Submitted:
26 May 2025
Posted:
27 May 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
- The nearest mean classifier (NMC) ([5]) which is a prototype-based classifier that assigns points according to the perpendicular bisector boundary between the centroids of two groups. This classifier has training time complexity.
- Fisher’s linear discriminant analysis (LDA, specifically refer to Fisher’s LDA in this study) is a variance-based classifier which can be trained in cubic time complexity . While faster implementations like spectral regression discriminant analysis (SRDA) ([6]) claim lower training time complexity, their efficiency depends on specific conditions, such as a sufficiently small iterative term and sparsity in the data. These constraints limit SRDA’s applicability in real-world classification tasks.
- Support vector machine (SVM) ([7]) with a linear kernel is a maximum-margin classifier, which has a training time complexity of . A fast implementation LIBLINEAR ([8]) (denoted as fast SVM) reduces to by using dual coordinate descent optimization. The iteration count k that depends on the optimization path leads to a quasi-quadratic overall complexity.
- Perceptron ([9]) is a misclassification-triggered ruled-based classifier. Its training time complexity is .
- Logistic regression (LR) ([10]) is a statistics-based classifier. It can be trained using either maximum likelihood estimation (MLE) or iteratively reweighted least squares, with time complexity of and respectively.
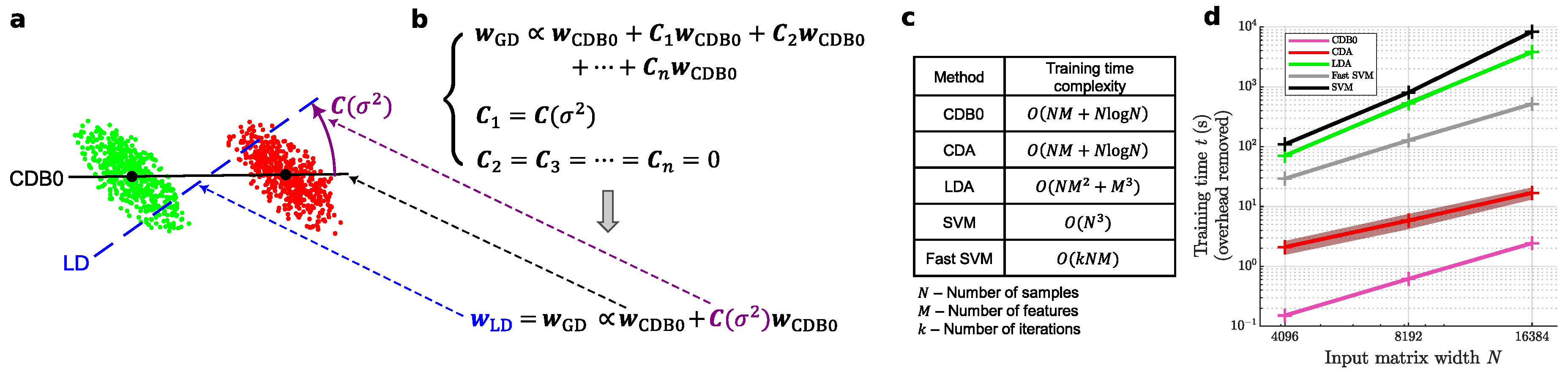
- A geometric theory framework for classifiers: This study introduces a geometric framework, geometric discriminant analysis (GDA), to unify certain linear classifiers under a common theoretical model. GDA leverages a special type of centroid discriminant basis (CDB0), a vector connecting the centroids of two classes, which serves as the foundation for constructing classifier decision boundaries. The GDA framework adjusts the CDB0 through geometric corrections under various constraints, enabling the derivation of classifiers with desirable properties. Notably, we show that: NMC is a special case of GDA, where geometric corrections are not applied to CDB0; linear discriminant analysis (LDA) is a special case of GDA, where the CDB0 is corrected by maximizing the projection variance ratio.
- A high-performance and scalable linear geometric classifier: Building on the GDA framework, we propose centroid discriminant analysis (CDA), a novel geometric classifier that iteratively adjusts the CDB through performance-dependent rotations on two-dimensional planes. These rotations are optimized via Bayesian optimization, enhancing the decision boundary’s adaptability while maintaining scalability. CDA exhibits quadratic training time complexity, outperforming LDA and SVM in terms of scalability and efficiency. Experimental evaluations on 27 real-world datasets, including standard image classification, medical imaging, and chemical property prediction, reveal that CDA consistently outperforms traditional linear methods such as LDA, SVM, and fast SVM in predictive performance, scalability, and stability.
- Nonlinear geometric classification via kernel method: For complex data where linear models are not enough, CDA supports nonlinear classification via kernel method. We demonstrated with challenging image and chemical datasets that kernel CDA improved over linear CDA and outperformed kernel SVM. Nonetheless, while kernel CDA offers greater expressiveness and improved capability, linear CDA remains highly valuable for real-world tasks due to its superior training efficiency, interpretability, and reduced risk of overfitting.
2. Geometric Discriminant Analysis (GDA)
3. Centroid Discriminant Analysis (CDA)
4. Experimental Evidences on Linear Classification of Real Data
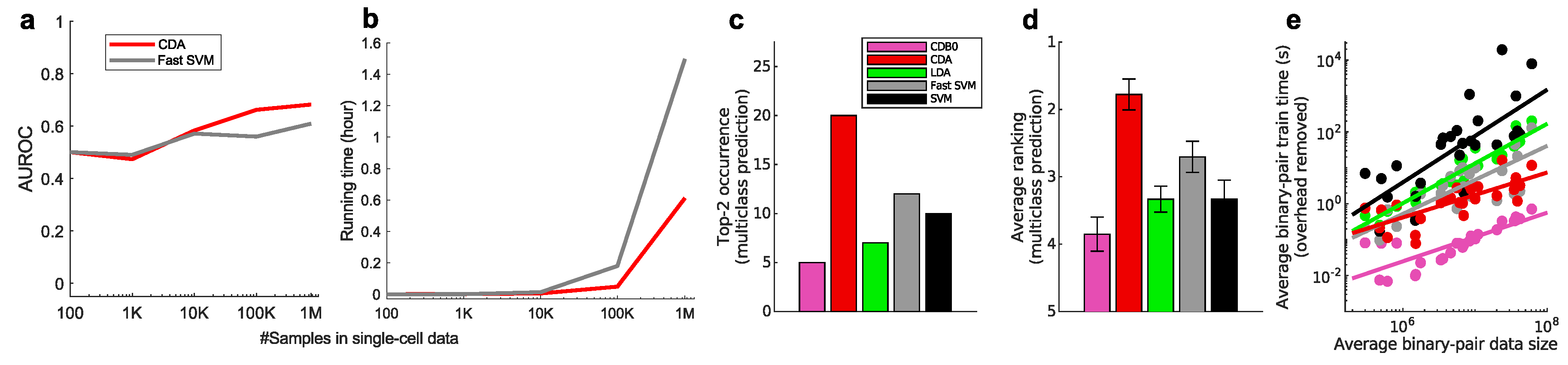
4.1. Algorithm Scalability of Linear Classifiers
4.2. Linear Classification Performance on Real Data
5. Preliminary Test on Nonlinear Kernel CDA
6. Conclusions and Discussions
Appendix A. GDA Demonstration with 2D Artificial Data
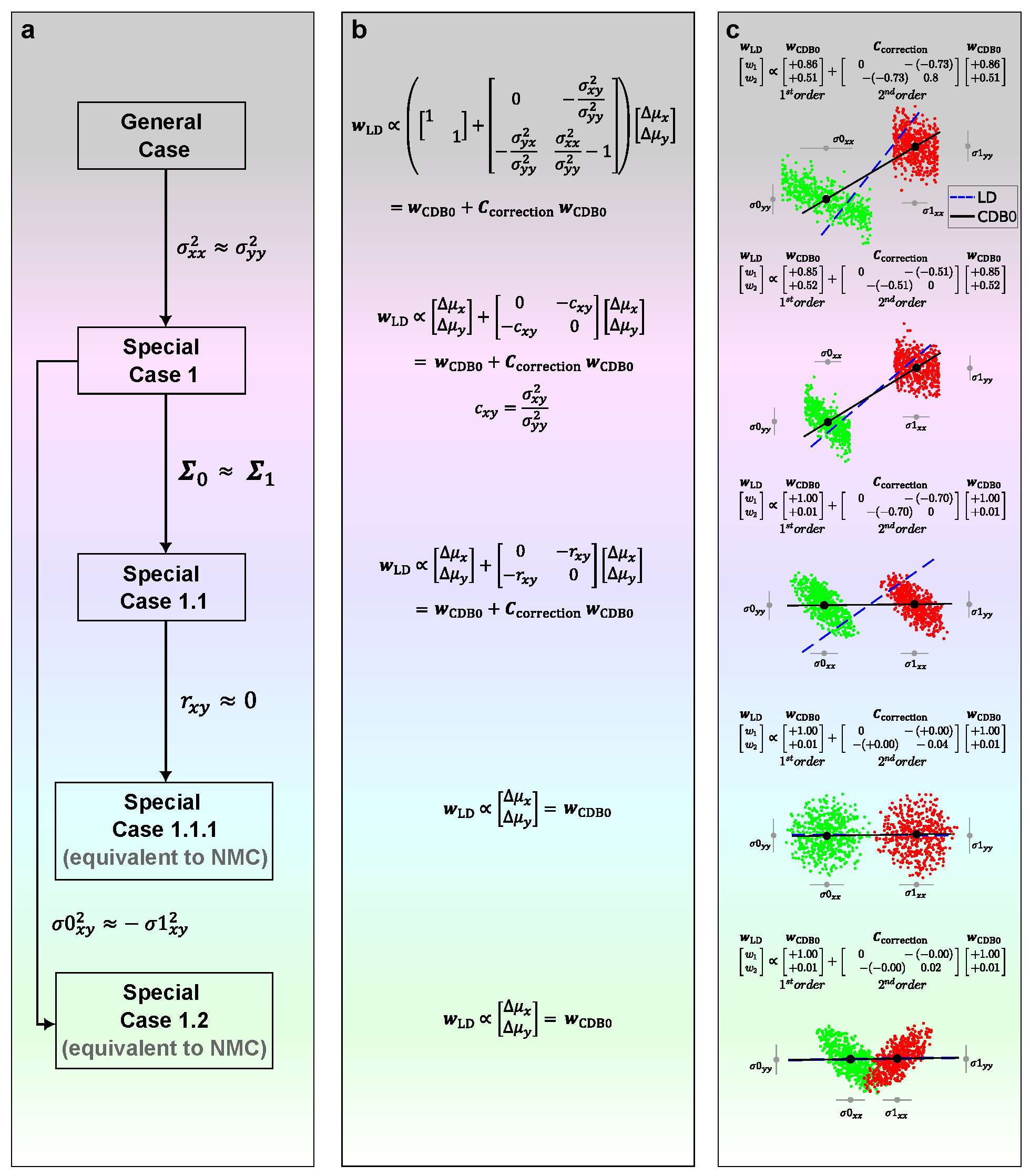
Appendix B. CDA Schematic Diagram
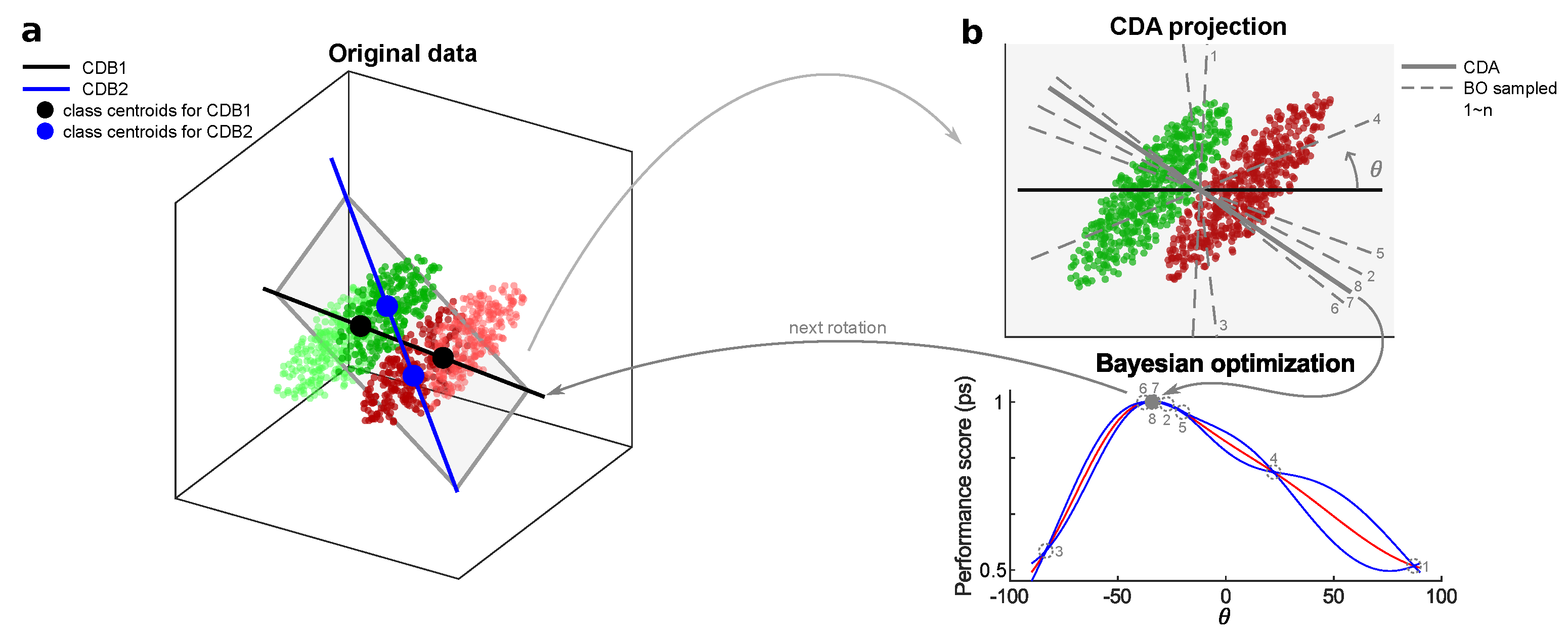
Appendix C. The CDA Algorithm
Appendix C.1. Bayesian Optimization
Appendix C.2. Refining CDA with Statistical Examination on 2D Plane
Appendix C.3. Linear CDA Pseudocode
| Algorithm A1 CDA Main Algorithm (CDA) |
|
| Algorithm A2 Update Sample Weights (updateSampleWeights) |
|
| Algorithm A3 Search Optimal Operating Point (searchOOP) |
|
| Algorithm A4 Approximate Optimal Line by BO (CdaRotation) |
|
| Algorithm A5 Evaluate the Line with Rotation Angle (evaluateRotation) |
|
| Algorithm A6 Refine on the Best Model (refineOnBestPlane) |
|
Appendix C.4. Nonlinear kernel CDA
Appendix D. Deriving LDA in the GDA theory for m-dimensions (m>2)
Appendix E. Deriving CDA in the GDA Theory
Appendix F. Classification Performance Evaluation
Individual Metrics
(a) AUROC
(b) AUPR
(c) F-score
(d) AC-score
(e) Performance-score (ps)
Appendix G. Supplemental Performances on Linear Classification of Large-Scale Data
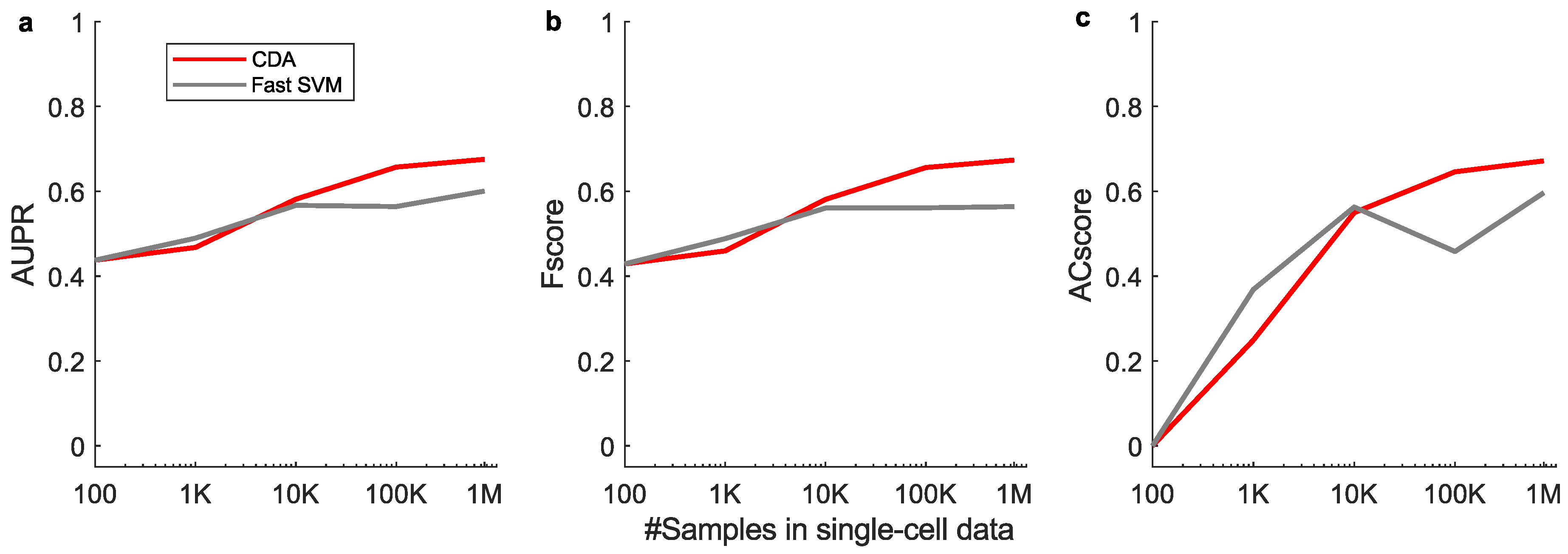
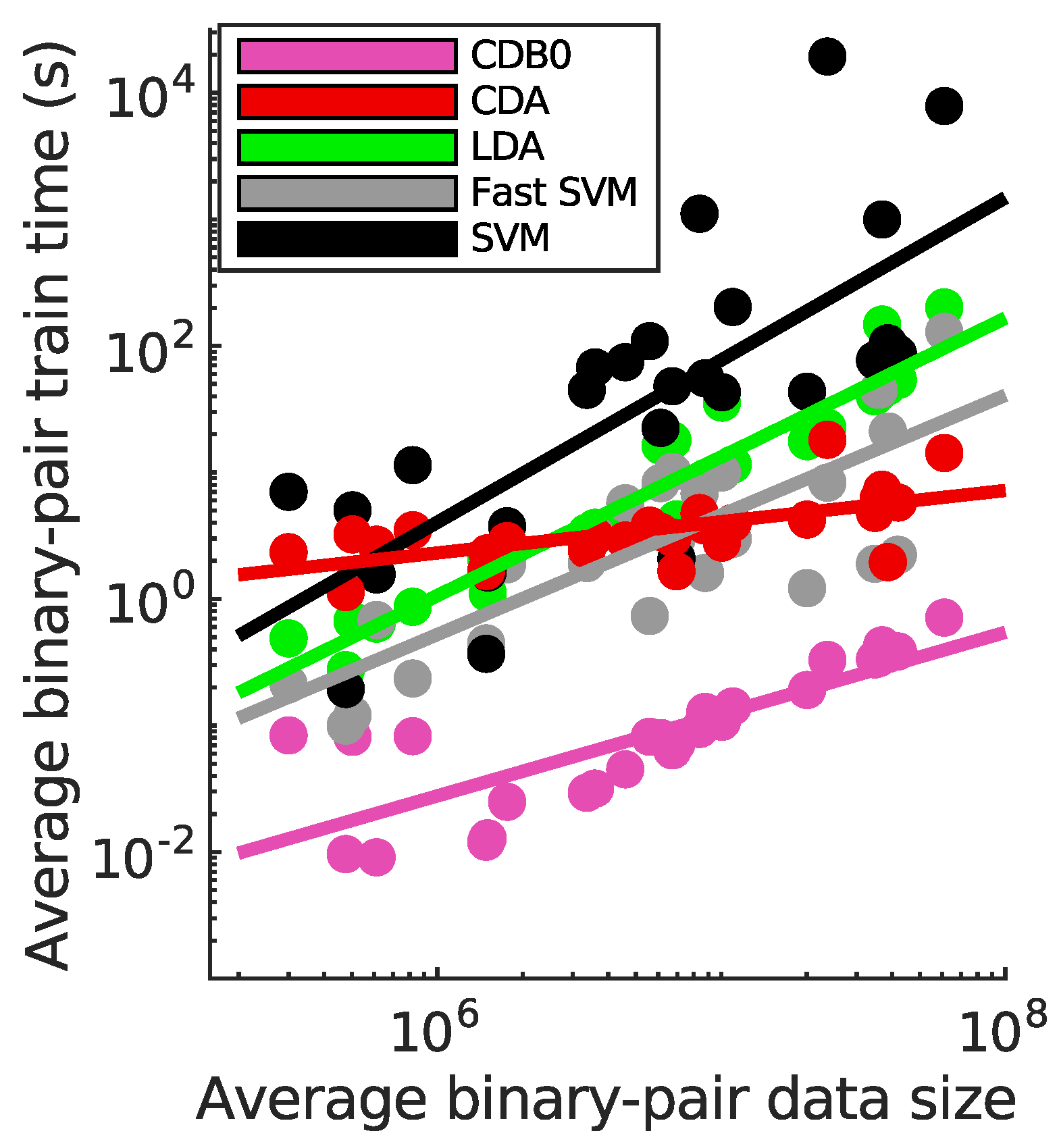
Appendix H. Supplemental Training Speed Results
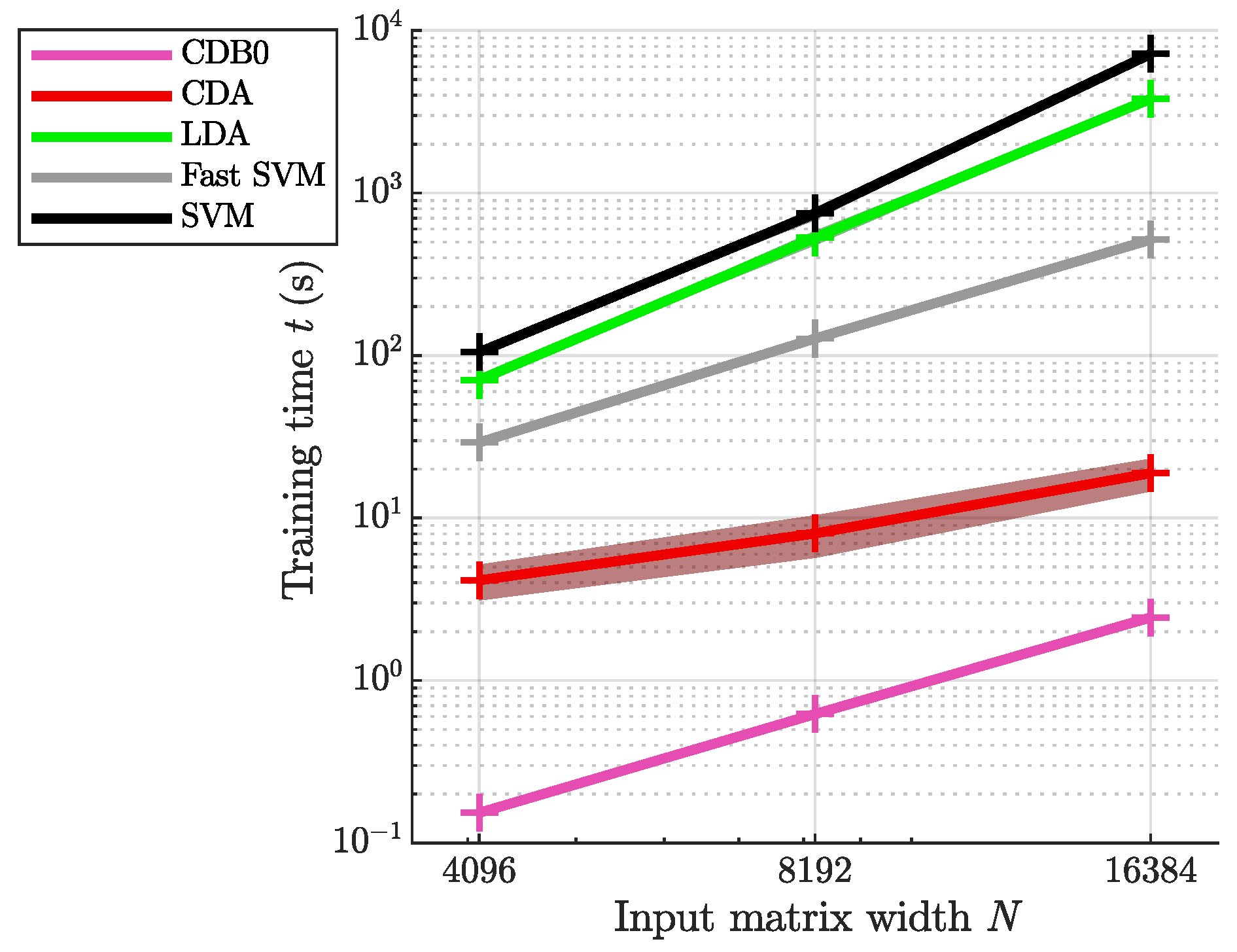
Appendix I. Implementation Details
Appendix I.1. Linear Classifiers
Appendix I.2. Nonlinear Classifiers
Appendix J. Dataset Description
| Dataset | #Samples | #Features | #Classes | Balancedness | Modality/source | Classification task |
| Standard images | ||||||
| MNIST | 70000 | 400 | 10 | imbalanced | image | digits |
| USPS | 9298 | 256 | 10 | imbalanced | image | digits |
| EMNIST | 145600 | 784 | 26 | balanced | image | letters |
| CIFAR10 | 60000 | 3072 | 10 | balanced | image | objects |
| SVHN | 99289 | 3072 | 10 | imbalanced | image | house numbers |
| flower | 3670 | 1200 | 5 | imbalanced | image | flowers |
| GTSRB | 26635 | 1200 | 43 | imbalanced | image | traffic signs |
| STL10 | 13000 | 2352 | 10 | balanced | image | objects |
| FMNIST | 70000 | 784 | 10 | balanced | image | fashion objects |
| Medical images | ||||||
| dermamnist | 10015 | 2352 | 7 | imbalanced | dermatoscope | dermal diseases |
| pneumoniamnist | 5856 | 784 | 2 | imbalanced | chest X-Ray | pneumonia |
| retinamnist | 1600 | 2352 | 5 | imbalanced | fundus camera | diabetic retinopathy |
| breastmnist | 780 | 784 | 2 | imbalanced | breast ultrasound | breast diseases |
| bloodmnist | 17092 | 2352 | 8 | imbalanced | blood cell microscope | blood diseases |
| organamnist | 58830 | 784 | 11 | imbalanced | abdominal CT | human organs |
| organcmnist | 23583 | 784 | 11 | imbalanced | abdominal CT | human organs |
| organsmnist | 25211 | 784 | 11 | imbalanced | abdominal CT | human organs |
| organmnist3d | 1472 | 21952 | 11 | imbalanced | abdominal CT | human organs |
| nodulemnist3d | 1633 | 21952 | 2 | imbalanced | chest CT | nodule malignancy |
| fracturemnist3d | 1370 | 21952 | 3 | imbalanced | chest CT | fracture types |
| adrenalmnist3d | 1584 | 21952 | 2 | imbalanced | shape from abdominal CT | adrenal gland mass |
| vesselmnist3d | 1908 | 21952 | 2 | imbalanced | shape from brain MRA | aneurysm |
| synapsemnist3d | 1759 | 21952 | 2 | imbalanced | electron microscope | excitatory/inhibitory |
| Chemical formula | ||||||
| bace | 1513 | 198 | 2 | imbalanced | chemical formula | BACE1 enzyme |
| BBBP | 2050 | 400 | 2 | imbalanced | chemical formula | blood-brain barrier permeability |
| clintox | 1484 | 339 | 2 | imbalanced | chemical formula | clinical toxicity |
| HIV | 41127 | 575 | 2 | imbalanced | chemical formula | HIV drug activity |
| Large-scale single-cell sequencing data | ||||||
| Mouse brain | 1306127 | 27998 | 10 | imbalanced | single-cell sequencing | cell type |
Appendix K. Supplemental Performances on Real Datasets of linear CDA
Appendix K.1. Binary Classification
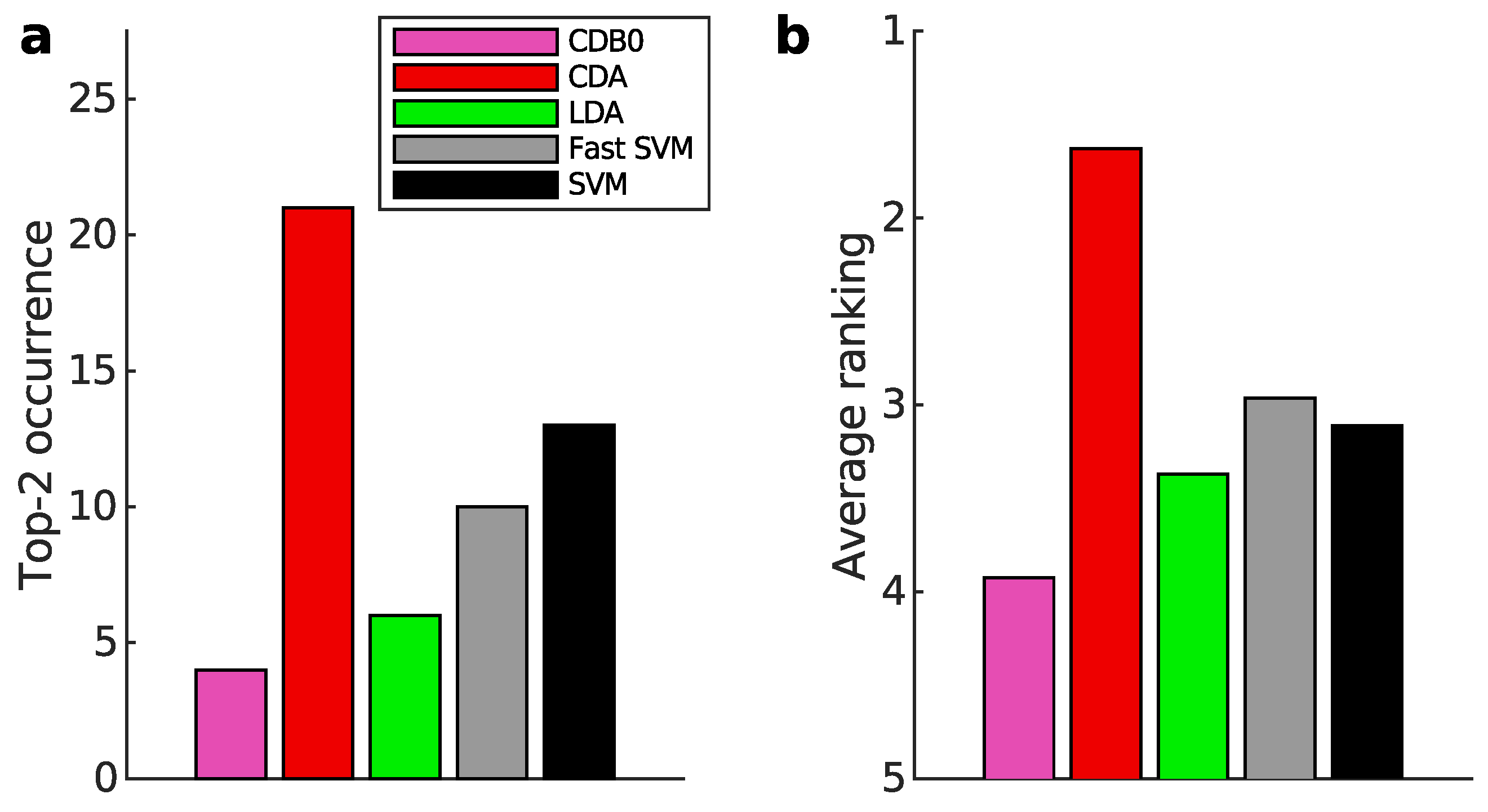
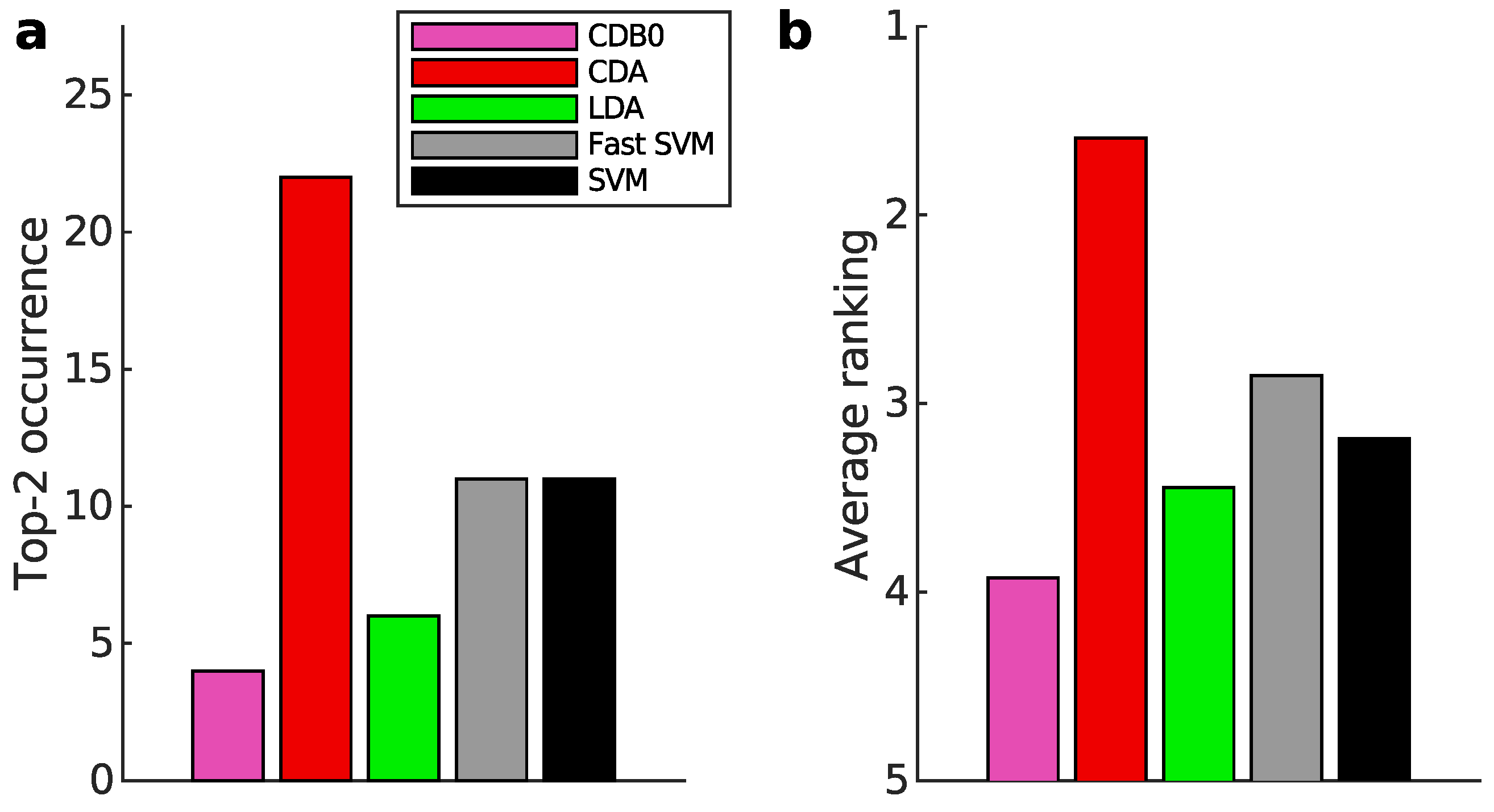
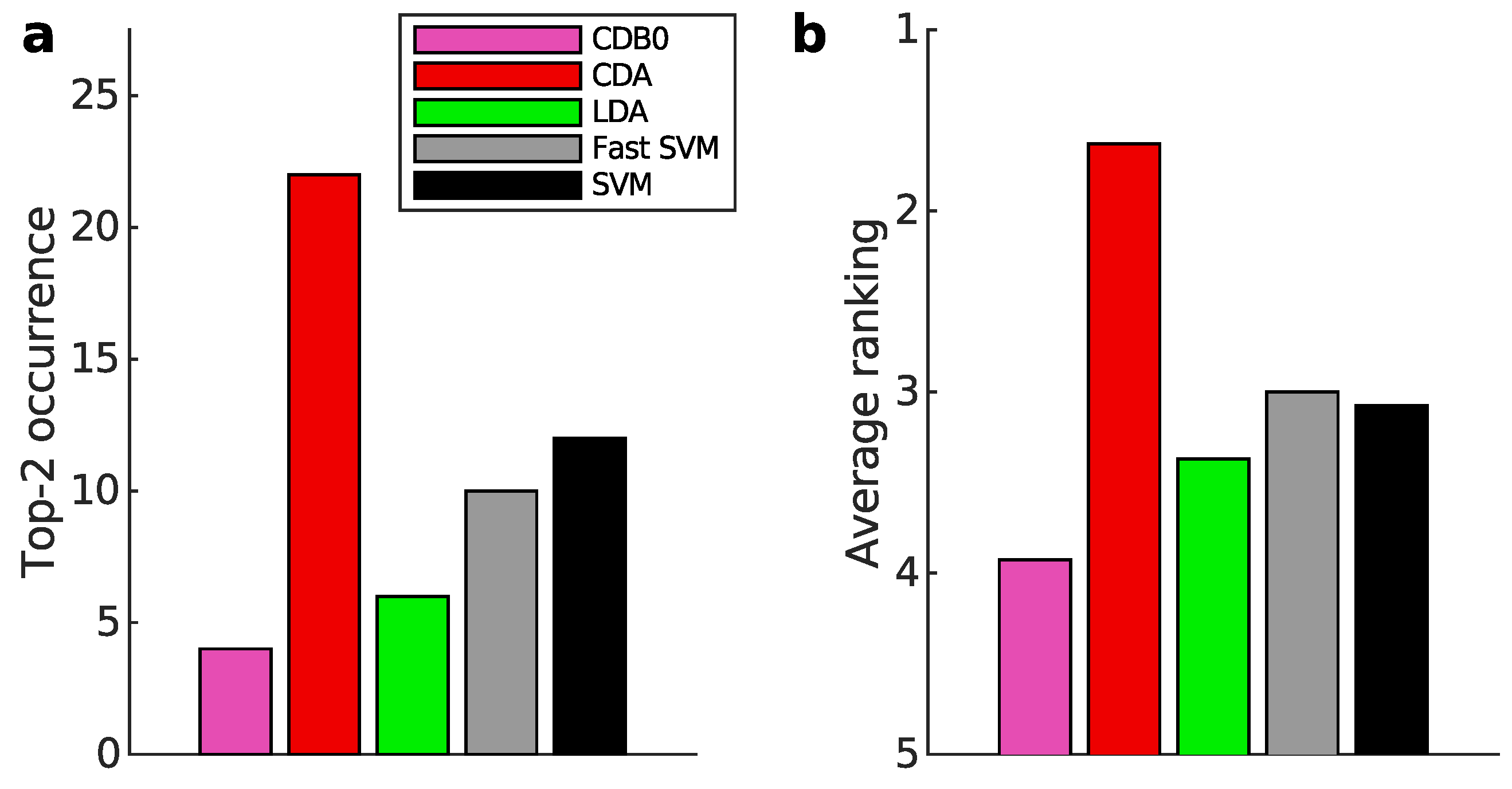
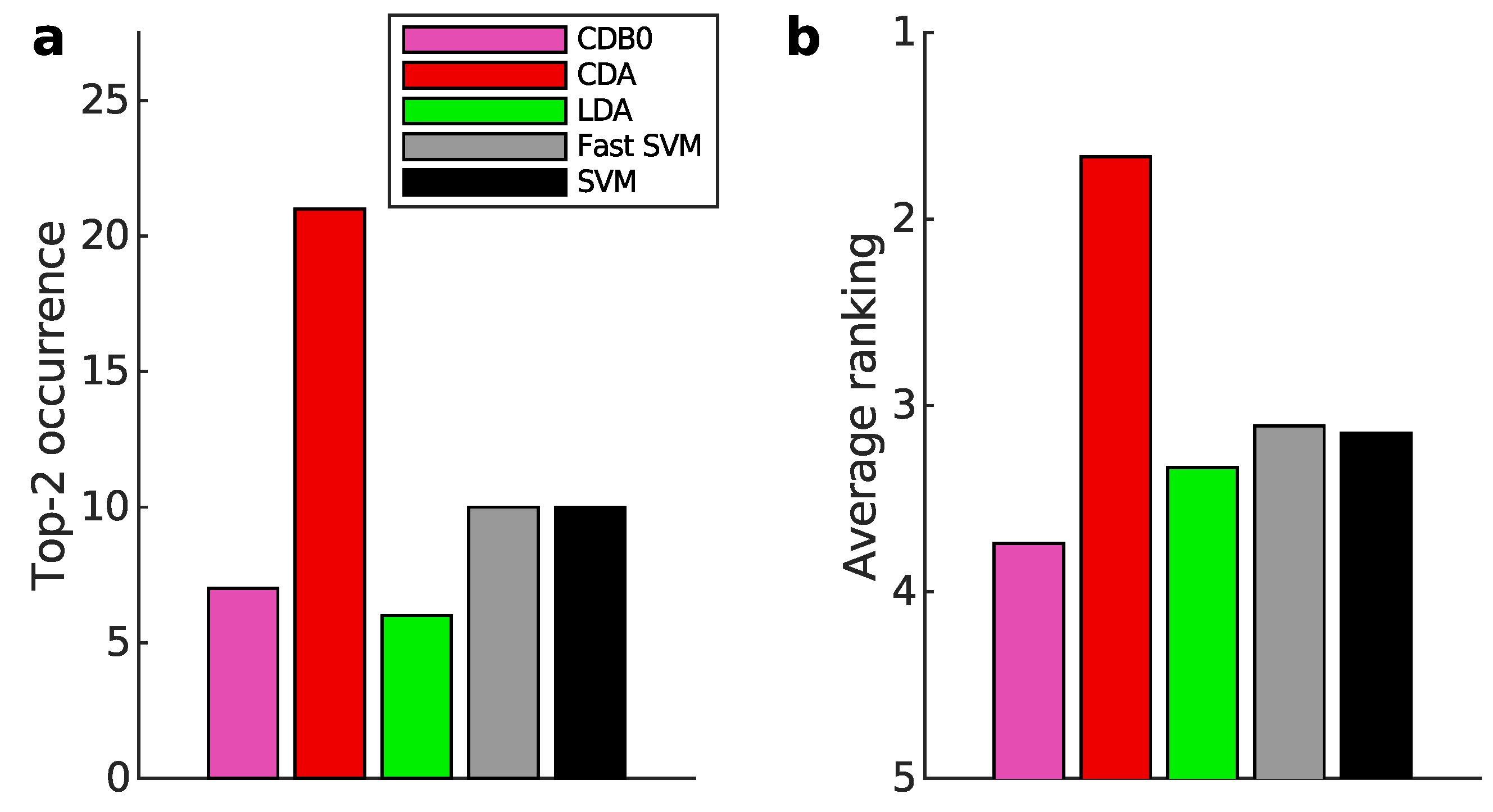
Appendix K.2. Multiclass Prediction
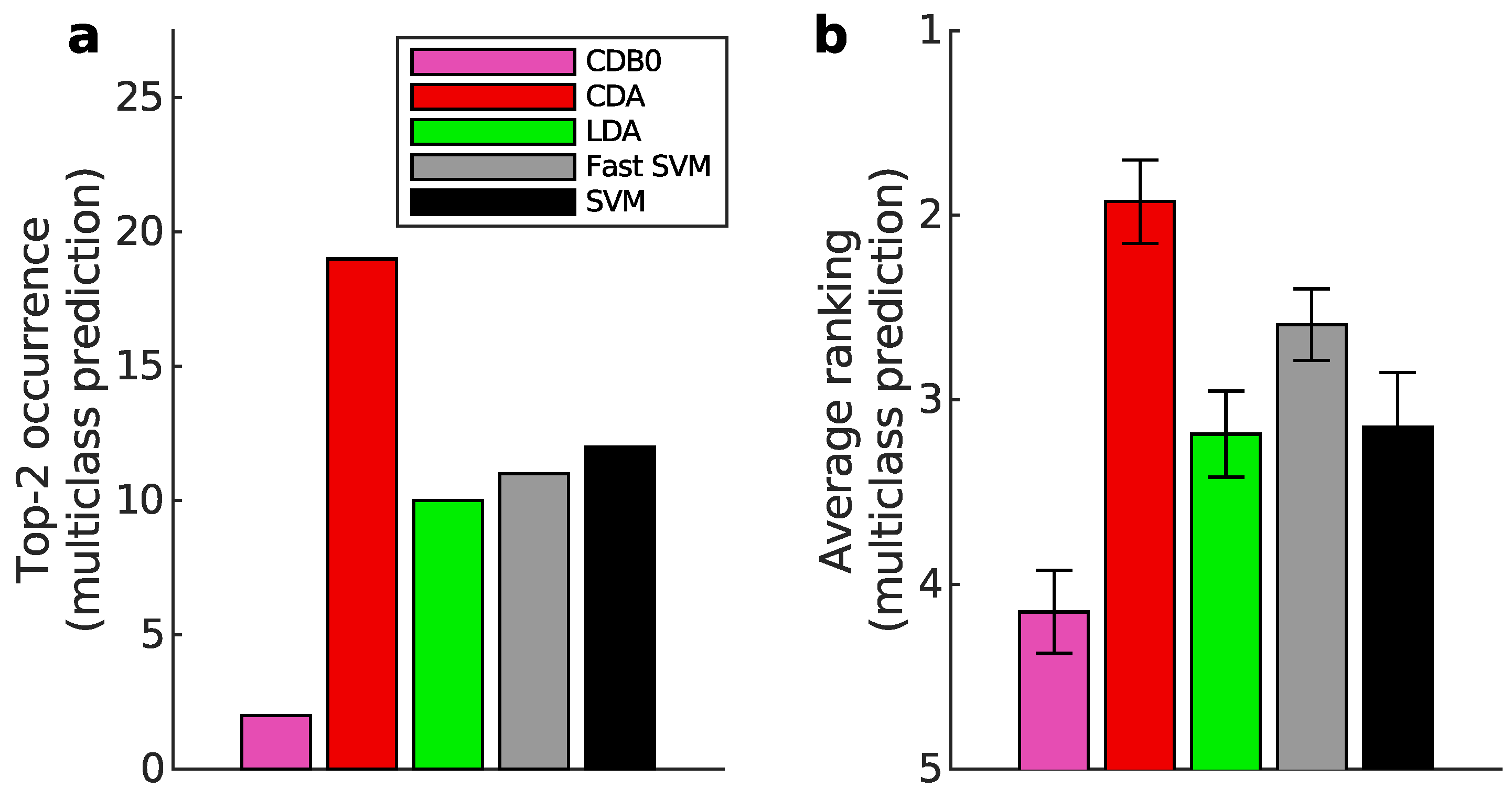
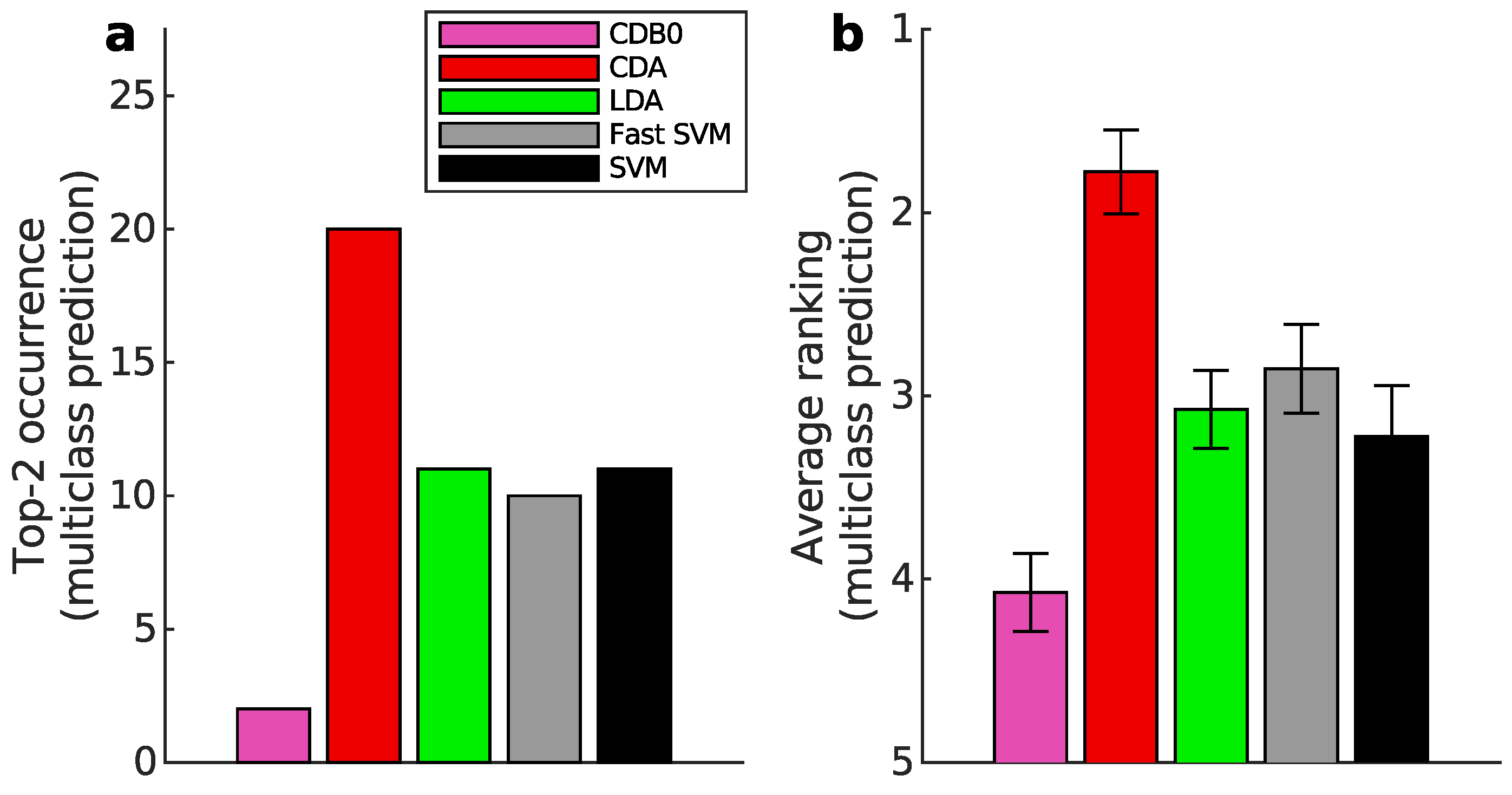
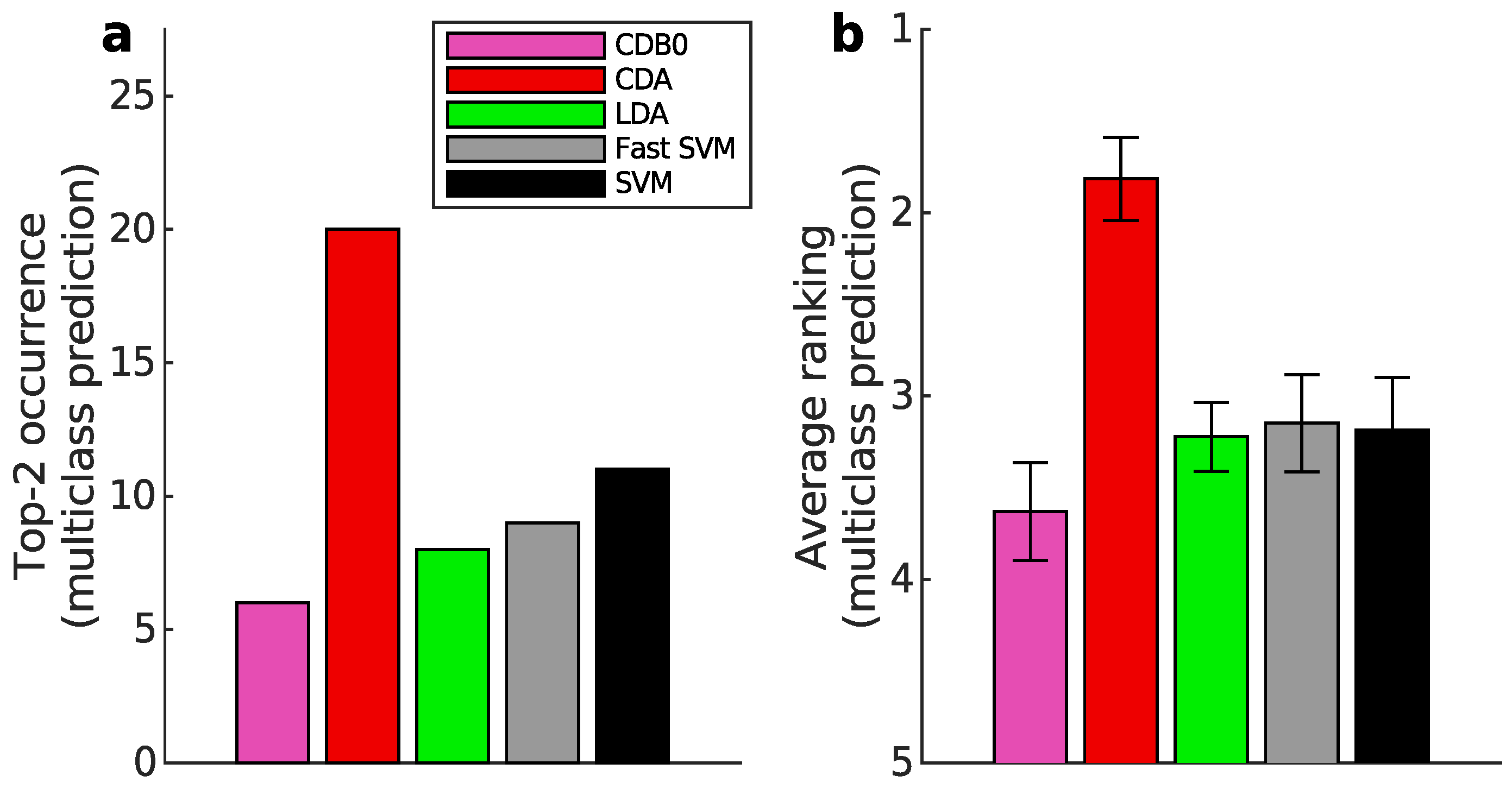
Appendix L. Full Classification Performance on Real Datasets of Linear CDA
Appendix L.1. Binary Classification
| Dataset | CDB0 | CDA | LDA | Fast SVM | SVM |
| Standard images | |||||
| MNIST | 0.957±0.004 | 0.985±0.002 | 0.981±0.002 | 0.985±0.002 | 0.986±0.002 |
| USPS | 0.966±0.005 | 0.989±0.001 | 0.982±0.002 | 0.99±0.002 | 0.99±0.002 |
| EMNIST | 0.928±0.003 | 0.972±0.001 | 0.964±0.001 | 0.97±0.001 | 0.97±0.001 |
| CIFAR10 | 0.696±0.01 | 0.797±0.01 | 0.741±0.01 | 0.754±0.01 | 0.787±0.01 |
| SVHN | 0.528±0.003 | 0.667±0.005 | 0.555±0.003 | 0.55±0.004 | 0.591±0.004 |
| flower | 0.703±0.02 | 0.739±0.02 | 0.571±0.01 | 0.71±0.03 | 0.71±0.03 |
| GTSRB | 0.767±0.003 | 0.972±0.001 | 0.942±0.002 | 0.995±0.0004 | 0.995±0.0004 |
| STL10 | 0.723±0.02 | 0.781±0.02 | 0.667±0.01 | 0.758±0.02 | 0.761±0.02 |
| FMNIST | 0.937±0.01 | 0.975±0.006 | 0.973±0.006 | 0.976±0.006 | 0.976±0.006 |
| Medical images | |||||
| dermamnist | 0.682±0.01 | 0.753±0.02 | 0.684±0.02 | 0.676±0.02 | 0.703±0.02 |
| pneumoniamnist | 0.837±0 | 0.933±0 | 0.912±0 | 0.941±0 | 0.872±0 |
| retinamnist | 0.63±0.03 | 0.662±0.04 | 0.616±0.02 | 0.631±0.03 | 0.61±0.02 |
| breastmnist | 0.66±0 | 0.763±0 | 0.703±0 | 0.726±0 | 0.71±0 |
| bloodmnist | 0.89±0.02 | 0.947±0.01 | 0.898±0.02 | 0.951±0.01 | 0.947±0.01 |
| organamnist | 0.897±0.009 | 0.948±0.008 | 0.95±0.008 | 0.928±0.01 | 0.894±0.02 |
| organcmnist | 0.89±0.01 | 0.925±0.01 | 0.908±0.01 | 0.895±0.01 | 0.886±0.02 |
| organsmnist | 0.831±0.01 | 0.886±0.01 | 0.866±0.01 | 0.842±0.02 | 0.814±0.02 |
| organmnist3d | 0.924±0.01 | 0.957±0.008 | 0.953±0.008 | 0.965±0.007 | 0.7±0.02 |
| nodulemnist3d | 0.715±0 | 0.781±0 | 0.732±0 | 0.687±0 | 0.654±0 |
| fracturemnist3d | 0.671±0.06 | 0.556±0.03 | 0.525±0.04 | 0.576±0.007 | 0.553±0.02 |
| adrenalmnist3d | 0.653±0 | 0.756±0 | 0.692±0 | 0.697±0 | 0.727±0 |
| vesselmnist3d | 0.605±0 | 0.685±0 | 0.681±0 | 0.61±0 | 0.612±0 |
| synapsemnist3d | 0.539±0 | 0.544±0 | 0.508±0 | 0.518±0 | NaN |
| Chemical formula | |||||
| bace | 0.621±0 | 0.705±0 | 0.684±0 | 0.618±0 | 0.61±0 |
| BBBP | 0.711±0 | 0.743±0 | 0.693±0 | 0.667±0 | 0.705±0 |
| clintox | 0.65±0 | 0.575±0 | 0.543±0 | 0.517±0 | 0.508±0 |
| HIV | 0.6±0 | 0.616±0 | 0.537±0 | 0.51±0 | 0.512±0 |
| Dataset | CDB0 | CDA | LDA | Fast SVM | SVM |
| Standard images | |||||
| MNIST | 0.957±0.004 | 0.985±0.002 | 0.981±0.002 | 0.985±0.002 | 0.986±0.002 |
| USPS | 0.966±0.005 | 0.989±0.001 | 0.983±0.002 | 0.990±0.002 | 0.990±0.002 |
| EMNIST | 0.928±0.003 | 0.972±0.001 | 0.964±0.001 | 0.970±0.001 | 0.970±0.001 |
| CIFAR10 | 0.697±0.01 | 0.797±0.01 | 0.741±0.01 | 0.762±0.01 | 0.787±0.01 |
| SVHN | 0.528±0.003 | 0.682±0.006 | 0.559±0.003 | 0.577±0.005 | 0.634±0.006 |
| flower | 0.704±0.02 | 0.741±0.02 | 0.571±0.01 | 0.712±0.03 | 0.712±0.03 |
| GTSRB | 0.757±0.003 | 0.973±0.001 | 0.934±0.002 | 0.995±0.0003 | 0.995±0.0003 |
| STL10 | 0.724±0.02 | 0.782±0.02 | 0.667±0.01 | 0.759±0.02 | 0.762±0.02 |
| FMNIST | 0.937±0.01 | 0.975±0.006 | 0.973±0.006 | 0.976±0.006 | 0.976±0.006 |
| Medical images | |||||
| dermamnist | 0.653±0.01 | 0.743±0.02 | 0.681±0.02 | 0.729±0.02 | 0.695±0.02 |
| pneumoniamnist | 0.817±0 | 0.931±0 | 0.922±0 | 0.937±0 | 0.871±0 |
| retinamnist | 0.614±0.03 | 0.649±0.03 | 0.612±0.02 | 0.641±0.02 | 0.607±0.02 |
| breastmnist | 0.653±0 | 0.759±0 | 0.690±0 | 0.743±0 | 0.706±0 |
| bloodmnist | 0.889±0.02 | 0.947±0.01 | 0.895±0.02 | 0.953±0.01 | 0.947±0.01 |
| organamnist | 0.902±0.009 | 0.950±0.007 | 0.951±0.008 | 0.929±0.01 | 0.895±0.02 |
| organcmnist | 0.901±0.009 | 0.931±0.009 | 0.908±0.01 | 0.893±0.01 | 0.883±0.02 |
| organsmnist | 0.839±0.01 | 0.892±0.01 | 0.867±0.01 | 0.840±0.02 | 0.813±0.02 |
| organmnist3d | 0.924±0.009 | 0.958±0.008 | 0.954±0.008 | 0.965±0.007 | 0.723±0.02 |
| nodulemnist3d | 0.7±0 | 0.771±0 | 0.745±0 | 0.695±0 | 0.637±0 |
| fracturemnist3d | 0.663±0.05 | 0.566±0.02 | 0.531±0.05 | 0.583±0.003 | 0.555±0.02 |
| adrenalmnist3d | 0.65±0 | 0.774±0 | 0.705±0 | 0.708±0 | 0.728±0 |
| vesselmnist3d | 0.582±0 | 0.671±0 | 0.694±0 | 0.627±0 | 0.659±0 |
| synapsemnist3d | 0.537±0 | 0.542±0 | 0.544±0 | 0.533±0 | NaN |
| Chemical formula | |||||
| bace | 0.620±0 | 0.704±0 | 0.685±0 | 0.643±0 | 0.615±0 |
| BBBP | 0.701±0 | 0.747±0 | 0.734±0 | 0.712±0 | 0.714±0 |
| clintox | 0.602±0 | 0.570±0 | 0.548±0 | 0.553±0 | 0.514±0 |
| HIV | 0.565±0 | 0.583±0 | 0.558±0 | 0.612±0 | 0.632±0 |
| Dataset | CDB0 | CDA | LDA | Fast SVM | SVM |
| Standard images | |||||
| MNIST | 0.957±0.004 | 0.985±0.002 | 0.981±0.002 | 0.985±0.002 | 0.986±0.002 |
| USPS | 0.966±0.005 | 0.989±0.001 | 0.983±0.002 | 0.99±0.002 | 0.99±0.002 |
| EMNIST | 0.928±0.003 | 0.972±0.001 | 0.964±0.001 | 0.97±0.001 | 0.97±0.001 |
| CIFAR10 | 0.696±0.01 | 0.797±0.01 | 0.741±0.01 | 0.747±0.01 | 0.787±0.01 |
| SVHN | 0.523±0.003 | 0.664±0.005 | 0.555±0.003 | 0.51±0.01 | 0.577±0.006 |
| flower | 0.701±0.02 | 0.738±0.02 | 0.57±0.01 | 0.709±0.03 | 0.709±0.03 |
| GTSRB | 0.743±0.003 | 0.972±0.001 | 0.931±0.002 | 0.995±0.0003 | 0.995±0.0003 |
| STL10 | 0.722±0.02 | 0.781±0.02 | 0.666±0.01 | 0.756±0.02 | 0.761±0.02 |
| FMNIST | 0.937±0.01 | 0.975±0.006 | 0.973±0.006 | 0.976±0.006 | 0.976±0.006 |
| Medical images | |||||
| dermamnist | 0.621±0.02 | 0.736±0.02 | 0.677±0.02 | 0.682±0.03 | 0.692±0.02 |
| pneumoniamnist | 0.812±0 | 0.931±0 | 0.921±0 | 0.937±0 | 0.871±0 |
| retinamnist | 0.594±0.02 | 0.639±0.03 | 0.61±0.02 | 0.611±0.04 | 0.606±0.02 |
| breastmnist | 0.651±0 | 0.759±0 | 0.684±0 | 0.739±0 | 0.705±0 |
| bloodmnist | 0.888±0.02 | 0.947±0.01 | 0.894±0.02 | 0.952±0.01 | 0.946±0.01 |
| organamnist | 0.899±0.01 | 0.949±0.008 | 0.951±0.008 | 0.923±0.02 | 0.893±0.02 |
| organcmnist | 0.896±0.01 | 0.929±0.009 | 0.908±0.01 | 0.892±0.02 | 0.882±0.02 |
| organsmnist | 0.834±0.01 | 0.89±0.01 | 0.866±0.01 | 0.834±0.02 | 0.811±0.02 |
| organmnist3d | 0.92±0.01 | 0.957±0.008 | 0.953±0.008 | 0.964±0.007 | 0.68±0.02 |
| nodulemnist3d | 0.695±0 | 0.769±0 | 0.743±0 | 0.694±0 | 0.622±0 |
| fracturemnist3d | 0.651±0.05 | 0.523±0.03 | 0.514±0.05 | 0.577±0.007 | 0.55±0.02 |
| adrenalmnist3d | 0.65±0 | 0.771±0 | 0.703±0 | 0.707±0 | 0.728±0 |
| vesselmnist3d | 0.56±0 | 0.669±0 | 0.693±0 | 0.623±0 | 0.638±0 |
| synapsemnist3d | 0.534±0 | 0.539±0 | 0.45±0 | 0.493±0 | NaN |
| Chemical formula | |||||
| bace | 0.619±0 | 0.704±0 | 0.684±0 | 0.569±0 | 0.607±0 |
| BBBP | 0.699±0 | 0.747±0 | 0.718±0 | 0.691±0 | 0.713±0 |
| clintox | 0.54±0 | 0.569±0 | 0.547±0 | 0.517±0 | 0.506±0 |
| HIV | 0.531±0 | 0.562±0 | 0.548±0 | 0.511±0 | 0.514±0 |
| Dataset | CDB0 | CDA | LDA | Fast SVM | SVM |
| Standard images | |||||
| MNIST | 0.957±0.004 | 0.985±0.002 | 0.981±0.002 | 0.985±0.002 | 0.986±0.002 |
| USPS | 0.966±0.005 | 0.989±0.001 | 0.982±0.002 | 0.99±0.002 | 0.99±0.002 |
| EMNIST | 0.927±0.003 | 0.972±0.001 | 0.964±0.001 | 0.97±0.001 | 0.97±0.001 |
| CIFAR10 | 0.694±0.01 | 0.796±0.01 | 0.74±0.01 | 0.725±0.02 | 0.787±0.01 |
| SVHN | 0.524±0.002 | 0.614±0.004 | 0.499±0.008 | 0.352±0.03 | 0.438±0.02 |
| flower | 0.698±0.02 | 0.733±0.02 | 0.567±0.01 | 0.701±0.03 | 0.7±0.03 |
| GTSRB | 0.753±0.003 | 0.97±0.002 | 0.941±0.002 | 0.995±0.0004 | 0.995±0.0004 |
| STL10 | 0.72±0.01 | 0.779±0.02 | 0.664±0.01 | 0.75±0.02 | 0.76±0.02 |
| FMNIST | 0.936±0.01 | 0.975±0.006 | 0.973±0.006 | 0.975±0.006 | 0.976±0.006 |
| Medical images | |||||
| dermamnist | 0.658±0.02 | 0.72±0.02 | 0.608±0.04 | 0.535±0.05 | 0.654±0.03 |
| pneumoniamnist | 0.837±0 | 0.932±0 | 0.908±0 | 0.94±0 | 0.868±0 |
| retinamnist | 0.62±0.03 | 0.639±0.05 | 0.567±0.03 | 0.513±0.07 | 0.561±0.03 |
| breastmnist | 0.641±0 | 0.751±0 | 0.698±0 | 0.682±0 | 0.691±0 |
| bloodmnist | 0.889±0.02 | 0.946±0.01 | 0.897±0.02 | 0.949±0.01 | 0.947±0.01 |
| organamnist | 0.892±0.01 | 0.946±0.008 | 0.949±0.008 | 0.919±0.02 | 0.882±0.02 |
| organcmnist | 0.881±0.01 | 0.92±0.01 | 0.907±0.01 | 0.893±0.02 | 0.886±0.02 |
| organsmnist | 0.822±0.01 | 0.88±0.01 | 0.862±0.01 | 0.833±0.02 | 0.801±0.02 |
| organmnist3d | 0.918±0.01 | 0.956±0.008 | 0.952±0.008 | 0.964±0.007 | 0.602±0.03 |
| nodulemnist3d | 0.707±0 | 0.773±0 | 0.693±0 | 0.636±0 | 0.651±0 |
| fracturemnist3d | 0.668±0.06 | 0.38±0.1 | 0.351±0.1 | 0.491±0.06 | 0.45±0.08 |
| adrenalmnist3d | 0.602±0 | 0.718±0 | 0.631±0 | 0.642±0 | 0.694±0 |
| vesselmnist3d | 0.547±0 | 0.615±0 | 0.58±0 | 0.43±0 | 0.405±0 |
| synapsemnist3d | 0.498±0 | 0.506±0 | 0.0612±0 | 0.189±0 | NaN |
| Chemical formula | |||||
| bace | 0.62±0 | 0.704±0 | 0.678±0 | 0.483±0 | 0.575±0 |
| BBBP | 0.693±0 | 0.715±0 | 0.603±0 | 0.553±0 | 0.66±0 |
| clintox | 0.634±0 | 0.365±0 | 0.238±0 | 0.0869±0 | 0.0868±0 |
| HIV | 0.471±0 | 0.465±0 | 0.159±0 | 0.0407±0 | 0.0473±0 |
Appendix L.2. Multiclass Prediction
| Dataset | CDB0 | CDA | LDA | Fast SVM | SVM |
| Standard images | |||||
| MNIST | 0.897±0.01 | 0.963±0.005 | 0.958±0.006 | 0.965±0.005 | 0.966±0.005 |
| USPS | 0.914±0.01 | 0.971±0.004 | 0.969±0.006 | 0.974±0.005 | 0.973±0.005 |
| EMNIST | 0.773±0.01 | 0.896±0.008 | 0.879±0.009 | 0.891±0.008 | 0.892±0.008 |
| CIFAR10 | 0.599±0.02 | 0.671±0.02 | 0.627±0.01 | 0.641±0.02 | 0.663±0.02 |
| SVHN | 0.522±0.006 | 0.638±0.01 | 0.531±0.007 | 0.536±0.01 | 0.558±0.01 |
| flower | 0.61±0.03 | 0.666±0.03 | 0.554±0.02 | 0.632±0.03 | 0.632±0.03 |
| GTSRB | 0.589±0.01 | 0.878±0.01 | 0.821±0.02 | 0.982±0.003 | 0.983±0.003 |
| STL10 | 0.607±0.02 | 0.655±0.02 | 0.596±0.02 | 0.648±0.02 | 0.653±0.02 |
| FMNIST | 0.836±0.03 | 0.917±0.02 | 0.92±0.02 | 0.924±0.02 | 0.924±0.02 |
| Medical images | |||||
| dermamnist | 0.614±0.03 | 0.658±0.03 | 0.588±0.02 | 0.595±0.03 | 0.622±0.02 |
| pneumoniamnist | 0.837±0 | 0.933±0 | 0.912±0 | 0.941±0 | 0.872±0 |
| retinamnist | 0.575±0.04 | 0.622±0.04 | 0.592±0.03 | 0.596±0.04 | 0.578±0.03 |
| breastmnist | 0.66±0 | 0.763±0 | 0.703±0 | 0.726±0 | 0.71±0 |
| bloodmnist | 0.789±0.04 | 0.88±0.03 | 0.817±0.03 | 0.882±0.03 | 0.881±0.03 |
| organamnist | 0.815±0.03 | 0.888±0.02 | 0.885±0.03 | 0.85±0.04 | 0.795±0.05 |
| organcmnist | 0.809±0.03 | 0.869±0.03 | 0.833±0.03 | 0.811±0.04 | 0.795±0.04 |
| organsmnist | 0.694±0.03 | 0.761±0.03 | 0.735±0.03 | 0.7±0.03 | 0.681±0.04 |
| organmnist3d | 0.867±0.03 | 0.913±0.02 | 0.903±0.03 | 0.924±0.02 | 0.636±0.03 |
| nodulemnist3d | 0.715±0 | 0.781±0 | 0.732±0 | 0.687±0 | 0.654±0 |
| fracturemnist3d | 0.622±0.04 | 0.518±0.01 | 0.554±0.04 | 0.574±0.004 | 0.554±0.02 |
| adrenalmnist3d | 0.653±0 | 0.756±0 | 0.9±0 | 0.928±0 | 0.947±0 |
| vesselmnist3d | 0.605±0 | 0.685±0 | 0.681±0 | 0.61±0 | 0.612±0 |
| synapsemnist3d | 0.539±0 | 0.544±0 | 0.508±0 | 0.518±0 | NaN |
| Chemical formula | |||||
| bace | 0.621±0 | 0.705±0 | 0.684±0 | 0.618±0 | 0.61±0 |
| BBBP | 0.711±0 | 0.743±0 | 0.693±0 | 0.667±0 | 0.705±0 |
| clintox | 0.65±0 | 0.575±0 | 0.543±0 | 0.517±0 | 0.508±0 |
| HIV | 0.6±0 | 0.616±0 | 0.537±0 | 0.51±0 | 0.512±0 |
| Dataset | CDB0 | CDA | LDA | Fast SVM | SVM |
| Standard images | |||||
| MNIST | 0.897±0.01 | 0.963±0.004 | 0.958±0.005 | 0.965±0.004 | 0.966±0.004 |
| USPS | 0.918±0.01 | 0.971±0.005 | 0.969±0.005 | 0.974±0.004 | 0.973±0.005 |
| EMNIST | 0.774±0.01 | 0.896±0.008 | 0.88±0.009 | 0.891±0.008 | 0.892±0.008 |
| CIFAR10 | 0.6±0.01 | 0.67±0.01 | 0.627±0.01 | 0.64±0.02 | 0.663±0.02 |
| SVHN | 0.528±0.009 | 0.666±0.01 | 0.533±0.006 | 0.546±0.005 | 0.597±0.005 |
| flower | 0.611±0.02 | 0.665±0.02 | 0.554±0.02 | 0.631±0.02 | 0.632±0.02 |
| GTSRB | 0.619±0.01 | 0.891±0.01 | 0.805±0.01 | 0.981±0.003 | 0.981±0.003 |
| STL10 | 0.604±0.02 | 0.653±0.02 | 0.598±0.02 | 0.648±0.02 | 0.651±0.02 |
| FMNIST | 0.836±0.03 | 0.916±0.02 | 0.92±0.02 | 0.923±0.02 | 0.924±0.02 |
| Medical images | |||||
| dermamnist | 0.592±0.02 | 0.645±0.02 | 0.603±0.02 | 0.608±0.03 | 0.613±0.02 |
| pneumoniamnist | 0.817±0 | 0.931±0 | 0.922±0 | 0.937±0 | 0.871±0 |
| retinamnist | 0.568±0.03 | 0.621±0.04 | 0.594±0.03 | 0.589±0.04 | 0.577±0.03 |
| breastmnist | 0.653±0 | 0.759±0 | 0.69±0 | 0.743±0 | 0.706±0 |
| bloodmnist | 0.785±0.04 | 0.878±0.03 | 0.818±0.03 | 0.89±0.03 | 0.88±0.03 |
| organamnist | 0.82±0.03 | 0.892±0.02 | 0.889±0.02 | 0.851±0.03 | 0.803±0.05 |
| organcmnist | 0.818±0.03 | 0.877±0.02 | 0.838±0.03 | 0.811±0.04 | 0.792±0.04 |
| organsmnist | 0.697±0.02 | 0.764±0.03 | 0.741±0.03 | 0.703±0.03 | 0.688±0.04 |
| organmnist3d | 0.867±0.02 | 0.913±0.02 | 0.904±0.03 | 0.924±0.02 | 0.668±0.04 |
| nodulemnist3d | 0.7±0 | 0.771±0 | 0.745±0 | 0.695±0 | 0.637±0 |
| fracturemnist3d | 0.617±0.04 | 0.526±0.02 | 0.553±0.04 | 0.578±0.004 | 0.555±0.02 |
| adrenalmnist3d | 0.65±0 | 0.774±0 | 0.911±0 | 0.939±0 | 0.949±0 |
| vesselmnist3d | 0.582±0 | 0.671±0 | 0.694±0 | 0.627±0 | 0.659±0 |
| synapsemnist3d | 0.537±0 | 0.542±0 | 0.544±0 | 0.533±0 | NaN |
| Chemical formula | |||||
| bace | 0.62±0 | 0.704±0 | 0.685±0 | 0.643±0 | 0.615±0 |
| BBBP | 0.701±0 | 0.747±0 | 0.734±0 | 0.712±0 | 0.714±0 |
| clintox | 0.602±0 | 0.57±0 | 0.548±0 | 0.553±0 | 0.514±0 |
| HIV | 0.565±0 | 0.583±0 | 0.558±0 | 0.612±0 | 0.632±0 |
| Dataset | CDB0 | CDA | LDA | Fast SVM | SVM |
| Standard images | |||||
| MNIST | 0.896±0.01 | 0.963±0.004 | 0.958±0.005 | 0.965±0.004 | 0.966±0.004 |
| USPS | 0.917±0.01 | 0.971±0.005 | 0.969±0.005 | 0.974±0.004 | 0.973±0.005 |
| EMNIST | 0.773±0.01 | 0.896±0.008 | 0.879±0.009 | 0.891±0.008 | 0.892±0.008 |
| CIFAR10 | 0.59±0.01 | 0.67±0.01 | 0.627±0.01 | 0.634±0.02 | 0.663±0.02 |
| SVHN | 0.509±0.006 | 0.649±0.01 | 0.529±0.005 | 0.526±0.006 | 0.55±0.01 |
| flower | 0.599±0.02 | 0.662±0.02 | 0.553±0.02 | 0.629±0.02 | 0.63±0.02 |
| GTSRB | 0.586±0.01 | 0.886±0.01 | 0.782±0.01 | 0.98±0.003 | 0.98±0.003 |
| STL10 | 0.601±0.02 | 0.653±0.02 | 0.597±0.02 | 0.646±0.02 | 0.651±0.02 |
| FMNIST | 0.835±0.03 | 0.916±0.02 | 0.919±0.02 | 0.923±0.02 | 0.924±0.02 |
| Medical images | |||||
| dermamnist | 0.571±0.02 | 0.639±0.02 | 0.599±0.02 | 0.602±0.03 | 0.61±0.02 |
| pneumoniamnist | 0.812±0 | 0.931±0 | 0.921±0 | 0.937±0 | 0.871±0 |
| retinamnist | 0.547±0.04 | 0.615±0.04 | 0.592±0.03 | 0.579±0.04 | 0.577±0.03 |
| breastmnist | 0.651±0 | 0.759±0 | 0.684±0 | 0.739±0 | 0.705±0 |
| bloodmnist | 0.779±0.04 | 0.877±0.03 | 0.817±0.03 | 0.887±0.03 | 0.88±0.03 |
| organamnist | 0.812±0.03 | 0.889±0.02 | 0.889±0.02 | 0.847±0.04 | 0.798±0.05 |
| organcmnist | 0.811±0.03 | 0.874±0.02 | 0.837±0.03 | 0.809±0.04 | 0.792±0.04 |
| organsmnist | 0.685±0.02 | 0.76±0.03 | 0.74±0.03 | 0.698±0.03 | 0.684±0.04 |
| organmnist3d | 0.862±0.02 | 0.913±0.02 | 0.903±0.03 | 0.922±0.02 | 0.627±0.04 |
| nodulemnist3d | 0.695±0 | 0.769±0 | 0.743±0 | 0.694±0 | 0.622±0 |
| fracturemnist3d | 0.601±0.04 | 0.486±0.02 | 0.547±0.04 | 0.575±0.002 | 0.552±0.02 |
| adrenalmnist3d | 0.65±0 | 0.771±0 | 0.911±0 | 0.939±0 | 0.949±0 |
| vesselmnist3d | 0.56±0 | 0.669±0 | 0.693±0 | 0.623±0 | 0.638±0 |
| synapsemnist3d | 0.534±0 | 0.539±0 | 0.45±0 | 0.493±0 | NaN |
| Chemical formula | |||||
| bace | 0.619±0 | 0.704±0 | 0.684±0 | 0.569±0 | 0.607±0 |
| BBBP | 0.699±0 | 0.747±0 | 0.718±0 | 0.691±0 | 0.713±0 |
| clintox | 0.54±0 | 0.569±0 | 0.547±0 | 0.517±0 | 0.506±0 |
| HIV | 0.531±0 | 0.562±0 | 0.548±0 | 0.511±0 | 0.514±0 |
| Dataset | CDB0 | CDA | LDA | Fast SVM | SVM |
| Standard images | |||||
| MNIST | 0.888±0.01 | 0.962±0.005 | 0.956±0.006 | 0.964±0.005 | 0.965±0.005 |
| USPS | 0.907±0.01 | 0.97±0.005 | 0.968±0.007 | 0.974±0.005 | 0.973±0.005 |
| EMNIST | 0.708±0.02 | 0.884±0.01 | 0.863±0.01 | 0.878±0.01 | 0.879±0.01 |
| CIFAR10 | 0.404±0.05 | 0.562±0.03 | 0.48±0.03 | 0.491±0.05 | 0.548±0.03 |
| SVHN | 0.223±0.05 | 0.478±0.03 | 0.241±0.04 | 0.223±0.06 | 0.247±0.05 |
| flower | 0.492±0.07 | 0.597±0.05 | 0.417±0.05 | 0.545±0.05 | 0.546±0.05 |
| GTSRB | 0.296±0.03 | 0.853±0.02 | 0.762±0.03 | 0.981±0.004 | 0.982±0.004 |
| STL10 | 0.425±0.05 | 0.523±0.05 | 0.413±0.04 | 0.512±0.05 | 0.521±0.05 |
| FMNIST | 0.801±0.05 | 0.909±0.02 | 0.912±0.02 | 0.916±0.02 | 0.917±0.02 |
| Medical images | |||||
| dermamnist | 0.439±0.09 | 0.527±0.07 | 0.352±0.07 | 0.348±0.1 | 0.462±0.06 |
| pneumoniamnist | 0.837±0 | 0.932±0 | 0.908±0 | 0.94±0 | 0.868±0 |
| retinamnist | 0.377±0.1 | 0.506±0.07 | 0.444±0.08 | 0.401±0.1 | 0.407±0.1 |
| breastmnist | 0.641±0 | 0.751±0 | 0.698±0 | 0.682±0 | 0.691±0 |
| bloodmnist | 0.742±0.05 | 0.864±0.04 | 0.784±0.04 | 0.865±0.03 | 0.866±0.03 |
| organamnist | 0.771±0.05 | 0.872±0.03 | 0.868±0.03 | 0.814±0.05 | 0.717±0.08 |
| organcmnist | 0.766±0.04 | 0.848±0.03 | 0.797±0.04 | 0.76±0.05 | 0.733±0.06 |
| organsmnist | 0.58±0.06 | 0.69±0.05 | 0.653±0.04 | 0.589±0.06 | 0.533±0.08 |
| organmnist3d | 0.845±0.04 | 0.902±0.03 | 0.89±0.03 | 0.914±0.03 | 0.434±0.09 |
| nodulemnist3d | 0.707±0 | 0.773±0 | 0.693±0 | 0.636±0 | 0.651±0 |
| fracturemnist3d | 0.569±0.09 | 0.279±0.05 | 0.425±0.2 | 0.5±0.07 | 0.451±0.1 |
| adrenalmnist3d | 0.602±0 | 0.718±0 | 0.894±0 | 0.924±0 | 0.946±0 |
| vesselmnist3d | 0.547±0 | 0.615±0 | 0.58±0 | 0.43±0 | 0.405±0 |
| synapsemnist3d | 0.498±0 | 0.506±0 | 0.0612±0 | 0.189±0 | NaN |
| Chemical formula | |||||
| bace | 0.62±0 | 0.704±0 | 0.678±0 | 0.483±0 | 0.575±0 |
| BBBP | 0.693±0 | 0.715±0 | 0.603±0 | 0.553±0 | 0.66±0 |
| clintox | 0.634±0 | 0.365±0 | 0.238±0 | 0.0869±0 | 0.0868±0 |
| HIV | 0.471±0 | 0.465±0 | 0.159±0 | 0.0407±0 | 0.0473±0 |
Appendix M. Computation Conditions
Appendix N. Licensing
Appendix O. Code Availability
References
- Varoquaux, G.; Raamana, P.R.; Engemann, D.A.; Hoyos-Idrobo, A.; Schwartz, Y.; Thirion, B. Assessing and tuning brain decoders: Cross-validation, caveats, and guidelines. NeuroImage 2017, 145. [CrossRef]
- Yuan, G.X.; Ho, C.H.; Lin, C.J. Recent advances of large-scale linear classification. Proceedings of the IEEE 2012, 100. [CrossRef]
- Schulz, M.A.; Yeo, B.T.; Vogelstein, J.T.; Mourao-Miranada, J.; Kather, J.N.; Kording, K.; Richards, B.; Bzdok, D. Different scaling of linear models and deep learning in UKBiobank brain images versus machine-learning datasets. Nature Communications 2020, 11. [CrossRef]
- Varoquaux, G.; Cheplygina, V. Machine learning for medical imaging: methodological failures and recommendations for the future. npj Digital Medicine 2022, 5. [CrossRef]
- Duda, R.O.; Hart, P.E.; Stork, D.G. Pattern classification, second edition. NY: Wiley-Interscience 2001.
- Cai, D.; He, X.; Han, J. Training linear discriminant analysis in linear time. In Proceedings of the Proceedings - International Conference on Data Engineering, 2008. [CrossRef]
- Cortes, C.; Vapnik, V. Support-Vector Networks. Machine Learning 1995, 20. [CrossRef]
- Fan, R.E.; Chang, K.W.; Hsieh, C.J.; Wang, X.R.; Lin, C.J. LIBLINEAR: A library for large linear classification. Journal of Machine Learning Research 2008, 9.
- Minsky, M.L.; Papert, S. Perceptrons, Expanded Edition An Introduction to Computational Geometry; MIT Press, 1969.
- Panda, N.R. A Review on Logistic Regression in Medical Research. National Journal of Community Medicine 2022, 13, 265–270. [CrossRef]
- Fisher, R.A. The use of multiple measurements in taxonomic problems. Annals of Eugenics 1936, 7. [CrossRef]
- Wu, Y.; Cannistraci, C.V. Accuracy Score for Evaluation of Classification on Imbalanced Data, 2025. Preprints, . [CrossRef]
- Frazier, P.I. Bayesian Optimization. In Proceedings of the INFORMS TutORials in Operations Research, 2018. [CrossRef]
- Allwein, E.L.; Schapire, R.E.; Singer, Y. Reducing multiclass to binary: A unifying approach for margin classifiers. Journal of Machine Learning Research 2001, 1.
- Acevedo, A.; Duran, C.; Kuo, M.J.; Ciucci, S.; Schroeder, M.; Cannistraci, C.V. Measuring Group Separability in Geometrical Space for Evaluation of Pattern Recognition and Dimension Reduction Algorithms. IEEE Access 2022, 10, 22441–22471. [CrossRef]
- Acevedo, A.; Wu, Y.; Traversa, F.L.; Cannistraci, C.V. Geometric separability of mesoscale patterns in embedding representation and visualization of multidimensional data and complex networks. PLOS Complex Systems 2024, 1, 1–28. [CrossRef]
- Krizhevsky, A. Learning Multiple Layers of Features from Tiny Images. Technical Report TR-2009, Computer Science Department, University of Toronto, Toronto 2009.
- 10x Genomics. 1.3 Million Brain Cells from E18 Mice. https://support.10xgenomics.com/single-cell-gene-expression/datasets/1.3.0/1M_neurons, 2017.
- Coates, A.; Lee, H.; Ng, A.Y. An analysis of single-layer networks in unsupervised feature learning. In Proceedings of the Journal of Machine Learning Research, 2011, Vol. 15.
- Cohen, G.; Afshar, S.; Tapson, J.; Schaik, A.V. EMNIST: Extending MNIST to handwritten letters. In Proceedings of the Proceedings of the International Joint Conference on Neural Networks, 2017, Vol. 2017-May. [CrossRef]
- Hull, J.J. A Database for Handwritten Text Recognition Research. IEEE Transactions on Pattern Analysis and Machine Intelligence 1994, 16. [CrossRef]
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proceedings of the IEEE 1998, 86. [CrossRef]
- Netzer, Y.; Wang, T. Reading digits in natural images with unsupervised feature learning. Nips 2011.
- Nilsback, M.E.; Zisserman, A. Automated flower classification over a large number of classes. In Proceedings of the Proceedings - 6th Indian Conference on Computer Vision, Graphics and Image Processing, ICVGIP 2008, 2008. [CrossRef]
- Stallkamp, J.; Schlipsing, M.; Salmen, J.; Igel, C. The German Traffic Sign Recognition Benchmark: A multi-class classification competition. In Proceedings of the Proceedings of the International Joint Conference on Neural Networks, 2011. [CrossRef]
- Xiao, H.; Rasul, K.; Vollgraf, R. Fashion-MNIST: a Novel Image Dataset for Benchmarking Machine Learning Algorithms, 2017. arXiv.
- Yang, J.; Shi, R.; Wei, D.; Liu, Z.; Zhao, L.; Ke, B.; Pfister, H.; Ni, B. MedMNIST v2 - A large-scale lightweight benchmark for 2D and 3D biomedical image classification. Scientific Data 2023, 10. [CrossRef]
- Wu, Z.; Ramsundar, B.; Feinberg, E.N.; Gomes, J.; Geniesse, C.; Pappu, A.S.; Leswing, K.; Pande, V. MoleculeNet: A benchmark for molecular machine learning. Chemical Science 2018, 9. [CrossRef]
| Dataset | Method | AUROC | AUPR | Fscore | ACscore |
| SVHN subset (image) |
CDA | 0.615±0.02 | 0.63±0.02 | 0.619±0.02 | 0.423±0.05 |
| nCDA | 0.777±0.01 | 0.782±0.01 | 0.78±0.01 | 0.731±0.02 | |
| SVM | 0.555±0.01 | 0.568±0.007 | 0.551±0.006 | 0.273±0.05 | |
| nSVM | 0.736±0.02 | 0.776±0.009 | 0.756±0.008 | 0.654±0.03 | |
| ClinTox (chemical) |
CDA | 0.567 | 0.561 | 0.56 | 0.351 |
| nCDA | 0.625 | 0.627 | 0.627 | 0.46 | |
| SVM | 0.565 | 0.578 | 0.575 | 0.294 | |
| nSVM | 0.500 | 0.481 | 0.480 | 0.000 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




