Submitted:
25 March 2025
Posted:
26 March 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Isolation and Identification of ESBL Organisms in WWTPs
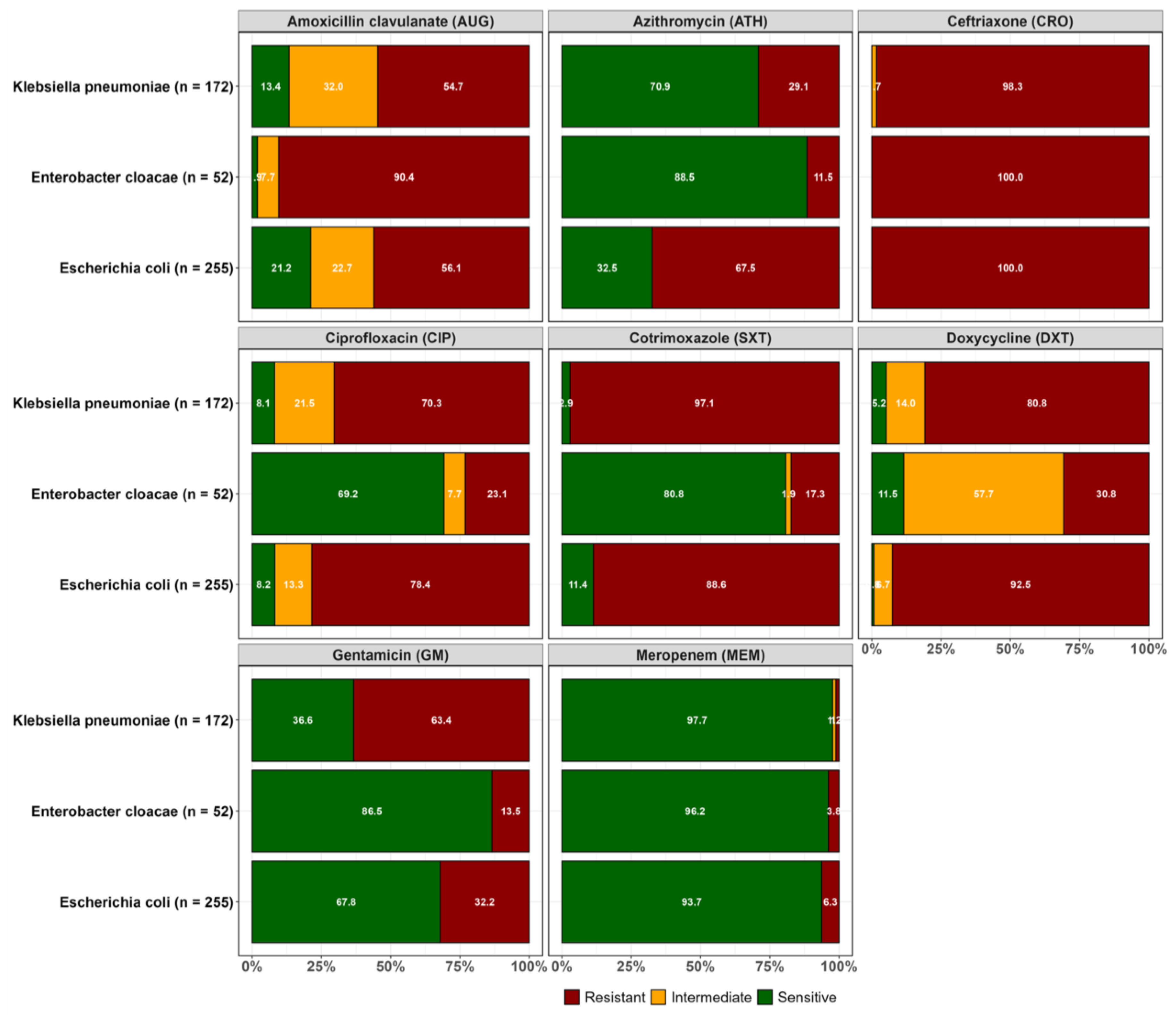
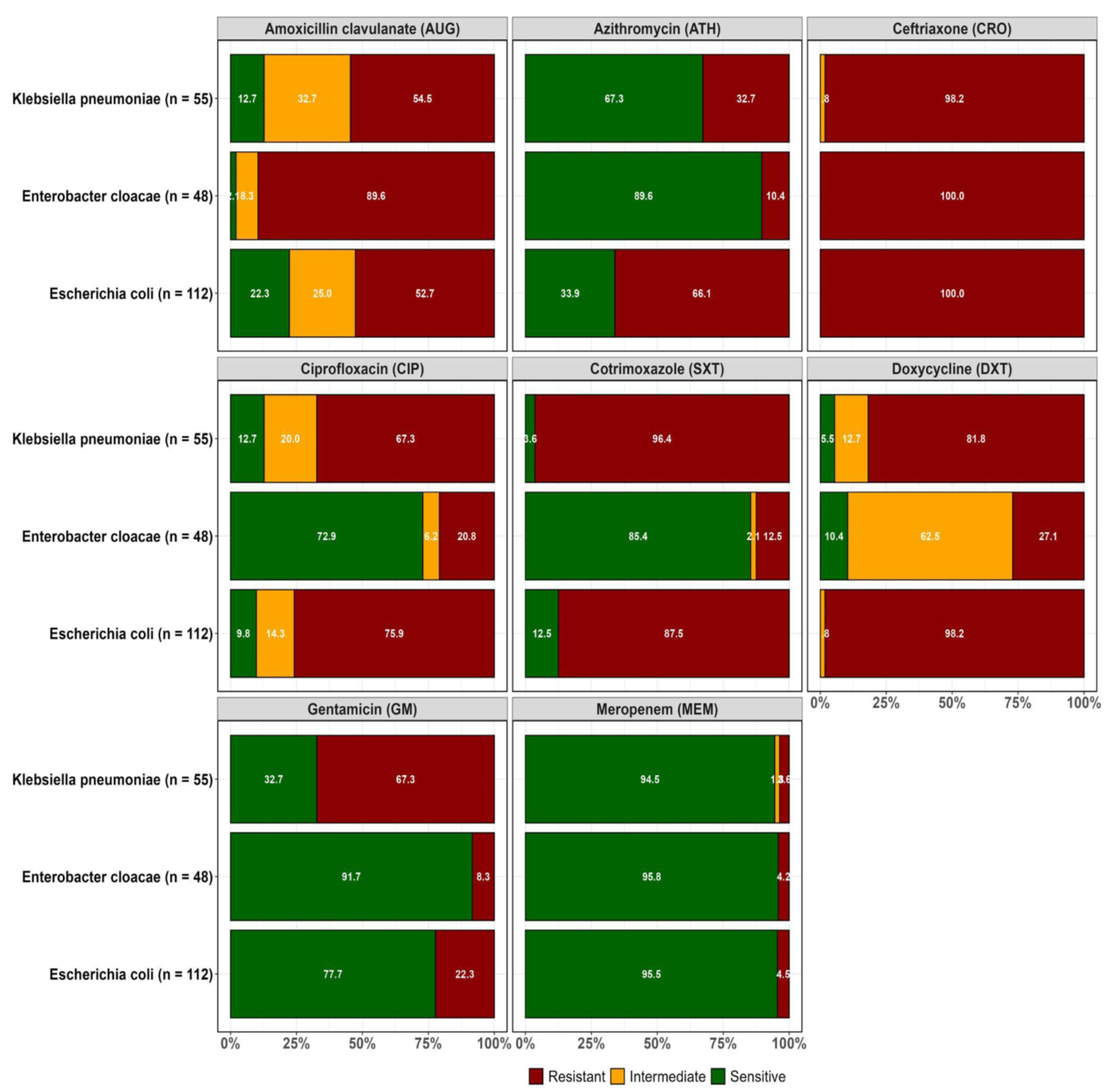
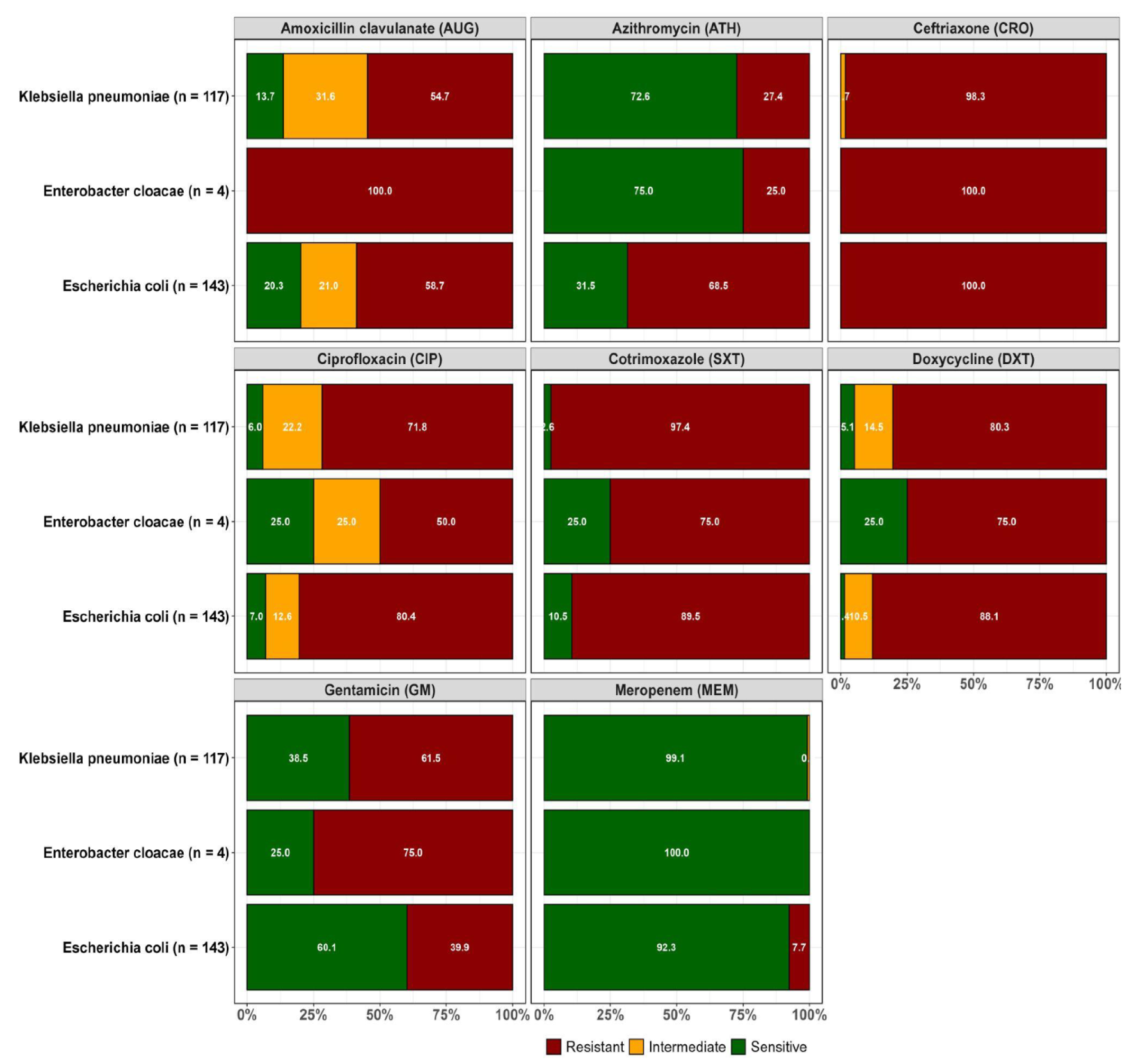
2.2. Resistance Profiles of Isolated ESBL-Producing E. coli, K. pneumoniae and Enterobacter cloacae
3. Discussion
4. Materials and Methods
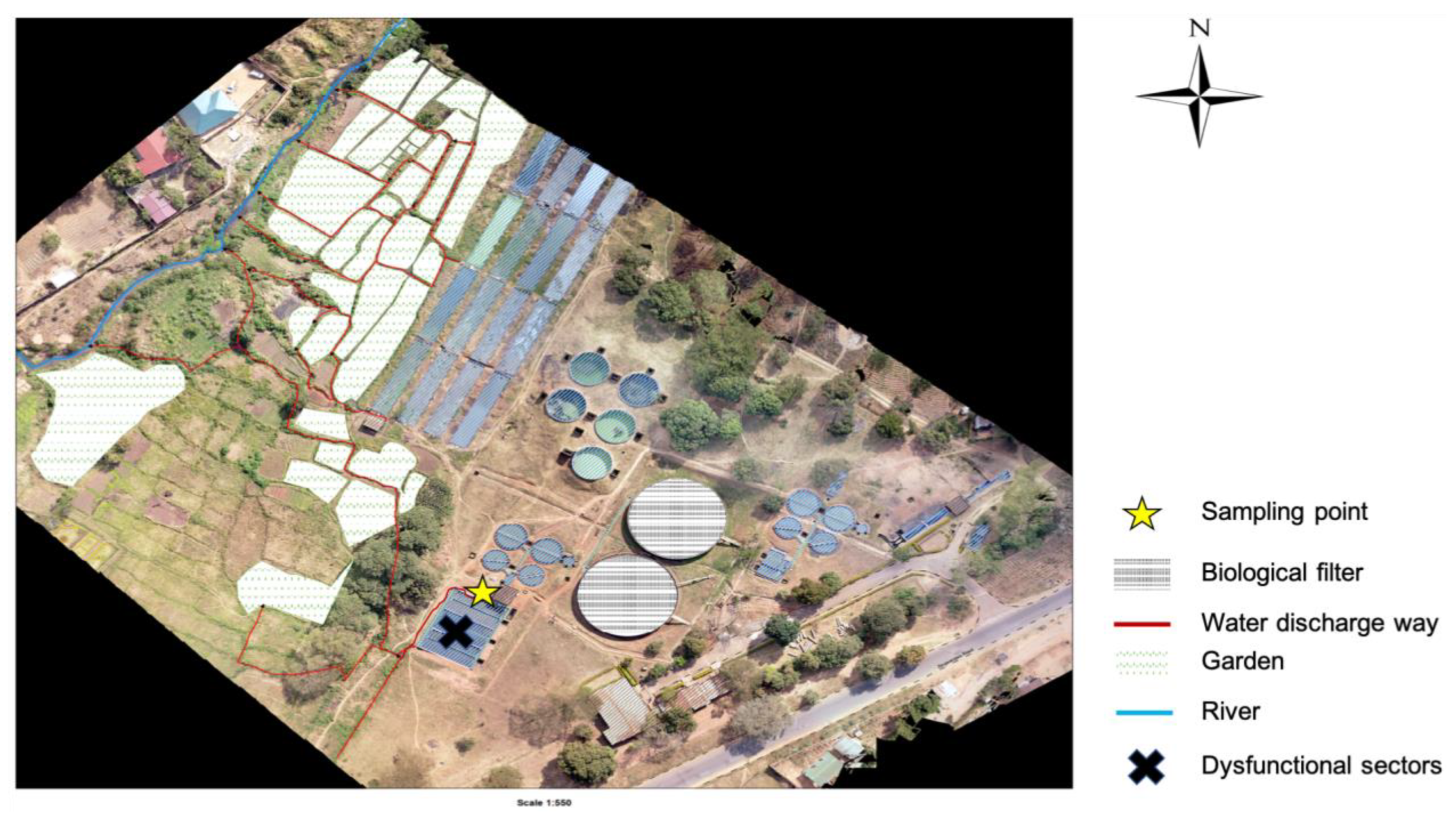
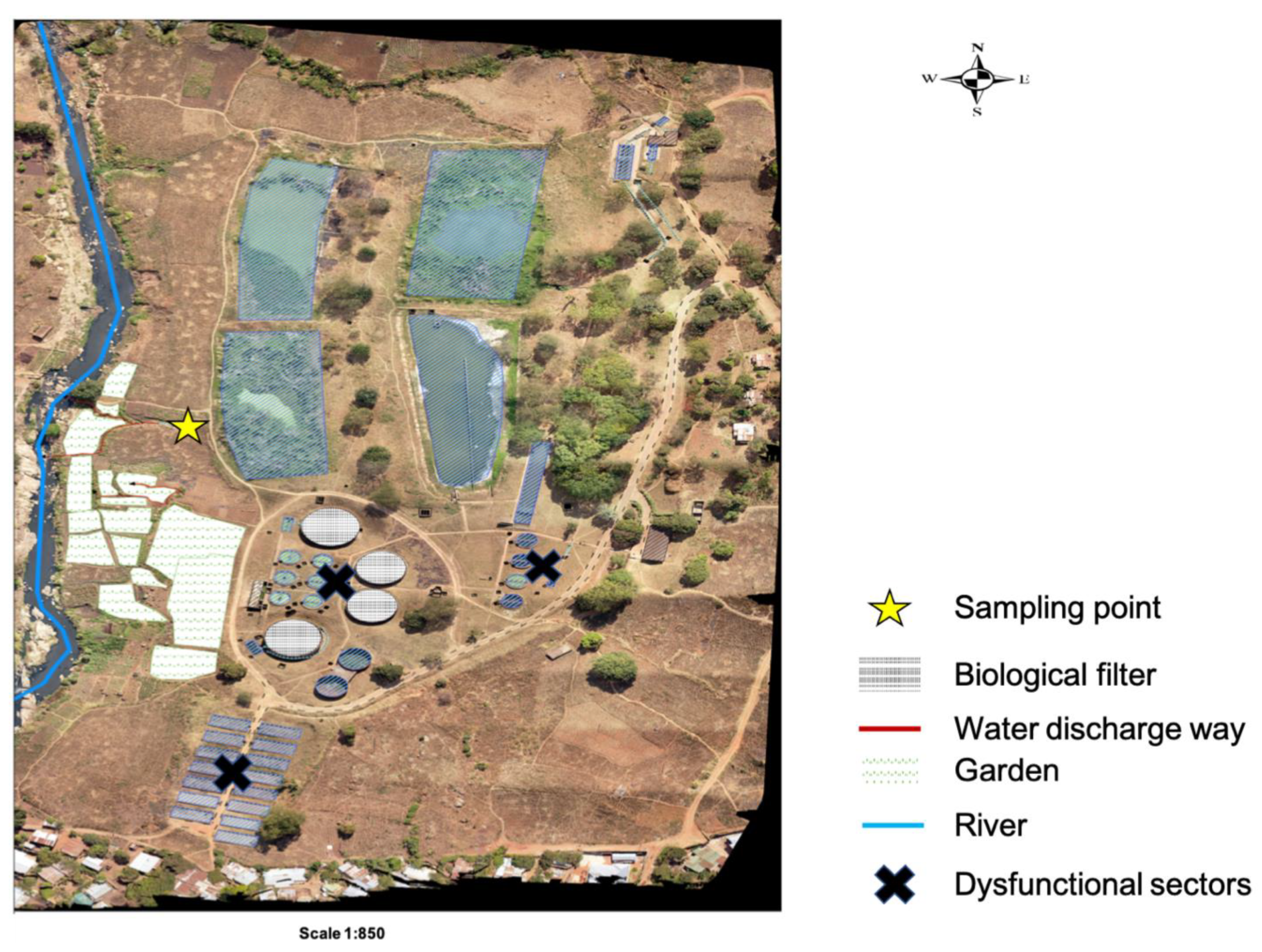
4.1. Study Sites
4.2. Sample Collection
4.3. Sample Processing, Culture and Bacterial Identification
4.4. Antimicrobial Susceptibility Testing
4.5. Multiple Antibiotic-Resistant Phenotype and Multiple Antibiotic Resistance Index
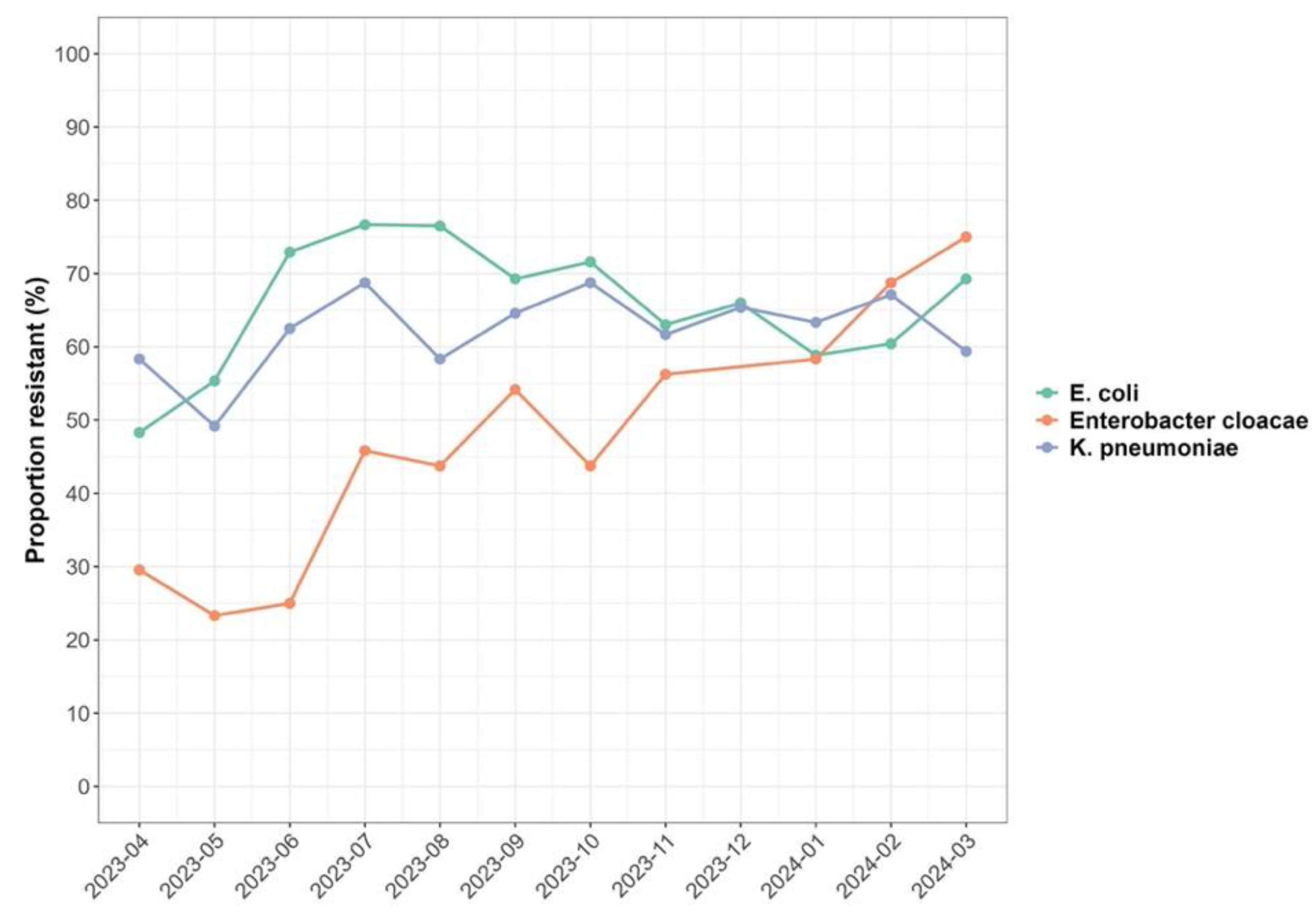
4.6. Temperature and Rainfall Data
4.7. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Singer AC, Shaw H, Rhodes V, et al. Review of antimicrobial resistance in the environment and its relevance to environmental regulators. Front Microbiol 2016; 7: 1–22.
- Andleeb S, Majid M, Sardar S. Environmental and public health effects of antibiotics and AMR/ARGs. 2020; 269–291.
- Thai PK, Xuan L, Ngan V, et al. Occurrence of antibiotic residues and antibiotic-resistant bacteria in effluents of pharmaceutical manufacturers and other sources around Hanoi, Vietnam. Sci Total Environ 2018; 645: 393–400.
- Chitescu CL, Lupoae M, Elisei AM. Pharmaceutical residues in the environment - new european integrated programs required. Rev Chim 2016; 67: 1008–1013.
- Papajov I, Gregov G, Papaj J, et al. Effect of wastewater treatment on bacterial community, antibiotic-resistant bacteria and endoparasites. Int J Environ Res Public Health; 19. [CrossRef]
- Mbanga J, Luther A, Abia K, et al. Longitudinal surveillance of antibiotic resistance in Escherichia coli and Enterococcus spp. from a wastewater treatment plant and its associated waters in KwaZulu-Natal, South Africa. Microb Drug Resist 2021; 27: 904–918.
- Moges F, Endris M, Belyhun Y, et al. Isolation and characterization of multiple drug resistance bacterial pathogens from waste water in hospital and non-hospital environments, northwest Ethiopia. BMC Res Notes 2014, 2014; 7: 2–7.
- Milakovi M, Vestergaard G, González-plaza JJ, et al. Antibiotic-manufacturing sites are hot-spots for the release and spread of antibiotic resistance genes and mobile genetic elements in receiving aquatic environments pollution from azithromycin-manufacturing promotes macrolide-resistance gene propagation a. Environ Int 2019; 123: 501–511.
- Max M. Antibiotics, antibiotic resistance and environment. Encyclopedia of the environnement 2019; 1–9.
- Rodríguez-molina D, Mang P, Schmitt H, et al. Do wastewater treatment plants increase antibiotic resistant bacteria or genes in the environment? protocol for a systematic review. Syst Rev 2019; 9: 1–8.
- Collaborators, AR. Articles Global burden of bacterial antimicrobial resistance in 2019 : a systematic analysis. 399. Epub ahead of print 2022. [CrossRef]
- Tam PI, Musicha P, Kawaza K, et al. Emerging resistance to empiric antimicrobial regimens for pediatric bloodstream infections in Malawi (1998-2017). Clinical infectious diseases 2019; 2–30.
- Lester R, Musicha P, Kawaza K, et al. Articles Effect of resistance to third-generation cephalosporins on morbidity and mortality from bloodstream infections in Blantyre , Malawi : a prospective cohort study. The Lancet Microbe; 3: e922–e930.
- Kayambankadzanja RK, Lihaka M, Barratt-due A, et al. The use of antibiotics in the intensive care unit of a tertiary hospital in Malawi. BMC Infect Dis 2020; 20: 1–7.
- Mankhomwa J, Tolhurst R, M’biya E, et al. A qualitative study of antibiotic use practices in intensive small-scale farming in urban and peri-urban Blantyre, Malawi: implications for antimicrobial resistance. Front Vet Sci 2022; 9: 1–16.
- Cocker D, Chidziwisano K, Mphasa M, et al. Articles investigating one health risks for human colonisation with extended spectrum β-lactamase-producing Escherichia coli and Klebsiella pneumoniae in Malawian households: a longitudinal cohort study. The Lancet Microbe 2023; 4: 534–543.
- Tegha G, Ciccone EJ, Krysiak R, et al. Genomic epidemiology of Escherichia coli isolates from a tertiary referral center in Lilongwe , Malawi. Microb Genomics 2021; 7: 1–12.
- Varela AR, Manaia CM. Human health implications of clinically relevant bacteria in wastewater habitats. Env Sci Pollut Res 2013; 20: 3550–3569.
- Navidinia, M. The clinical importance of emerging ESKAPE pathogens in nosocomial infections. J Paramed Sci 2016; 7: 43–57.
- Oliveira DMP De, Forde BM, Kidd TJ, et al. Antimicrobial resistance in ESKAPE pathogens. Clin Microbiol Rev 2020; 33: 1–49.
- Musicha P, Feasey NA, Cain AK, et al. Genomic landscape of extended-spectrum β-lactamase resistance in Escherichia coli from an urban African setting. J Antimicrob Chemother 2017; 72: 1602–1609.
- Musicha P, Msefula CL, Mather AE, et al. Genomic analysis of Klebsiella pneumoniae isolates from Malawi reveals acquisition of multiple ESBL determinants across diverse lineages. J Antimicrob Chemother 2019; 74: 1223–1232.
- Nzima B, Adegoke AA, Ofon UA, et al. Resistotyping and extended-spectrum beta-lactamase genes among Escherichia coli from wastewater treatment plants and recipient surface water for reuse in South Africa. New Microbes New Infect 2020; 38: 1–7.
- Mutuku C, Melegh S, Kovacs K, Urban P, Virág E, Heninger R, Herczeg R, Sonnevend A, Gyenesei A, Fekete C et al. Characterization of β -lactamases and multidrug resistance mechanisms in enterobacterales from hospital effluents and wastewater treatment plant. Antibiotics 2022; 11: 1–21.
- Verburg I, Garc S, Hern L, et al. Abundance and antimicrobial resistance of three bacterial species along a complete wastewater pathway. Microorganisms 2019; 7: 1–15.
- Sekizuka T, Tanaka R, Hashino M, et al. Comprehensive genome and plasmidome analysis of antimicrobial resistant bacteria in wastewater treatment plant effluent of Tokyo. Antibiotics 2022; 11: 1–16.
- Alouache S, Estepa V, Messai Y, et al. Characterization of esbls and associated quinolone resistance in Escherichia coli and Klebsiella pneumoniae isolates from an urban wastewater plant in Algeria. Microb Drug Resist 2013; 00: 1–9.
- Mwapasa T, Mphasa M, Cocker D, et al. Community exposure assessment to anti-microbial resistance (AMR); case study of Malawi. In: UNC Water and Health, p. 1.
- Ministry of Health (Mw). Malawi standard treatment guidelines. 6th ed., https://www.differentiatedservicedelivery.org/wp-content/uploads/MSTG-6th-Edition-2023-Final-Draft-CC-gn-2-edditi_230719_133059.pdf (2023).
- Afunwa RA, Ezeanyinka J, Afunwa EC, et al. Multiple antibiotic resistant index of gram-negative bacteria from bird droppings in two commercial poultries in Enugu , Nigeria. Open J Med Microbiol 2020; 10: 171–181.
- Sibande GT, Banda NPK, Moya T, et al. Antibiotic guideline adherence by Clinicians in medical wards at Queen Elizabeth Central Hospital (QECH), Blantyre Malawi. Malawi Med J 2022; 34: 3–8.
- Macpherson EE, Mankhomwa J, Dixon J, et al. Household antibiotic use in Malawi : A cross- sectional survey from urban and peri-urban Blantyre. PLOS Glob Public Heal 2023; 1–13.
- Raven KE, Ludden C, Gouliouris T, et al. Genomic surveillance of Escherichia coli in municipal wastewater treatment plants as an indicator of clinically relevant pathogens and their resistance genes. Microbiol Soc 2019; 5: 1–10.
- Malawi national action plan on antimicrobial resistance: review of progress in the human health sector. Geneva: World Health Organization, 2021. Epub ahead of print 2021. DOI: licenses/by-nc-sa/3.0/igo.
- Conforti S, Holschneider A, Sylvestre É, et al. Monitoring ESBL- Escherichia coli in Swiss wastewater between November 2021 and November 2022: insights into population carriage . mSphere; 9. Epub ahead of print 2024. [CrossRef]
- MacFadden DR, McGough SF, Fisman D, et al. Antibiotic resistance increases with local temperature. Nat Clim Chang 2018; 8: 510–514.
- Kraslawski A, Avramenko Y. Comparison of pollutant levels in effluent from wastewater treatment plants in Blantyre, Malawi. Int J Water Resour Environ Eng 2010; 2: 79–86.
- Feasey NA, Gaskell K, Wong V, et al. Rapid emergence of multidrug resistant, H58-lineage salmonella typhi in Blantyre. PLoS Negl Trop Dis 2015; 9: 1–13.
- Teshome A, Alemayehu T, Deriba W, et al. Antibiotic resistance profile of bacteria isolated from wastewater systems in eastern Ethiopia. J Environ Public Health 2020; 2020: 10.
- CLSI. Performance standards for antimicrobial susceptibility testing. 30th ed. Wayne, PA: Clinical and Laboratory Standards Institute, 2020.
| Species of ESBL organisms isolated from each sample | Number of samples per site | ||
| Blantyre, N = 138 | Soche, N = 143 | Overall, N = 281 | |
| E. coli only | 4/138 (2.9%) |
3/143 (2.1%) |
7/281 (2.5%) |
| Enterobacter cloacae only | 16/138 (11.6%) |
0/143 (0.0%) |
16/281 (5.7%) |
| Enterobacter asberiae only | 1/138 (0.7%) |
0/143 (0.0%) |
1/281 (0.4%) |
| E.coli & Enterobacter cloacae | 32/138 (23.2%) |
4/143 (2.8%) |
36/281 (12.8%) |
| E. coli &K.pneumoniae pneumoniae | 55/138 (39.9%) |
117/143 (81.8%) |
172/281 (61.2%) |
| E. coli &K. pneumoniae ozaenae | 3/138 (2.2%) |
2/143 (1.4%) |
5/281 (1.8%) |
| Proteus vulgaris | 1/138 (0.7%) |
0/143 (0.0%) |
1/281 (0.4%) |
| E. coli & Rahnella aqualitis | 0/138 (0.0%) |
1/143 (0.7%) |
1/281 (0.4%) |
| E. coli & Raoultella ornithinolytica | 0/138 (0.0%) |
2/143 (1.4%) |
2/281 (0.7%) |
| E. coli & Low discrimination profiles (Aeromonas, other Klebsiella pneumoniae subspp, Pantoea and Vibrio species) | 19/138 (13.8%) |
14/143 (9.8%) |
33/281 (11.7%) |
| Other ESBL growth but no pink or blue colonies | 8/138 (5.8%) |
0/143 (0.0%) |
8/281 (2.8%) |
| MAR PHENOTYPE |
E. coli (n = 255) |
K. pneumoniae (n = 172) | E. cloacae (n = 52) | Total | MAR Index |
|---|---|---|---|---|---|
| AUG-MEM-SXT-DXT-CIP-GM-ATH-CRO | 6 | 0 | 0 | 6 | 1 |
| AUG-SXT-DXT-CIP-GM-ATH-CRO | 45 | 7 | 3 | 55 | 0.9 |
| AUG-SXT-DXT-GM-ATH-CRO | 1 | 4 | 0 | 5 | 0.8 |
| AUG-SXT-GM-ATH-CRO | 0 | 1 | 0 | 1 | 0.6 |
| SXT-GM-ATH-CRO | 0 | 1 | 0 | 1 | 0.5 |
| SXT-ATH-CRO | 1 | 1 | 1 | 3 | 0.4 |
| SXT-CIP-GM-CRO | 0 | 1 | 0 | 1 | 0.5 |
| SXT-CIP-GM-ATH-CRO | 0 | 1 | 0 | 1 | 0.6 |
| SXT-DXT-CIP-GM-ATH-CRO | 14 | 9 | 0 | 23 | 0.8 |
| SXT-CIP-ATH-CRO | 1 | 0 | 0 | 1 | 0.5 |
| SXT-CIP-CRO | 0 | 2 | 0 | 2 | 0.4 |
| SXT-DXT-CRO | 9 | 6 | 0 | 15 | 0.4 |
| SXT-DXT-CIP-CRO | 16 | 11 | 1 | 28 | 0.5 |
| AUG-SXT-DXT-CIP-CRO | 10 | 8 | 0 | 18 | 0.6 |
| AUG-SXT-DXT-CRO | 7 | 2 | 1 | 10 | 0.5 |
| AUG-SXT-CRO | 1 | 0 | 0 | 1 | 0.4 |
| AUG-MEM-SXT-DXT-ATH-CRO | 1 | 0 | 0 | 1 | 0.8 |
| AUG-SXT-DXT-CIP-GM-CRO | 3 | 40 | 2 | 45 | 0.8 |
| AUG-SXT-DXT-GM-CRO | 0 | 7 | 1 | 8 | 0.6 |
| DXT-CIP-GM-CRO | 1 | 0 | 1 | 2 | 0.5 |
| SXT-DXT-GM-CRO | 1 | 7 | 0 | 8 | 0.5 |
| SXT-DXT-CIP-GM-CRO | 6 | 10 | 0 | 16 | 0.6 |
| AUG-MEM-SXT-DXT-CIP-ATH-CRO | 6 | 0 | 0 | 6 | 0.9 |
| SXT-DXT-GM-ATH-CRO | 1 | 2 | 0 | 3 | 0.6 |
| AUG-SXT-CIP-ATH-CRO | 2 | 0 | 0 | 2 | 0.6 |
| AUG-MEM-SXT-CIP-GM-CRO | 0 | 1 | 0 | 1 | 0.8 |
| AUG-MEM-DXT-CIP-ATH-CRO | 0 | 0 | 1 | 1 | 0.7 |
| AUG-MEM-DXT-CIP-CRO | 0 | 0 | 1 | 1 | 0.6 |
| AUG-CIP-CRO | 0 | 1 | 2 | 3 | 0.4 |
| SXT-DXT-ATH-CRO | 4 | 6 | 0 | 10 | 0.5 |
| AUG-DXT-CRO | 1 | 0 | 4 | 5 | 0.4 |
| AUG-SXT-CIP-GM-CRO | 1 | 10 | 0 | 11 | 0.6 |
| AUG-MEM-SXT-DXT-GM-ATH-CRO | 1 | 0 | 0 | 1 | 0.9 |
| AUG-CIP-GM-CRO | 1 | 0 | 0 | 1 | 0.5 |
| AUG-MEM-CRO | 1 | 0 | 0 | 1 | 0.4 |
| SXT-DXT-CIP-ATH-CRO | 30 | 10 | 0 | 40 | 0.6 |
| DXT-CIP-ATH-CRO | 3 | 2 | 0 | 5 | 0.5 |
| DXT-CIP-CRO | 4 | 1 | 0 | 5 | 0.4 |
| DXT-GM-ATH-CRO | 1 | 0 | 0 | 1 | 0.5 |
| AUG-DXT-CIP-CRO | 1 | 0 | 0 | 1 | 0.5 |
| AUG-DXT-CIP-ATH-CRO | 0 | 0 | 1 | 1 | 0.6 |
| Univariable Poisson Model | Multivariable Poisson Model | |||||
| Characteristic | IRR1 | 95% CI1 | p-value | IRR1 | 95% CI1 | p-value |
| Minimum Temperature | 1.02 | 1.00, 1.04 | 0.080 | 1.04 | 1.01, 1.08 | 0.019 |
| Maximum Temperature | 1.00 | 0.98, 1.02 | 0.821 | 1.00 | 0.97, 1.02 | 0.637 |
| Total rainfall in Chichiri | 1.00 | 1.00, 1.00 | 0.015 | 1.00 | 1.00, 1.00 | 0.561 |
| Total rainfall in Mpemba | 1.00 | 1.00, 1.00 | 0.133 | 1.00 | 1.00, 1.00 | 0.564 |
| Wastewater treatment plant | <0.001 | <0.001 | ||||
| Blantyre | — | — | — | — | ||
| Soche | 1.41 | 1.30, 1.53 | 1.33 | 1.22, 1.46 | ||
| Organism | <0.001 | <0.001 | ||||
| E. coli | — | — | — | — | ||
| Enterobacter cloacae | 0.27 | 0.23, 0.32 | 0.30 | 0.25, 0.35 | ||
| K. pneumoniae | 0.75 | 0.69, 0.82 | 0.71 | 0.65, 0.78 | ||
| Antibiotic | <0.001 | <0.001 | ||||
| Amoxicillin clavulanate (AUG) | — | — | — | — | ||
| Azithromycin (ATH) | 1.01 | 0.85, 1.20 | 0.95 | 0.79, 1.14 | ||
| Ceftriaxone (CRO) | 1.62 | 1.40, 1.88 | 1.74 | 1.48, 2.04 | ||
| Ciprofloxacin (CIP) | 1.33 | 1.14, 1.56 | 1.33 | 1.13, 1.58 | ||
| Cotrimoxazole (SXT) | 1.58 | 1.35, 1.84 | 1.52 | 1.29, 1.79 | ||
| Doxycycline (DXT) | 1.40 | 1.20, 1.63 | 1.47 | 1.25, 1.73 | ||
| Gentamicin (GM) | 0.86 | 0.71, 1.03 | 0.86 | 0.71, 1.04 | ||
| Meropenem (MEM) | 0.35 | 0.21, 0.53 | 0.31 | 0.19, 0.48 | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




