Submitted:
28 March 2025
Posted:
01 April 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
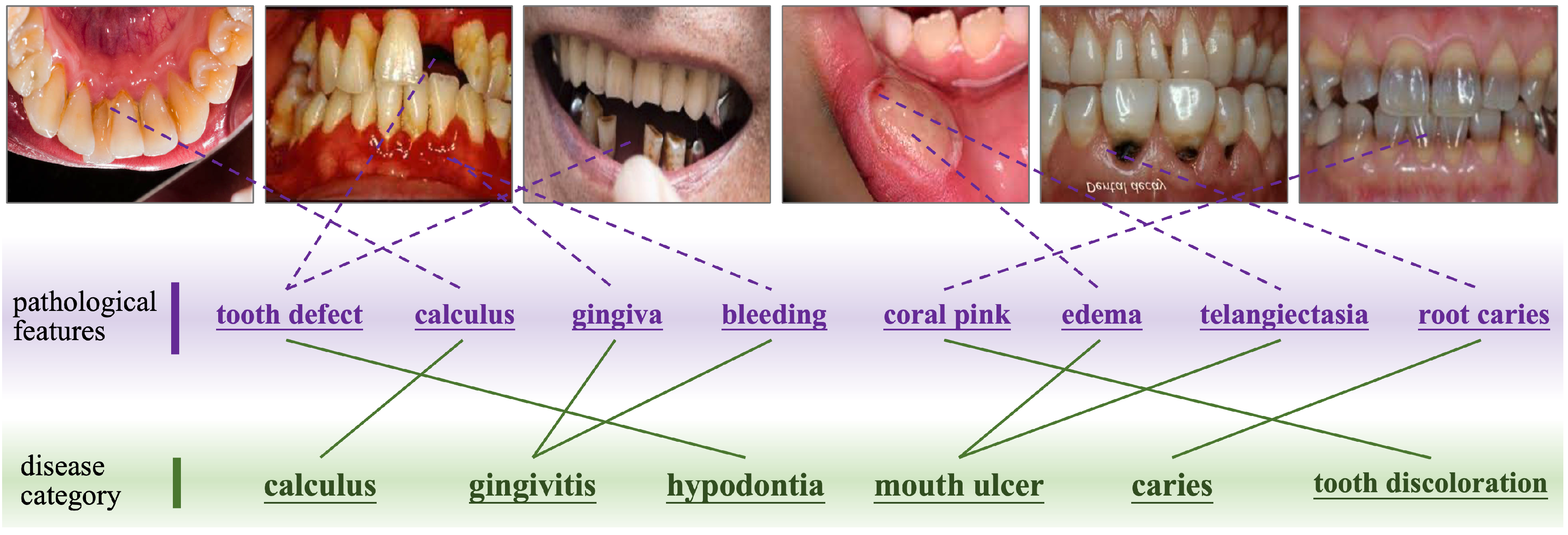
- Compositional Disease Representation. DyCoP decomposes complex diseases into reusable componential prototypes representing fundamental visual patterns shared across oral pathologies (e.g., "calculus deposits" or "enamel demineralization"). This decomposition enables efficient knowledge transfer, reducing reliance on large datasets.
- Dynamic Prototype Assembly. A sparse activation mechanism adaptively assembles disease-specific signatures from relevant prototypes based on input image characteristics. This flexibility accommodates intra-class variations across disease stages and presentations.
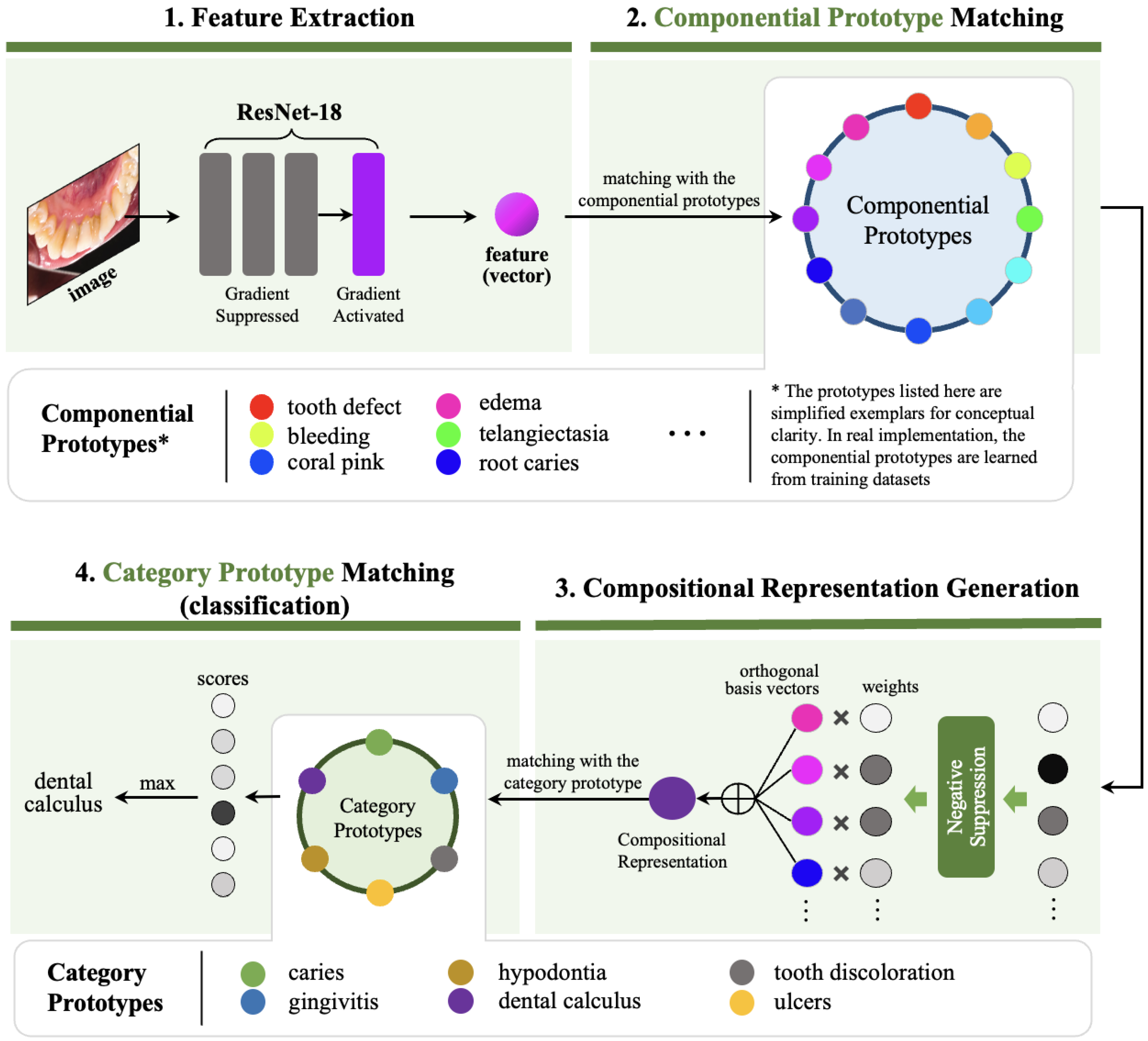
- Strategic Preservation of Pre-trained Knowledge. By performing gradient suppression to lower-level convolutional layers during training, DyCoP retains generalized visual feature extraction capabilities from pre-trained models while fine-tuning higher layers for oral disease specifics. This balances domain adaptation with data efficiency.
2. Related Work
2.1. Oral Disease Recognition
2.2. Prototype Learning for Image Analysis
3. Methodology
3.1. Feature Extraction
3.2. Componential Prototype Matching
3.3. Dynamic Compositional Representation Generation
3.4. Category Prototype Matching
4. Experiments
4.1. Experimental Setup
4.1.1. Dataset
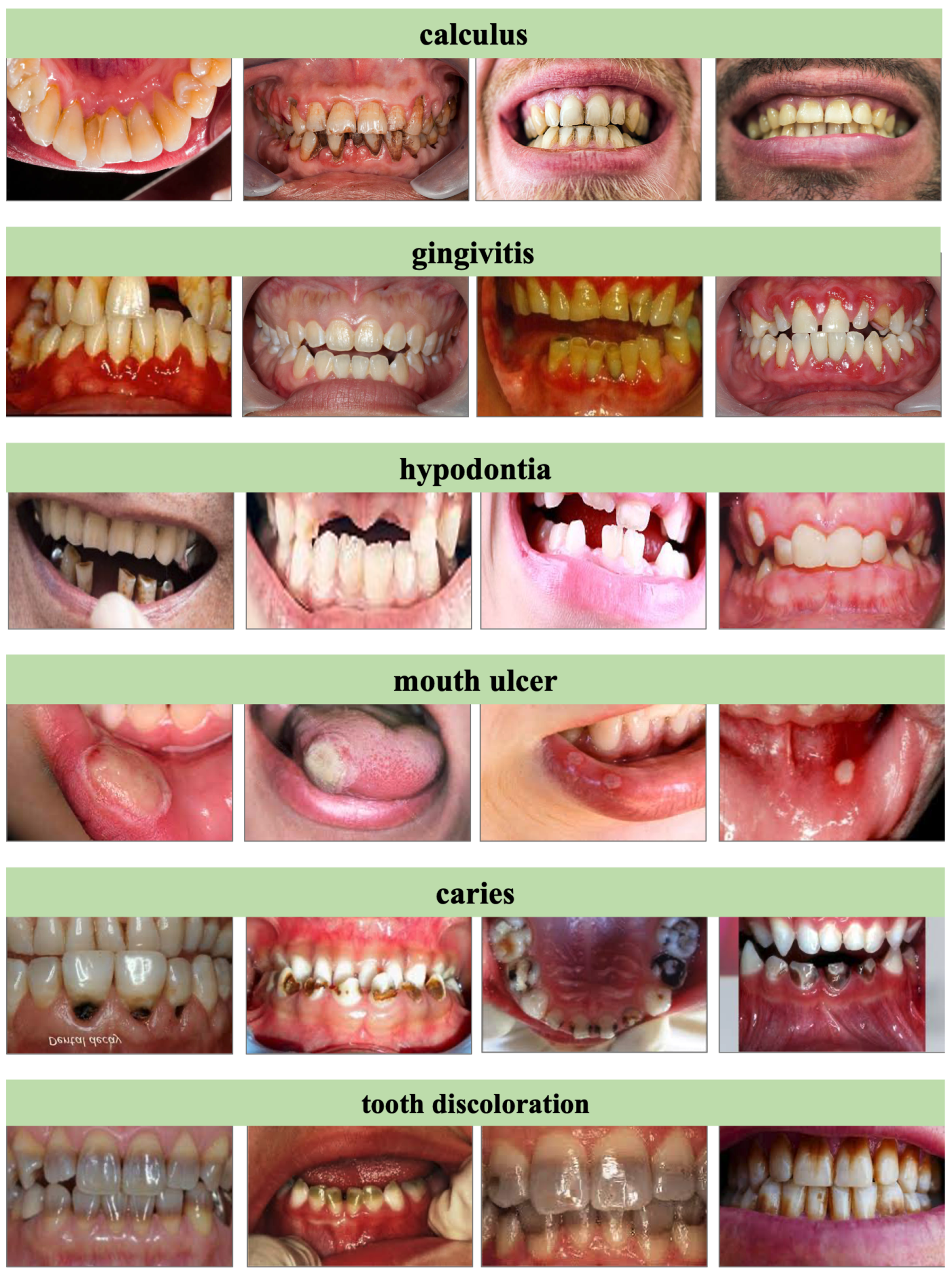
- Caries: Images depicting tooth decay, cavities, or carious lesions.
- Gingivitis: Cases of inflamed or infected gum tissue.
- Hypodontia: Evidence of congenital or acquired absence of one or more teeth.
- Mouth Ulcers: Visible canker sores or ulcerative lesions in oral mucosa.
- Calculus: Examples of dental calculus or tartar buildup on tooth surfaces.
- Tooth Discoloration: Manifestations of intrinsic or extrinsic tooth staining.
4.1.2. Implemental Details
4.1.3. Compared Methods
4.1.4. Evaluation Metrics
4.2. Experimental Results and Analysis
4.2.1. Performance Comparison with Other Models
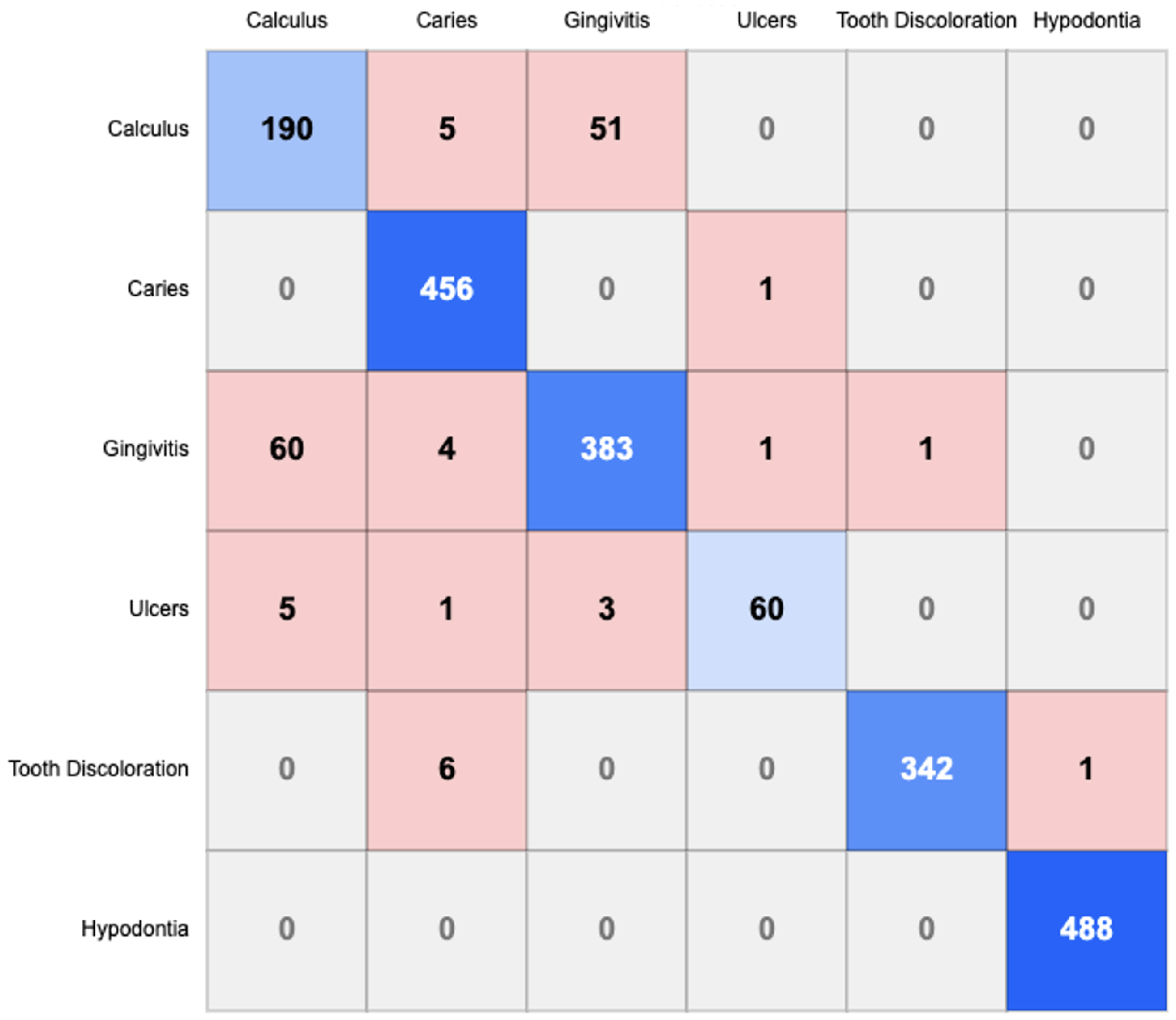
4.2.2. Confusion Matrix Analysis
- Calculus-Gingivitis Confusion: A substantial bidirectional misclassification exists between Calculus and Gingivitis, with 51 instances of Calculus misclassified as Gingivitis and 60 instances of Gingivitis misclassified as Calculus. This suggests morphological similarities between these conditions that challenge accurate differentiation.
- Caries Classification: Caries demonstrates minimal confusion with other categories, indicating that its visual manifestations are sufficiently distinctive for accurate identification by the model.
- Tooth Discoloration: Six instances of Tooth Discoloration are misclassified as Caries, indicating potential visual similarities in certain presentations of these conditions.
- Hypodontia Recognition: Hypodontia shows negligible confusion with other conditions, achieving near-perfect classification with 488 correct identifications and no false negatives, suggesting highly distinctive visual characteristics.
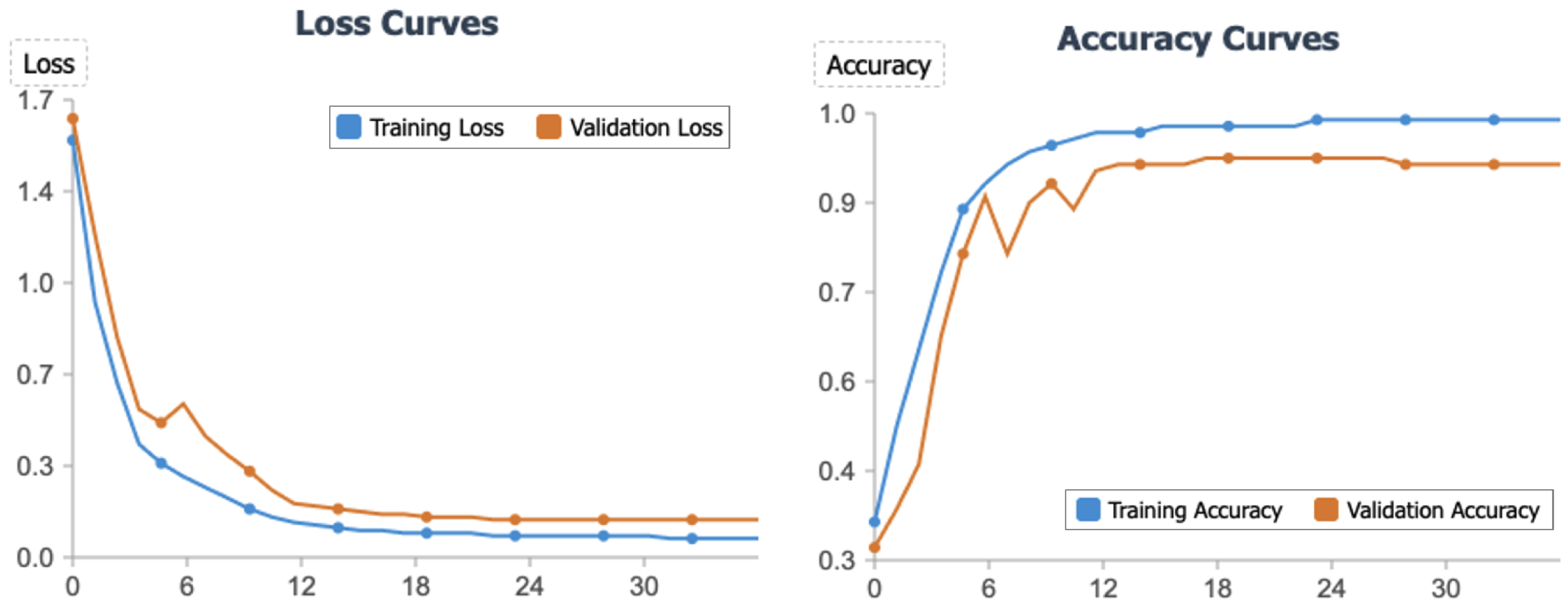
4.3. Convergence Analysis
4.4. Ablation Study
4.4.1. Overall Performance Gain from Our Multiple Designs
4.4.2. Effectiveness of Main Designs
- Gradient Suppression: Eliminating the gradient suppression mechanism reduces accuracy by 2.2% (from 93.3% to 91.1%) and macro-F1 score by 1.8% (from 91.9% to 90.1%). This decline underscores the importance of gradient suppression in mitigating overfitting and bolstering the model’s generalization capacity.
-
Componential Prototypes: Removing componential prototypes results in a notable decrease of 3.0% in accuracy (from 93.3% to 90.3%) and 2.8% in macro-F1 score (from 91.9% to 89.1%). This indicates that componential prototypes are instrumental in capturing fine-grained features essential for distinguishing diverse oral disease manifestations. Furthermore, we investigated the impact of varying the number of componential prototypes on model performance:
- -
- Increased Prototype Number: Doubling the number of componential prototypes to 1024 yields a slight performance reduction compared to the default configuration, with accuracy decreasing by 1.2% (from 93.3% to 92.1%) and macro-F1 score by 0.6% (from 91.9% to 91.3%). This suggests that an excess of prototypes may introduce redundancy, potentially capturing noise rather than meaningful patterns.
- -
- Reduced Prototype Number: Halving the number of componential prototypes to 256 results in an accuracy drop of 1.7% (from 93.3% to 91.6%) and a macro-F1 score reduction of 1.4% (from 91.9% to 90.5%). This indicates that a minimum threshold of prototypes is required to adequately represent the diversity of features necessary for precise oral disease classification.
- Dynamic Compositional Representation: The absence of dynamic compositional representation leads to the most pronounced performance drop, with accuracy decreasing by 3.5% (from 93.3% to 89.8%) and macro-F1 score by 3.8% (from 91.9% to 88.1%). This substantial degradation emphasizes the pivotal role of dynamic representations in generating discriminative features that effectively differentiate between oral disease categories.
5. Conclusions
Author Contributions: Wenyuan Wang
References
- Zhao, Y.; Zhang, L.; Liu, Y.; Meng, D.; Cui, Z.; Gao, C.; Gao, X.; Lian, C.; Shen, D. Two-stream graph convolutional network for intra-oral scanner image segmentation. IEEE Transactions on Medical Imaging 2021, 41, 826–835. [Google Scholar] [CrossRef]
- Dixit, S.; Kumar, A.; Srinivasan, K. A current review of machine learning and deep learning models in oral cancer diagnosis: Recent technologies, open challenges, and future research directions. Diagnostics 2023, 13, 1353. [Google Scholar] [CrossRef] [PubMed]
- López-Cortés, X.A.; Matamala, F.; Venegas, B.; Rivera, C. Machine-learning applications in oral cancer: A systematic review. Applied Sciences 2022, 12, 5715. [Google Scholar] [CrossRef]
- Rashid, J.; Qaisar, B.; Faheem, M.; Akram, A.; Amin, R.; Hamid, M. Mouth and oral disease classification using InceptionResNetV2 method. Multimedia Tools and Applications 2024, 83, 33903–33921. [Google Scholar] [CrossRef]
- Can, Z.; Isik, S.; Anagun, Y. CVApool: using null-space of CNN weights for the tooth disease classification. Neural Computing and Applications 2024, 36, 16567–16579. [Google Scholar] [CrossRef]
- Kang, J.; Le, V.; Lee, D.; Kim, S. Diagnosing oral and maxillofacial diseases using deep learning. Scientific Reports 2024, 14, 2497. [Google Scholar] [CrossRef]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2017, pp.
- Sandler, M.; Howard, A.; Zhu, M.; Zhmoginov, A.; Chen, L.C. MobileNetV2: Inverted Residuals and Linear Bottlenecks. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), June 2018. [Google Scholar]
- Huo, S.; Zhou, Y.; Chen, K.; Xiang, W. Skim-and-scan transformer: A new transformer-inspired architecture for video-query based video moment retrieval. Expert Systems with Applications 2025, 270, 126525. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2016, pp.
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. In Proceedings of the Proceedings of the International Conference on Learning Representations (ICLR); 2015. [Google Scholar]
- Zhou, Y.; Wang, H.; Huo, S.; Wang, B. Hierarchical full-attention neural architecture search based on search space compression. Knowledge-Based Systems 2023, 269, 110507. [Google Scholar] [CrossRef]
- Zhou, Y.; Hao, J.; Huo, S.; Wang, B.; Ge, L.; Kung, S.Y. Automatic metric search for few-shot learning. IEEE Transactions on Neural Networks and Learning Systems 2023. [Google Scholar] [CrossRef]
- Adeoye, J.; Su, Y. Artificial intelligence in salivary biomarker discovery and validation for oral diseases. Oral Diseases 2024, 30, 23–37. [Google Scholar] [CrossRef]
- Zhou, M.; Jie, W.; Tang, F.; Zhang, S.; Mao, Q.C.; et al. L. Deep learning algorithms for classification and detection of recurrent aphthous ulcerations using oral clinical photographic images. Journal of Dental Sciences 2024, 19, 254–260. [Google Scholar] [CrossRef]
- Mira, E.; Sapri, A.; Aljehanı, R.; Jambı, B.; Bashir, T.E.; et al. E.K. Early diagnosis of oral cancer using image processing and artificial intelligence. Fusion: Practice and Applications 2024, 14, 293–308. [Google Scholar]
- Babu, P.; Rai, A.; Ramesh, J.; Nithyasri, A.; Sangeetha, S.P.; et al. K. An explainable deep learning approach for oral cancer detection. Journal of Electrical Engineering and Technology 2024, 19, 1837–1848. [Google Scholar] [CrossRef]
- Rokhshad, R.; Mohammad-Rahimi, H.; Price, J.; Shoorgashti, R.; Abbasiparashkouh, Z.M.; et al. E. Artificial intelligence for classification and detection of oral mucosa lesions on photographs: a systematic review and meta-analysis. Clinical Oral Investigations 2024, 28, 88. [Google Scholar] [CrossRef]
- Zanini, L.; Rubira-Bullen, I.; Nunes, F. A systematic review on caries detection classification and segmentation from x-ray images: methods datasets evaluation and open opportunities. Journal of Imaging Informatics in Medicine 2024, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Montagnoli, D.; Leite, V.; Godoy, Y.; Lafetá, V.; Junior, E.A.; et al. C. Can predictive factors determine the time to treatment initiation for oral and oropharyngeal cancer? A classification and regression tree analysis. Plos One 2024, 19, e0302370. [Google Scholar] [CrossRef]
- Deo, B.; Pal, M.; Panigrahi, P.; Pradhan, A. An ensemble deep learning model with empirical wavelet transform feature for oral cancer histopathological image classification. International Journal of Data Science and Analytics 2024, 1–18. [Google Scholar] [CrossRef]
- Ansari, A.; Singh, A.; Singh, M.; Kukreja, V. Enhancing Skin Disease Classification: A Hybrid CNN-SVM Model Approach. In Proceedings of the Proceedings of the 2024 International Conference on Automation and Computation (AUTOCOM), March 2024, pp.
- Holtfreter, B.; Kuhr, K.; Tonetti, M.; Sanz, M.K.; et al. K. ACES: A new framework for the application of the 2018 periodontal status classification scheme to epidemiological survey data. Journal of Clinical Periodontology 2024, 51, 512–521. [Google Scholar] [CrossRef]
- Liu, C.; Eim, I.; Kim, J. High accuracy handwritten Chinese character recognition by improved feature matching method. In Proceedings of the Proceedings of the Fourth International Conference on Document Analysis and Recognition (ICDAR).
- Yang, H.; Zhang, X.; Yin, F.; Liu, C. Robust classification with convolutional prototype learning. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2018, pp.
- Snell, J.; Swersky, K.; Zemel, R. Prototypical networks for few-shot learning. In Proceedings of the Advances in Neural Information Processing Systems (NeurIPS); 2017; pp. 4077–4087. [Google Scholar]
- Chen, Y.; Zhu, X.; Gong, S. Semi-supervised deep learning with memory. In Proceedings of the Proceedings of the European Conference on Computer Vision (ECCV), 2018, pp.
- Han, J.; Luo, P.; Wang, X. Deep self-learning from noisy labels. In Proceedings of the Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV), 2019, pp.
- Kuo, C.; Ma, C.; Huang, J.; Kira, Z. Manifold graph with learned prototypes for semi-supervised image classification. arXiv preprint 2019, arXiv:1906.05202. [Google Scholar]
- Zhou, L.; Chen, H.; Wei, Y.; Li, X. Mining confident supervision by prototypes discovering and annotation selection for weakly supervised semantic segmentation. Neurocomputing 2022, 501, 420–435. [Google Scholar] [CrossRef]
- Zhou, T.; Zhang, M.; Zhao, F.; Li, J. Regional semantic contrast and aggregation for weakly supervised semantic segmentation. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2022, pp.
- Yang, B.; Liu, C.; Li, B.; Jiao, J.; Ye, Q. Prototype mixture models for few-shot semantic segmentation. In Proceedings of the Proceedings of the European Conference on Computer Vision (ECCV).
- Du, Y.; Fu, Z.; Liu, Q.; Wang, Y. Weakly Supervised Semantic Segmentation by Pixel-to-Prototype Contrast. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2022, pp.
- Chen, Q.; Yang, L.; Lai, J.; Xie, X. Self-supervised Image-specific Prototype Exploration for Weakly Supervised Semantic Segmentation. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2022, pp.
- Cheng, P.; Lin, L.; Lyu, J.; Huang, Y.; Luo, W.; Tang, X. Prior: Prototype representation joint learning from medical images and reports. In Proceedings of the Proceedings of the IEEE International Conference on Computer Vision (ICCV), 2023, pp.
- Song, G.; Liu, Y.; Wang, X. Revisiting the Sibling Head in Object Detector. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2020, pp.
- Rezatofighi, H.; Tsoi, N.; Gwak, J.; Sadeghian, A.; Reid, I.; Savarese, S. Generalized intersection over union: A metric and a loss for bounding box regression. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2019, pp.
- Carion, N.; Massa, F.; Synnaeve, G.; Usunier, N.; Kirillov, A.; Zagoruyko, S. End-to-end object detection with transformers. In Proceedings of the Proceedings of the European Conference on Computer Vision (ECCV).
- Fan, H.; Lin, L.; Yang, F.; Chu, P.; Deng, G.; Yu, S.; Bai, H.; Xu, Y.; Liao, C.; Ling, H. Lasot: A high-quality benchmark for large-scale single object tracking. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2019, pp.
- Huang, L.; Zhao, X.; Huang, K. Got-10k: A large high-diversity benchmark for generic object tracking in the wild. IEEE Transactions on Pattern Analysis and Machine Intelligence 2019, 43, 1562–1577. [Google Scholar] [CrossRef] [PubMed]
- Tan, M.; Le, Q.V. EfficientNet: Rethinking Model Scaling for Convolutional Neural Networks. In Proceedings of the Proceedings of the International Conference on Machine Learning (ICML); 2019. [Google Scholar]
- Cai, H.; Li, J.; Hu, M.; Gan, C.; Han, S. EfficientViT: Lightweight Multi-Scale Attention for High-Resolution Dense Prediction. In Proceedings of the Proceedings of the IEEE International Conference on Computer Vision (ICCV); 2023. [Google Scholar]
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A.A. Inception-v4, Inception-ResNet and the Impact of Residual Connections on Learning. In Proceedings of the Proceedings of the AAAI Conference on Artificial Intelligence (AAAI); 2017. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the Inception architecture for computer vision. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2016, pp.
- Chen, C.F.R.; Zhu, Y.; Sukthankar, R. CrossViT: Cross-Attention Multi-Scale Vision Transformer for Image Classification. In Proceedings of the Proceedings of the IEEE International Conference on Computer Vision (ICCV); 2021. [Google Scholar]
- Touvron, H.; Cord, M.; Douze, M.; Massa, F.; Sablayrolles, A.; Jégou, H. Training data-efficient image transformers and distillation through attention. In Proceedings of the Proceedings of the International Conference on Machine Learning (ICML); 2021. [Google Scholar]
- Han, K.; Xiao, A.; Wu, E.; et al. . Transformer in Transformer. arXiv 2021, arXiv:2103.00112. [Google Scholar]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; et al. . An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale. In Proceedings of the Proceedings of the International Conference on Learning Representations (ICLR); 2021. [Google Scholar]
| Model | Accuracy | Macro-Precision | Macro-Recall | Macro-F1 Score |
|---|---|---|---|---|
| DenseNet121 | 89.6% | 89.7% | 89.6% | 89.6% |
| MobileNet-v2 | 89.4% | 89.8% | 89.4% | 89.6% |
| ResNet50 | 82.7% | 82.8% | 82.7% | 82.2% |
| VGG16 | 64.5% | 62.1% | 64.5% | 61.2% |
| VGG19 | 50.3% | 48.9% | 50.3% | 44.2% |
| EfficientNet-B0 | 90.8% | 90.9% | 90.8% | 90.4% |
| EfficientNet-B5 | 86.8% | 87.5% | 86.8% | 86.9% |
| EfficientViT-B3 | 91.0% | 89.5% | 90.3% | 89.8% |
| EfficientViT-L3 | 90.5% | 89.5% | 90.0% | 89.7% |
| EfficientViT-M3 | 89.5% | 89.1% | 88.7% | 88.8% |
| InceptionResnet-v2 | 92.5% | 91.1% | 91.9% | 91.3% |
| Inception-v3 | 84.3% | 84.7% | 84.3% | 84.5% |
| CrossViT | 79.5% | 78.8% | 77.6% | 78.1% |
| DEIT | 87.3% | 86.2% | 85.2% | 85.6% |
| TNTTransformer | 76.4% | 75.1% | 73.3% | 73.8% |
| Vision Transformer | 75.6% | 75.7% | 74.6% | 74.9% |
| DyCoP (Ours) | 93.3% | 92.5% | 91.2% | 91.9% |
| Variant | Accuracy | Macro-Precision | Macro-Recall | Macro-F1 Score |
|---|---|---|---|---|
| DyCoP (Full) | 93.3% | 92.5% | 91.2% | 91.9% |
| DyCoP w/o all designs (ResNet-18) | 83.3% | 81.4% | 85.0% | 83.2% |
| DyCoP w/o Componential Prototypes | 90.3% | 89.7% | 88.6% | 89.1% |
| w/ 1024 (2×) Componential Prototypes | 92.1% | 91.5% | 91.0% | 91.3% |
| w/ 256 (half) Componential Prototypes | 91.6% | 90.9% | 90.2% | 90.5% |
| DyCoP w/o Dynamic Comp. Rep. | 89.8% | 88.9% | 87.4% | 88.1% |
| DyCoP w/o Gradient Suppression | 91.1% | 90.8% | 89.5% | 90.1% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




