Submitted:
21 January 2025
Posted:
23 January 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.2. Methods
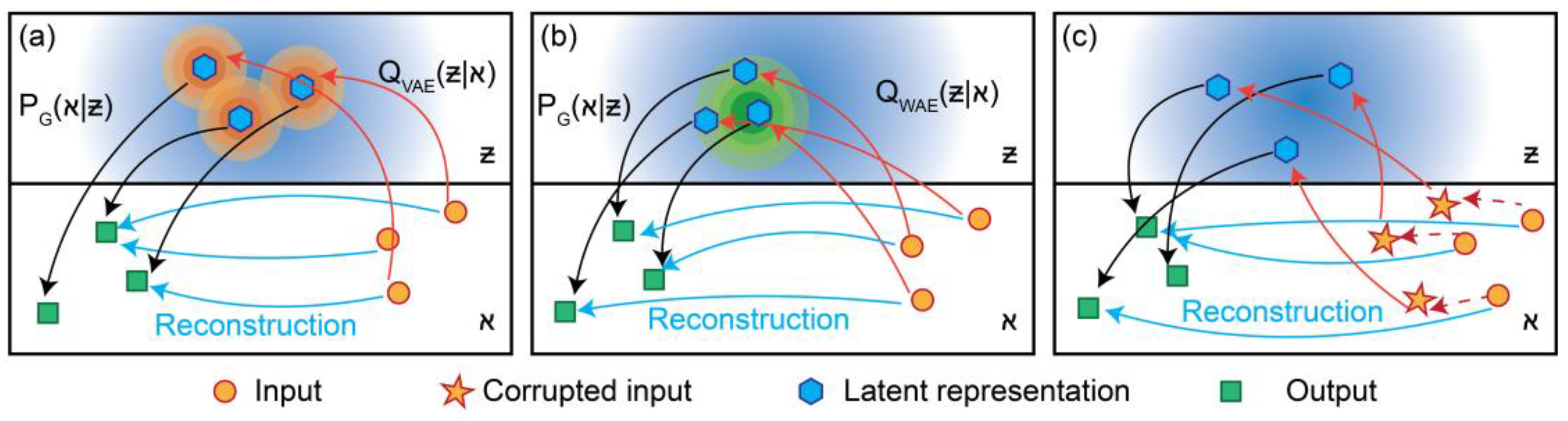
2.2.1. Variational Autoencoder (VAE) Model
2.2.2. Wasserstein Autoencoder (WAE) Model
2.2.3. Denoising Autoencoder (DAE) Model
2.3. Postprocessing and Analysis of Data
2.4. Code Availability
3. Results and Discussion
4. Conclusions
Acknowledgments
References
- Laio, A.; Parrinello, M. Escaping Free-Energy Minima. PNAS 2002, 99, 12562–12566. [Google Scholar] [CrossRef] [PubMed]
- Bussi, G.; Laio, A.; Parrinello, M. Equilibrium Free Energies from Nonequilibrium Metadynamics. Phys. Rev. Lett. 2006, 96, 090601. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Straub, J.E.; Keyes, T. Statistical-Temperature Monte Carlo and Molecular Dynamics Algorithms. Phys. Rev. Lett. 2006, 97, 050601. [Google Scholar] [CrossRef] [PubMed]
- Pang, Y.T.; Miao, Y.; Wang, Y.; McCammon, J.A. Gaussian Accelerated Molecular Dynamics in NAMD. J. Chem. Theory Comput. 2017, 13, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Bourlard, H.; Kamp, Y. Auto-Association by Multilayer Perceptrons and Singular Value Decomposition. Biol. Cybern. 1988, 59, 291–294. [Google Scholar] [CrossRef] [PubMed]
- Hinton, G.E.; Salakhutdinov, R.R. Reducing the Dimensionality of Data with Neural Networks. Science 2006, 313, 504–507. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.-J.; Zhang, N.-J.; Wei, T.; Chen, H.-F. Enhancing Conformational Sampling for Intrinsically Disordered and Ordered Proteins by Variational Autoencoder. International Journal of Molecular Sciences 2023, 24, 6896. [Google Scholar] [CrossRef] [PubMed]
- Bousquet, O.; Gelly, S.; Tolstikhin, I.; Simon-Gabriel, C.-J.; Schoelkopf, B. From Optimal Transport to Generative Modeling: The VEGAN Cookbook 2017.
- Tolstikhin, I.; Bousquet, O.; Gelly, S.; Schoelkopf, B. Wasserstein Auto-Encoders 2019.
- Vincent, P.; Larochelle, H.; Bengio, Y.; Manzagol, P.-A. Extracting and Composing Robust Features with Denoising Autoencoders. In Proceedings of the Proceedings of the 25th international conference on Machine learning - ICML ’08; ACM Press: Helsinki, Finland, 2008; pp. 1096–1103. [Google Scholar]
- Ojeda-May, P. Exploring the Dynamics of Shikimate Kinase through Molecular Mechanics. Biophysica 2022, 2, 194–202. [Google Scholar] [CrossRef]
- Kullback, S.; Leibler, R.A. On Information and Sufficiency. Ann. Math. Statist. 1951, 22, 79–86. [Google Scholar] [CrossRef]
- Gretton, A.; Borgwardt, K.M.; Rasch, M.J.; Schölkopf, B.; Smola, A. A Kernel Two-Sample Test. Journal of Machine Learning Research 2012, 13, 723–773. [Google Scholar]
- Vayer, T.; Gribonval, R. Controlling Wasserstein Distances by Kernel Norms with Application to Compressive Statistical Learning. Journal of Machine Learning Research 2023, 24, 1–51. [Google Scholar]
- Anand, N.; Achim, T. Protein Structure and Sequence Generation with Equivariant Denoising Diffusion Probabilistic Models 2022.
- Watson, J.L.; Juergens, D.; Bennett, N.R.; Trippe, B.L.; Yim, J.; Eisenach, H.E.; Ahern, W.; Borst, A.J.; Ragotte, R.J.; Milles, L.F.; et al. De Novo Design of Protein Structure and Function with RFdiffusion. Nature 2023, 620, 1089–1100. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual Molecular Dynamics. Journal of Molecular Graphics 1996, 14, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Johannissen, L.O.; Hay, S. Predicting New Protein Conformations from Molecular Dynamics Simulation Conformational Landscapes and Machine Learning. Proteins: Structure, Function, and Bioinformatics 2021, 89, 915–921. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Li, Z.; Tong, H.; Lu, Z.; Zhang, N.; Wei, T.; Chen, H.-F. Phanto-IDP: Compact Model for Precise Intrinsically Disordered Protein Backbone Generation and Enhanced Sampling. Briefings in Bioinformatics 2024, 25. [Google Scholar] [CrossRef] [PubMed]
- Yim, J.; Trippe, B.L.; Bortoli, V.D.; Mathieu, E.; Doucet, A.; Barzilay, R.; Jaakkola, T. SE(3) Diffusion Model with Application to Protein Backbone Generation 2023.
| Model | SpCC TrS | SpCC TsS | MSE TrS (Å2) | MSE TsS (Å2) |
|---|---|---|---|---|
| VAE | 0.997 | 0.992 | 0.64 | 1.60 |
| WAE | 0.996 | 0.992 | 0.69 | 1.59 |
| DAE | 0.996 | 0.993 | 0.66 | 1.49 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




