Submitted:
02 April 2025
Posted:
03 April 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. The Outline of the p53 Pathway
3. The Current Picture of Interactions Between Bacterial Infections and the p53 Pathway
4. The p53 and Inflammasomes
5. Genes of Bacteriostatic Proteins Activated by p53
6. p53 Promotes the Detection and Destruction of Bacteria
7. TNFRSF14, p53, and Bacteria
8. p53 Promotes the Presentation of Bacterial Metabolites to Lymphocytes
9. Concluding Remarks
Funding
Acknowledgments
Conflicts of Interest
References
- Levine, A.J. The many faces of p53: something for everyone. J. Mol. Cell Biol. 2019, 11, 524–530. [Google Scholar] [CrossRef] [PubMed]
- Nagata, J.; Kijima, H.; Takagi, A.; Ito, M.; Goto, K.; Yamazaki, H.; Nakamura, M.; Mine, T.; Ueyama, Y. Helicobacter pylori induces chronic active gastritis in p53-knockout mice. Int. J. Mol. Med. 2004, 13, 773–777. [Google Scholar] [CrossRef] [PubMed]
- Carlsen, L.; Zhang, S.; Tian, X.; De La Cruz, A.; George, A.; Arnoff, T.E.; El-Deiry, W.S. The role of p53 in anti-tumor immunity and response to immunotherapy. Front. Mol. Biosci. 2023, 10, 1148389. [Google Scholar] [CrossRef]
- Łasut-Szyszka, B.; Rusin, M. The Wheel of p53 Helps to Drive the Immune System. Int. J. Mol. Sci. 2023, 24, 7645. [Google Scholar] [CrossRef] [PubMed]
- Łasut-Szyszka, B.; Gdowicz-Kłosok, A.; Małachowska, B.; Krześniak, M.; Będzińska, A.; Gawin, M.; Pietrowska, M.; Rusin, M. Transcriptomic and proteomic study of cancer cell lines exposed to actinomycin D and nutlin-3a reveals numerous, novel candidates for p53-regulated genes. Chem. Biol. Interact. 2024, 392, 110946. [Google Scholar] [CrossRef]
- Timofeev, O.; Stiewe, T. Rely on Each Other: DNA Binding Cooperativity Shapes p53 Functions in Tumor Suppression and Cancer Therapy. Cancers (Basel) 2021, 13, 2422. [Google Scholar] [CrossRef]
- Wang, H.; Guo, M.; Wei, H.; Chen, Y. Targeting p53 pathways: mechanisms, structures, and advances in therapy. Signal Transduct. Target. Ther. 2023, 8, 92. [Google Scholar] [CrossRef]
- Fischer, M.; Sammons, M.A. Determinants of p53 DNA binding, gene regulation, and cell fate decisions. Cell Death Differ. 2024, 31, 836–843. [Google Scholar] [CrossRef]
- Rusin, M. The p53 protein - not only the guardian of the genome. Postepy Biochem. 2024, 70, 71–87. [Google Scholar] [CrossRef]
- Liu, Y.; Su, Z.; Tavana, O.; Gu, W. Understanding the complexity of p53 in a new era of tumor suppression. Cancer Cell 2024, 42, 946–967. [Google Scholar] [CrossRef]
- Lane, D.P. Cancer. p53, guardian of the genome. Nature 1992, 358, 15–16. [Google Scholar] [CrossRef] [PubMed]
- Harford, J.B. A Second Career for p53 as A Broad-Spectrum Antiviral? Viruses 2023, 15, 2377. [Google Scholar] [CrossRef]
- Kocik, J.; Machula, M.; Wisniewska, A.; Surmiak, E.; Holak, T.A.; Skalniak, L. Helping the Released Guardian: Drug Combinations for Supporting the Anticancer Activity of HDM2 (MDM2) Antagonists. Cancers (Basel) 2019, 11, 1014. [Google Scholar] [CrossRef] [PubMed]
- Uxa, S.; Bernhart, S.H.; Mages, C.F.S.; Fischer, M.; Kohler, R.; Hoffmann, S.; Stadler, P.F.; Engeland, K.; Müller, G.A. DREAM and RB cooperate to induce gene repression and cell-cycle arrest in response to p53 activation. Nucleic Acids Res. 2019, 47, 9087–9103. [Google Scholar] [CrossRef] [PubMed]
- Schwab, K.; Riege, K.; Coronel, L.; Stanko, C.; Förste, S.; Hoffmann, S.; Fischer, M. p53 target ANKRA2 cooperates with RFX7 to regulate tumor suppressor genes. Cell Death Discov. 2024, 10, 376. [Google Scholar] [CrossRef]
- Kruse, J.P.; Gu, W. Modes of p53 regulation. Cell 2009, 137, 609–622. [Google Scholar] [CrossRef]
- Smeenk, L.; van Heeringen, S.J.; Koeppel, M.; Gilbert, B.; Janssen-Megens, E.; Stunnenberg, H.G.; Lohrum, M. Role of p53 serine 46 in p53 target gene regulation. PLoS One 2011, 6, e17574. [Google Scholar] [CrossRef] [PubMed]
- Liebl, M.C.; Hofmann, T.G. Cell Fate Regulation upon DNA Damage: p53 Serine 46 Kinases Pave the Cell Death Road. Bioessays 2019, 41, e1900127. [Google Scholar] [CrossRef]
- Childs, B.G.; Baker, D.J.; Kirkland, J.L.; Campisi, J.; van Deursen, J.M. Senescence and apoptosis: dueling or complementary cell fates? EMBO Rep. 2014, 15, 1139–1153. [Google Scholar] [CrossRef]
- Łasut-Szyszka B, Gdowicz-Kłosok A, Krześniak M, Głowala-Kosińska M, Będzińska A, Rusin M. Strong activation of p53 by actinomycin D and nutlin-3a overcomes the resistance of cancer cells to the pro-apoptotic activity of the FAS ligand. Apoptosis. 2024 29(9-10):1515-1528. [CrossRef]
- Liu, Y.; Gu, W. The complexity of p53-mediated metabolic regulation in tumor suppression. Semin. Cancer Biol. 2022, 85, 4–32. [Google Scholar] [CrossRef]
- Andrysik, Z.; Galbraith, M.D.; Guarnieri, A.L.; Zaccara, S.; Sullivan, K.D.; Pandey, A.; MacBeth, M.; Inga, A.; Espinosa, J.M. Identification of a core TP53 transcriptional program with highly distributed tumor suppressive activity. Genome Res. 2017, 27, 1645–1657. [Google Scholar] [CrossRef]
- Tatavosian, R.; Donovan, M.G.; Galbraith, M.D.; Duc, H.N.; Szwarc, M.M.; Joshi, M.U.; Frieman, A.; Bilousova, G.; Cao, Y.; Smith, K.P.; Song, K.; Rachubinski, A.L.; Andrysik, Z.; Espinosa, J.M. Cell differentiation modifies the p53 transcriptional program through a combination of gene silencing and constitutive transactivation. Cell Death Differ. 2023, 30, 952–965. [Google Scholar] [CrossRef]
- Fischer, M. Census and evaluation of p53 target genes. Oncogene 2017, 36, 3943–3956. [Google Scholar] [CrossRef] [PubMed]
- Fischer, M.; Schwarz, R.; Riege, K.; DeCaprio, J.A.; Hoffmann, S. TargetGeneReg 2.0: a comprehensive web-atlas for p53, p63, and cell cycle-dependent gene regulation. NAR Cancer 2022, 4, zcac009. [Google Scholar] [CrossRef] [PubMed]
- Chi, S.W. Structural insights into the transcription-independent apoptotic pathway of p53. BMB Rep. 2014, 47, 167–172. [Google Scholar] [CrossRef]
- Jiang, P.; Du, W.; Wang, X.; Mancuso, A.; Gao, X.; Wu, M.; Yang, X. p53 regulates biosynthesis through direct inactivation of glucose-6-phosphate dehydrogenase. Nat. Cell Biol. 2011, 13, 310–316. [Google Scholar] [CrossRef]
- Liu, M.; Jia, Y.; Xie, Y.; Zhang, C.; Ma, J.; Sun, C.; Ju, J. Identification of the Actinomycin D Biosynthetic Pathway from Marine-Derived Streptomyces costaricanus SCSIO ZS0073. Mar. Drugs 2019, 17, 240. [Google Scholar] [CrossRef]
- Shacter, E.; Beecham, E.J.; Covey, J.M.; Kohn, K.W.; Potter, M. Activated neutrophils induce prolonged DNA damage in neighboring cells. Carcinogenesis 1988, 9, 2297–2304. [Google Scholar] [CrossRef]
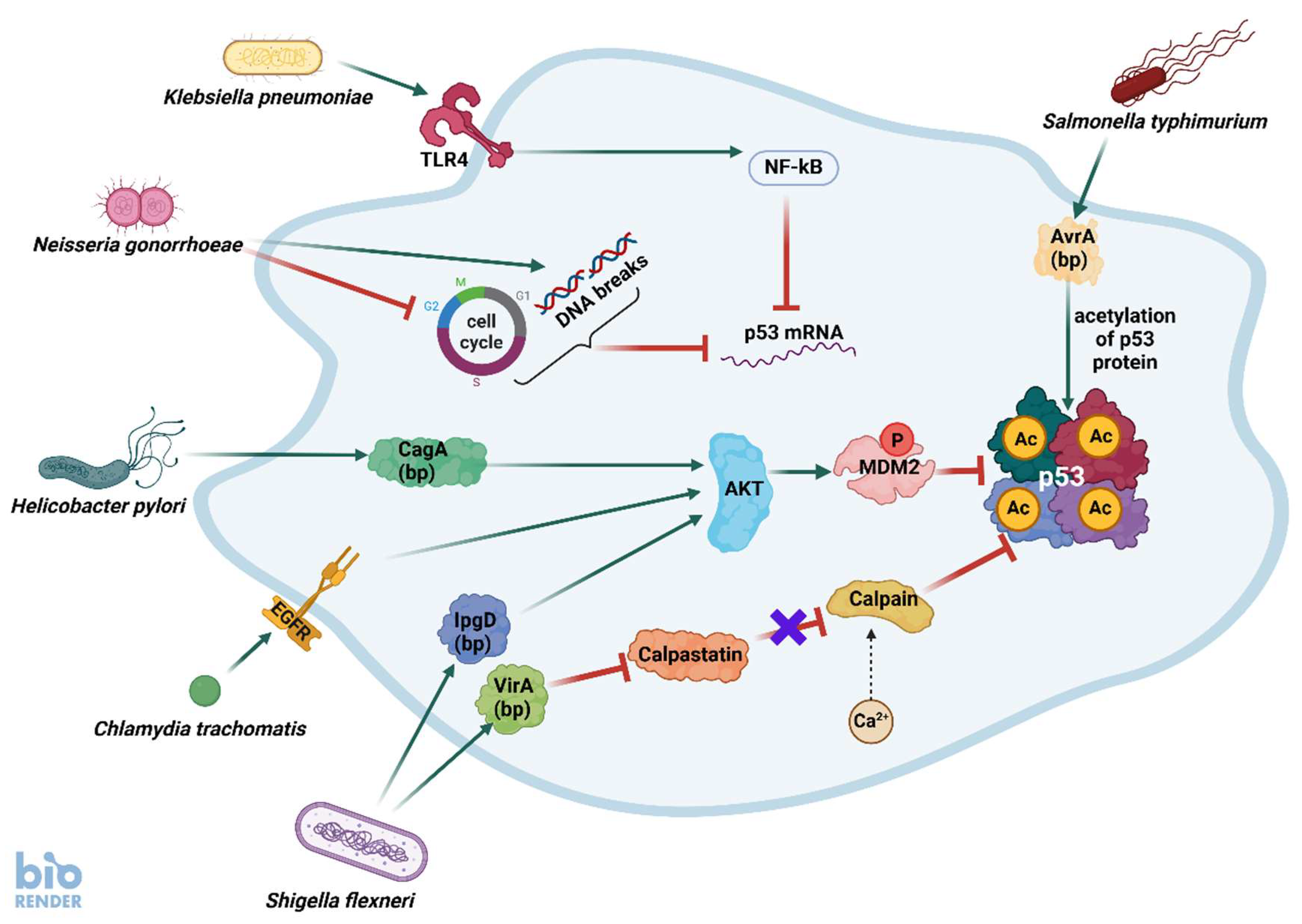
- Siegl, C.; Rudel, T. Modulation of p53 during bacterial infections. Nat. Rev. Microbiol. 2015, 13, 741–748. [Google Scholar] [CrossRef]
- Zaika, A.I.; Wei, J.; Noto, J.M.; Peek, R.M. Microbial Regulation of p53 Tumor Suppressor. PLoS Pathog. 2015, 11, e1005099. [Google Scholar] [CrossRef]
- Toller, I.M.; Neelsen, K.J.; Steger, M.; Hartung, M.L.; Hottiger, M.O.; Stucki, M.; Kalali, B.; Gerhard, M.; Sartori, A.A.; Lopes, M.; Müller, A. Carcinogenic bacterial pathogen Helicobacter pylori triggers DNA double-strand breaks and a DNA damage response in its host cells. Proc. Natl. Acad. Sci. U.S.A. 2011, 108, 14944–14949. [Google Scholar] [CrossRef] [PubMed]
- Vielfort, K.; Söderholm, N.; Weyler, L.; Vare, D.; Löfmark, S.; Aro, H. Neisseria gonorrhoeae infection causes DNA damage and affects the expression of p21, p27 and p53 in non-tumor epithelial cells. J. Cell Sci. 2013, 126, 339–347. [Google Scholar] [CrossRef]
- Aschtgen, M.S.; Fragkoulis, K.; Sanz, G.; Normark, S.; Selivanova, G.; Henriques-Normark, B.; Peuget, S. Enterobacteria impair host p53 tumor suppressor activity through mRNA destabilization. Oncogene 2022, 41, 2173–2186. [Google Scholar] [CrossRef]
- Fathima, F.; Subramaniyan, Y.; Rai, A.; Rekha, P.D. Enterococcus faecalis co-cultured with oral cancer cells exhibits higher virulence and promotes cancer cell survival, proliferation, and migration: an in vitro study. J. Med. Microbiol. 2024, 73, 11. [Google Scholar] [CrossRef]
- Wu, S.; Ye, Z.; Liu, X.; Zhao, Y.; Xia, Y.; Steiner, A.; Petrof, E.O.; Claud, E.C.; Sun, J. Salmonella typhimurium infection increases p53 acetylation in intestinal epithelial cells. Am. J. Physiol. Gastrointest. Liver Physiol. 2010, 298, G784–G794. [Google Scholar] [CrossRef] [PubMed]
- González, E.; Rother, M.; Kerr, M.; et al. Chlamydia infection depends on a functional MDM2-p53 axis. Nat. Commun. 2014, 5, 5201. [Google Scholar] [CrossRef]
- Bhardwaj, V.; Noto, J.M.; Wei, J.; Andl, C.; El-Rifai, W.; Peek, R.M.; Zaika, A.I. Helicobacter pylori Bacteria Alter the p53 Stress Response via ERK-HDM2 Pathway. Oncotarget 2015, 6, 1531–1543. [Google Scholar] [CrossRef]
- Bao, J.; He, Y.; Yang, C.; Lu, N.; Li, A.; Gao, S.; Hosyanto, F.F.; Tang, J.; Si, J.; Tang, X.; Fu, H.; Xu, L. Inhibition of Mycobacteria Proliferation in Macrophages by Low Cisplatin Concentration through Phosphorylated p53-Related Apoptosis Pathway. PLoS ONE 2023, 18, e0281170. [Google Scholar] [CrossRef]
- Madenspacher, J.H.; Azzam, K.M.; Gowdy, K.M.; Malcolm, K.C.; Nick, J.A.; Dixon, D.; Aloor, J.J.; Draper, D.W.; Guardiola, J.J.; Shatz, M.; Menendez, D.; Lowe, J.; Lu, J.; Bushel, P.; Li, L.; Merrick, B.A.; Resnick, M.A.; Fessler, M.B. p53 Integrates host defense and cell fate during bacterial pneumonia. J. Exp. Med. 2013, 210, 891–904. [Google Scholar] [CrossRef]
- Pal S, Bhattacharjee A, Ali A, Mandal NC, Mandal SC, Pal M. Chronic inflammation and cancer: potential chemoprevention through nuclear factor kappa B and p53 mutual antagonism. J. Inflamm. (Lond.) 2014, 9;11:23. [CrossRef]
- Jarczak, D.; Nierhaus, A. Cytokine Storm—Definition, Causes, and Implications. Int. J. Mol. Sci. 2022, 23, 11740. [Google Scholar] [CrossRef]
- Wang S, Liu P, Wei J, Zhu Z, Shi Z, Shao D, Ma Z. Tumor suppressor p53 protects mice against Listeria monocytogenes infection. Sci Rep. 2016, 6:33815. [CrossRef]
- Lim, Y.J.; Lee, J.; Choi, J.A.; Cho, S.N.; Son, S.H.; Kwon, S.J.; Son, J.W.; Song, C.H. M1 macrophage dependent-p53 regulates the intracellular survival of mycobacteria. Apoptosis 2020, 25, 42–55. [Google Scholar] [CrossRef]
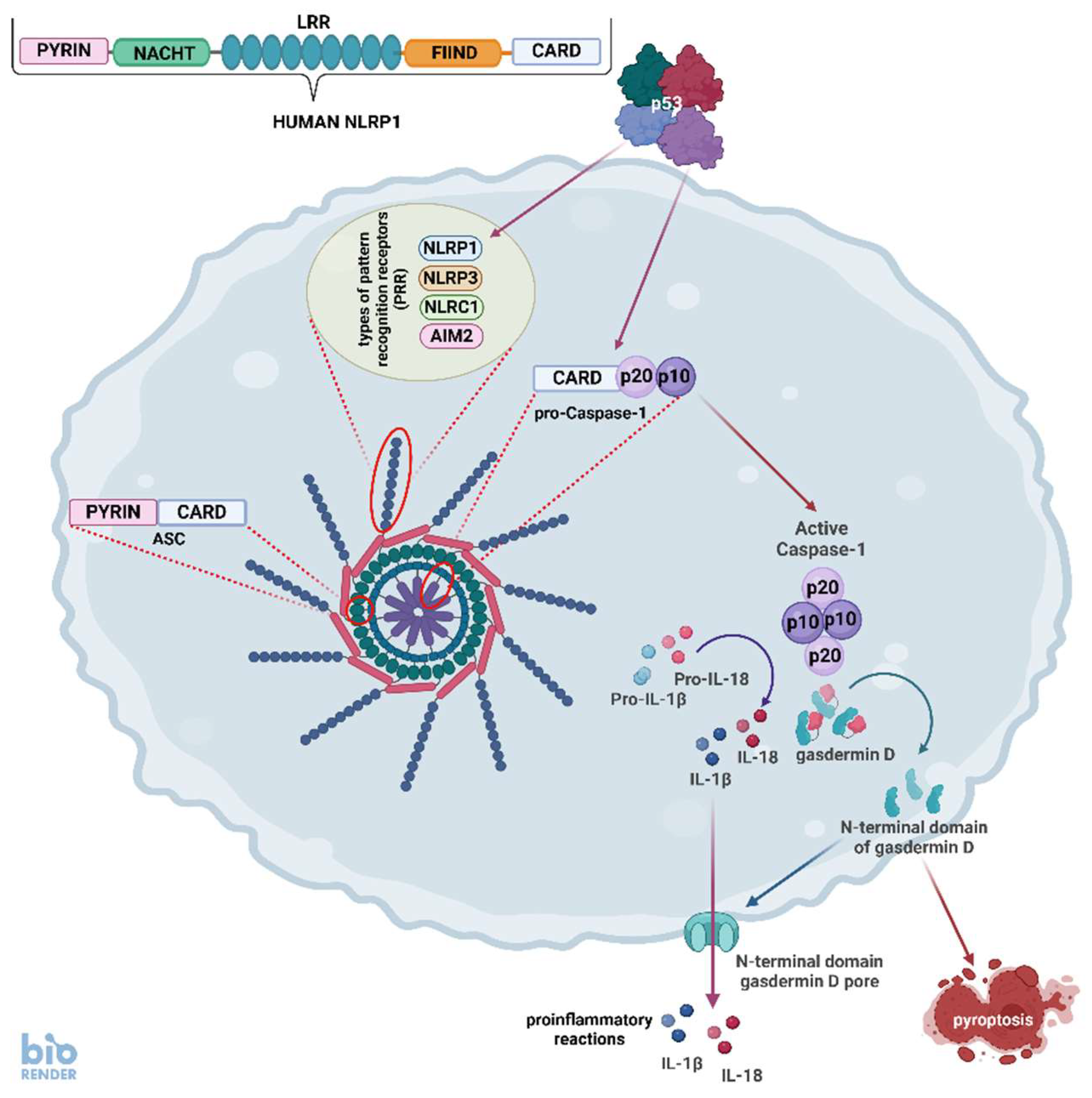
- Martinon, F.; Burns, K.; Tschopp, J. The inflammasome: a molecular platform triggering activation of inflammatory caspases and processing of proIL-beta. Mol. Cell 2002, 10, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Deng, Z.; Lu, L.; Li, B.; Shi, X.; Jin, H.; Hu, W. The Roles of Inflammasomes in Cancer. Front. Immunol. 2023, 14, 1195572. [Google Scholar] [CrossRef]
- Krześniak, M.; Zajkowicz, A.; Gdowicz-Kłosok, A.; Głowala-Kosińska, M.; Łasut-Szyszka, B.; Rusin, M. Synergistic activation of p53 by actinomycin D and nutlin-3a is associated with the upregulation of crucial regulators and effectors of innate immunity. Cell Signal. 2020, 69, 109552. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Radha, V.; Furukawa, Y.; Swarup, G. Direct transcriptional activation of human caspase-1 by tumor suppressor p53. J. Biol. Chem. 2001, 276, 10585–10588. [Google Scholar] [CrossRef] [PubMed]
- Schlereth K, Beinoraviciute-Kellner R, Zeitlinger MK, Bretz AC, Sauer M, Charles JP, Vogiatzi F, Leich E, Samans B, Eilers M, Kisker C, Rosenwald A, Stiewe T. DNA binding cooperativity of p53 modulates the decision between cell-cycle arrest and apoptosis. Mol. Cell. 2010, 38(3):356-68. [CrossRef]
- Deng, Z.; Matsuda, K.; Tanikawa, C.; Lin, J.; Furukawa, Y.; Hamamoto, R.; Nakamura, Y. Late Cornified Envelope Group I, a Novel Target of p53, Regulates PRMT5 Activity. Neoplasia 2014, 16, 656–664. [Google Scholar] [CrossRef]
- Taura M, Eguma A, Suico MA, Shuto T, Koga T, Komatsu K, Komune T, Sato T, Saya H, Li JD, Kai H. p53 regulates Toll-like receptor 3 expression and function in human epithelial cell lines. Mol Cell Biol. 2008, 28(21):6557-67. [CrossRef]
- Xu Z, Kombe Kombe AJ, Deng S, Zhang H, Wu S, Ruan J, Zhou Y, Jin T. NLRP inflammasomes in health and disease. Mol Biomed. 2024, 5(1):14. [CrossRef]
- Wang T, Yazdi AS, Panayotova-Dimitrova D. Comparison of Different Keratinocyte Cell Line Models for Analysis of NLRP1 Inflammasome Activation. Biomolecules. 2024, 14(11):1427. [CrossRef]
- Pinilla M, Mazars R, Vergé R, Gorse L, Paradis M, Suire B, Santoni K, Robinson KS, Toh GA, Prouvensier L, Leon-Icaza SA, Hessel A, Péricat D, Murris M, Guet-Revillet H, Henras A, Buyck J, Ravet E, Zhong FL, Cougoule C, Planès R, Meunier E. EEF2-inactivating toxins engage the NLRP1 inflammasome and promote epithelial barrier disruption. J. Exp. Med. 2023, 220(10):e20230104. [CrossRef]
- Robinson KS, Toh GA, Firdaus MJ, Tham KC, Rozario P, Lim CK, Toh YX, Lau ZH, Binder SC, Mayer J, Bonnard C, Schmidt FI, Common JEA, Zhong FL. Diphtheria toxin activates ribotoxic stress and NLRP1 inflammasome-driven pyroptosis. J. Exp. Med. 2023, 220(10):e20230105. [CrossRef]
- Song LL, Alimirah F, Panchanathan R, Xin H, Choubey D. Expression of an IFN-inducible cellular senescence gene, IFI16, is up-regulated by p53. Mol Cancer Res. 2008, 6(11):1732-41. [CrossRef]
- Zhang F, Yuan Y, Ma F. Function and Regulation of Nuclear DNA Sensors During Viral Infection and Tumorigenesis. Front Immunol. 2021, 11:624556. [CrossRef]
- Cruz Díaz, L.A.; Flores Miramontes, M.G.; Chávez Hurtado, P.; Allen, K.; Gonzalez Ávila, M.; Prado Montes de Oca, E. Ascorbic Acid, Ultraviolet C Rays, and Glucose but Not Hyperthermia Are Elicitors of Human β-Defensin 1 mRNA in Normal Keratinocytes. Biomed Res. Int. 2015, 2015, 714580. [Google Scholar] [CrossRef]
- Pitkänen K, Haapajärvi T, Laiho M. U.V.C.-induction of p53 activation and accumulation is dependent on cell cycle and pathways involving protein synthesis and phosphorylation. Oncogene. 1998, 16(4):459-69. [CrossRef]
- Ryan LK, Diamond G. Modulation of Human β-Defensin-1 Production by Viruses. Viruses. 2017, 9(6):153. [CrossRef]
- Prado-Montes de Oca E. Human beta-defensin 1: a restless warrior against allergies, infections and cancer. Int. J. Biochem. Cell Biol. 2010, 42(6):800-4. [CrossRef]
- Duits, L.A.; Ravensbergen, B.; Rademaker, M.; Hiemstra, P.S.; Nibbering, P.H. Expression of Beta-Defensin 1 and 2 mRNA by Human Monocytes, Macrophages and Dendritic Cells. Immunology 2002, 106, 517–525. [Google Scholar] [CrossRef]
- Pazgier M, Hoover DM, Yang D, Lu W, Lubkowski J. Human beta-defensins. Cell Mol. Life Sci. 2006, 63(11):1294-313. [CrossRef]
- Álvarez, Á.H.; Martínez Velázquez, M.; Prado Montes de Oca, E. Human β-defensin 1 update: Potential clinical applications of the restless warrior. Int. J. Biochem. Cell Biol. 2018, 104, 133–137. [Google Scholar] [CrossRef] [PubMed]
- Moser, C.; Weiner, D.J.; Lysenko, E.; Bals, R.; Weiser, J.N.; Wilson, J.M. Beta-defensin 1 contributes to pulmonary innate immunity in mice. Infect. Immun. 2002, 70(6), 3068–3072. [Google Scholar] [CrossRef]
- Diao, R.; Fok, K.L.; Chen, H.; Yu, M.K.; Duan, Y.; Chung, C.M.; Li, Z.; Wu, H.; Li, Z.; Zhang, H.; et al. Deficient Human β-Defensin 1 Underlies Male Infertility Associated with Poor Sperm Motility and Genital Tract Infection. Sci. Transl. Med. 2014, 6, 249ra108. [Google Scholar] [CrossRef] [PubMed]
- Kypriotou, M.; Huber, M.; Hohl, D. The human epidermal differentiation complex: cornified envelope precursors, S100 proteins and the ‘fused genes’ family. Exp. Dermatol. 2012, 21, 643–649. [Google Scholar] [CrossRef] [PubMed]
- Niehues, H.; van der Krieken, D.A.; Ederveen, T.H.A.; Jansen, P.A.M.; van Niftrik, L.; Mesman, R.; Netea, M.G.; Smits, J.P.H.; Schalkwijk, J.; van den Bogaard, E.H.; Zeeuwen, P.L.J.M. Antimicrobial late cornified envelope proteins: The psoriasis risk factor deletion of LCE3B/C genes affects microbiota composition. J. Invest. Dermatol. 2022, 142(7), 1947–1955.e6. [Google Scholar] [CrossRef]
- Niehues H, Rikken G, Kersten FFJ, Eeftens JM, van Vlijmen-Willems IMJJ, Rodijk-Olthuis D, Jansen PAM, Hendriks WJAJ, Ederveen THA, Schalkwijk J, van den Bogaard EH, Zeeuwen PLJM. CYSRT1: An antimicrobial epidermal protein that can interact with late cornified envelope proteins. J. Invest. Dermatol. 2023, 143(8):1498-1508.e7. [CrossRef]
- Archer, N.K.; Dilolli, M.N.; Miller, L.S. Pushing the Envelope in Psoriasis: Late Cornified Envelope Proteins Possess Antimicrobial Activity. J. Investig. Dermatol. 2017, 137, 2257-2259. [CrossRef]
- Small DM, Doherty DF, Dougan CM, Weldon S, Taggart CC. The role of whey acidic protein four-disulfide-core proteins in respiratory health and disease. Biol Chem. 2017, 398(4):425-440. [CrossRef]
- Dochez, V.; Caillon, H.; Vaucel, E.; Dimet, J.; Winer, N.; Ducarme, G. Biomarkers and Algorithms for Diagnosis of Ovarian Cancer: CA125, HE4, RMI and ROMA, a Review. J. Ovarian Res. 2019, 12, 28. [Google Scholar] [CrossRef]
- Hua, L.; Liu, Y.; Zhen, S.; Wan, D.; Cao, J.; Gao, X. Expression and biochemical characterization of recombinant human epididymis protein 4. Protein Expr. Purif. 2014, 102, 52–62. [Google Scholar] [CrossRef]
- Watt AP, Sharp JA, Lefevre C, Nicholas KR. WFDC2 is differentially expressed in the mammary gland of the tammar wallaby and provides immune protection to the mammary gland and the developing pouch young. Dev Comp Immunol. 2012, 36(3):584-90. [CrossRef]
- Parikh K, Antanaviciute A, Fawkner-Corbett D, Jagielowicz M, Aulicino A, Lagerholm C, Davis S, Kinchen J, Chen HH, Alham NK, Ashley N, Johnson E, Hublitz P, Bao L, Lukomska J, Andev RS, Björklund E, Kessler BM, Fischer R, Goldin R, Koohy H, Simmons A. Colonic epithelial cell diversity in health and inflammatory bowel disease. Nature. 2019, 567(7746):49-55. [CrossRef]
- Dougherty, G.W.; Ostrowski, L.E.; Nöthe-Menchen, T.; Raidt, J.; Schramm, A.; Olbrich, H.; Yin, W.; Sears, P.R.; Dang, H.; Smith, A.J.; et al. Recessively Inherited Deficiency of Secreted WFDC2 (HE4) Causes Nasal Polyposis and Bronchiectasis. Am. J. Respir. Crit. Care Med. 2024, 210, 63–76. [Google Scholar] [CrossRef]
- Wang Y, Li Z, Lu J. Single-cell RNA sequencing reveals the epithelial cell, fibroblast, and key gene alterations in chronic rhinosinusitis with nasal polyps. Sci Rep. 2024 14: 2270. [CrossRef] [PubMed]
- Andrade, A.D.; Almeida, P.G.C.; Mariani, N.A.P.; Freitas, G.A.; Kushima, H.; Filadelpho, A.L.; Spadella, M.A.; Avellar, M.C.W.; Silva, E.J.R. Lipopolysaccharide-Induced Epididymitis Modifies the Transcriptional Profile of Wfdc Genes in Mice. Biol. Reprod. 2021, 104, 144–158. [Google Scholar] [CrossRef]
- Miñoza, J.M.A.; Rico, J.A.; Zamora, P.R.F.; Bacolod, M.; Laubenbacher, R.; Dumancas, G.G.; de Castro, R. Biomarker discovery for meta-classification of melanoma metastatic progression using transfer learning. Genes (Basel) 2022, 13(12), 2303. [Google Scholar] [CrossRef]
- Chen, Y.H.; Wu, K.H.; Wu, H.P. Unraveling the Complexities of Toll-like Receptors: From Molecular Mechanisms to Clinical Applications. Int. J. Mol. Sci. 2024, 25, 5037. [Google Scholar] [CrossRef] [PubMed]
- Groskreutz, D.J.; Monick, M.M.; Powers, L.S.; Yarovinsky, T.O.; Look, D.C.; Hunninghake, G.W. Respiratory syncytial virus induces TLR3 protein and protein kinase R, leading to increased double-stranded RNA responsiveness in airway epithelial cells. J. Immunol. 2006, 176, 1733–1740. [Google Scholar] [CrossRef]
- Heinrich, A.; Haarmann, H.; Zahradnik, S.; Frenzel, K.; Schreiber, F.; Klassert, T.E.; Heyl, K.A.; Endres, A.S.; Schmidtke, M.; Hofmann, J.; Slevogt, H. Moraxella catarrhalis decreases antiviral innate immune responses by down-regulation of TLR3 via inhibition of p53 in human bronchial epithelial cells. FASEB J. 2016, 30, 2426–2434. [Google Scholar] [CrossRef]
- Campos, P.C.; Gomes, M.T.; Guimarães, E.S.; Guimarães, G.; Oliveira, S.C. TLR7 and TLR3 Sense Brucella abortus RNA to Induce Proinflammatory Cytokine Production but They Are Dispensable for Host Control of Infection. Front. Immunol. 2017, 8, 28. [Google Scholar] [CrossRef] [PubMed]
- Derbigny, W.A.; Shobe, L.R.; Kamran, J.C.; Toomey, K.S.; Ofner, S. Identifying a Role for Toll-Like Receptor 3 in the Innate Immune Response to Chlamydia muridarum Infection in Murine Oviduct Epithelial Cells. Infect. Immun. 2012, 80, 254–265. [Google Scholar] [CrossRef]
- Grigoryeva, L.S.; Cianciotto, N.P. Human macrophages utilize a wide range of pathogen recognition receptors to recognize Legionella pneumophila, including Toll-Like Receptor 4 engaging Legionella lipopolysaccharide and the Toll-like Receptor 3 nucleic-acid sensor. PLoS Pathog. 2021, 17, e1009781. [Google Scholar] [CrossRef]
- Echchannaoui, H.; Frei, K.; Schnell, C.; Leib, S.L.; Zimmerli, W.; Landmann, R. Toll-like receptor 2-deficient mice are highly susceptible to Streptococcus pneumoniae meningitis because of reduced bacterial clearing and enhanced inflammation. J. Infect. Dis. 2002, 186, 798–806. [Google Scholar] [CrossRef]
- Takeuchi O, Hoshino K, Akira S. Cutting edge: TLR2-deficient and MyD88-deficient mice are highly susceptible to Staphylococcus aureus infection. J Immunol. 2000, 165(10):5392-6. [CrossRef]
- Sugawara I, Yamada H, Li C, Mizuno S, Takeuchi O, Akira S. Mycobacterial infection in TLR2 and TLR6 knockout mice. Microbiol Immunol. 2003;47(5):327-36. [CrossRef]
- Wooten RM, Ma Y, Yoder RA, Brown JP, Weis JH, Zachary JF, Kirschning CJ, Weis JJ. Toll-like receptor 2 is required for innate, but not acquired, host defense to Borrelia burgdorferi. J Immunol. 2002, 168(1):348-55. [CrossRef]
- Mayer, A.K.; Muehmer, M.; Mages, J.; Gueinzius, K.; Hess, C.; Heeg, K.; Bals, R.; Lang, R.; Dalpke, A.H. Differential recognition of TLR-dependent microbial ligands in human bronchial epithelial cells. J. Immunol. 2007, 178, 3134–3142. [Google Scholar] [CrossRef]
- Melmed, G.; Thomas, L.S.; Lee, N.; Tesfay, S.Y.; Lukasek, K.; Michelsen, K.S.; Zhou, Y.; Hu, B.; Arditi, M.; Abreu, M.T. Human intestinal epithelial cells are broadly unresponsive to Toll-like receptor 2-dependent bacterial ligands: Implications for host-microbial interactions in the gut. J. Immunol. 2003, 170(3), 1406–1415. [Google Scholar] [CrossRef] [PubMed]
- Bune, A.J.; Hayman, A.R.; Evans, M.J.; Cox, T.M. Mice Lacking Tartrate-Resistant Acid Phosphatase (Acp5) Have Disordered Macrophage Inflammatory Responses and Reduced Clearance of the Pathogen, Staphylococcus aureus. Immunology 2001, 102, 103–113. [Google Scholar] [CrossRef]
- Tanner L, Bergwik J, Bhongir RKV, Puthia M, Lång P, Ali MN, Welinder C, Önnerfjord P, Erjefält JS, Palmberg L, Andersson G, Egesten A. Tartrate resistant acid phosphatase 5 (TRAP5) mediates immune cell recruitment in a murine model of pulmonary bacterial infection. Front Immunol. 2022, 13:1079775. [CrossRef]
- Mavlankar, A.; Sharma, M.; Ansari, A.; Singh, P. Comparative host transcriptomics as a tool to identify candidate biomarkers for immune reactions in leprosy using meta-analysis. Indian J. Dermatol. Venereol. Leprol. 2024, 90, 731–741. [Google Scholar] [CrossRef] [PubMed]
- Galluzzi, L.; Baehrecke, E.H.; Ballabio, A.; Boya, P.; Bravo-San Pedro, J.M.; Cecconi, F.; Choi, A.M.; Chu, C.T.; Codogno, P.; Colombo, M.I.; Cuervo, A.M.; Debnath, J.; Deretic, V.; Dikic, I.; Eskelinen, E.L.; Fimia, G.M.; Fulda, S.; Gewirtz, D.A.; Green, D.R.; Hansen, M.; Harper, J.W.; Jäättelä, M.; Johansen, T.; Juhasz, G.; Kimmelman, A.C.; Kraft, C.; Ktistakis, N.T.; Kumar, S.; Levine, B.; Lopez-Otin, C.; Madeo, F.; Martens, S.; Martinez, J.; Melendez, A.; Mizushima, N.; Münz, C.; Murphy, L.O.; Penninger, J.M.; Piacentini, M.; Reggiori, F.; Rubinsztein, D.C.; Ryan, K.M.; Santambrogio, L.; Scorrano, L.; Simon, A.K.; Simon, H.U.; Simonsen, A.; Tavernarakis, N.; Tooze, S.A.; Yoshimori, T.; Yuan, J.; Yue, Z.; Zhong, Q.; Kroemer, G. Molecular definitions of autophagy and related processes. EMBO J. 2017, 36, 1811–1836. [Google Scholar] [CrossRef] [PubMed]
- Chew, T.S.; O’Shea, N.R.; Sewell, G.W.; Oehlers, S.H.; Mulvey, C.M.; Crosier, P.S.; Godovac-Zimmermann, J.; Bloom, S.L.; Smith, A.M.; Segal, A.W. Optineurin Deficiency in Mice Contributes to Impaired Cytokine Secretion and Neutrophil Recruitment in Bacteria-Driven Colitis. Dis. Model. Mech. 2015, 8, 817–829. [Google Scholar] [CrossRef]
- Slowicka K, Vereecke L, Mc Guire C, Sze M, Maelfait J, Kolpe A, Saelens X, Beyaert R, van Loo G. Optineurin deficiency in mice is associated with increased sensitivity to Salmonella but does not affect proinflammatory NF-κB signaling. Eur J Immunol. 2016, 46(4):971-80. [CrossRef]
- Wild P, Farhan H, McEwan DG, Wagner S, Rogov VV, Brady NR, Richter B, Korac J, Waidmann O, Choudhary C, Dötsch V, Bumann D, Dikic I. Phosphorylation of the autophagy receptor optineurin restricts Salmonella growth. Science. 2011, 333(6039):228-33. [CrossRef]
- Qiu Y, Wang J, Li H, Yang B, Wang J, He Q, Weng Q. Emerging views of OPTN (optineurin) function in the autophagic process associated with disease. Autophagy. 2022, 18(1):73-85. [CrossRef]
- Demerlé, C.; Gorvel, L.; Olive, D. BTLA-HVEM Couple in Health and Diseases: Insights for Immunotherapy in Lung Cancer. Front. Oncol. 2021, 11, 682007. [Google Scholar] [CrossRef]
- Šedý JR, Ramezani-Rad P. HVEM network signaling in cancer. Adv. Cancer Res. 2019;142:145-186. [CrossRef]
- Shui JW, Larange A, Kim G, Vela JL, Zahner S, Cheroutre H, Kronenberg M. HVEM signalling at mucosal barriers provides host defence against pathogenic bacteria. Nature. 2012, 488(7410):222-5. [CrossRef]
- Ananthakrishnan, A.N.; Oxford, E.C.; Nguyen, D.D.; Sauk, J.; Yajnik, V.; Xavier, R.J. Genetic Risk Factors for Clostridium difficile Infection in Ulcerative Colitis. Aliment. Pharmacol. Ther. 2013, 38, 522–530. [Google Scholar] [CrossRef]
- Cheng, T.Y.; Liu, Y.J.; Yan, H.; Xi, Y.B.; Duan, L.Q.; Wang, Y.; Zhang, T.T.; Gu, Y.M.; Wang, X.D.; Wu, C.X.; Gao, S. Tumor Cell-Intrinsic BTLA Receptor Inhibits the Proliferation of Tumor Cells via ERK1/2. Cells 2022, 11, 4021. [Google Scholar] [CrossRef]
- Boice, M.; Salloum, D.; Mourcin, F.; Sanghvi, V.; Amin, R.; Oricchio, E.; Jiang, M.; Mottok, A.; Denis-Lagache, N.; Ciriello, G.; Tam, W.; Teruya-Feldstein, J.; de Stanchina, E.; Chan, W.C.; Malek, S.N.; Ennishi, D.; Brentjens, R.J.; Gascoyne, R.D.; Cogné, M.; Tarte, K.; Wendel, H.G. Loss of the HVEM Tumor Suppressor in Lymphoma and Restoration by Modified CAR-T Cells. Cell 2016, 167, 405–418.e13. [Google Scholar] [CrossRef]
- Hu, X. The role of the BTLA-HVEM complex in the pathogenesis of breast cancer. Breast Cancer 2024, 31, 358–370. [Google Scholar] [CrossRef]
- McWilliam, H.E.G.; Villadangos, J.A. MR1 antigen presentation to MAIT cells and other MR1-restricted T cells. Nat. Rev. Immunol. 2024, 24, 178–192. [Google Scholar] [CrossRef]
- López-Rodríguez, J.C.; Barral, P. Mucosal associated invariant T cells: Powerhouses of the lung. Immunol. Lett. 2024, 269, 106910. [Google Scholar] [CrossRef]
- Dolton, G.; Thomas, H.; Tan, L.R.; Rius, R.C.; Doetsch, S.; Ionescu, G.A.; Cardo, L.F.; Crowther, M.D.; Behiry, E.; Morin, T.; et al. MHC-Related Protein 1-Restricted Recognition of Cancer via a Semi-Invariant TCR-α Chain. J. Clin. Invest. 2025, 135, e181895. [Google Scholar] [CrossRef] [PubMed]
- Howson, L.J.; Awad, W.; von Borstel, A.; Lim, H.J.; McWilliam, H.E.G.; Sandoval-Romero, M.L.; Majumdar, S.; Hamzeh, A.R.; Andrews, T.D.; McDermott, D.H.; Murphy, P.M.; Le Nours, J.; Mak, J.Y.W.; Liu, L.; Fairlie, D.P.; McCluskey, J.; Villadangos, J.A.; Cook, M.C.; Turner, S.J.; Davey, M.S.; Ojaimi, S.; Rossjohn, J. Absence of mucosal-associated invariant T cells in a person with a homozygous point mutation in MR1. Sci. Immunol. 2020, 5, eabc9492. [Google Scholar] [CrossRef] [PubMed]
- Sugimoto C, Fujita H, Wakao H. Harnessing the Power of Mucosal-Associated Invariant T (MAIT) Cells in Cancer Cell Therapy. Biomedicines. 2022, 10(12):3160. [CrossRef]
- Lepore, M.; Kalinichenko, A.; Calogero, S.; Kumar, P.; Paleja, B.; Schmaler, M.; Narang, V.; Zolezzi, F.; Poidinger, M.; Mori, L.; De Libero, G. Functionally diverse human T cells recognize non-microbial antigens presented by MR1. Elife 2017, 6, e24476. [Google Scholar] [CrossRef]
- Crowther, M.D.; Dolton, G.; Legut, M.; Caillaud, M.E.; Lloyd, A.; Attaf, M.; Galloway, S.A.E.; Rius, C.; Farrell, C.P.; Szomolay, B.; et al. Genome-Wide CRISPR-Cas9 Screening Reveals Ubiquitous T Cell Cancer Targeting via the Monomorphic MHC Class I-Related Protein MR1. Nat. Immunol. 2020, 21, 178–185. [Google Scholar] [CrossRef]
- Yan J, Allen S, McDonald E, Das I, Mak JYW, Liu L, Fairlie DP, Meehan BS, Chen Z, Corbett AJ, Varelias A, Smyth MJ, Teng MWL. MAIT Cells Promote Tumor Initiation, Growth, and Metastases via Tumor MR1. Cancer Discov. 2020, 10(1):124-141. [CrossRef]
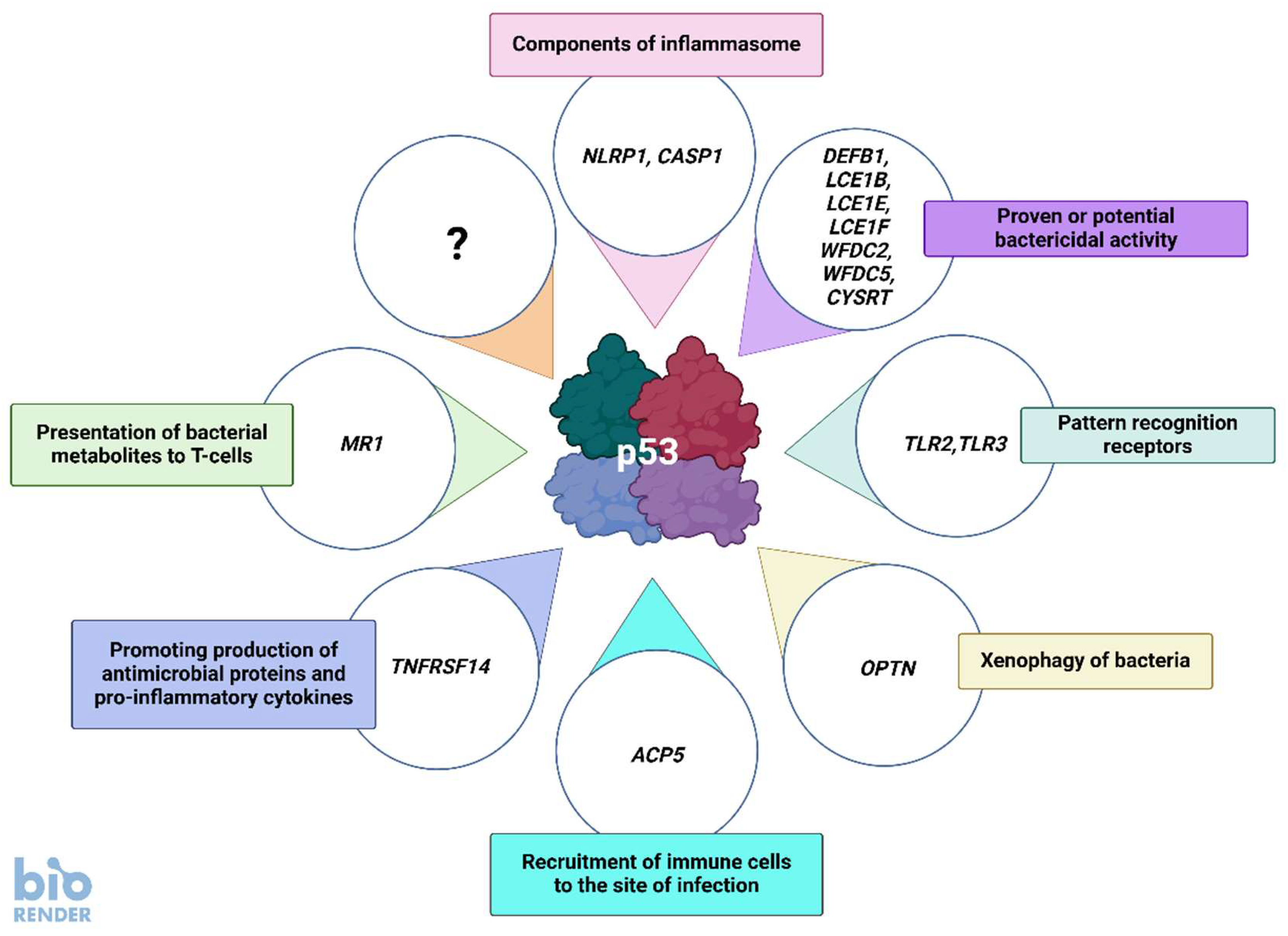
| Gene Name | p53 Expression Score (max. 57)* | Activated by Actinomycin D and Nutlin-3a in Cell Lines# | p53 Bound Promoter or Enhancer* | Regulation by p53 Detected in Individual Study | Antibacterial Function |
|---|---|---|---|---|---|
| NLRP1 | 33 | A549, NCI-H460, U-2 OS, | yes | Krześniak et el. [47] |
Pattern recognition receptor (PRR) of inflammasome able to detect bacteria |
| CASP1 | 15 | A549, NCI-H460, U-2 OS, A375 | yes | Gupta et al. [48], Schlereth et al. [49] | Common element of classic inflammasomes, activates cytokines and induces pyroptosis, activated by PRR recognizing either bacteria or viruses |
| DEFB1 | 3 | A549, NCI-H460, | no | no | Extracellular bactericidal activity, antimicrobial defense of epithelia |
| LCE1B | 24 | A549, NCI-H460, U-2 OS, A375 | yes | Deng et al. [50] | Constitutively expressed in epidermis, antimicrobial activity inferred from the function of related proteins from LCE3 group |
| LCE1E | 18 | A549, NCI-H460, U-2 OS, A375 | yes | Deng et al. [50] | Constitutively expressed in epidermis, antimicrobial activity inferred from the function of related proteins from LCE3 group |
| LCE1F | 17 | A549, NCI-H460, U-2 OS, A375 | no | Deng et al. [50] | Constitutively expressed in epidermis, antimicrobial activity inferred from the function of related proteins from LCE3 group |
| CYSRT1 | 34 | A549, NCI-H460, U-2 OS, A375 | yes | no | Constitutively expressed in stratum corneum of epidermis, where may contribute to antimicrobial host defenses |
| WFDC2 | 11 | A549, U-2 OS | no | no | Extracellular protease inhibitor with antimicrobial activity of poorly studied mechanism |
| WFDC5 | 2 | A549, NCI-H460 | no | no | Plausible antimicrobial activity similar to WFDC2 |
| TLR2 | 2 | A549, NCI-H460 | no | no | Pattern recognition receptor, expressed on cell surface, heterodimerizes with TLR1 and TLR6, recognizes bacterial molecules, activates NF-κB transcription factors and promotes the expression of pro-inflammatory cytokines |
| TLR3 | 26 | A549, NCI-H460, U-2 OS, A375 | yes | Taura et al. [51] | Pattern recognition receptor, detects double- stranded RNA derived from viruses, and probably from bacteria, and signals through IRF3 and IRF7 transcription factors activating expression of type I interferons |
| ACP5 | 10 | A549, NCI-H460, U-2 OS, A375 |
no | no | May help to recruit immune cells (macrophages, neutrophils) to the site of infection |
| OPTN | 31 | A549, NCI-H460, A375 | yes | no | May participate in autophagy (xenophagy) of bacteria; indirectly helps to recruit neutrophils to the site of infection |
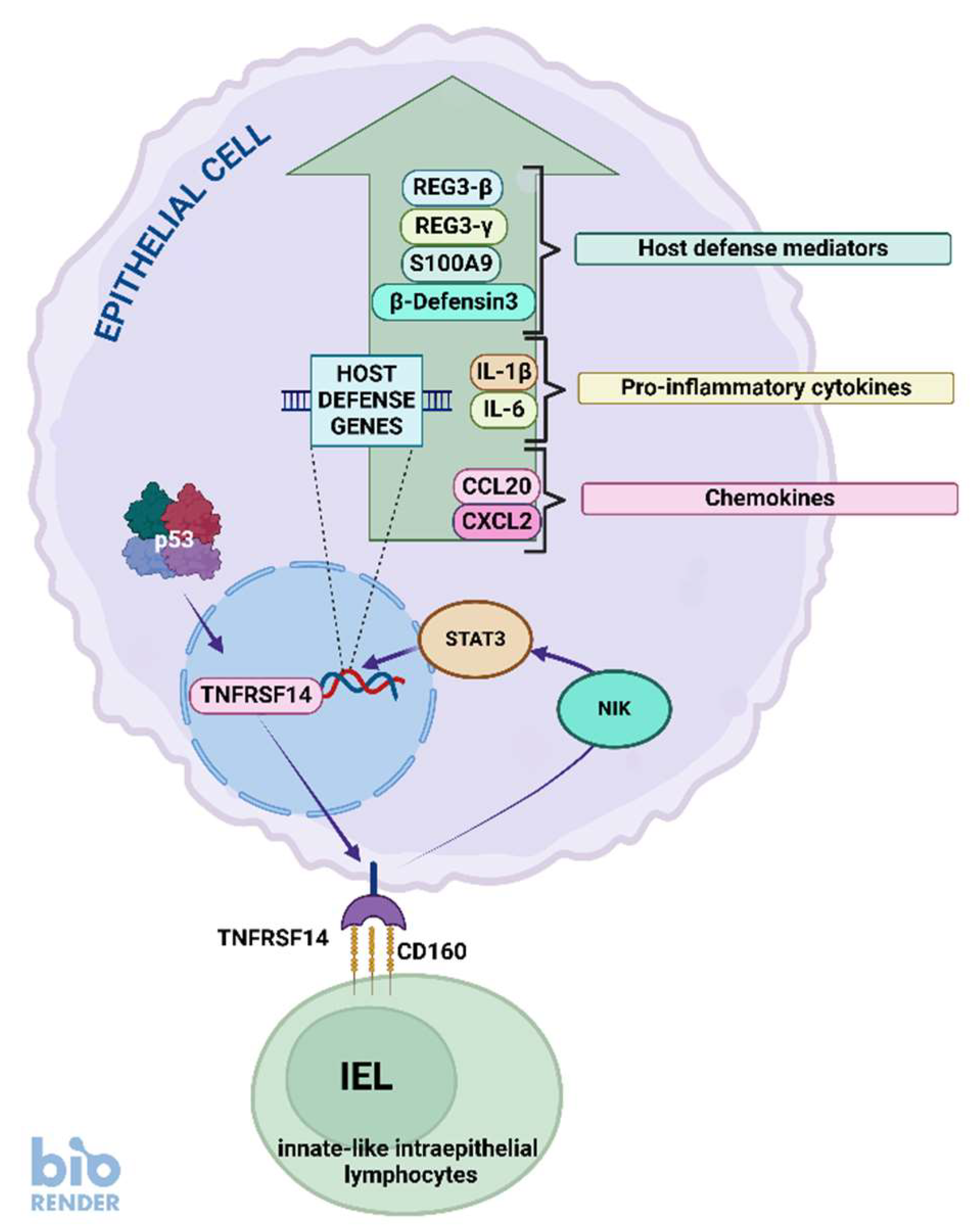
| TNFRSF14 | 34 | A549, NCI-H460, U-2 OS, A375 | no | no | Member tumor necrosis factor receptor superfamily; may also function as a ligand. Complicated role in regulation of immunity and cell growth; signaling receptor on epithelial cells for CD160 ligand expressed on intraepithelial lymphocytes, triggering the production of antimicrobial proteins and pro-inflammatory cytokines |
| MR1 | 40 | A549, NCI-H460, U-2 OS, A375 | yes | no | Major Histocompatibility Complex Class I-Related Gene Protein; the antigen-presenting molecule specialized in displaying microbial metabolites to T-cell receptors present on specialized lymphocytes called innate-type mucosal-associated invariant T (MAIT) cells |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




