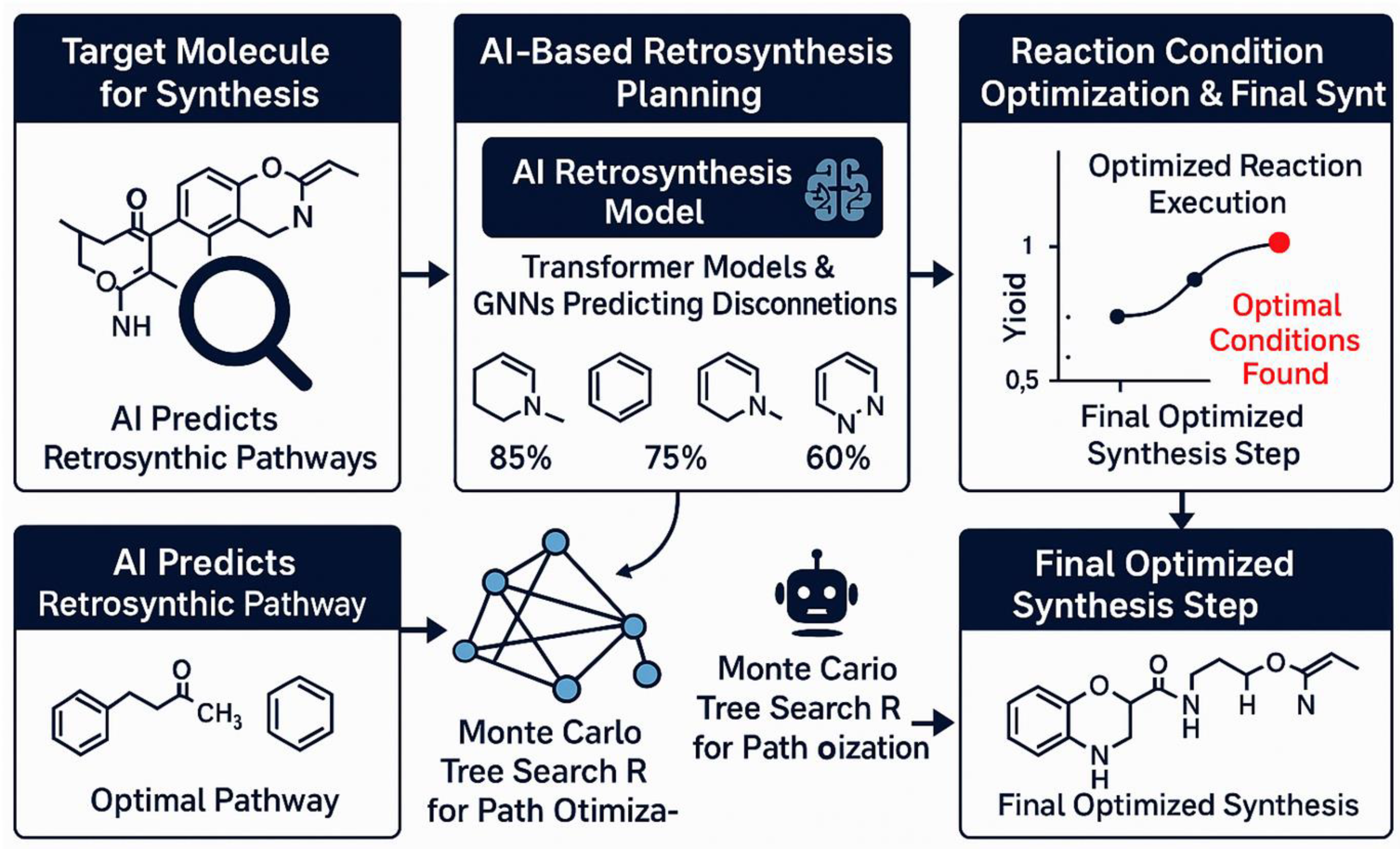
5. AI-Driven Synthesis Planning and Retrosynthesis
6. AI for Pharmacokinetics and Toxicity Prediction
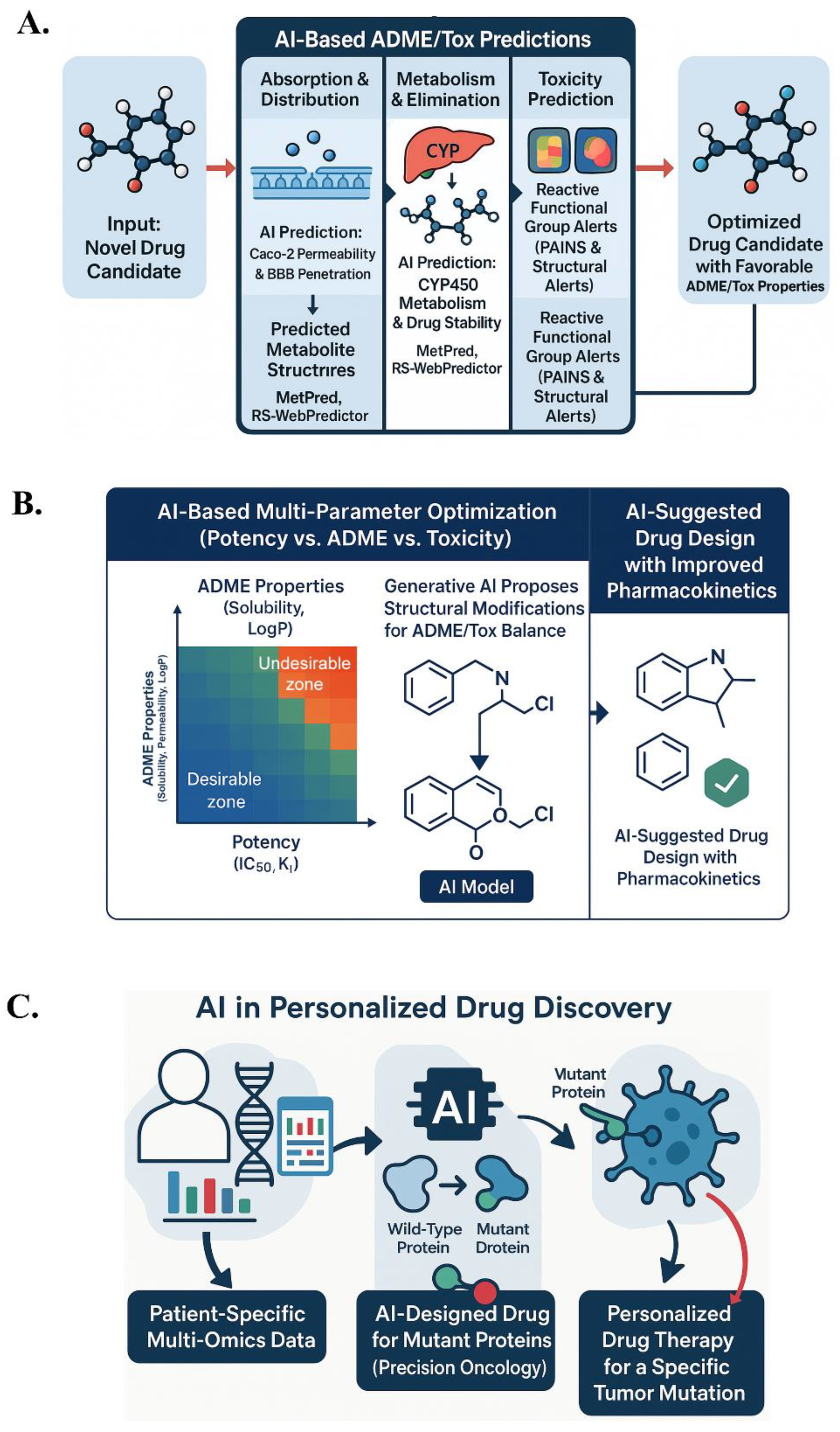
6.1. ADME/Tox Predictions
- In vitro cytotoxicity using Tox21 challenge data.
- Organ toxicity (hepatotoxicity, cardiotoxicity), including hERG channel inhibition, predicted by ML models.
- Genotoxicity and carcinogenicity predictions using Ames test data or animal studies.
- Reactive functional group alerts: AI identifies substructures causing nonspecific reactivity or toxicity, learning broader patterns of reactivity beyond known PAINS.
6.2. Personalized Drug Discovery
7. Experimental Validation and AI-Augmented Pipelines
7.1. Wet Lab Validation: Case Studies of AI-Designed Drugs and Challenges in Translation
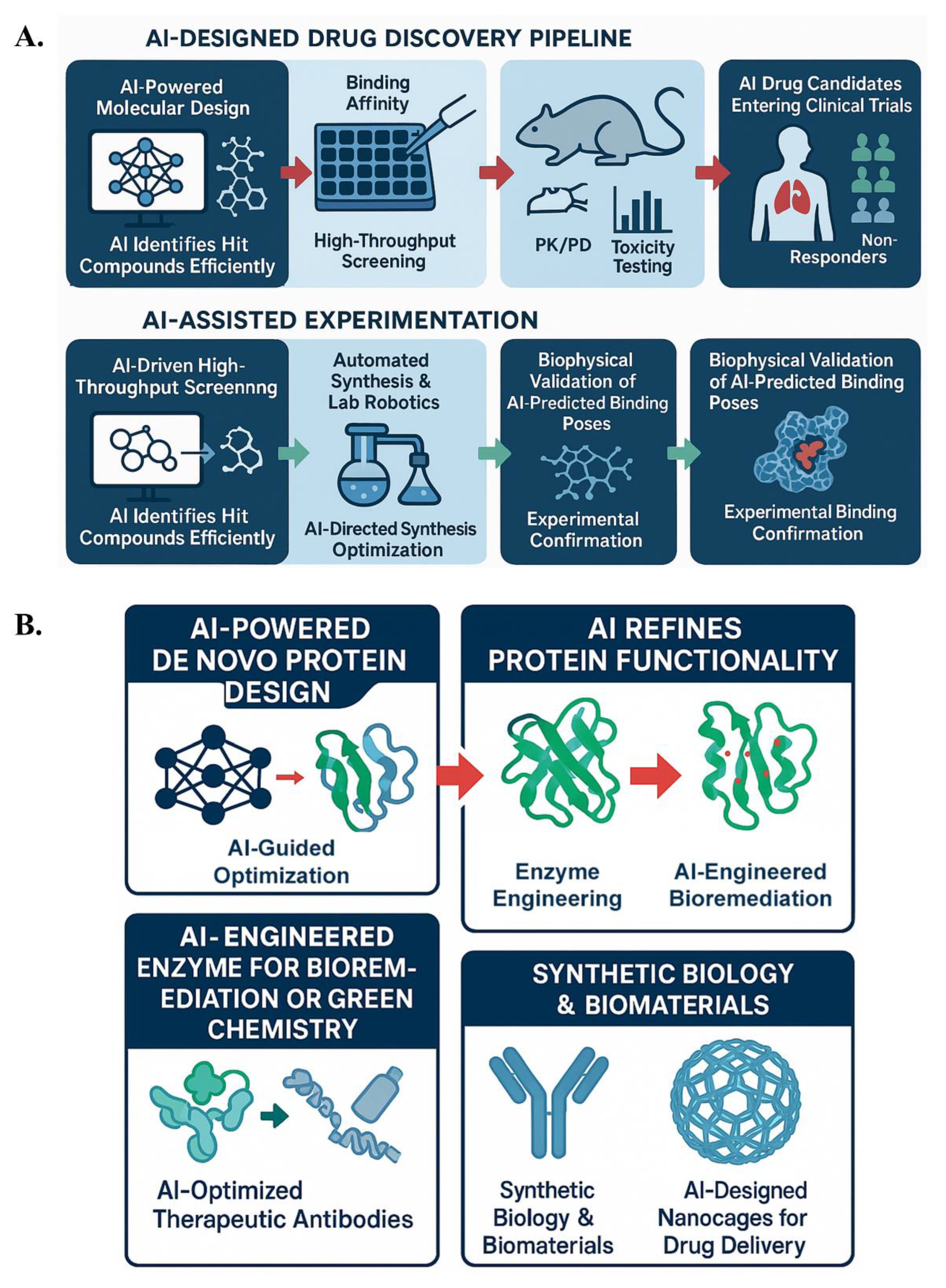
7.2. Protein Engineering in Biotechnology: AI-Augmented Enzyme and Pathway Design
8. Future Perspectives: AI-Designed Medicines & Autonomous Discovery
8.1. Fusion with Quantum Computing
8.2. Ethics and Regulation of AI-Designed Drugs
8.3. Personalized Drug Design Ethics
9. Conclusions
Funding
Author Contributions
Conflicts of Interest
Declaration of generative AI and AI-assisted technologies in the writing process
References
- Hinkson IV, Madej B, Stahlberg EA. Accelerating Therapeutics for Opportunities in Medicine: A Paradigm Shift in Drug Discovery. Front Pharmacol. 2020 Jun 30;11:770. [CrossRef]
- Vijayan RSK, Kihlberg J, Cross JB, Poongavanam V. Enhancing preclinical drug discovery with artificial intelligence. Drug Discov Today. 2022 Apr;27(4):967–84. [CrossRef]
- Sun D, Gao W, Hu H, Zhou S. Why 90% of clinical drug development fails and how to improve it? Acta Pharm Sin B. 2022 Jul;12(7):3049–62. [CrossRef]
- Das U, Banerjee S, Sarkar M. Bibliometric analysis of circular RNA cancer vaccines and their emerging impact. Vacunas. 2025 Mar;500391. [CrossRef]
- Boyd NK, Teng C, Frei CR. Brief Overview of Approaches and Challenges in New Antibiotic Development: A Focus On Drug Repurposing. Front Cell Infect Microbiol. 2021;11:684515. [CrossRef]
- Singh S, Gupta H, Sharma P, Sahi S. Advances in Artificial Intelligence (AI)-assisted approaches in drug screening. Artif Intell Chem. 2024 Jun;2(1):100039. [CrossRef]
- Mouchlis VD, Afantitis A, Serra A, Fratello M, Papadiamantis AG, Aidinis V, et al. Advances in de Novo Drug Design: From Conventional to Machine Learning Methods. Int J Mol Sci. 2021 Feb 7;22(4):1676. [CrossRef]
- Das U, Chanda T, Kumar J, Peter A. Discovery of Natural MCL1 Inhibitors using Pharmacophore modelling, QSAR, Docking, ADMET, Molecular Dynamics, and DFT Analysis [Internet]. 2024 [cited 2025 Jan 9]. Available from: http://biorxiv.org/lookup/doi/10.1101/2024.10.14.618373. [CrossRef]
- Das U, Chandramouli L, Uttarkar A, Kumar J, Niranjan V. Discovery of natural compounds as novel FMS-like tyrosine kinase-3 (FLT3) therapeutic inhibitors for the treatment of acute myeloid leukemia: An in-silico approach. Asp Mol Med. 2025 Jun;5:100058. [CrossRef]
- Gangwal A, Ansari A, Ahmad I, Azad AK, Kumarasamy V, Subramaniyan V, et al. Generative artificial intelligence in drug discovery: basic framework, recent advances, challenges, and opportunities. Front Pharmacol. 2024;15:1331062. [CrossRef]
- Mroz AM, Posligua V, Tarzia A, Wolpert EH, Jelfs KE. Into the Unknown: How Computation Can Help Explore Uncharted Material Space. J Am Chem Soc. 2022 Oct 19;144(41):18730–43. [CrossRef]
- Han R, Yoon H, Kim G, Lee H, Lee Y. Revolutionizing Medicinal Chemistry: The Application of Artificial Intelligence (AI) in Early Drug Discovery. Pharm Basel Switz. 2023 Sep 6;16(9):1259. [CrossRef]
- Ivanenkov YA, Polykovskiy D, Bezrukov D, Zagribelnyy B, Aladinskiy V, Kamya P, et al. Chemistry42: An AI-Driven Platform for Molecular Design and Optimization. J Chem Inf Model. 2023 Feb 13;63(3):695–701. [CrossRef]
- Zeng X, Wang F, Luo Y, Kang S gu, Tang J, Lightstone FC, et al. Deep generative molecular design reshapes drug discovery. Cell Rep Med. 2022 Dec;3(12):100794. [CrossRef]
- Giordano D, Biancaniello C, Argenio MA, Facchiano A. Drug Design by Pharmacophore and Virtual Screening Approach. Pharm Basel Switz. 2022 May 23;15(5):646. [CrossRef]
- Çatalkaya S, Sabancı N, Yavuz SÇ, Sarıpınar E. The effect of stereoisomerism on the 4D-QSAR study of some dipeptidyl boron derivatives. Comput Biol Chem. 2020 Feb;84:107190. [CrossRef]
- Farghali H, Kutinová Canová N, Arora M. The potential applications of artificial intelligence in drug discovery and development. Physiol Res. 2021 Dec 30;70(Suppl4):S715–22. [CrossRef]
- Loeffler HH, He J, Tibo A, Janet JP, Voronov A, Mervin LH, et al. Reinvent 4: Modern AI–driven generative molecule design. J Cheminformatics. 2024 Feb 21;16(1):20. [CrossRef]
- Das U, Banerjee S, Sarkar M, Muhammad L F, Soni TK, Saha M, et al. Circular RNA vaccines: Pioneering the next-gen cancer immunotherapy. Cancer Pathog Ther. 2024 Dec;S2949713224000892. [CrossRef]
- Jiang Y, Yu Y, Kong M, Mei Y, Yuan L, Huang Z, et al. Artificial Intelligence for Retrosynthesis Prediction. Engineering. 2023 Jun;25:32–50.
- Ananikov VP. Top 20 influential AI-based technologies in chemistry. Artif Intell Chem. 2024 Dec;2(2):100075. [CrossRef]
- Liu Y, Yang Z, Yu Z, Liu Z, Liu D, Lin H, et al. Generative artificial intelligence and its applications in materials science: Current situation and future perspectives. J Materiomics. 2023 Jul;9(4):798–816. [CrossRef]
- Ochiai T, Inukai T, Akiyama M, Furui K, Ohue M, Matsumori N, et al. Variational autoencoder-based chemical latent space for large molecular structures with 3D complexity. Commun Chem. 2023 Nov 16;6(1):249. [CrossRef]
- Asperti A, Trentin M. Balancing Reconstruction Error and Kullback-Leibler Divergence in Variational Autoencoders. IEEE Access. 2020;8:199440–8. [CrossRef]
- Zheng W, Li J, Zhang Y. Desirable molecule discovery via generative latent space exploration. Vis Inform. 2023 Dec;7(4):13–21. [CrossRef]
- Abram KJ, McCloskey D. In Search of Disentanglement in Tandem Mass Spectrometry Datasets. Biomolecules. 2023 Sep 4;13(9):1343. [CrossRef]
- Sousa T, Correia J, Pereira V, Rocha M. Generative Deep Learning for Targeted Compound Design. J Chem Inf Model. 2021 Nov 22;61(11):5343–61. [CrossRef]
- Yang N, Wu H, Zeng K, Li Y, Bao S, Yan J. Molecule generation for drug design: A graph learning perspective. Fundam Res. 2024 Dec;S2667325824005259. [CrossRef]
- Vafaii H, Yates JL, Butts DA. Hierarchical VAEs provide a normative account of motion processing in the primate brain [Internet]. 2023 [cited 2025 Mar 30]. Available from: http://biorxiv.org/lookup/doi/10.1101/2023.09.27.559646. [CrossRef]
- Jang H, Seo S, Park S, Kim BJ, Choi GW, Choi J, et al. De novo drug design through gradient-based regularized search in information-theoretically controlled latent space. J Comput Aided Mol Des. 2024 Dec;38(1):32, s10822-024-00571–3. [CrossRef]
- Zhang Y, Li J, Chao X. ChemNav: An interactive visual tool to navigate in the latent space for chemical molecules discovery. Vis Inform. 2024 Dec;8(4):60–70. [CrossRef]
- Sharma P, Kumar M, Sharma HK, Biju SM. Generative adversarial networks (GANs): Introduction, Taxonomy, Variants, Limitations, and Applications. Multimed Tools Appl. 2024 Mar 26;83(41):88811–58. [CrossRef]
- Wu B, Li L, Cui Y, Zheng K. Cross-Adversarial Learning for Molecular Generation in Drug Design. Front Pharmacol. 2022 Jan 21;12:827606. [CrossRef]
- Tripathi S, Augustin AI, Dunlop A, Sukumaran R, Dheer S, Zavalny A, et al. Recent advances and application of generative adversarial networks in drug discovery, development, and targeting. Artif Intell Life Sci. 2022 Dec;2:100045. [CrossRef]
- Kucera T, Togninalli M, Meng-Papaxanthos L. Conditional generative modeling for de novo protein design with hierarchical functions. Wren J, editor. Bioinformatics. 2022 Jun 27;38(13):3454–61. [CrossRef]
- Putin E, Asadulaev A, Vanhaelen Q, Ivanenkov Y, Aladinskaya AV, Aliper A, et al. Adversarial Threshold Neural Computer for Molecular de Novo Design. Mol Pharm. 2018 Oct 1;15(10):4386–97. [CrossRef]
- Feng Y, Yang Y, Deng W, Chen H, Ran T. SyntaLinker-Hybrid: A deep learning approach for target specific drug design. Artif Intell Life Sci. 2022 Dec;2:100035. [CrossRef]
- De Cao N, Kipf T. MolGAN: An implicit generative model for small molecular graphs. 2018 [cited 2025 Mar 31]; Available from: https://arxiv.org/abs/1805.11973.
- Iglesias G, Talavera E, Díaz-Álvarez A. A survey on GANs for computer vision: Recent research, analysis and taxonomy. Comput Sci Rev. 2023 May;48:100553. [CrossRef]
- Méndez-Lucio O, Baillif B, Clevert DA, Rouquié D, Wichard J. De novo generation of hit-like molecules from gene expression signatures using artificial intelligence. Nat Commun. 2020 Jan 3;11(1):10. [CrossRef]
- Jiang J, Ke L, Chen L, Dou B, Zhu Y, Liu J, et al. Transformer technology in molecular science. WIREs Comput Mol Sci. 2024 Jul;14(4):e1725. [CrossRef]
- Chithrananda S, Grand G, Ramsundar B. ChemBERTa: Large-Scale Self-Supervised Pretraining for Molecular Property Prediction [Internet]. arXiv; 2020 [cited 2025 Mar 31]. Available from: https://arxiv.org/abs/2010.09885.
- Mswahili ME, Jeong YS. Transformer-based models for chemical SMILES representation: A comprehensive literature review. Heliyon. 2024 Oct;10(20):e39038. [CrossRef]
- Luong KD, Singh A. Application of Transformers in Cheminformatics. J Chem Inf Model. 2024 Jun 10;64(11):4392–409. [CrossRef]
- Yoshimori A, Bajorath J. DeepAS – Chemical language model for the extension of active analogue series. Bioorg Med Chem. 2022 Jul;66:116808. [CrossRef]
- Madani A, Krause B, Greene ER, Subramanian S, Mohr BP, Holton JM, et al. Large language models generate functional protein sequences across diverse families. Nat Biotechnol. 2023 Aug;41(8):1099–106. [CrossRef]
- Sumida KH, Núñez-Franco R, Kalvet I, Pellock SJ, Wicky BIM, Milles LF, et al. Improving Protein Expression, Stability, and Function with ProteinMPNN. J Am Chem Soc. 2024 Jan 24;146(3):2054–61. [CrossRef]
- Chandra A, Tünnermann L, Löfstedt T, Gratz R. Transformer-based deep learning for predicting protein properties in the life sciences. eLife. 2023 Jan 18;12:e82819. [CrossRef]
- Cerchia C, Lavecchia A. New avenues in artificial-intelligence-assisted drug discovery. Drug Discov Today. 2023 Apr;28(4):103516. [CrossRef]
- Ramos MC, Collison CJ, White AD. A review of large language models and autonomous agents in chemistry. Chem Sci. 2025;16(6):2514–72. [CrossRef]
- Parigi M, Martina S, Caruso F. Quantum-Noise-Driven Generative Diffusion Models. Adv Quantum Technol. 2024 Jul 15;2300401. [CrossRef]
- Soleymani F, Paquet E, Viktor HL, Michalowski W. Structure-based protein and small molecule generation using EGNN and diffusion models: A comprehensive review. Comput Struct Biotechnol J. 2024 Dec;23:2779–97. [CrossRef]
- Xu C, Liu R, Yao Y, Huang W, Li Z, Luo HB. 3D-EDiffMG: 3D equivariant diffusion-driven molecular generation to accelerate drug discovery. J Pharm Anal. 2025 Mar;101257. [CrossRef]
- Alakhdar A, Poczos B, Washburn N. Diffusion Models in De Novo Drug Design. J Chem Inf Model. 2024 Oct 14;64(19):7238–56. [CrossRef]
- Xu M, Yu L, Song Y, Shi C, Ermon S, Tang J. GeoDiff: a Geometric Diffusion Model for Molecular Conformation Generation [Internet]. arXiv; 2022 [cited 2025 Mar 31]. Available from: https://arxiv.org/abs/2203.02923.
- Watson JL, Juergens D, Bennett NR, Trippe BL, Yim J, Eisenach HE, et al. De novo design of protein structure and function with RFdiffusion. Nature. 2023 Aug 31;620(7976):1089–100. [CrossRef]
- Corso G, Stärk H, Jing B, Barzilay R, Jaakkola T. DiffDock: Diffusion Steps, Twists, and Turns for Molecular Docking [Internet]. arXiv; 2022 [cited 2025 Mar 31]. Available from: https://arxiv.org/abs/2210.01776.
- Wei YH. VAEs and GANs: Implicitly Approximating Complex Distributions with Simple Base Distributions and Deep Neural Networks -- Principles, Necessity, and Limitations [Internet]. arXiv; 2025 [cited 2025 Mar 31]. Available from: https://arxiv.org/abs/2503.01898.
- Wu AN, Stouffs R, Biljecki F. Generative Adversarial Networks in the built environment: A comprehensive review of the application of GANs across data types and scales. Build Environ. 2022 Sep;223:109477. [CrossRef]
- Jiang J, Chen L, Ke L, Dou B, Zhang C, Feng H, et al. A review of transformers in drug discovery and beyond. J Pharm Anal. 2024 Aug;101081. [CrossRef]
- Chen M, Mei S, Fan J, Wang M. Opportunities and challenges of diffusion models for generative AI. Natl Sci Rev. 2024 Nov 14;11(12):nwae348. [CrossRef]
- Gupta R, Tiwari S, Chaudhary P. Generative AI Techniques and Models. In: Generative AI: Techniques, Models and Applications [Internet]. Cham: Springer Nature Switzerland; 2025 [cited 2025 Mar 31]. p. 45–64. (Lecture Notes on Data Engineering and Communications Technologies; vol. 241). Available from: https://link.springer.com/10.1007/978-3-031-82062-5_3.
- Li C, Zhang T, Du X, Zhang Y, Xie H. Generative AI models for different steps in architectural design: A literature review. Front Archit Res. 2025 Jun;14(3):759–83. [CrossRef]
- Shu D, Li Z, Barati Farimani A. A physics-informed diffusion model for high-fidelity flow field reconstruction. J Comput Phys. 2023 Apr;478:111972. [CrossRef]
- Connor MC, Canal GH, Rozell CJ. Variational Autoencoder with Learned Latent Structure [Internet]. arXiv; 2020 [cited 2025 Mar 31]. Available from: https://arxiv.org/abs/2006.10597.
- Chen N, Klushyn A, Ferroni F, Bayer J, van der Smagt P. Learning Flat Latent Manifolds with VAEs. 2020 [cited 2025 Mar 31]; Available from: https://arxiv.org/abs/2002.04881.
- Chandra R, Horne RI, Vendruscolo M. Bayesian Optimization in the Latent Space of a Variational Autoencoder for the Generation of Selective FLT3 Inhibitors. J Chem Theory Comput. 2024 Jan 9;20(1):469–76. [CrossRef]
- Yang X, Wang Y, Byrne R, Schneider G, Yang S. Concepts of Artificial Intelligence for Computer-Assisted Drug Discovery. Chem Rev. 2019 Sep 25;119(18):10520–94. [CrossRef]
- Trunz E, Weinmann M, Merzbach S, Klein R. Efficient structuring of the latent space for controllable data reconstruction and compression. Graph Vis Comput. 2022 Dec;7:200059. [CrossRef]
- Shen C, Krenn M, Eppel S, Aspuru-Guzik A. Deep molecular dreaming: inverse machine learning for de-novo molecular design and interpretability with surjective representations. Mach Learn Sci Technol. 2021 Sep 1;2(3):03LT02. [CrossRef]
- Prykhodko O, Johansson SV, Kotsias PC, Arús-Pous J, Bjerrum EJ, Engkvist O, et al. A de novo molecular generation method using latent vector based generative adversarial network. J Cheminformatics. 2019 Dec;11(1):74. [CrossRef]
- Rossi E, Wheeler JM, Sebastiani M. High-speed nanoindentation mapping: A review of recent advances and applications. Curr Opin Solid State Mater Sci. 2023 Oct;27(5):101107. [CrossRef]
- Bilodeau C, Jin W, Jaakkola T, Barzilay R, Jensen KF. Generative models for molecular discovery: Recent advances and challenges. WIREs Comput Mol Sci. 2022 Sep;12(5):e1608. [CrossRef]
- Guo J, Schwaller P. Directly optimizing for synthesizability in generative molecular design using retrosynthesis models. Chem Sci. 2025;10.1039.D5SC01476J. [CrossRef]
- Wang J, Zhu F. ExSelfRL: An exploration-inspired self-supervised reinforcement learning approach to molecular generation. Expert Syst Appl. 2025 Jan;260:125410. [CrossRef]
- Nakamura S, Yasuo N, Sekijima M. Molecular optimization using a conditional transformer for reaction-aware compound exploration with reinforcement learning. Commun Chem. 2025 Feb 8;8(1):40. [CrossRef]
- Korn M, Ehrt C, Ruggiu F, Gastreich M, Rarey M. Navigating large chemical spaces in early-phase drug discovery. Curr Opin Struct Biol. 2023 Jun;80:102578. [CrossRef]
- Anstine DM, Isayev O. Generative Models as an Emerging Paradigm in the Chemical Sciences. J Am Chem Soc. 2023 Apr 26;145(16):8736–50. [CrossRef]
- Świechowski M, Godlewski K, Sawicki B, Mańdziuk J. Monte Carlo Tree Search: a review of recent modifications and applications. Artif Intell Rev. 2023 Mar;56(3):2497–562. [CrossRef]
- Park J, Ahn J, Choi J, Kim J. Mol-AIR: Molecular Reinforcement Learning with Adaptive Intrinsic Rewards for Goal-Directed Molecular Generation. J Chem Inf Model. 2025 Mar 10;65(5):2283–96. [CrossRef]
- Zhavoronkov A, Ivanenkov YA, Aliper A, Veselov MS, Aladinskiy VA, Aladinskaya AV, et al. Deep learning enables rapid identification of potent DDR1 kinase inhibitors. Nat Biotechnol. 2019 Sep;37(9):1038–40. [CrossRef]
- Greenstein BL, Elsey DC, Hutchison GR. Determining best practices for using genetic algorithms in molecular discovery. J Chem Phys. 2023 Sep 7;159(9):091501. [CrossRef]
- McCall J. Genetic algorithms for modelling and optimisation. J Comput Appl Math. 2005 Dec;184(1):205–22. [CrossRef]
- Kim M, Gu J, Yuan Y, Yun T, Liu Z, Bengio Y, et al. Offline Model-Based Optimization: Comprehensive Review [Internet]. arXiv; 2025 [cited 2025 Mar 31]. Available from: https://arxiv.org/abs/2503.17286.
- Schulam P, Muslea I. Improving the Exploration/Exploitation Trade-Off in Web Content Discovery. In: Companion Proceedings of the ACM Web Conference 2023 [Internet]. Austin TX USA: ACM; 2023 [cited 2025 Mar 31]. p. 1183–9. Available from: https://dl.acm.org/doi/10.1145/3543873.3587574.
- Gupta P, Ding B, Guan C, Ding D. Generative AI: A systematic review using topic modelling techniques. Data Inf Manag. 2024 Jun;8(2):100066. [CrossRef]
- Abeer ANMN, Urban NM, Weil MR, Alexander FJ, Yoon BJ. Multi-objective latent space optimization of generative molecular design models. Patterns. 2024 Oct;5(10):101042. [CrossRef]
- Menon D, Ranganathan R. A Generative Approach to Materials Discovery, Design, and Optimization. ACS Omega. 2022 Aug 2;7(30):25958–73. [CrossRef]
- Aal E Ali RS, Meng J, Khan MEI, Jiang X. Machine learning advancements in organic synthesis: A focused exploration of artificial intelligence applications in chemistry. Artif Intell Chem. 2024 Jun;2(1):100049.
- Vogt M. Exploring chemical space — Generative models and their evaluation. Artif Intell Life Sci. 2023 Dec;3:100064. [CrossRef]
- Rehman AU, Li M, Wu B, Ali Y, Rasheed S, Shaheen S, et al. Role of Artificial Intelligence in Revolutionizing Drug Discovery. Fundam Res. 2024 May;S266732582400205X. [CrossRef]
- Magar R, Wang Y, Barati Farimani A. Crystal twins: self-supervised learning for crystalline material property prediction. Npj Comput Mater. 2022 Nov 10;8(1):231. [CrossRef]
- Wang J, Guan J, Zhou S. Molecular property prediction by contrastive learning with attention-guided positive sample selection. Wren J, editor. Bioinformatics. 2023 May 4;39(5):btad258. [CrossRef]
- Yang X, Wang Y, Lin Y, Zhang M, Liu O, Shuai J, et al. A Multi-Task Self-Supervised Strategy for Predicting Molecular Properties and FGFR1 Inhibitors. Adv Sci. 2025 Feb 8;2412987. [CrossRef]
- Cafiero M. Transformer-Decoder GPT Models for Generating Virtual Screening Libraries of HMG-Coenzyme A Reductase Inhibitors: Effects of Temperature, Prompt Length, and Transfer-Learning Strategies. J Chem Inf Model. 2024 Nov 25;64(22):8464–80. [CrossRef]
- Chen S, Guo W. Auto-Encoders in Deep Learning—A Review with New Perspectives. Mathematics. 2023 Apr 7;11(8):1777. [CrossRef]
- Korshunova M, Huang N, Capuzzi S, Radchenko DS, Savych O, Moroz YS, et al. Generative and reinforcement learning approaches for the automated de novo design of bioactive compounds. Commun Chem. 2022 Oct 18;5(1):129. [CrossRef]
- Popova M, Isayev O, Tropsha A. Deep reinforcement learning for de novo drug design. Sci Adv. 2018 Jul 6;4(7):eaap7885. [CrossRef]
- Tan RK, Liu Y, Xie L. Reinforcement learning for systems pharmacology-oriented and personalized drug design. Expert Opin Drug Discov. 2022 Aug;17(8):849–63. [CrossRef]
- Dodds M, Guo J, Löhr T, Tibo A, Engkvist O, Janet JP. Sample efficient reinforcement learning with active learning for molecular design. Chem Sci. 2024;15(11):4146–60. [CrossRef]
- Reiser P, Neubert M, Eberhard A, Torresi L, Zhou C, Shao C, et al. Graph neural networks for materials science and chemistry. Commun Mater. 2022 Nov 26;3(1):93. [CrossRef]
- Abate C, Decherchi S, Cavalli A. Graph neural networks for conditional de novo drug design. WIREs Comput Mol Sci. 2023 Jul;13(4):e1651. [CrossRef]
- Zheng S, Lei Z, Ai H, Chen H, Deng D, Yang Y. Deep scaffold hopping with multimodal transformer neural networks. J Cheminformatics. 2021 Nov 13;13(1):87. [CrossRef]
- Hu C, Li S, Yang C, Chen J, Xiong Y, Fan G, et al. ScaffoldGVAE: scaffold generation and hopping of drug molecules via a variational autoencoder based on multi-view graph neural networks. J Cheminformatics. 2023 Oct 4;15(1):91. [CrossRef]
- Wu KE, Yang KK, Van Den Berg R, Alamdari S, Zou JY, Lu AX, et al. Protein structure generation via folding diffusion. Nat Commun. 2024 Feb 5;15(1):1059. [CrossRef]
- Sarumi OA, Heider D. Large language models and their applications in bioinformatics. Comput Struct Biotechnol J. 2024 Dec;23:3498–505. [CrossRef]
- Valentini G, Malchiodi D, Gliozzo J, Mesiti M, Soto-Gomez M, Cabri A, et al. The promises of large language models for protein design and modeling. Front Bioinforma. 2023;3:1304099. [CrossRef]
- Nana Teukam YG, Kwate Dassi L, Manica M, Probst D, Schwaller P, Laino T. Language models can identify enzymatic binding sites in protein sequences. Comput Struct Biotechnol J. 2024 Dec;23:1929–37. [CrossRef]
- Liu J, Yang M, Yu Y, Xu H, Wang T, Li K, et al. Advancing bioinformatics with large language models: components, applications and perspectives. ArXiv. 2025 Jan 31;arXiv:2401.04155v2.
- Bzdok D, Thieme A, Levkovskyy O, Wren P, Ray T, Reddy S. Data science opportunities of large language models for neuroscience and biomedicine. Neuron. 2024 Mar;112(5):698–717. [CrossRef]
- Hie BL, Shanker VR, Xu D, Bruun TUJ, Weidenbacher PA, Tang S, et al. Efficient evolution of human antibodies from general protein language models. Nat Biotechnol. 2024 Feb;42(2):275–83. [CrossRef]
- Kim J, McFee M, Fang Q, Abdin O, Kim PM. Computational and artificial intelligence-based methods for antibody development. Trends Pharmacol Sci. 2023 Mar;44(3):175–89. [CrossRef]
- Luo S, Su Y, Peng X, Wang S, Peng J, Ma J. Antigen-Specific Antibody Design and Optimization with Diffusion-Based Generative Models for Protein Structures [Internet]. 2022 [cited 2025 Mar 31]. Available from: http://biorxiv.org/lookup/doi/10.1101/2022.07.10.499510.
- Dewaker V, Morya VK, Kim YH, Park ST, Kim HS, Koh YH. Revolutionizing oncology: the role of Artificial Intelligence (AI) as an antibody design, and optimization tools. Biomark Res. 2025 Mar 29;13(1):52. [CrossRef]
- Yang J, Li FZ, Arnold FH. Opportunities and Challenges for Machine Learning-Assisted Enzyme Engineering. ACS Cent Sci. 2024 Feb 28;10(2):226–41. [CrossRef]
- Zhou J, Huang M. Navigating the landscape of enzyme design: from molecular simulations to machine learning. Chem Soc Rev. 2024;53(16):8202–39. [CrossRef]
- Orsi E, Schada Von Borzyskowski L, Noack S, Nikel PI, Lindner SN. Automated in vivo enzyme engineering accelerates biocatalyst optimization. Nat Commun. 2024 Apr 24;15(1):3447. [CrossRef]
- Baum ZJ, Yu X, Ayala PY, Zhao Y, Watkins SP, Zhou Q. Artificial Intelligence in Chemistry: Current Trends and Future Directions. J Chem Inf Model. 2021 Jul 26;61(7):3197–212. [CrossRef]
- Arya SS, Dias SB, Jelinek HF, Hadjileontiadis LJ, Pappa AM. The convergence of traditional and digital biomarkers through AI-assisted biosensing: A new era in translational diagnostics? Biosens Bioelectron. 2023 Sep;235:115387. [CrossRef]
- Stärk H, Ganea OE, Pattanaik L, Barzilay R, Jaakkola T. EquiBind: Geometric Deep Learning for Drug Binding Structure Prediction. 2022 [cited 2025 Mar 31]; Available from: https://arxiv.org/abs/2202.05146.
- Ketata MA, Laue C, Mammadov R, Stärk H, Wu M, Corso G, et al. DiffDock-PP: Rigid Protein-Protein Docking with Diffusion Models [Internet]. arXiv; 2023 [cited 2025 Mar 31]. Available from: https://arxiv.org/abs/2304.03889.
- Yang C, Chen EA, Zhang Y. Protein-Ligand Docking in the Machine-Learning Era. Mol Basel Switz. 2022 Jul 18;27(14):4568. [CrossRef]
- Cao D, Chen M, Zhang R, Wang Z, Huang M, Yu J, et al. SurfDock is a surface-informed diffusion generative model for reliable and accurate protein–ligand complex prediction. Nat Methods. 2025 Feb;22(2):310–22. [CrossRef]
- B Fortela DL, Mikolajczyk AP, Carnes MR, Sharp W, Revellame E, Hernandez R, et al. Predicting Molecular Docking of Per- and Polyfluoroalkyl Substances to Blood Protein Using Generative Artificial Intelligence Algorithm Diffdock. BioTechniques. 2024 Jan;76(1):14–26. [CrossRef]
- Wang Y, Jiao Q, Wang J, Cai X, Zhao W, Cui X. Prediction of protein-ligand binding affinity with deep learning. Comput Struct Biotechnol J. 2023;21:5796–806. [CrossRef]
- Wang DD, Wu W, Wang R. Structure-based, deep-learning models for protein-ligand binding affinity prediction. J Cheminformatics. 2024 Jan 3;16(1):2. [CrossRef]
- Zhang S, Jin Y, Liu T, Wang Q, Zhang Z, Zhao S, et al. SS-GNN: A Simple-Structured Graph Neural Network for Affinity Prediction. ACS Omega. 2023 Jun 27;8(25):22496–507. [CrossRef]
- Wang H. Prediction of protein–ligand binding affinity via deep learning models. Brief Bioinform. 2024 Jan 22;25(2):bbae081. [CrossRef]
- Wang R, Fang X, Lu Y, Wang S. The PDBbind Database: Collection of Binding Affinities for Protein−Ligand Complexes with Known Three-Dimensional Structures. J Med Chem. 2004 Jun 1;47(12):2977–80. [CrossRef]
- Weidman JD, Sajjan M, Mikolas C, Stewart ZJ, Pollanen J, Kais S, et al. Quantum computing and chemistry. Cell Rep Phys Sci. 2024 Sep;5(9):102105. [CrossRef]
- Morawietz T, Artrith N. Machine learning-accelerated quantum mechanics-based atomistic simulations for industrial applications. J Comput Aided Mol Des. 2021 Apr;35(4):557–86. [CrossRef]
- Doga H, Raubenolt B, Cumbo F, Joshi J, DiFilippo FP, Qin J, et al. A Perspective on Protein Structure Prediction Using Quantum Computers. J Chem Theory Comput. 2024 May 14;20(9):3359–78. [CrossRef]
- How ML, Cheah SM. Forging the Future: Strategic Approaches to Quantum AI Integration for Industry Transformation. AI. 2024 Jan 29;5(1):290–323. [CrossRef]
- Liu X, Jiang S, Duan X, Vasan A, Liu C, Tien C chan, et al. Binding Affinity Prediction: From Conventional to Machine Learning-Based Approaches [Internet]. arXiv; 2024 [cited 2025 Mar 31]. Available from: https://arxiv.org/abs/2410.00709.
- Yan J, Ye Z, Yang Z, Lu C, Zhang S, Liu Q, et al. Multi-task bioassay pre-training for protein-ligand binding affinity prediction. Brief Bioinform. 2023 Nov 22;25(1):bbad451. [CrossRef]
- Schwaller P, Gaudin T, Lányi D, Bekas C, Laino T. “Found in Translation”: predicting outcomes of complex organic chemistry reactions using neural sequence-to-sequence models. Chem Sci. 2018;9(28):6091–8. [CrossRef]
- Jackson I, Jesus Saenz M, Ivanov D. From natural language to simulations: applying AI to automate simulation modelling of logistics systems. Int J Prod Res. 2024 Feb 16;62(4):1434–57. [CrossRef]
- Sinha S, Lee YM. Challenges with developing and deploying AI models and applications in industrial systems. Discov Artif Intell. 2024 Aug 16;4(1):55. [CrossRef]
- Hong S, Zhuo HH, Jin K, Shao G, Zhou Z. Retrosynthetic planning with experience-guided Monte Carlo tree search. Commun Chem. 2023 Jun 10;6(1):120. [CrossRef]
- Lai H, Kannas C, Hassen AK, Granqvist E, Westerlund AM, Clevert DA, et al. Multi-objective synthesis planning by means of Monte Carlo Tree search. Artif Intell Life Sci. 2025 Jun;7:100130. [CrossRef]
- Terven J. Deep Reinforcement Learning: A Chronological Overview and Methods. AI. 2025 Feb 24;6(3):46. [CrossRef]
- Nambiar AMK, Breen CP, Hart T, Kulesza T, Jamison TF, Jensen KF. Bayesian Optimization of Computer-Proposed Multistep Synthetic Routes on an Automated Robotic Flow Platform. ACS Cent Sci. 2022 Jun 22;8(6):825–36. [CrossRef]
- Schilter O, Gutierrez DP, Folkmann LM, Castrogiovanni A, García-Durán A, Zipoli F, et al. Combining Bayesian optimization and automation to simultaneously optimize reaction conditions and routes. Chem Sci. 2024;15(20):7732–41. [CrossRef]
- Tachibana R, Zhang K, Zou Z, Burgener S, Ward TR. A Customized Bayesian Algorithm to Optimize Enzyme-Catalyzed Reactions. ACS Sustain Chem Eng. 2023 Aug 21;11(33):12336–44. [CrossRef]
- Omotehinwa TO, Lawrence MO, Oyewola DO, Dada EG. Bayesian optimization of one-dimensional convolutional neural networks (1D CNN) for early diagnosis of Autistic Spectrum Disorder. J Comput Math Data Sci. 2024 Dec;13:100105. [CrossRef]
- Kwon Y, Lee D, Kim JW, Choi YS, Kim S. Exploring Optimal Reaction Conditions Guided by Graph Neural Networks and Bayesian Optimization. ACS Omega. 2022 Dec 13;7(49):44939–50. [CrossRef]
- Parrot M, Tajmouati H, Da Silva VBR, Atwood BR, Fourcade R, Gaston-Mathé Y, et al. Integrating synthetic accessibility with AI-based generative drug design. J Cheminformatics. 2023 Sep 19;15(1):83. [CrossRef]
- Retchin M, Wang Y, Takaba K, Chodera JD. DrugGym: A testbed for the economics of autonomous drug discovery [Internet]. 2024 [cited 2025 Mar 31]. Available from: http://biorxiv.org/lookup/doi/10.1101/2024.05.28.596296.
- D. Segall M. Multi-Parameter Optimization: Identifying High Quality Compounds with a Balance of Properties. Curr Drug Metab. 2012 Mar 1;18(9):1292–310.
- Wager TT, Hou X, Verhoest PR, Villalobos A. Central Nervous System Multiparameter Optimization Desirability: Application in Drug Discovery. ACS Chem Neurosci. 2016 Jun 15;7(6):767–75. [CrossRef]
- Joshi-Barr S, Wampole M. Artificial Intelligence for Drug Toxicity and Safety. In: Hock FJ, Pugsley MK, editors. Drug Discovery and Evaluation: Safety and Pharmacokinetic Assays [Internet]. Cham: Springer International Publishing; 2024 [cited 2025 Mar 31]. p. 2637–71. Available from: https://link.springer.com/10.1007/978-3-031-35529-5_134.
- Burki T. A new paradigm for drug development. Lancet Digit Health. 2020 May;2(5):e226–7. [CrossRef]
- Shanehsazzadeh A, McPartlon M, Kasun G, Steiger AK, Sutton JM, Yassine E, et al. Unlocking de novo antibody design with generative artificial intelligence [Internet]. 2023 [cited 2025 Mar 31]. Available from: http://biorxiv.org/lookup/doi/10.1101/2023.01.08.523187.
- Visan AI, Negut I. Integrating Artificial Intelligence for Drug Discovery in the Context of Revolutionizing Drug Delivery. Life Basel Switz. 2024 Feb 7;14(2):233. [CrossRef]
- Guan S, Wang G. Drug discovery and development in the era of artificial intelligence: From machine learning to large language models. Artif Intell Chem. 2024 Jun;2(1):100070. [CrossRef]
- Schneider G. Automating drug discovery. Nat Rev Drug Discov. 2018 Feb;17(2):97–113. [CrossRef]
- Atomwise AIMS Program. AI is a viable alternative to high throughput screening: a 318-target study. Sci Rep. 2024 Apr 2;14(1):7526.
- Dhudum R, Ganeshpurkar A, Pawar A. Revolutionizing Drug Discovery: A Comprehensive Review of AI Applications. Drugs Drug Candidates. 2024 Feb 13;3(1):148–71. [CrossRef]
- Qiu X, Li H, Ver Steeg G, Godzik A. Advances in AI for Protein Structure Prediction: Implications for Cancer Drug Discovery and Development. Biomolecules. 2024 Mar 12;14(3):339. [CrossRef]
- Qin Y, Chen Z, Peng Y, Xiao Y, Zhong T, Yu X. Deep learning methods for protein structure prediction. MedComm – Future Med. 2024 Sep;3(3):e96. [CrossRef]
- Xu Y, Liu X, Cao X, Huang C, Liu E, Qian S, et al. Artificial intelligence: A powerful paradigm for scientific research. The Innovation. 2021 Nov;2(4):100179. [CrossRef]
- Sliwoski G, Kothiwale S, Meiler J, Lowe EW. Computational methods in drug discovery. Pharmacol Rev. 2014;66(1):334–95. [CrossRef]
- Khakzad H, Igashov I, Schneuing A, Goverde C, Bronstein M, Correia B. A new age in protein design empowered by deep learning. Cell Syst. 2023 Nov;14(11):925–39. [CrossRef]
- Fu C, Chen Q. The future of pharmaceuticals: Artificial intelligence in drug discovery and development. J Pharm Anal. 2025 Feb;101248. [CrossRef]
- Wang X, Xu K, Tan Y, Liu S, Zhou J. Possibilities of Using De Novo Design for Generating Diverse Functional Food Enzymes. Int J Mol Sci. 2023 Feb 14;24(4):3827. [CrossRef]
- Bhisetti G, Fang C. Artificial Intelligence–Enabled De Novo Design of Novel Compounds that Are Synthesizable. In: Heifetz A, editor. Artificial Intelligence in Drug Design [Internet]. New York, NY: Springer US; 2022 [cited 2025 Mar 31]. p. 409–19. (Methods in Molecular Biology; vol. 2390). Available from: https://link.springer.com/10.1007/978-1-0716-1787-8_17.
- Shi Y, Hu H. AI accelerated discovery of self-assembling peptides. Biomater Transl. 2023;4(4):291–3. [CrossRef]
- Ding N, Yuan Z, Ma Z, Wu Y, Yin L. AI-Assisted Rational Design and Activity Prediction of Biological Elements for Optimizing Transcription-Factor-Based Biosensors. Mol Basel Switz. 2024 Jul 26;29(15):3512. [CrossRef]
- Divine R, Dang HV, Ueda G, Fallas JA, Vulovic I, Sheffler W, et al. Designed proteins assemble antibodies into modular nanocages. Science. 2021 Apr 2;372(6537):eabd9994. [CrossRef]
- Tom G, Schmid SP, Baird SG, Cao Y, Darvish K, Hao H, et al. Self-Driving Laboratories for Chemistry and Materials Science. Chem Rev. 2024 Aug 28;124(16):9633–732. [CrossRef]
- Blunt NS, Camps J, Crawford O, Izsák R, Leontica S, Mirani A, et al. Perspective on the Current State-of-the-Art of Quantum Computing for Drug Discovery Applications. J Chem Theory Comput. 2022 Dec 13;18(12):7001–23. [CrossRef]
- Ur Rasool R, Ahmad HF, Rafique W, Qayyum A, Qadir J, Anwar Z. Quantum Computing for Healthcare: A Review. Future Internet. 2023 Feb 27;15(3):94. [CrossRef]
- Outeiral C, Strahm M, Shi J, Morris GM, Benjamin SC, Deane CM. The prospects of quantum computing in computational molecular biology. WIREs Comput Mol Sci. 2021 Jan;11(1):e1481. [CrossRef]
- Serrano DR, Luciano FC, Anaya BJ, Ongoren B, Kara A, Molina G, et al. Artificial Intelligence (AI) Applications in Drug Discovery and Drug Delivery: Revolutionizing Personalized Medicine. Pharmaceutics. 2024 Oct 14;16(10):1328. [CrossRef]
- Cheong BC. Transparency and accountability in AI systems: safeguarding wellbeing in the age of algorithmic decision-making. Front Hum Dyn. 2024 Jul 3;6:1421273. [CrossRef]
- Choudhury A, Asan O. Role of Artificial Intelligence in Patient Safety Outcomes: Systematic Literature Review. JMIR Med Inform. 2020 Jul 24;8(7):e18599. [CrossRef]
- Alizadehsani R, Oyelere SS, Hussain S, Jagatheesaperumal SK, Calixto RR, Rahouti M, et al. Explainable Artificial Intelligence for Drug Discovery and Development: A Comprehensive Survey. IEEE Access. 2024;12:35796–812. [CrossRef]
- Kapustina O, Burmakina P, Gubina N, Serov N, Vinogradov V. User-friendly and industry-integrated AI for medicinal chemists and pharmaceuticals. Artif Intell Chem. 2024 Dec;2(2):100072. [CrossRef]
- Taherdoost H, Ghofrani A. AI’s role in revolutionizing personalized medicine by reshaping pharmacogenomics and drug therapy. Intell Pharm. 2024 Oct;2(5):643–50. [CrossRef]
- Saini JPS, Thakur A, Yadav D. AI-driven innovations in pharmaceuticals: optimizing drug discovery and industry operations. RSC Pharm. 2025;10.1039.D4PM00323C. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



