Submitted:
15 February 2025
Posted:
17 February 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
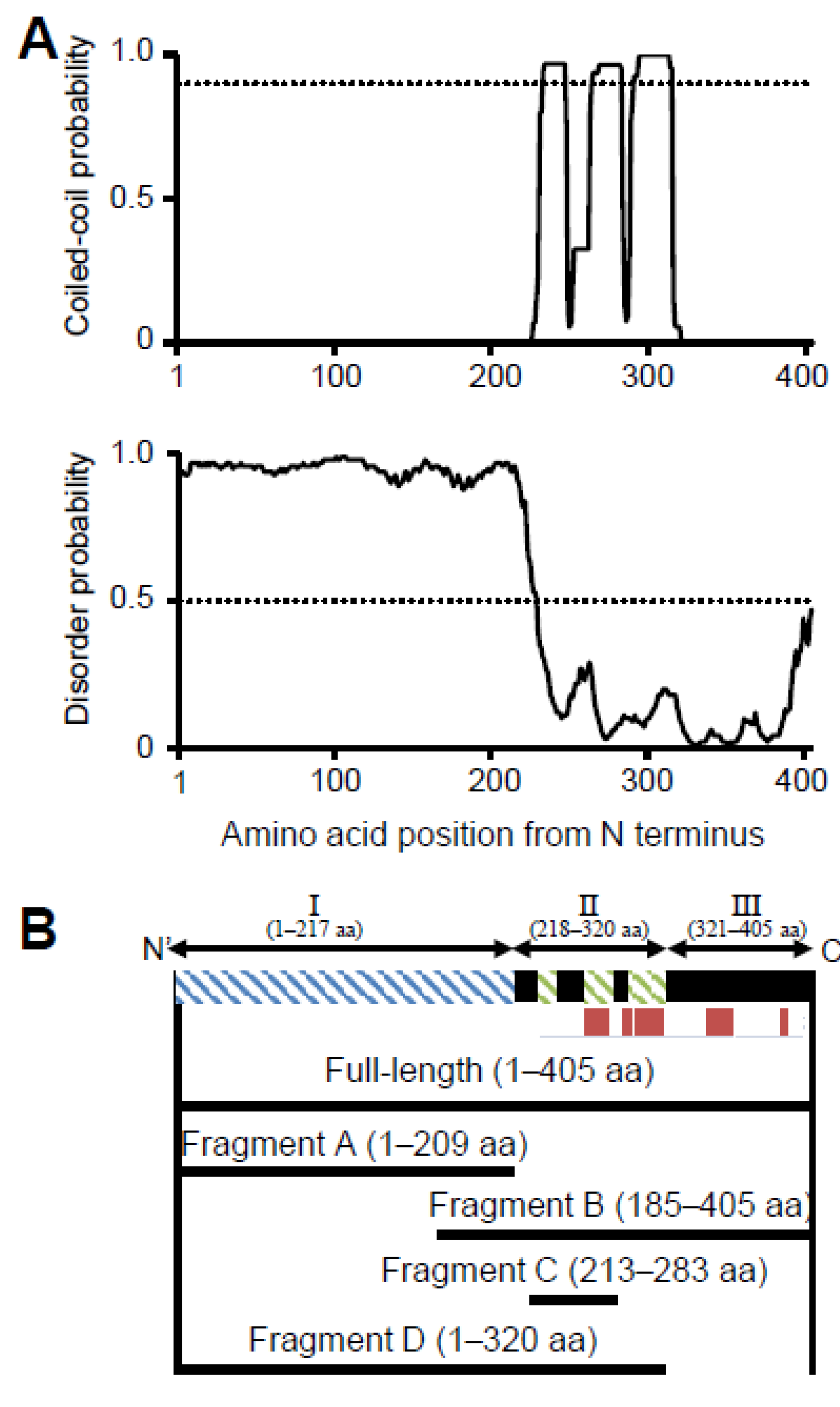
2.1. Sequence Analyses
2.2. P65 Protein Expression and Isolation
2.3. Preparation of P65 Protein Fragments
2.4. Size Exclusion Chromatography and Analytical Centrifugation
2.5. Far-Ultraviolet (UV) Circular Dichroism (CD) Spectroscopy
2.6. Small-Angle X-Ray Scattering (SAXS) Measurements
2.7. SAXS Data Analysis
2.8. Negative Staining EM
3. Results
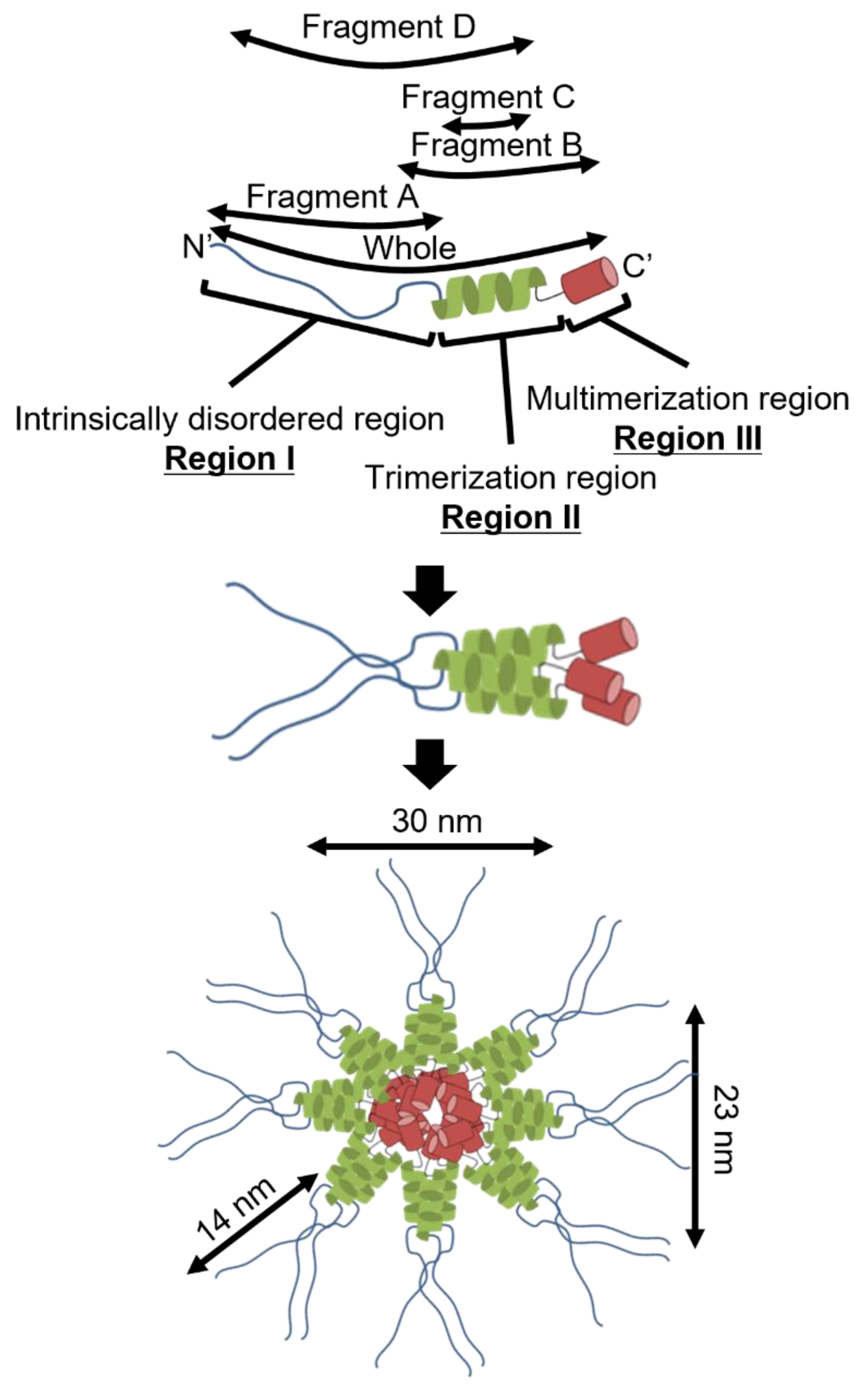
3.1. P65 Can Be Divided into Three Regions Based on Amino Acid Sequence
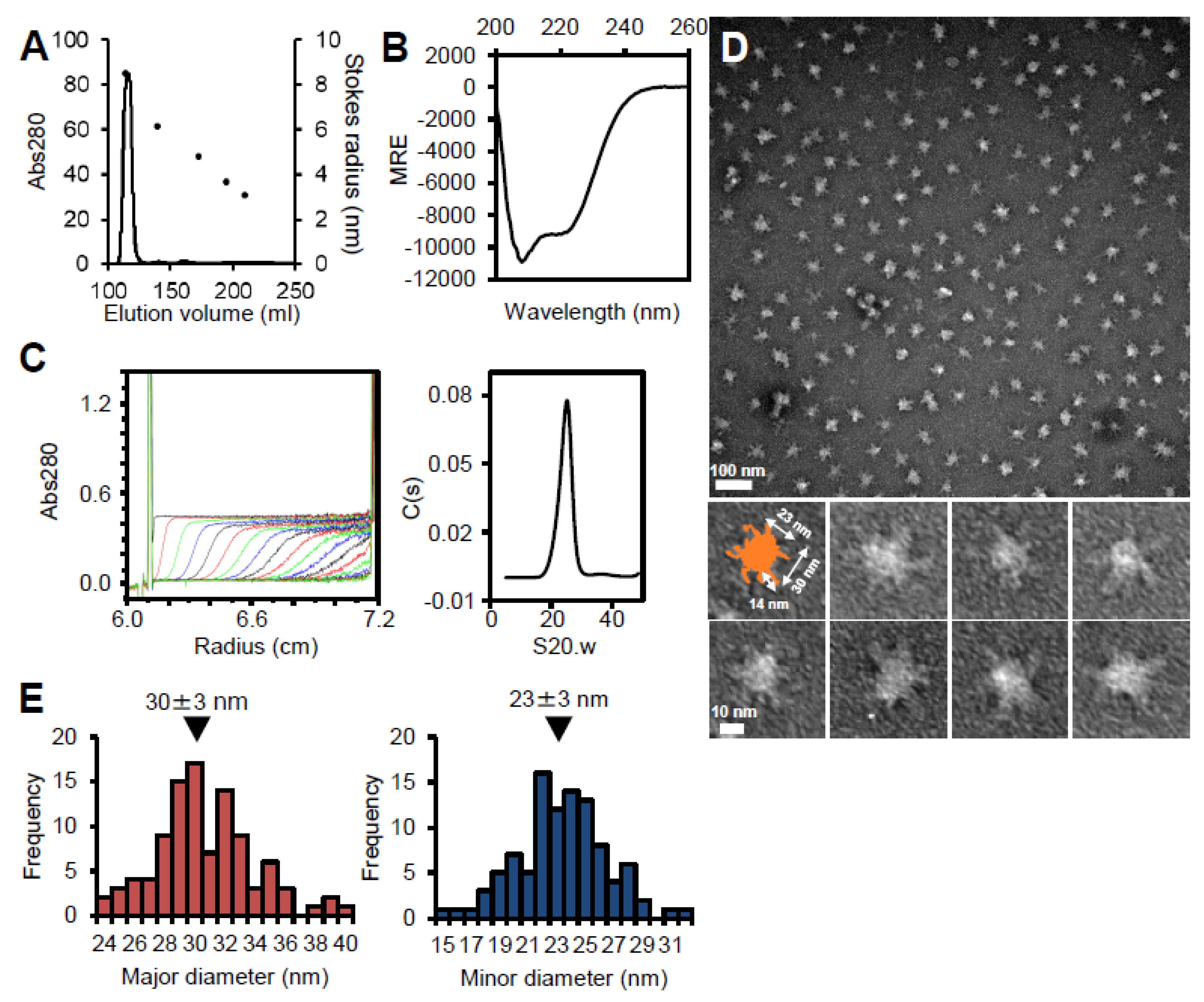
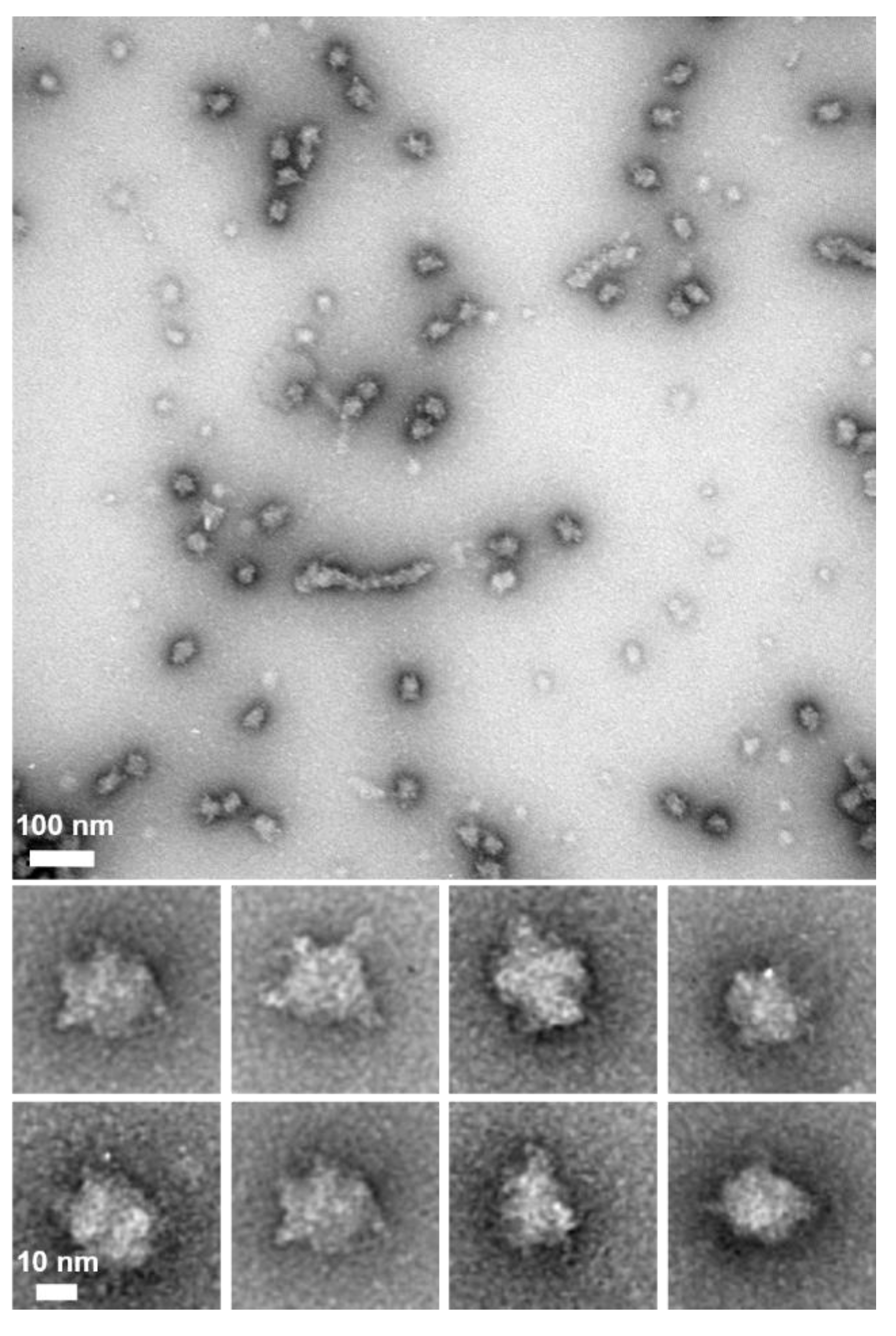
3.2. P65 Forms a Large Complex
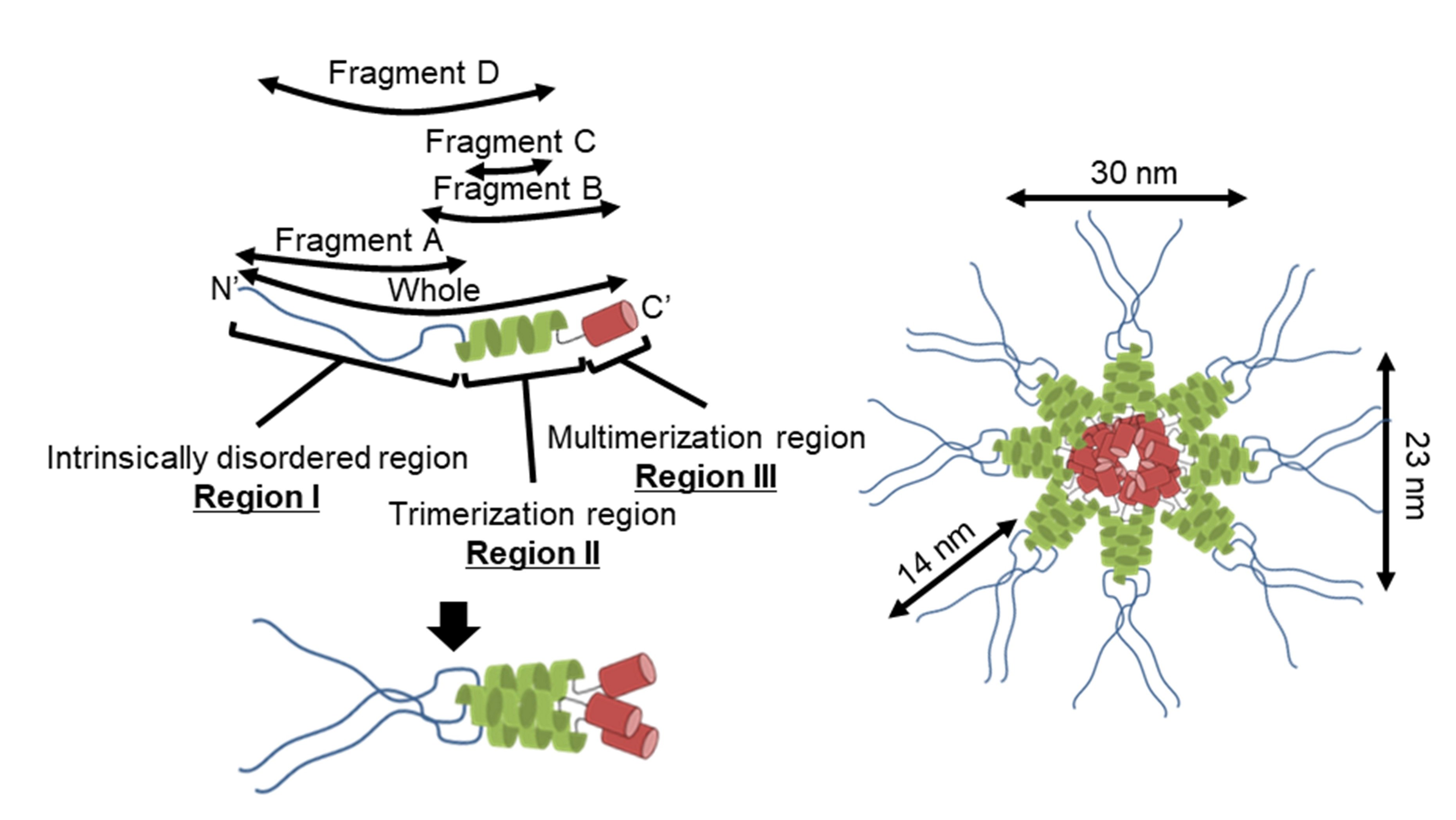
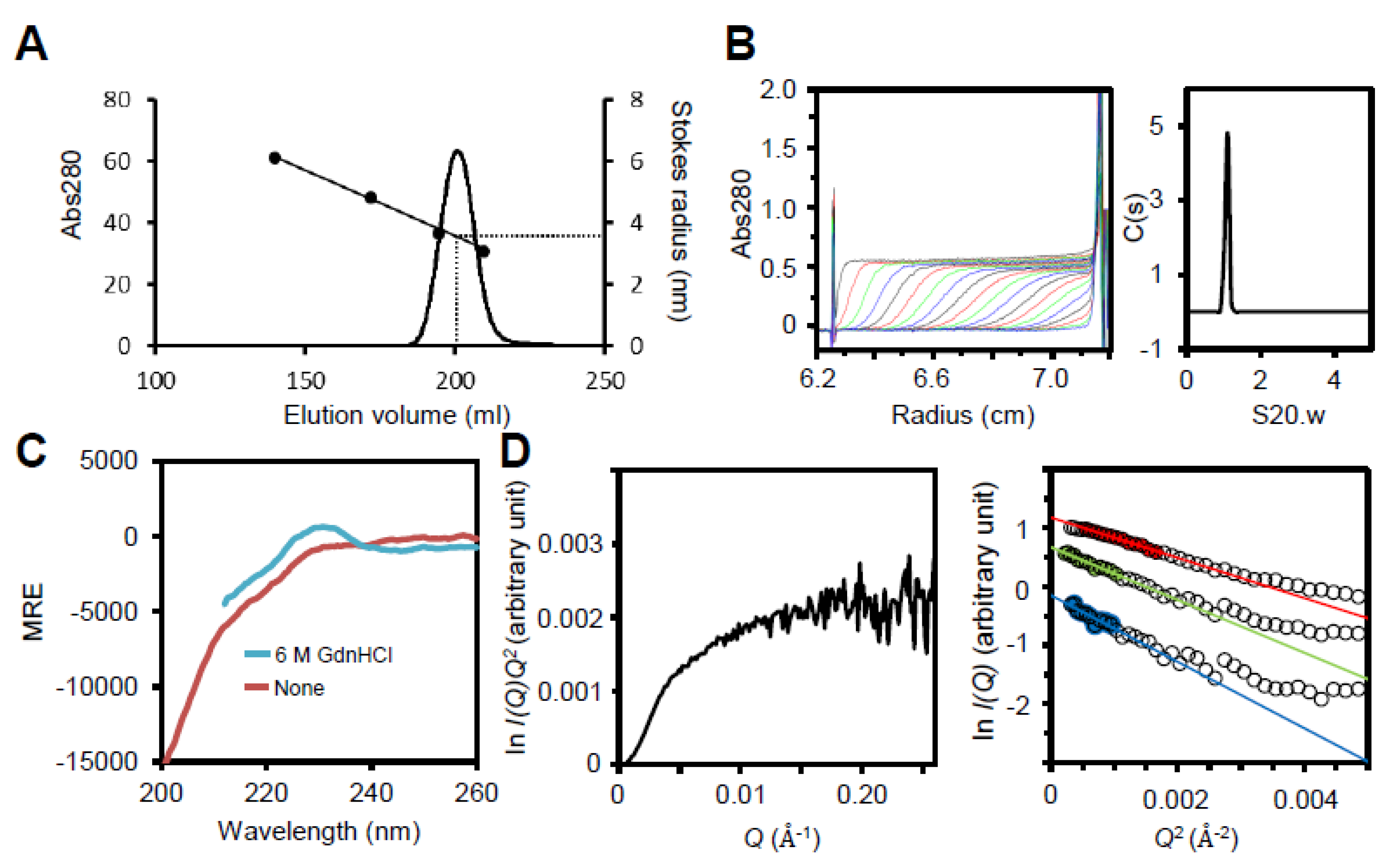
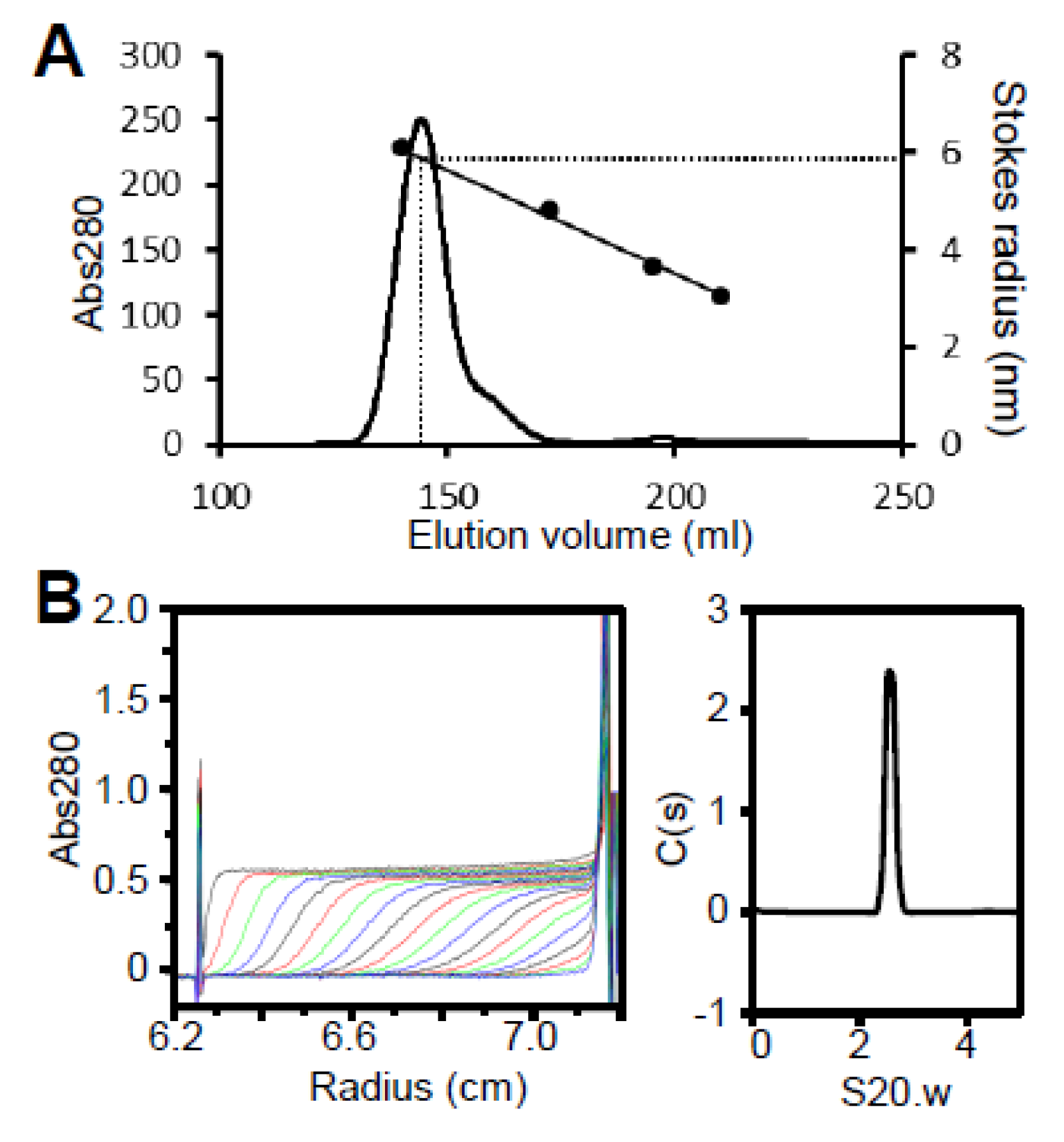
3.3. Region I (Mostly Corresponding to Fragment A) Is Intrinsically Disordered
3.4. Region I (Fragment A) Forms the Flexible Spines and Regions II–III (Fragment B) form Core of the P65 Star-like Particle
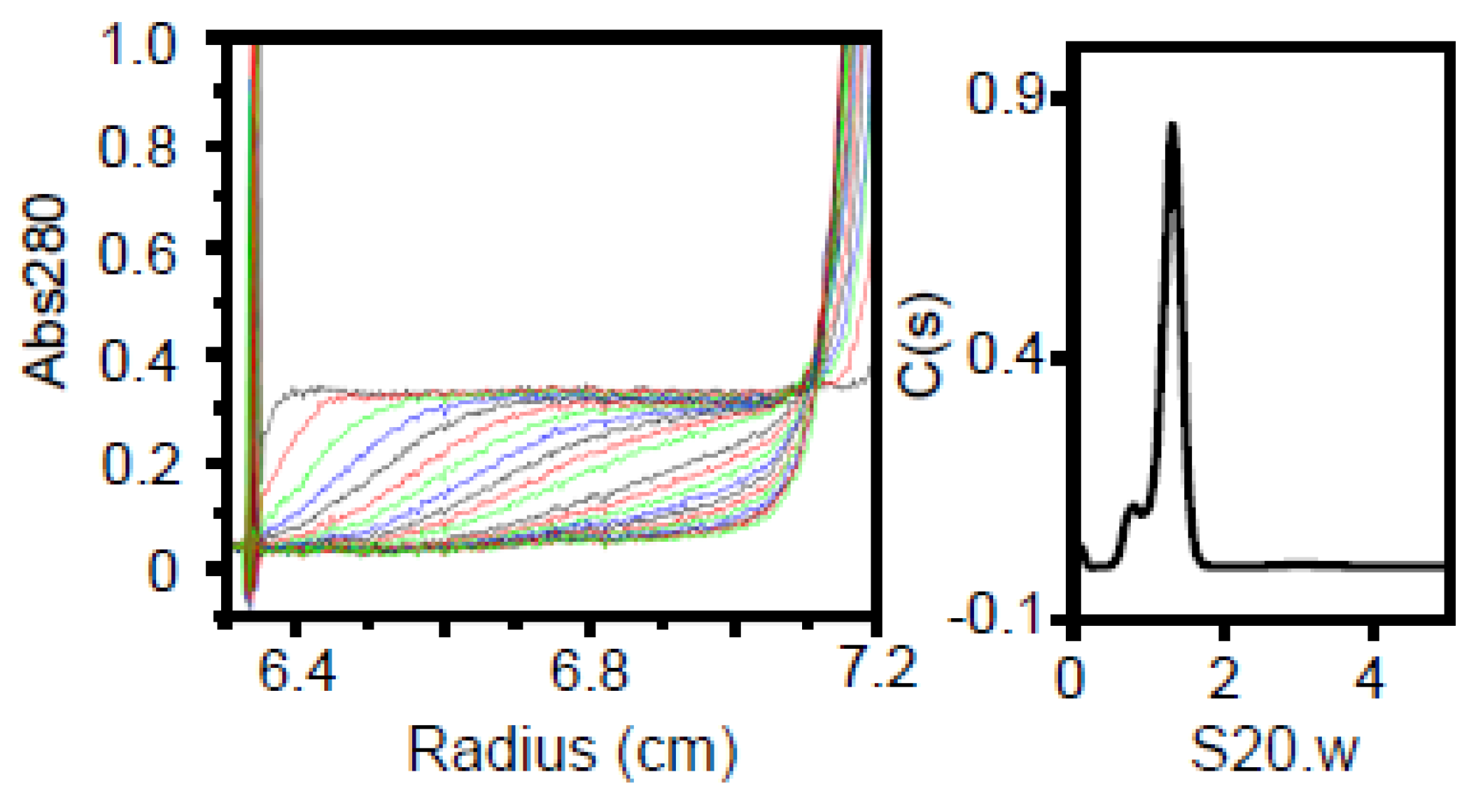
3.5. Region II (Fragment C) Forms a Trimer
3.6. Regions I–II (Fragment D) Forms a Trimer
4. Discussion
4.1. Implication for the Star-like Complex Formation
4.2. Assignment of the Star-like Complex in the Gliding Machinery
Supplementary Materials
Funding
Acknowledgments
References
- Gupta, R.S.; Oren, A. Necessity and rationale for the proposed name changes in the classification of Mollicutes species. Reply to: 'Recommended rejection of the names Malacoplasma gen. nov., Mesomycoplasma gen. nov., Metamycoplasma gen. nov., Metamycoplasmataceae fam. nov., Mycoplasmoidaceae fam. nov., Mycoplasmoidales ord. nov., Mycoplasmoides gen. nov., Mycoplasmopsis gen. nov. [Gupta, Sawnani, Adeolu, Alnajar and Oren 2018] and all proposed species comb. nov. placed therein', by M. Balish et al. (Int J Syst Evol Microbiol, 2019;69:3650-3653). Int J Syst Evol Microbiol 2020, 70, 1431–1438. [Google Scholar] [CrossRef] [PubMed]
- Razin, S. , Yogev, D., AND Naot, Y. Molecular biology and pathogenicity of mycoplasmas. Microbiol. Mol. Biol. Rev. 1998, 62, 1094–1156. [Google Scholar] [CrossRef] [PubMed]
- Razin, S.; Hayflick, L. Highlights of mycoplasma research--an historical perspective. Biologicals. 2010, 38, 183–190. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Huang, J.M.; Zhou, Y.L.; Almeida, A.; Finn, R.D.; Danchin, A.; He, L.S. Phylogenomics of expanding uncultured environmental Tenericutes provides insights into their pathogenicity and evolutionary relationship with Bacilli. BMC Genomics 2020, 21, 408. [Google Scholar] [CrossRef]
- Miyata, M.; Hamaguchi, T. Integrated Information and Prospects for Gliding Mechanism of the Pathogenic Bacterium Mycoplasma pneumoniae. Front. Microbiol. 2016, 7, 960. [Google Scholar] [CrossRef]
- Mizutani, M.; Sasajima, Y.; Miyata, M. Force and stepwise movements of gliding motility in human pathogenic bacterium Mycoplasma pneumoniae. Front Microbiol 2021, 12, 747905. [Google Scholar] [CrossRef]
- Nakane, D. , and Miyata, M. Cytoskeletal asymmetrical-dumbbell structure of a gliding mycoplasma, Mycoplasma gallisepticum, revealed by negative-staining electron microscopy. J. Bacteriol. 2009, 191, 3256–3264. [Google Scholar] [CrossRef]
- Nakane, D.; Murata, K.; Kenri, T.; Shibayama, K.; Nishizaka, T. Molecular ruler of the attachment organelle in Mycoplasma pneumoniae. PLoS Pathog 2021, 17, e1009621. [Google Scholar] [CrossRef]
- Radestock, U. , and Bredt, W. Motility of Mycoplasma pneumoniae. J Bacteriol. 1977, 129, 1495–1501. [Google Scholar] [CrossRef]
- Seto, S. , Layh-Schmitt, G., Kenri, T., and Miyata, M. Visualization of the attachment organelle and cytadherence proteins of Mycoplasma pneumoniae by immunofluorescence microscopy. J Bacteriol. 2001, 183, 1621–1630. [Google Scholar] [CrossRef]
- Prince, O.A. , Krunkosky, T.M., and Krause, D.C. In vitro spatial and temporal analysis of Mycoplasma pneumoniae colonization of human airway epithelium. Infect. Immun. 2014, 82, 579–586. [Google Scholar] [CrossRef] [PubMed]
- Krunkosky, T.M. , Jordan, J.L., Chambers, E., and Krause, D.C. Mycoplasma pneumoniae host-pathogen studies in an air-liquid culture of differentiated human airway epithelial cells. Microb. Pathog. 2007, 42, 98–103. [Google Scholar] [CrossRef]
- Miyata, M.; Robinson, R.C.; Uyeda, T.Q.P.; Fukumori, Y.; Fukushima, S.I.; Haruta, S.; Homma, M.; Inaba, K.; Ito, M.; Kaito, C.; et al. Tree of motility - A proposed history of motility systems in the tree of life. Genes Cells 2020, 25, 6–21. [Google Scholar] [CrossRef] [PubMed]
- Miyata, M.; Hamaguchi, T. Prospects for the gliding mechanism of Mycoplasma mobile. Curr. Opin. Microbiol. 2016, 29, 15–21. [Google Scholar] [CrossRef]
- Miyata, M. Unique centipede mechanism of Mycoplasma gliding. Annu. Rev. Microbiol. 2010, 64, 519–537. [Google Scholar] [CrossRef]
- Miyata, M. Centipede and inchworm models to explain Mycoplasma gliding. Trends Microbiol. 2008, 16, 6–12. [Google Scholar] [CrossRef]
- Seybert, A.; Gonzalez-Gonzalez, L.; Scheffer, M.P.; Lluch-Senar, M.; Mariscal, A.M.; Querol, E.; Matthaeus, F.; Pinol, J.; Frangakis, A.S. Cryo-electron tomography analyses of terminal organelle mutants suggest the motility mechanism of Mycoplasma genitalium. Mol Microbiol 2018, 108, 319–329. [Google Scholar] [CrossRef]
- Krause, D.C.; Chen, S.; Shi, J.; Jensen, A.J.; Sheppard, E.S.; Jensen, G.J. Electron cryotomography of Mycoplasma pneumoniae mutants correlates terminal organelle architectural features and function. Mol Microbiol 2018, 108, 306–318. [Google Scholar] [CrossRef]
- Kawamoto, A.; Matsuo, L.; Kato, T.; Yamamoto, H.; Namba, K.; Miyata, M. Periodicity in attachment organelle revealed by electron cryotomography suggests conformational changes in gliding mechanism of Mycoplasma pneumoniae. MBio 2016, 7, e00243–00216. [Google Scholar] [CrossRef]
- Nakane, D.; Kenri, T.; Matsuo, L.; Miyata, M. Systematic structural analyses of attachment organelle in Mycoplasma pneumoniae. PLoS Pathog 2015, 11, e1005299. [Google Scholar] [CrossRef]
- Vizarraga, D.; Kawamoto, A.; Matsumoto, U.; Illanes, R.; Perez-Luque, R.; Martin, J.; Mazzolini, R.; Bierge, P.; Pich, O.Q.; Espasa, M.; et al. Immunodominant proteins P1 and P40/P90 from human pathogen Mycoplasma pneumoniae. Nat Commun 2020, 11, 5188. [Google Scholar] [CrossRef] [PubMed]
- Aparicio, D.; Scheffer, M.P.; Marcos-Silva, M.; Vizarraga, D.; Sprankel, L.; Ratera, M.; Weber, M.S.; Seybert, A.; Torres-Puig, S.; Gonzalez-Gonzalez, L.; et al. Structure and mechanism of the Nap adhesion complex from the human pathogen Mycoplasma genitalium. Nat Commun 2020, 11, 2877. [Google Scholar] [CrossRef] [PubMed]
- Kenri, T.; Kawakita, Y.; Kudo, H.; Matsumoto, U.; Mori, S.; Furukawa, Y.; Tahara, Y.O.; Shibayama, K.; Hayashi, Y.; Arai, M.; et al. Production and characterization of recombinant P1 adhesin essential for adhesion, gliding, and antigenic variation in the human pathogenic bacterium, Mycoplasma pneumoniae. Biochem Biophys Res Commun 2019, 508, 1050–1055. [Google Scholar] [CrossRef] [PubMed]
- Aparicio, D.; Torres-Puig, S.; Ratera, M.; Querol, E.; Pinol, J.; Pich, O.Q.; Fita, I. Mycoplasma genitalium adhesin P110 binds sialic-acid human receptors. Nat Commun 2018, 9, 4471. [Google Scholar] [CrossRef]
- Nakane, D.; Adan-Kubo, J.; Kenri, T.; Miyata, M. Isolation and characterization of P1 adhesin, a leg protein of the gliding bacterium Mycoplasma pneumoniae. J. Bacteriol. 2011, 193, 715–722. [Google Scholar] [CrossRef]
- Sprankel, L.; Scheffer, M.P.; Manger, S.; Ermel, U.H.; Frangakis, A.S. Cryo-electron tomography reveals the binding and release states of the major adhesion complex from Mycoplasma genitalium. PLoS Pathog 2023, 19, e1011761. [Google Scholar] [CrossRef]
- Kawakita, Y.; Kinoshita, M.; Furukawa, Y.; Tulum, I.; Tahara, Y.O.; Katayama, E.; Namba, K.; Miyata, M. Structural study of MPN387, an essential protein for gliding motility of a human-pathogenic bacterium, Mycoplasma pneumoniae. J Bacteriol 2016, 198, 2352–2359. [Google Scholar] [CrossRef]
- Seto, S.; Kenri, T.; Tomiyama, T.; Miyata, M. Involvement of P1 adhesin in gliding motility of Mycoplasma pneumoniae as revealed by the inhibitory effects of antibody under optimized gliding conditions. J. Bacteriol. 2005, 187, 1875–1877. [Google Scholar] [CrossRef]
- Kasai, T.; Nakane, D.; Ishida, H.; Ando, H.; Kiso, M.; Miyata, M. Role of binding in Mycoplasma mobile and Mycoplasma pneumoniae gliding analyzed through inhibition by synthesized sialylated compounds. J Bacteriol 2013, 195, 429–435. [Google Scholar] [CrossRef]
- Proft, T. , Hilbert, H., Layh-Schmitt, G., and Herrmann, R. The proline-rich P65 protein of Mycoplasma pneumoniae is a component of the Triton X-100-insoluble fraction and exhibits size polymorphism in the strains M129 and FH. J Bacteriol. 1995, 177, 3370–3378. [Google Scholar] [CrossRef]
- Hasselbring, B.M. , Sheppard, E.S., and Krause, D.C. P65 truncation impacts P30 dynamics during Mycoplasma pneumoniae gliding. J Bacteriol. 2012, 194, 3000–3007. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.Y. , Jordan, J.L., and Krause, D.C. Domain analysis of protein P30 in Mycoplasma pneumoniae cytadherence and gliding motility. J Bacteriol. 2011, 193, 1726–1733. [Google Scholar] [CrossRef] [PubMed]
- Jordan, J.L.; Berry, K.M.; Balish, M.F.; Krause, D.C. Stability and subcellular localization of cytadherence-associated protein P65 in Mycoplasma pneumoniae. J Bacteriol 2001, 183, 7387–7391. [Google Scholar] [CrossRef]
- Lupas, A.; Van Dyke, M.; Stock, J. Predicting coiled coils from protein sequences. Science 1991, 252, 1162–1164. [Google Scholar] [CrossRef]
- Ward, J.J. , Sodhi, J.S. McGuffin, L.J., Buxton, B.F., and Jones, D.T. Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J. Mol. Biol. 2004, 337, 635–645. [Google Scholar] [CrossRef]
- Drozdetskiy, A.; Cole, C.; Procter, J.; Barton, G.J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res 2015, 43, W389–394. [Google Scholar] [CrossRef]
- Altschul, S.F. , Gish, W., Miller, W., Myers, E.W., and Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Notredame, C. , Higgins, D.G., and Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 2000, 302, 205–217. [Google Scholar] [CrossRef]
- Erickson, H.P. Size and shape of protein molecules at the nanometer level determined by sedimentation, gel filtration, and electron microscopy. Biol. Proced. Online. 2009, 11, 32–51. [Google Scholar] [CrossRef]
- le Maire, M. , Arnou, B., Olesen, C., Georgin, D., Ebel, C., and Moller, J.V. Gel chromatography and analytical ultracentrifugation to determine the extent of detergent binding and aggregation, and Stokes radius of membrane proteins using sarcoplasmic reticulum Ca2+-ATPase as an example. Nat. Protoc. 2008, 3, 1782–1795. [Google Scholar] [CrossRef]
- Adan-Kubo, J. , Yoshii, S.H., Kono, H., and Miyata, M. Molecular structure of isolated MvspI, a variable surface protein of the fish pathogen Mycoplasma mobile. J Bacteriol. 2012, 194, 3050–3057. [Google Scholar] [CrossRef] [PubMed]
- Schuck, P. Size distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and Lamm equation modeling. Biophys. J. 2000, 78, 1606–1619. [Google Scholar] [CrossRef] [PubMed]
- Louis-Jeune, C. , Andrade-Navarro, M.A., and Perez-Iratxeta, C. Prediction of protein secondary structure from circular dichroism using theoretically derived spectra. Proteins. 2012, 80, 374–381. [Google Scholar] [CrossRef] [PubMed]
- Hammersley, A. ESRF Internal report, ESRF97HA02T, “FIT2D: An introduction and overview”. 1997.
- Kataoka, M. , Nishii, I., Fujisawa, T., Ueki, T., Tokunaga, F., and Goto, Y. Structural characterization of the molten globule and native states of apomyoglobin by solution X-ray scattering. J. Mol. Biol. 1995, 249, 215–228. [Google Scholar] [CrossRef]
- Toyonaga, T.; Kato, T.; Kawamoto, A.; Kodera, N.; Hamaguchi, T.; Tahara, Y.O.; Ando, T.; Namba, K.; Miyata, M. Chained structure of dimeric F1-like ATPase in Mycoplasma mobile gliding machinery. mBio 2021, 12, e0141421. [Google Scholar] [CrossRef]
- Nonaka, T. , Adan-Kubo, J., and Miyata, M. Triskelion structure of the Gli521 protein, involved in the gliding mechanism of Mycoplasma mobile. J Bacteriol. 2010, 192, 636–642. [Google Scholar] [CrossRef]
- Crick, F. The packing of a-helices: simple coiled-coils. Acta Cryst. 1953, 6, 689–697. [Google Scholar] [CrossRef]
- Townend, R.K., T. F., Timasheff, S.N., Fasman, G.D., and Davidson, B. The circular dichroism of the beta structure of poly-L-lysine. Biochem. Biophys. Res. Commun. 1966, 23, 163–169. [Google Scholar] [CrossRef]
- Cantor, C.R., and Schimmel, P.R. Biophysical chemistry part II: Techniques for the study of biological structure and function, W.H.Freeman and Co., San Francisco; 1980.
- Arai, M. , Kondrashkina, E., Kayatekin, C., Matthews, C.R., Iwakura, M.;, a.B., O. Microsecond hydrophobic collapse in the folding of Escherichia coli dihydrofolate reductase, an alpha/beta-type protein. J. Mol. Biol. 2007, 368, 219–229. [Google Scholar] [CrossRef]
- Glatter, O., and Kratky, O. Small angle X ray scattering. Academic Press, London. 1982. [CrossRef]
- Kohn, J.E. , Millett, I.S., Jacob, J., Zagrovic, B., Dillon, T.M., Cingel, N., Dothager, R.S., Seifert, S., Thiyagarajan, P., Sosnick, T.R., Hasan, M.Z., Pande, V.S., Ruczinski, I., Doniach, S., and Plaxco, K.W. Random-coil behavior and the dimensions of chemically unfolded proteins. Proc. Natl. Acad. Sci. USA. 2005, 101, 12491–12496. [Google Scholar] [CrossRef]
- Mirdita, M.; Schutze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: making protein folding accessible to all. Nat Methods 2022, 19, 679–682. [Google Scholar] [CrossRef] [PubMed]
- Henderson, G.P. , and Jensen, G.J. Three-dimensional structure of Mycoplasma pneumoniae's attachment organelle and a model for its role in gliding motility. Mol. Microbiol. 2006, 60, 376–385. [Google Scholar] [CrossRef] [PubMed]
- Gobel, U. , Speth, V., Bredt, W. Filamentous structures in adherent Mycoplasma pneumoniae cells treated with nonionic detergents. J Cell Biol. 1981, 91, 537–543. [Google Scholar] [CrossRef] [PubMed]
- Meng, K.E. , and Pfister, R.M. Intracellular structures of Mycoplasma pneumoniae revealed after membrane removal. J Bacteriol. 1980, 144, 390–399. [Google Scholar] [CrossRef]
- Kenri, T. , Seto, S., Horino, A., Sasaki, Y., Sasaki, T., and Miyata, M. Use of fluorescent-protein tagging to determine the subcellular localization of Mycoplasma pneumoniae proteins encoded by the cytadherence regulatory locus. J Bacteriol. 2004, 186, 6944–6955. [Google Scholar] [CrossRef]
- Hasselbring, B.M. , Jordan, J.L., and Krause, D.C. Mutant analysis reveals a specific requirement for protein P30 in Mycoplasma pneumoniae gliding motility. J Bacteriol. 2005, 187, 6281–6289. [Google Scholar] [CrossRef]
- Garcia-Morales, L.; Gonzalez-Gonzalez, L.; Querol, E.; Pinol, J. A minimized motile machinery for Mycoplasma genitalium. Mol Microbiol 2016, 100, 125–138. [Google Scholar] [CrossRef]
- Fisseha, M. , Gohlmann, H.W., Herrmann, R., and Krause, D.C. Identification and complementation of frameshift mutations associated with loss of cytadherence in Mycoplasma pneumoniae. J Bacteriol. 1999, 181, 4404–4410. [Google Scholar] [CrossRef]
- Dyson, H.J. , and Wright, P.E. Intrinsically unstructured proteins and their functions. Nat. Rev. Mol. Cell Biol. 2005, 6, 197–208. [Google Scholar] [CrossRef]
| Fragment | Amino Acid Sequence | Gel Filtration |
Analytical Ultracentrifugation | |||||
|---|---|---|---|---|---|---|---|---|
| Mass (Da) | Stokes Radius as Spherical Monomer (nm) | Stokes Radius (nm) | S20.w | Stokes Radius (nm) | Dimension (nm) | Mass (kDa) | Assembly | |
| Whole | 49,203 | 2.9 | n/a | 22.2 | 32.9 | n/a | 3.10 × 103 | Multimer |
| A | 23,868 | 2.2 | 3.4 | 0.97 | 6.4 | 48.7 × 1.1 | 25.7 | Monomer |
| C | 8,631 | 1.5 | n/a | 1.30 | 5.2 | 3.8 × 2.4 | 25.6 | Trimer |
| D | 39,080 | 2.7 | 5.9 | 2.56 | 10.7 | 21.3 × 2.1 | 105 | Trimer |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




