Submitted:
25 February 2025
Posted:
26 February 2025
You are already at the latest version
Abstract
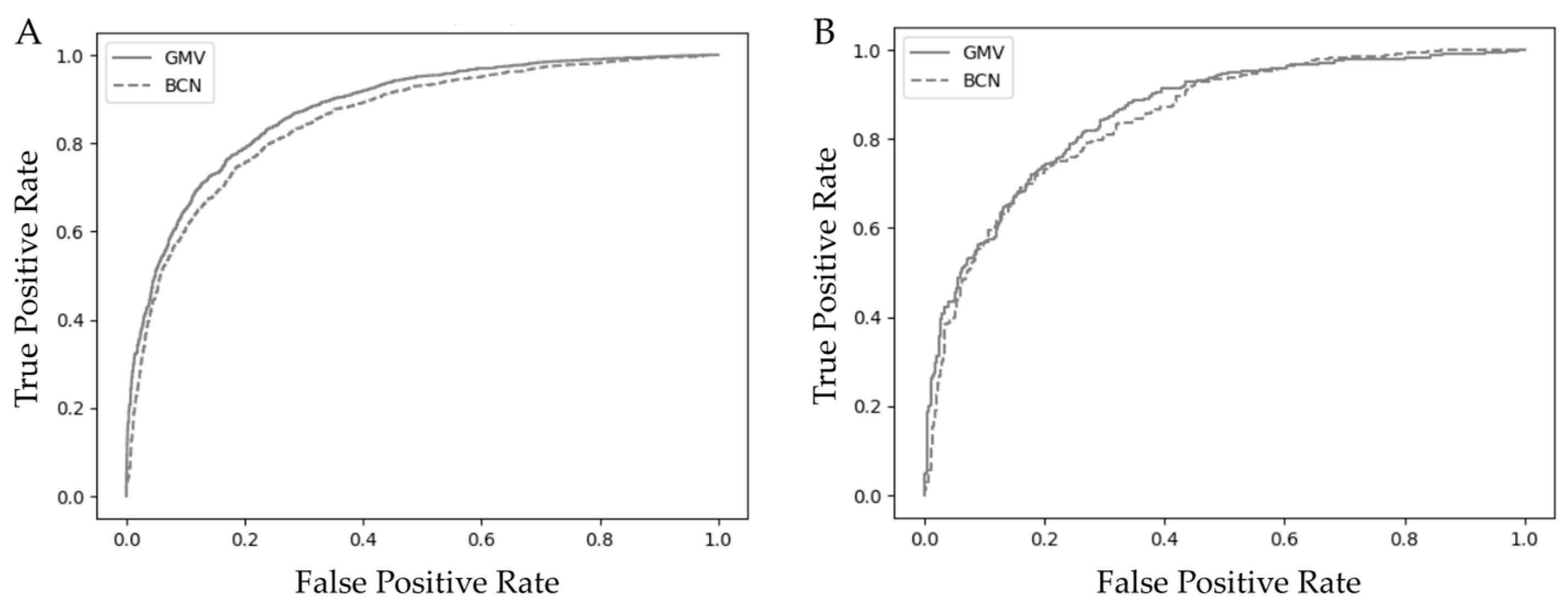
This study compares machine learning and logistic regression algorithms in developing a predictive model for sPCa using the seven predictive variables from the Barcelona (BCN-MRI) predictive model. A cohort of 5,005 men suspected of having PCa who underwent MRI, and targeted and/or systematic biopsies was used for training, validation and testing. A Feedforward Neural Network (FNN) SimpleNet-based model (GMV) and a logistic regression-based model (BCN) were developed. The models were evaluated for discrimination ability, net benefit and clinical utility. Both models demonstrated strong predictive performance. The GMV model achieved an area under the curve of 0.88 in training and 0.85 in test cohorts (95% CI: 0.83-0.90), while the BCN model reached 0.85 and 0.84 (95% CI: 0.82-0.87), respectively (p > 0.05). The GMV model exhibited superior recall, making it more suitable for clinical scenarios prioritizing sensitivity, whereas the BCN model demonstrated higher precision and specificity, optimizing the reduction of unnecessary biopsies. Both models provided net benefit over biopsying all men, reducing unnecessary procedures by 27.5-29% and 27-27.5% of prostate biopsies at 95% sensitivity, respectively (p > 0.05). Both, machine learning and logistic regression-based models exhibited high and comparable similar clinical performance in sPCa detection using a limited dataset.
Keywords:
1. Introduction
2. Materials and Methods
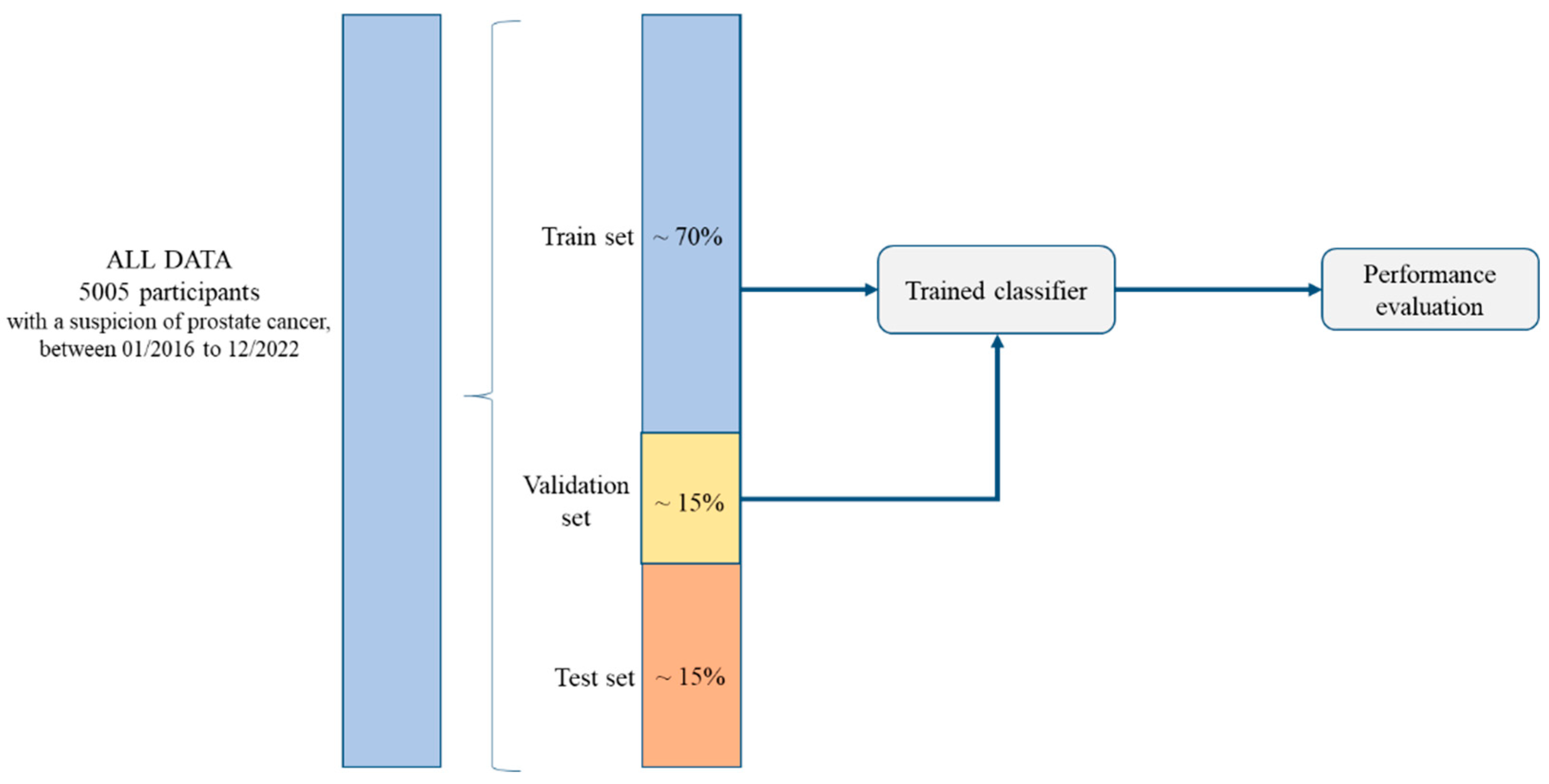
2.1. Study Design and Participants
2.2. Diagnostic Approach for Significant Prostate Cancer
2.3. Predictive Variables Included in the Models and Outcome Variable
2.4. Algorithms Used for Model Development
2.5. Statistical Analyses, Algorithm Performance, and Interpretation
3. Results
3.1. Participant Characteristics
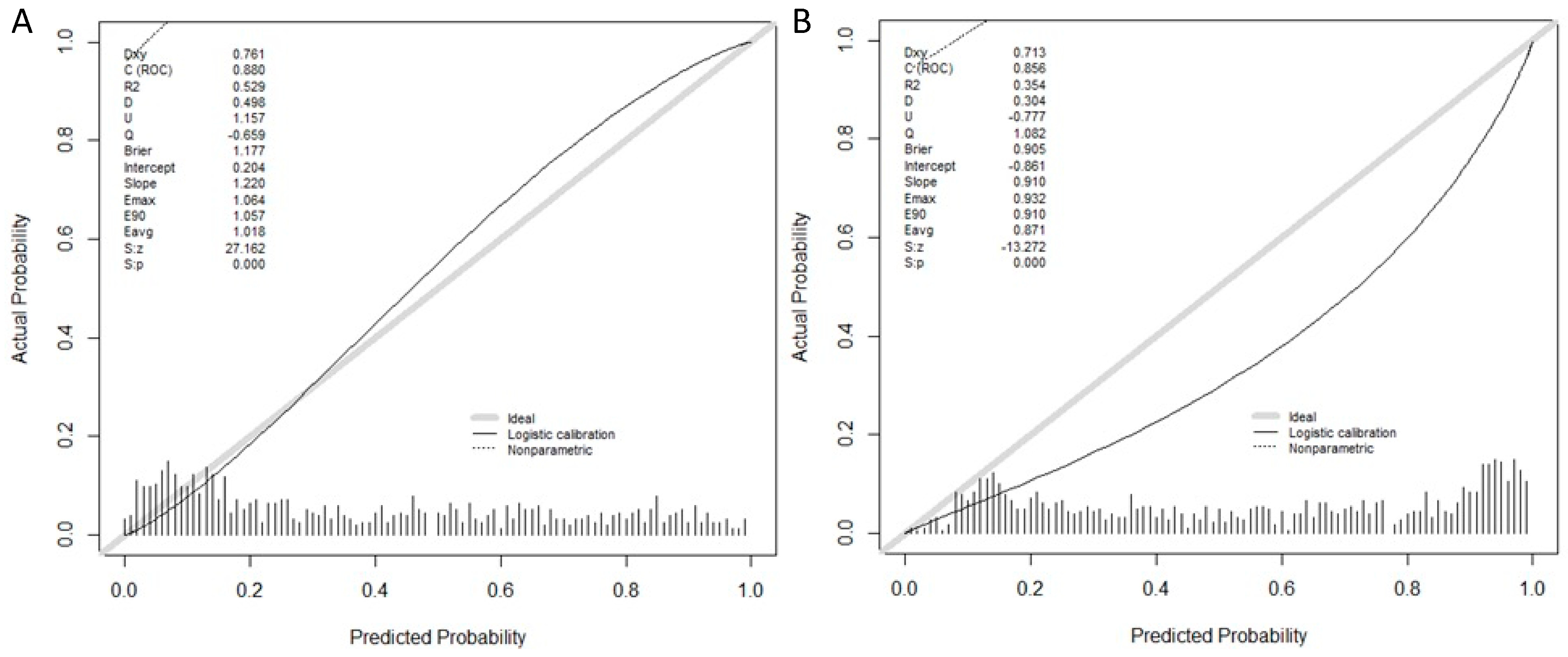
2.2. Calibration and Validation of the GMV and BCN Predictive Models
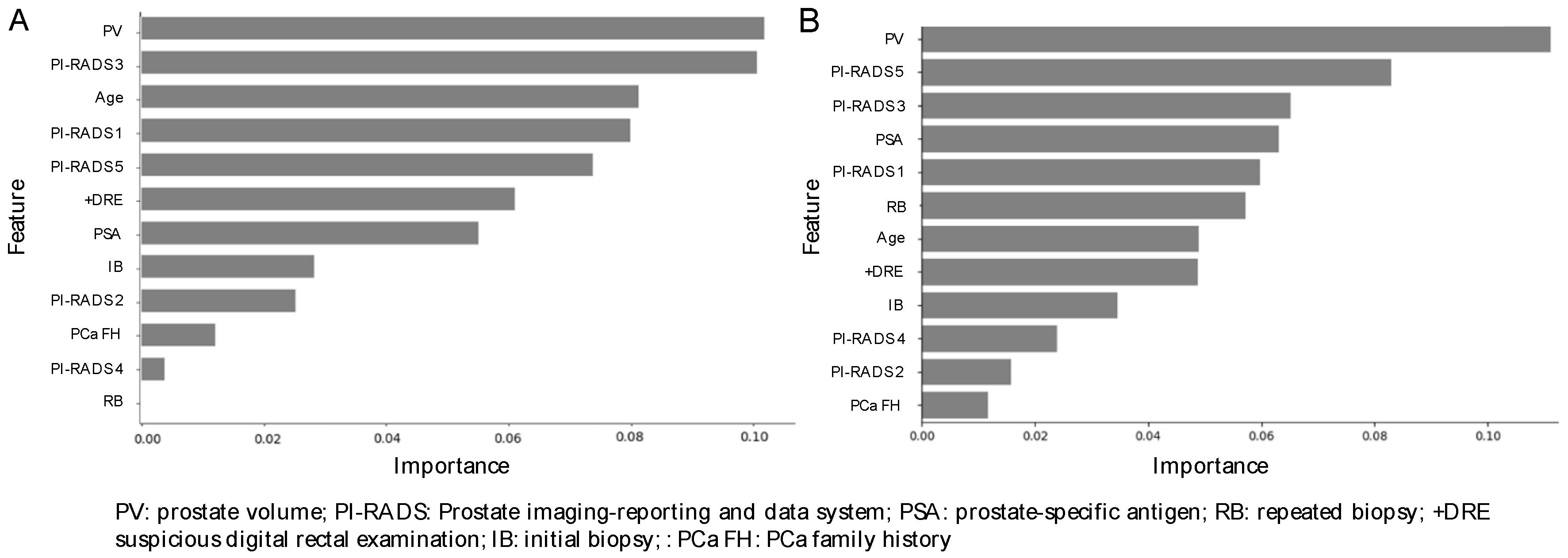
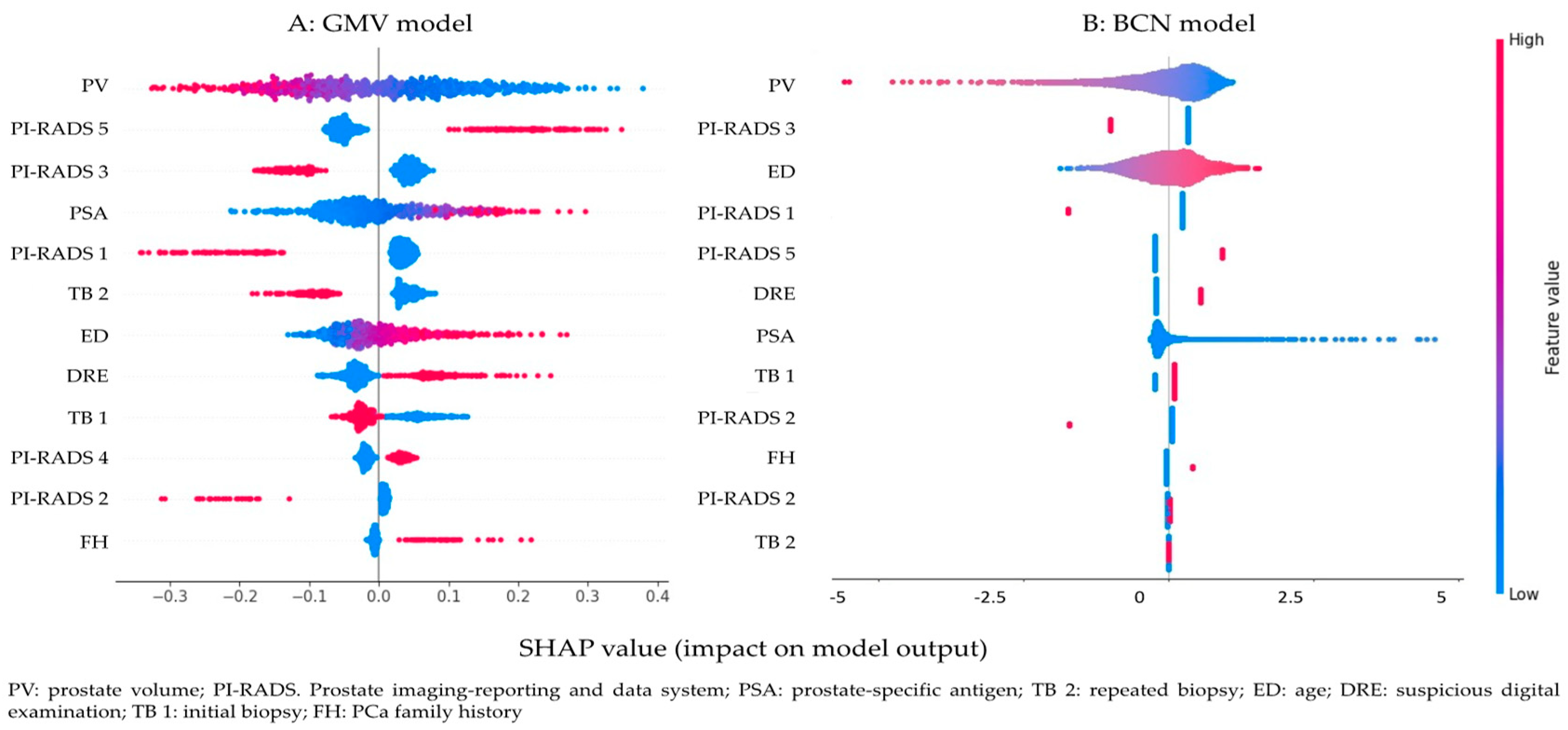
3.3. Variable Importance Interpretation with SHapley Additive exPlanations (SHAP)
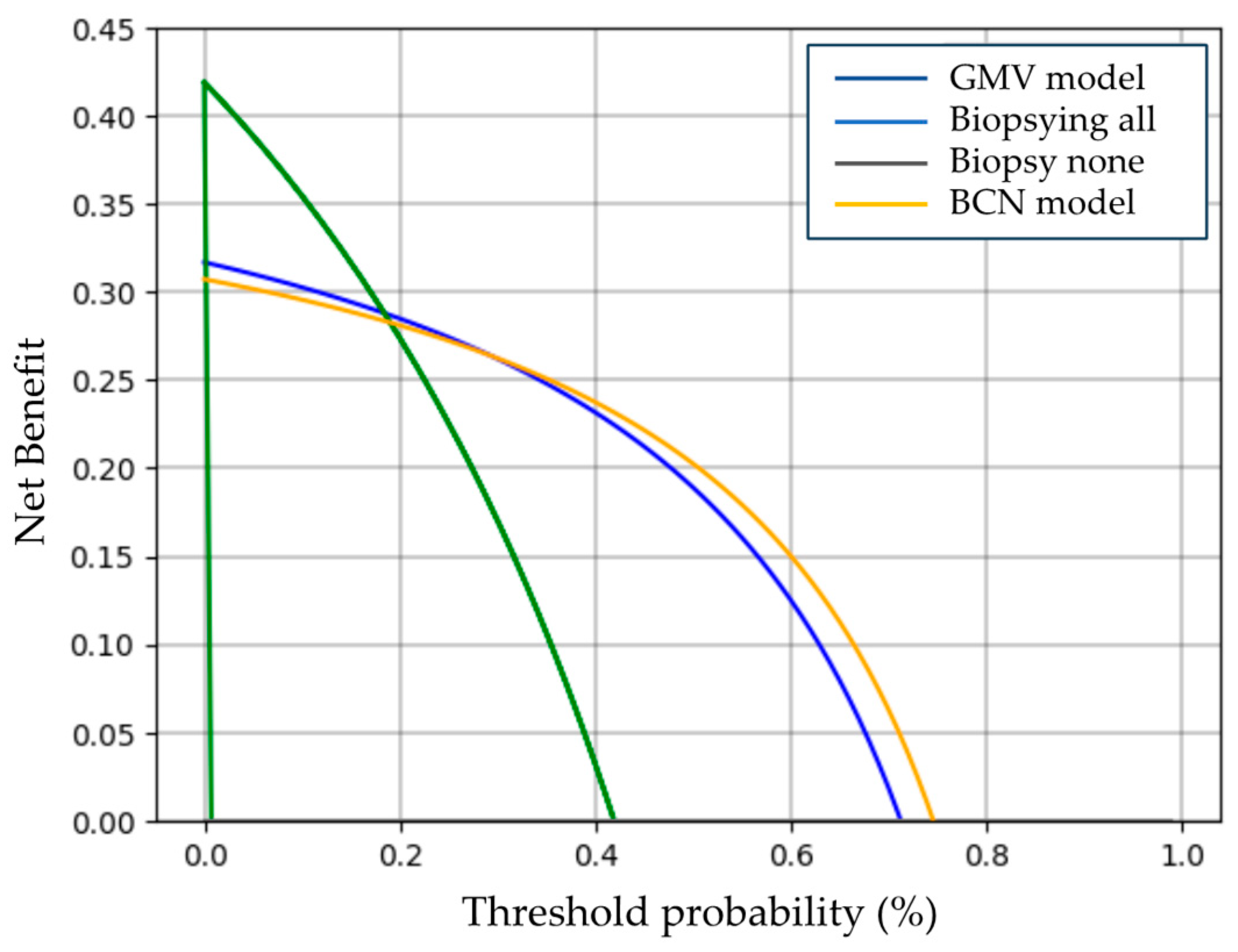
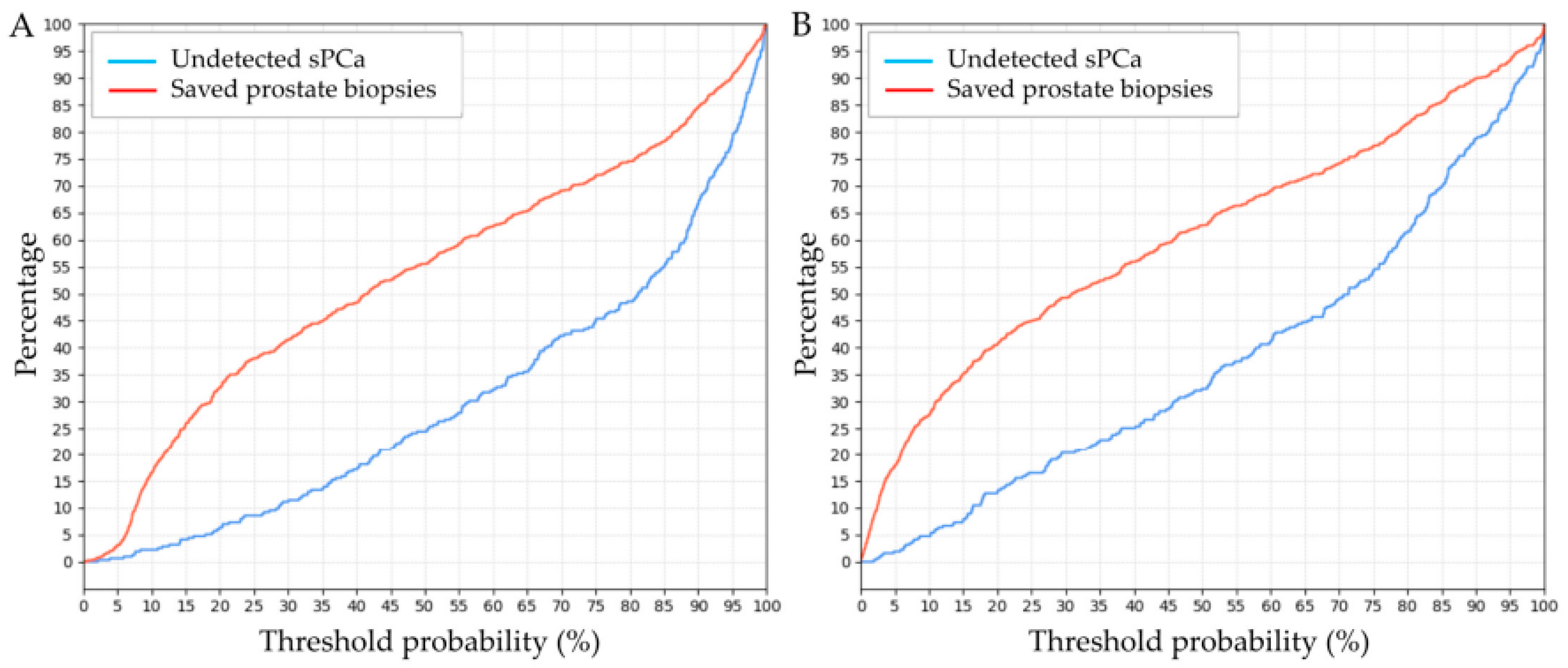
3.4. Clinical Comparison of GMV and BCN Predictive Models for sPCa Detection
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| Characteristic | Development cohort | Validation cohort | Test cohort | |
|---|---|---|---|---|
| Number of men | 4,254 | 639 | 751 | |
| Mean age at biopsy, years (SD) | 68 (8.3) | 68 (8.3) | 68 (8.2) | |
| Mean serum PSA, ng/mL (SD) | 13.4 (75.0) | 12.7 (35.5) | 12.8 (47.6) | |
| Suspicious DRE, n (%) | 1,210 (28.5) | 192 (30.1) | 217 (28.9) | |
| PCa family history, n (%) | 304 (7.2) | 51 (8.0) | 48 (6.4) | |
| Mean prostate volume, mL (SD) | 61.5 (32.3) | 62.0 (33.1) | 64.6 (35.8) | |
| Previous negative prostate biopsy, n (%) | 1,281 (30.2) | 216 (33.9) | 224 (29.9) | |
| PI-RADS version used | 2 | 2.1 | 2 | |
| Mean number of suspicious lesions | 2 | 2 | 2 | |
| PI-RADS score of index lesion, n (%) | ||||
| 1 | 470 (11.1) | 71 (11.2) | 104 (13.9) | |
| 2 | 148 (3.5) | 15 (2.4) | 27 (3.6) | |
| 3 | 1,053 (24.8) | 161 (25.2) | 197 (26.3) | |
| 4 | 1,743 (41.0) | 261 (40.9) | 272 (36.3) | |
| 5 | 840 (19.8) | 131 (20.6) | 151 (20.2) | |
| sPCa detection, n (%) | 1,782 (41.9) | 268 (42.0) | 315 (42.0) | |
| SD: standard deviation; PSA: prostatic specific antigen; DRE: digital rectal examination; PI-RADS: Prostate Imaging-Reporting and Data System; PCa: prostate cancer; sPCa: significant PCa. | ||||
References
- Van Poppel, H.; Albreht, T.; Basu, P.; Hogenhout, R.; Collen, S.; Roobol, M. Serum PSA-based early detection of prostate cancer in Europe and globally: Past, present and future. Nat. Rev. Urol. 2022, 19, 562–572. [Google Scholar] [CrossRef]
- Schroder, F.H.; Hugosson, J.; Roobol, M.J.; Tammela, T.L.; Ciatto, S.; Nelen, V.; Kwiatkowski, M.; Lujan, M.; Lilja, H.; Zappa, M.; et al. Screening and prostate-cancer mortality in a randomized European study. N. Engl. J. Med. 2009, 360, 1320–1328. [Google Scholar] [CrossRef] [PubMed]
- Frånlund, M.; Månsson, M.; Godtman, R.A.; Aus, G.; Holmberg, E.; Kollberg, K.S.; Lodding, P.; Pihl, C.G.; Stranne, J.; Lilja, H.; et al. Results from 22 years of Followup in the Göteborg Randomized Population-Based Prostate Cancer Screening Trial. J. Urol. 2022, 208, 292–300. [Google Scholar] [CrossRef] [PubMed]
- Kasivisvanathan, V.; Rannikko, A.S.; Borghi, M.; Panebianco, V.; Mynderse, L.A.; Vaarala, M.H.; Briganti, A.; Budäus, L.; Hellawell, G.; Hindley, R.G.; et al. MRI-Targeted or Standard Biopsy for Prostate-Cancer Diagnosis. N. Engl. J. Med. 2018, 378, 1767–1777. [Google Scholar] [CrossRef] [PubMed]
- Turkbey, B.; Rosenkrantz, A.B.; Haider, M.A.; Padhani, A.R.; Villeirs, G.; Macura, K.J.; Tempany, C.M.; Choyke, P.L.; Cornud, F.; Margolis, D.J.; et al. Prostate Imaging Reporting and Data System Version 2.1: 2019 Update of Prostate Imaging Reporting and Data System Version 2. Eur. Urol. 2019, 76, 340–351. [Google Scholar] [CrossRef]
- Westphalen, A.C.; McCulloch, C.E.; Anaokar, J.M.; Arora, S.; Barashi, N.S.; Barentsz, J.O.; Bathala, T.K.; Bittencourt, L.K.; Booker, M.T.; Braxton, V.G.; et al. Variability of the Positive Predictive Value of PI-RADS for Prostate MRI across 26 Centers: Experience of the Society of Abdominal Radiology Prostate Cancer Disease-focused Panel. Radiology 2020, 296, 76–84. [Google Scholar] [CrossRef]
- Girometti, R.; Peruzzi, V.; Polizzi, P.; De Martino, M.; Cereser, L.; Casarotto, L.; Pizzolitto, S.; Isola, M.; Crestani, A. Giannarini, G. et al. Case-by-case combination of the prostate imaging reporting and data system version 2.1 with the Likert score to reduce the false-positives of prostate MRI: a proof-of-concept study. Abdom. Radiol. 2024, 49, 4273–4285. [Google Scholar] [CrossRef]
- Morote, J.; Borque-Fernando, A.; Triquell, M.; Celma, A.; Regis, L.; Mast, R.; de Torres, I.M.; Semidey, M.E.; Abascal, J.M.; Servian, P.; et al. Comparative Analysis of PSA Density and an MRI-Based Predictive Model to Improve the Selection of Candidates for Prostate Biopsy. Cancers (Basel) 2022, 11, 2374. [Google Scholar] [CrossRef]
- Haj-Mirzaian, A.; Burk, K.S.; Lacson, R.; Glazer, D.I.; Saini, S.; Kibel, A.S.; Khorasani, R. Magnetic Resonance Imaging, Clinical, and Biopsy Findings in Suspected Prostate Cancer: A Systematic Review and Meta-Analysis. JAMA Netw Open 2024, 7, e244258. [Google Scholar] [CrossRef]
- Eyrich, N.W.; Morgan, T.M.; Tosoian, J.J. Biomarkers for detection of clinically significant prostate cancer: contemporary clinical data and future directions. Translational Andrology and Urology 2021, 10, 3091–3103. [Google Scholar] [CrossRef]
- Li, K.; Wang, Q.; Tang, X.; Akakuru, O.U.; Li, R.; Wang, Y.; Zhang, R.; Jiang, Z.; Yang, Z. Advances in Prostate Cancer Biomarkers and Probes. Cyborg Bionic Syst. 2024, 5, 0129. [Google Scholar]
- Morote, J.; Borque-Fernando, A.; Triquell, M.; Celma, A.; Regis, L.; Escobar, M.; Mast, R.; de Torres, I.M.; Semidey, M.E.; Abascal, J.M.; et al. The Barcelona Predictive Model of Clinically Significant Prostate Cancer. Cancers (Basel) 2022, 14, 1589. [Google Scholar] [PubMed]
- Davis, S.E.; Greevy, R.A.; Lasko, T.A.; Walsh, C.G.; Matheny, M.E. Comparison of Prediction Model Performance Updating Protocols: Using a Data-Driven Testing Procedure to Guide Updating. AMIA Ann. Symp. Proc. 2020, 2019, 1002–1010. [Google Scholar]
- Aladwani, M.; Lophatananon, A.; Ollier, W.; Muir, K. Prediction models for prostate cancer to be used in the primary care setting: a systematic review. BMJ Open 2020, 10, e034661. [Google Scholar] [CrossRef]
- Riaz, I.B.; Harmon, S.; Chen, Z.; Naqvi, S.A.A.; Cheng, L. Applications of Artificial Intelligence in Prostate Cancer Care: A Path to Enhanced Efficiency and Outcomes. American Society of Clinical Oncology Educational Book. Ed. Wolters Kluwer 2024, 44, 1548–8748. [Google Scholar]
- Nandi, A.; Xhafa, F. A federated learning method for real-time emotion state classification from multi-modal streaming. Methods 2022, 204, 340–347. [Google Scholar]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning. MIT Press 2016. http://www.deeplearningbook.org (accessed on 23 December 2024). (accessed on 23 December 2024).
- Jui, J.J.; Molla, M.M.I.; Alam, M.K.; Ferdowsi, A. Prostate Cancer Prediction Using Feedforward Neural Network Trained with Particle Swarm Optimizer. In: M.D. Zain, Z.; Sulaiman, M.H.; Mohamed, A.I.; Bakar, M.S.; Ramli, M.S. (eds) Proceedings of the 6th International Conference on Electrical, Control and Computer Engineering. Lecture Notes in Electrical Engineering, 2022; 842. Springer, Singapore. [CrossRef]
- Matsukawa, A.; Yanagisawa, T.; Bekku, K.; Kardoust Parizi, M.; Laukhtina, E.; Klemm, J.; Chiujdea, S.; Mori, K.; Kimura, S.; Fazekas, T.; et al. Comparing the Performance of Digital Rectal Examination and Prostate-specific Antigen as a Screening Test for Prostate Cancer: A Systematic Review and Meta-analysis. Eur. Urol. Oncol. 2024, 7, 697–704. [Google Scholar]
- Barentsz, J.O.; Richenberg, J.; Clements, R.; Choyke, P.; Verma, S.; Villeirs, G.; Rouviere, O.; Logager, V.; Futterer, J.J. ESUR prostate MR guidelines 2012. Eur. Radiol. 2012, 22, 746–757. [Google Scholar]
- Weinreb, J.C.; Barentsz, J.O.; Choyke, P.L.; Cornud, F.; Haider, M.A.; Macura, K.J.; Margolis, D.; Schnall, M.D.; Shtern, F.; Tempany, C.M.; et al. PI-RADS Prostate Imaging – Reporting and Data System: 2015, Version 2. Eur. Urol. 2016, 69, 16–40. [Google Scholar]
- Khoo, C.C.; Eldred-Evans, D.; Peters, M.; van Son, M.; van Rossum, P.S.N.; Connor, M.J.; Hosking-Jervis, F.; Tanaka, M.B.; Reddy, D.; Bass, E.; et al. A Comparison of Prostate Cancer Detection between Visual Estimation (Cognitive Registration) and Image Fusion (Software Registration) Targeted Transperineal Prostate Biopsy. J Urol. 2021, 205, 1075–1081. [Google Scholar]
- Wu, Q.; Tu, X.; Zhang, C.; Ye, J.; Lin, T.; Liu, Z.; Yang, L.; Qiu, S.; Bao, Y.; Wei, Q. Transperineal magnetic resonance imaging targeted biopsy versus transrectal route in the detection of prostate cancer: a systematic review and meta-analysis. Prostate Cancer Prostatic Dis. 2024, 27, 212–221. [Google Scholar] [CrossRef] [PubMed]
- Epstein, J.I.; Egevad, L.; Amin, M.B.; Delahunt, B.; Srigley, J.R.; Humphrey, P.A.; Grading, C. The 2014 International Society of Urological Pathology (ISUP) Consensus Conference on Gleason Grading of Prostatic Carcinoma: Definition of Grading Patterns and Proposal for a New Grading System. Am. J. Surg. Pathol. 2016, 40, 244–252. [Google Scholar] [CrossRef]
- Moore, C.M.; Kasivisvanathan, V.; Eggener, S.; Emberton, M.; Fütterer, J.J.; Gill, I.S.; Grubb Iii, R.L.; Hadaschik, B.; Klotz, L.; Margolis, D.J.; et al. Standards of reporting for MRI-targeted biopsy studies (START) of the prostate: recommendations from an International Working Group. Eur. Urol. 2013, 64, 544–552. [Google Scholar] [CrossRef]
- Anjum, M.; Khan, K.; Ahmad, W.; Ahmad, A.; Amin, M. N.; Nafees, A.; et al. New SHapley Additive ExPlanations (SHAP) approach to evaluate the raw materials interactions of steel-fiber-reinforced concrete. Materials (Basel) 2022, 15, 6261. [Google Scholar] [CrossRef]
- Stephan, C.; Xu, C.; Finne, P.; Cammann, H.; Meyer, H.A.; Lein, M.; Jung, K.; Stenman, U.H. Comparison of two different artificial neural networks for prostate biopsy indication in two different patient populations. Urology 2007, 70, 596–601. [Google Scholar] [CrossRef] [PubMed]
- Jansen, F.H.; van Schaik, R.H.; Kurstjens, J.; Horninger, W.; Klocker, H.; Bektic, J.; Wildhagen, M.F.; Roobol, M.J.; Bangma, C.H.; Bartsch, G. Prostate-specific antigen (PSA) isoform p2PSA in combination with total PSA and free PSA improves diagnostic accuracy in prostate cancer detection. Eur. Urol. 2010, 57, 921–927. [Google Scholar]
- Ecke, T.H.; Bartel, P.; Hallmann, S.; Koch, S.; Ruttloff, J.; Cammann, H.; Lein, M.; Schrader, M.; Miller, K.; Stephan, C. Outcome prediction for prostate cancer detection rate with artificial neural network (ANN) in daily routine. Urol. Oncol. 2012, 30, 139–144. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, T.; Hattori-Kato, M.; Okuno, Y.; Iwai, S.; Mikami, K. Prediction of prostate cancer by deep learning with multilayer artificial neural network. Can. Urol. Assoc. J. 2019, 13, E145–E150. [Google Scholar] [CrossRef]
- Checcucci, E.; Rosati, S.; De Cillis, S.; Giordano, N.; Volpi, G.; Granato, S.; Zamengo, D.; Verri, P.; Amparore, D.; De Luca, S.; et al. Machine-Learning-Based Tool to Predict Target Prostate Biopsy Outcomes: An Internal Validation Study. J. Clin. Med. 2023, 17, 4358. [Google Scholar] [CrossRef]
- Michaely, H.J.; Aringhieri, G.; Cioni, D.; Neri, E. Current Value of Biparametric Prostate MRI with Machine-Learning or Deep-Learning in the Detection, Grading, and Characterization of Prostate Cancer: A Systematic Review. Diagnostics (Basel). 2022, 12, 799. [Google Scholar] [CrossRef]
- Huynh, L.M.; Hwang, Y.; Taylor, O.; Baine, M.J. The Use of MRI-Derived Radiomic Models in Prostate Cancer Risk Stratification: A Critical Review of Contemporary Literature. Diagnostics (Basel). 2023, 13, 1128. [Google Scholar]
- Antolin, A.; Roson, N.; Mast, R.; Arce, J.; Almodovar, R.; Cortada, R.; Maceda, A.; Escobar, M.; Trilla, E.; Morote, J. The Role of Radiomics in the Prediction of Clinically Significant Prostate Cancer in the PI-RADS v2 and v2.1 Era: A Systematic Review. Cancers (Basel). 2024, 16, 2951. [Google Scholar] [PubMed]
- Boesen, L.; Thomsen, F.B.; Nørgaard, N.; Løgager, V.; Balslev, I.; Bisbjerg, R.; Thomsen, H.S.; Jakobsen, H. A predictive model based on biparametric magnetic resonance imaging and clinical parameters for improved risk assessment and selection of biopsy-naïve men for prostate biopsies. Prostate Cancer Prostatic Diseases 2019, 22, 609–616. [Google Scholar] [PubMed]
- Thompson, I.M.; Ankerst, D.P.; Chi, C.; Goodman, P.J.; Tangen, C.M.; Lucia, M.S.; Feng, Z.; Parnes, H.L.; Coltman, C.A. Jr. Assessing prostate cancer risk: results from the Prostate Cancer Prevention Trial. J. Natl. Cancer Inst. 2006, 98, 529–34. [Google Scholar] [PubMed]
- Morote, J.; Borque-Fernando, Á.; Triquell, M.; Campistol, M.; Servian, P.; Abascal, J.M.; Planas, J.; Méndez, O.; Esteban, L.M.; Trilla, E. Comparison of Rotterdam and Barcelona Magnetic Resonance Imaging Risk Calculators for Predicting Clinically Significant Prostate Cancer. Eur. Urol. Open Sci. 2023, 53, 46–54. [Google Scholar]
- Ogbonnaya, C.N.; Alsaedi, B.S.O.; Alhussaini, A.J.; Hislop, R.; Pratt, N.; Steele, J.D.; Kernohan, N.; Nabi, G. Radiogenomics Map-Based Molecular and Imaging Phenotypical Characterization in Localized Prostate Cancer Using Pre-Biopsy Biparametric MR Imaging. Int. J. Mol. Sci. 2024, 25, 5379. [Google Scholar]
- Zheng, B.; Mo, F.; Shi, X.; Li, W.; Shen, Q.; Zhang, L.; Liao, Z.; Fan, C.; Liu, Y.; Zhong, J.; et al. An Automatic Deep-Radiomics Framework for Prostate Cancer Diagnosis and Stratification in Patients with Serum Prostate-Specific Antigen of 4.0-10.0 ng/mL: A Multicenter Retrospective Study. Acad Radiol. 2025; in press. [Google Scholar] [CrossRef]
| Characteristic | sPCa | nsPCa | Odds Ratio (95%CI) | p Value |
|---|---|---|---|---|
| Number of men (%) | 2,097 (41.9) | 2,908 (58.1) | - | - |
| Mean age, years (SD) | 70 (8.2) | 66 (7.6) | 1.07 (1.06-1.08) | <0.001 |
| Mean serum PSA, ng/mL (SD) | 20 (109) | 8.4 (9.6) | 1.04 (1.03-1.05) | <0.001 |
| PCa family history, n (%) | ||||
| No | 1,930 (92%) | 2,723 (93.6%) | - | Ref. |
| Yes | 167 (8%) | 185 (6.4%) | 1.27 (1.02-1.58) | 0.033 |
| Type of prostate biopsy, n (%) | ||||
| Initial | 1594 (76%) | 1906 (65.5%) | - | Ref. |
| Repeated | 503 (24%) | 1002 (34.5%) | 0.6 (0.53-0.68) | <0.001 |
| DRE, n (%) | ||||
| Normal | 1161 (55.4%) | 2417 (83.1%) | - | Ref. |
| Suspicious | 936 (44.6%) | 491 (16.9%) | 3.97 (3.49-4.52) | <0.001 |
| Prostate volume (mL) | 51.9 (27.6) | 69.1 (34.4) | 0.98 (0.98-0.98) | <0.001 |
| PI-RADS score, n (%) | ||||
| 1 | 60 (2.9%) | 514 (17.7%) | - | Ref. |
| 2 | 23 (1.1%) | 152 (5.2%) | 1.3 (0.76-2.14) | 0.322 |
| 3 | 206 (9.8%) | 1044 (35.9%) | 1.69 (1.25-2.31) | 0.001 |
| 4 | 991 (47.3%) | 1024 (35.2%) | 8.29 (6.31-11.08) | <0.001 |
| 5 | 817 (39%) | 174 (6%) | 40.22 (29.61-55.48) | <0.001 |
| CI: confidence interval; SD: standard deviation; PCa: prostate cancer; sPCa: significant PCa; nsPCa: non-significant PCa; PSA: prostate-specific antigen; DRE: digital rectal examination; PI-RADS: Prostate Imaging-Reporting and Data System. | ||||
| Metric | Training set (n = 4,254) | Validation set (n = 631) | Test set (n = 751) | |||
|---|---|---|---|---|---|---|
| GMV model | BCN model | GMV model | BCN model | GMV model | BCN model | |
| AUC (95% CI) |
0.88 (0.87, 0.90) |
0.85 (0.84, 0.86) |
0.88 (0.86, 0.91) |
0.86 (0.85, 0.87) |
0.85 (0.83, 0.88) |
0.84 (0.82, 0.86) |
| Precision | 0.7171 | 0.7435 | 0.7184 | 0.7426 | 0.7126 | 0.7607 |
| Recall | 0.8266 | 0.7435 | 0.8284 | 0.7537 | 0.7556 | 0.6762 |
| Specificity | 0.765 | 0.8151 | 0.7655 | 0.8113 | 0.7798 | 0.8463 |
| Accuracy | 0.7908 | 0.7851 | 0.7919 | 0.7872 | 0.7696 | 0.7750 |
| F1 score | 0.768 | 0.7435 | 0.7695 | 0.7481 | 0.7334 | 0.7160 |
| Kappa score | 0.5792 | 0.5587 | 0.5815 | 0.5639 | 0.5309 | 0.5307 |
| MCC | 0.5841 | 0.5587 | 0.5864 | 0.5639 | 0.5316 | 0.5332 |
| AUC: area under the curve; MCC: Matthew´s correlation coefficient. | ||||||
| Threshold (%) |
GMVmodel | BCN model | |||
|---|---|---|---|---|---|
| Saved biopsies (%) | Undetected sPCa (%) | Saved biopsies (%) | Undetected sPCa (%) | ||
| 5 | 3.5 | 0.5 | 18 | 2 | |
| 6 | 5 | 0.75 | 20 | 2.5 | |
| 7 | 9.5 | 1 | 23 | 3.5 | |
| 8 | 10 | 2 | 26 | 4.75 | |
| 9 | 14 | 2 | 27 | 5 | |
| 10 | 17 | 2 | 27.5 | 5 | |
| 11 | 19 | 2.5 | 30 | 6 | |
| 12 | 20 | 2.5 | 32.5 | 6.5 | |
| 13 | 22 | 2.6 | 33.5 | 6.5 | |
| 14 | 23 | 2.6 | 34 | 8 | |
| 15 | 26 | 4.5 | 35 | 8.5 | |
| 16 | 27.5 | 5 | 36 | 10.5 | |
| 17 | 29 | 5 | 37.5 | 10.5 | |
| 18 | 30 | 5.1 | 39.5 | 13 | |
| 19 | 30 | 5.1 | 40 | 13 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




