Submitted:
07 January 2025
Posted:
08 January 2025
You are already at the latest version
Abstract
Keywords:
1. Background
2. Methods
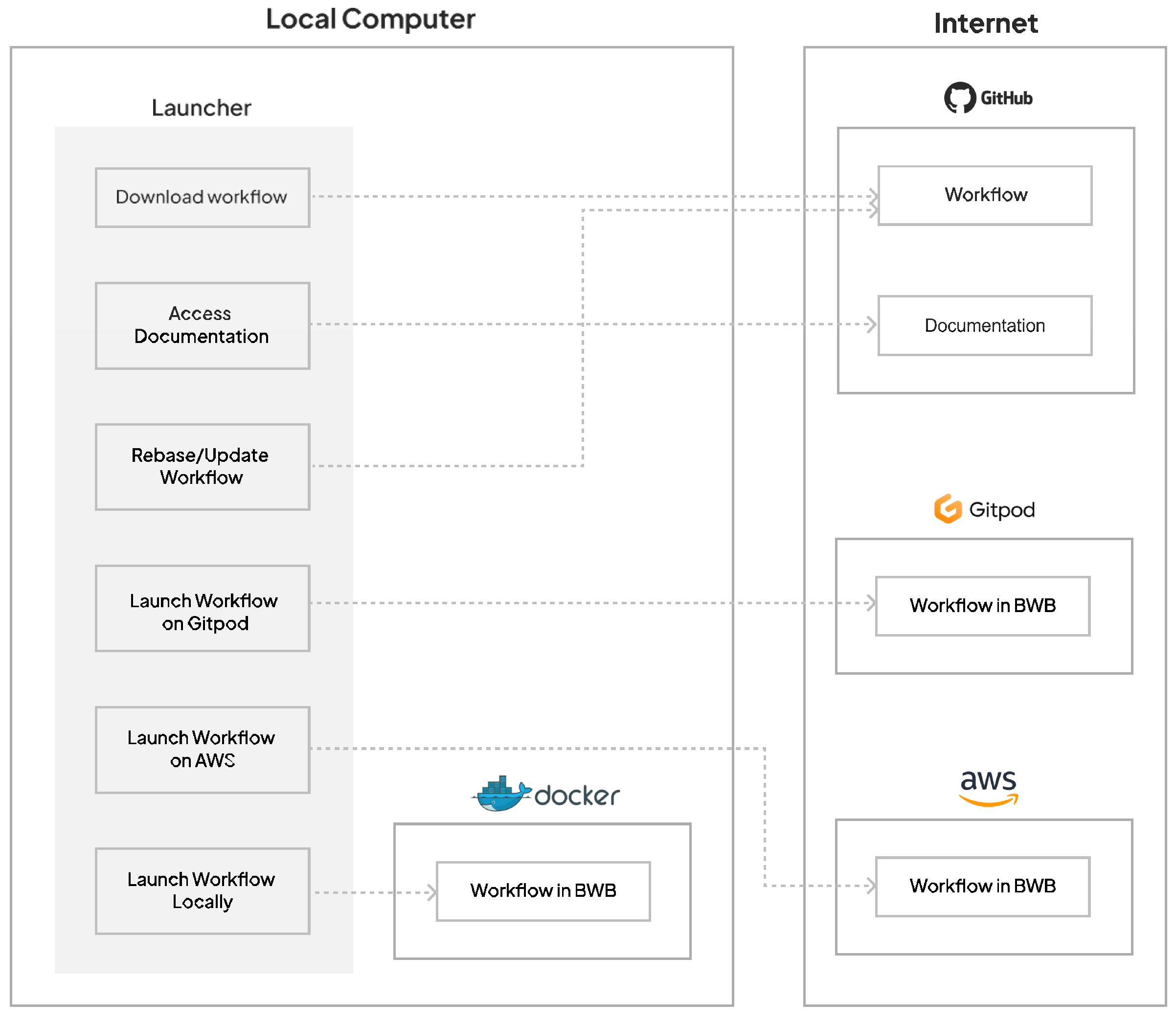
2.1. Launching Bwb and Managing Workflows
2.2. Implementation
2.2.1. Reactjs
2.2.2. Neutralinojs
2.2.3. GitHub Actions
2.2.4. Updating and Launching Workflows
2.3. Testing Across Operating Systems
3. Installation of the Biodepot Launcher Application
- Download binaries.zip from "Project home page" link in the "Availability of source code and requirements" section.
- Unzip the binaries.zip file on your machine. Inside the unzipped folder, there will be binaries following a naming convention that ends with the OS and architecture of your CPU.
- Once the binary is selected and unzipped, the binary can be run with a simple double-click.
- For macOS: When opening the binary for the first time, there will be a security prompt. While the prompt is showing, navigate to the Security and Privacy menu in the System Settings and click the "Open Anyway" button for the application. This allows the binary to be opened.
4. Usage
4.1. Starting Screen
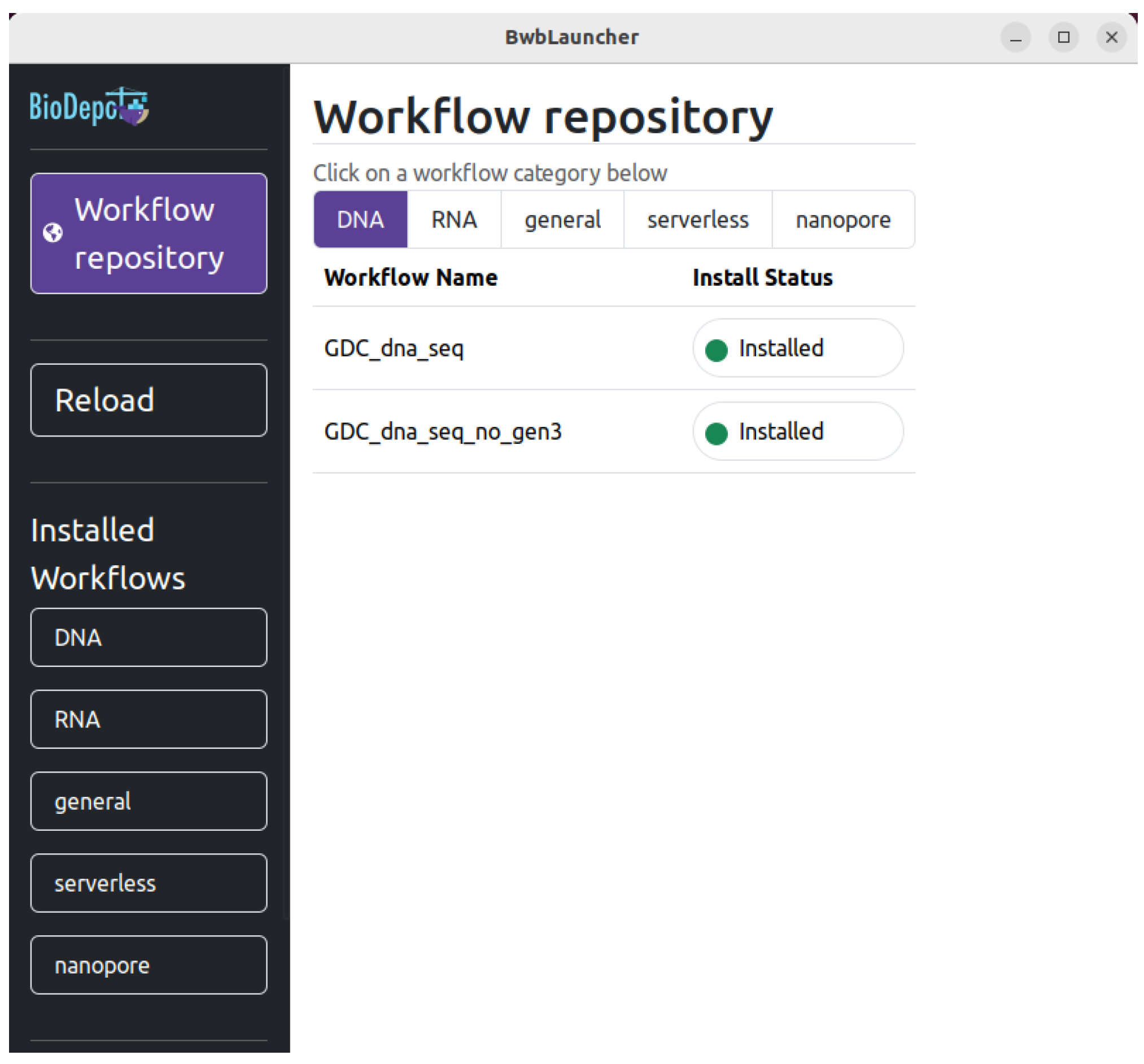
4.2. Installing Workflows
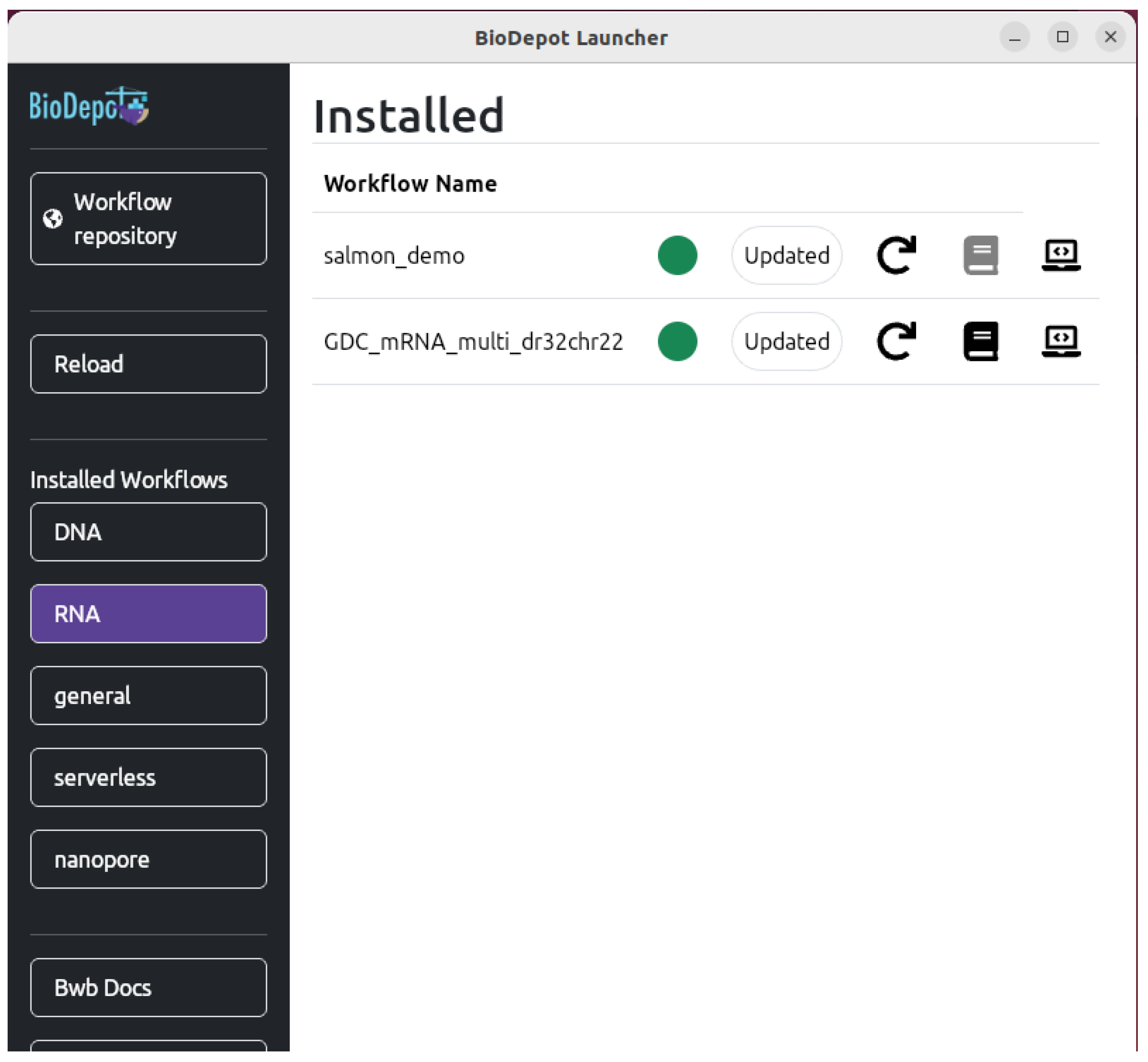
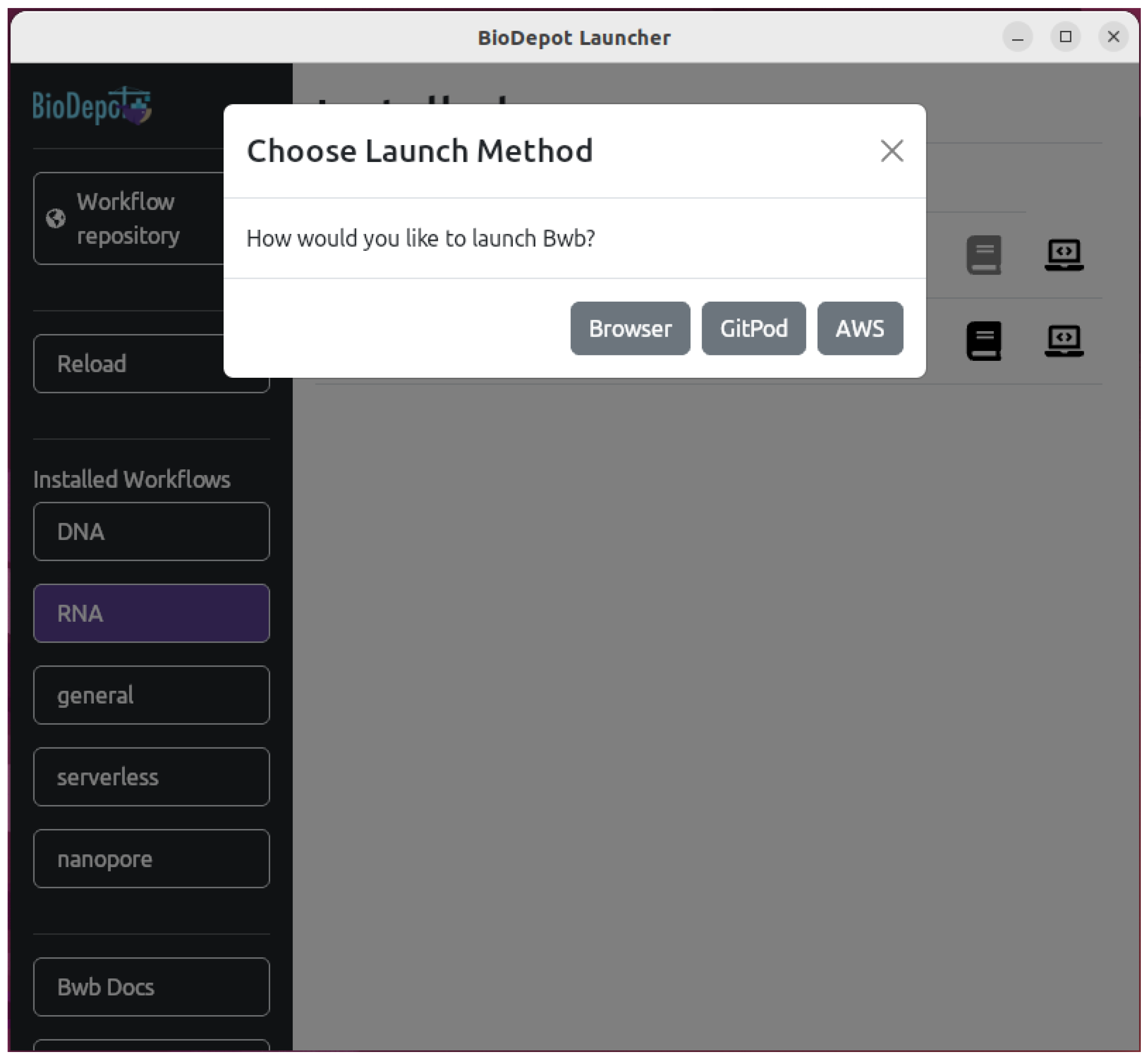
4.3. Deploying Bwb Locally
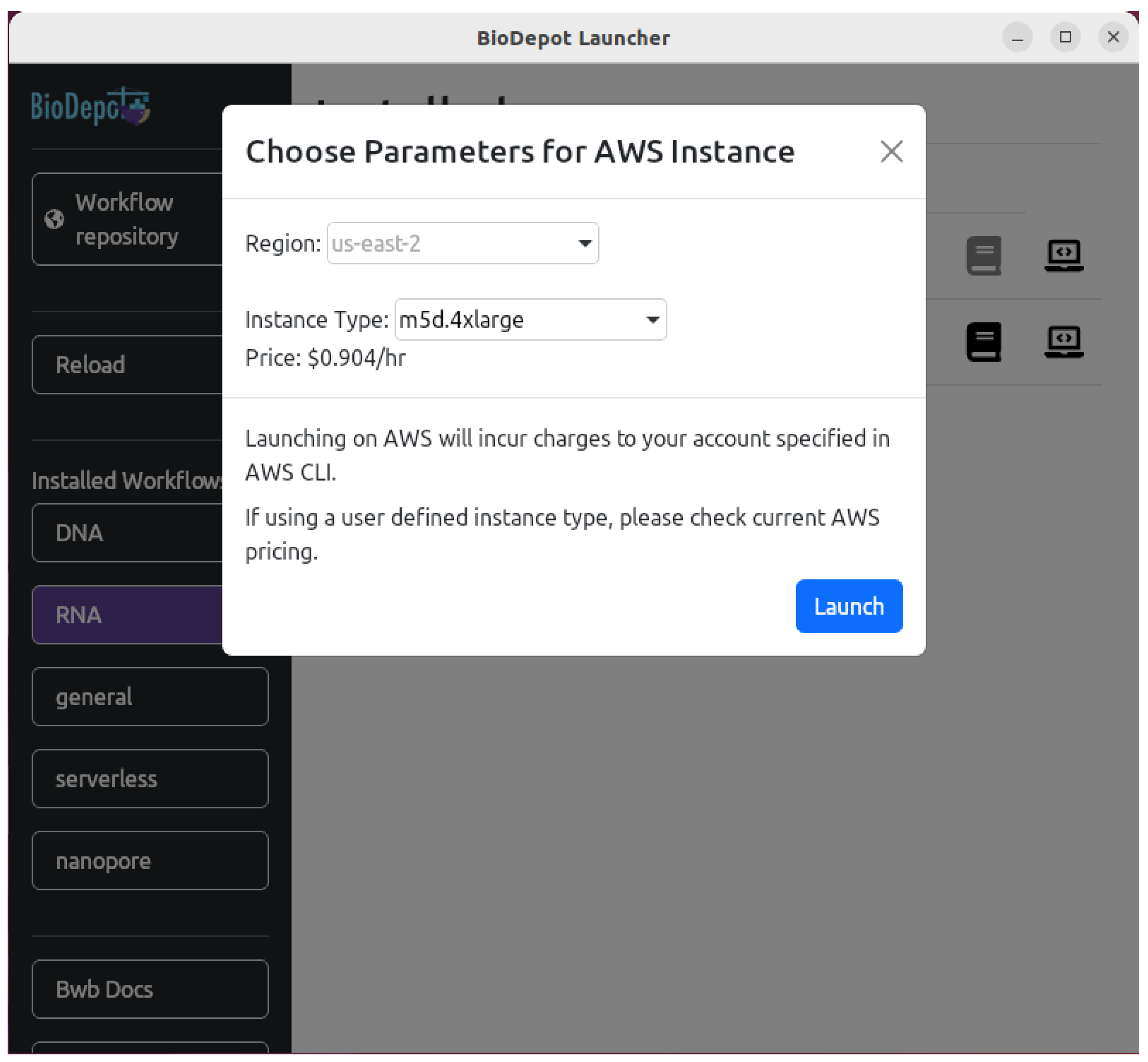
4.4. Deploying Bwb on the Cloud
5. Conclusions
6. Availability of Source Code and Requirements
- Project title: Biodepot Launcher
- Project home page: https://github.com/BioDepot/biodepot-launcher
- Training portal: https://biodepot.github.io/training/running_bwb/bwb_launcher/
- Demo video: https://www.youtube.com/watch?v=8EeKYQkQxxU
- Operating system(s): Ubuntu Linux, Windows 10/11 WSL2 Ubuntu, macOS (M-Series)
- Programming language: Javascript, React
- Other requirements: Docker, Bwb, launcher-utils, AWS CLI (Optional), Docker Machine (Optional)
- License: MIT
- Available from bio.tools with ID biotools:biodepot_launcher at https://bio.tools/biodepot_launcher and SciCrunch.org with RRID:SCR_02613.
7. Declarations
7.1. List of Abbreviations
7.2. Ethical Approval
7.3. Competing Interests
7.4. Funding
7.5. Author’s Contributions
8. Acknowledgements
References
- Hung, L.H.; Hu, J.; Meiss, T.; Ingersoll, A.; Lloyd, W.; Kristiyanto, D.; Xiong, Y.; Sobie, E.; Yeung, K.Y. Building containerized workflows using the BioDepot-workflow-builder. Cell systems 2019, 9, 508–514.
- Biodepot-workflow-builder (Bwb) GitHub repository. https://github.com/BioDepot/BioDepot-workflow-builder.
- Hung, L.H.; Fukuda, B.; Schmitz, R.; Hoang, V.; Lloyd, W.; Yeung, K.Y. Accessible, interactive and cloud-enabled genomic workflows integrated with the NCI Genomic Data Commons. bioRxiv 2022, pp. 2022–08.
- Reddy, S.; Hung, L.H.; Sala-Torra, O.; Radich, J.P.; Yeung, C.C.; Yeung, K.Y. A graphical, interactive and GPU-enabled workflow to process long-read sequencing data. BMC genomics 2021, 22, 1–8. [CrossRef]
- Yeung, C.C.; Sala-Torra, O.; Reddy, S.; Hung, L.H.; Radich, J.; Yeung, K.Y. Ultrarapid Targeted Nanopore Sequencing for Fusion Detection of Leukemias. medRxiv 2022, pp. 2022–06.
- Hung, L.H.; Straw, E.; Reddy, S.; Schmitz, R.; Colburn, Z.; Yeung, K.Y. Cloud-enabled Biodepot workflow builder integrates image processing using Fiji with reproducible data analysis using Jupyter notebooks. Scientific Reports 2022, 12, 14920. [CrossRef]
- GitPod. https://gitpod.io/.
- GitHub. https://github.com/.
- React. https://react.dev/.
- Suranga, S. Neutralino.js. https://neutralino.js.org/.
- Docker Machine: GitLab’s fork. https://gitlab-docker-machine-downloads.s3.amazonaws.com/main/index.html.
- Rancher Machine. https://github.com/rancher/machine.
- Docker. https://www.docker.com/.
- AWS CLI. https://aws.amazon.com/cli/.
- Bioinformatics pipelines: DNA-Seq analysis. https://docs.gdc.cancer.gov/Data/Bioinformatics_Pipelines/DNA_Seq_Variant_Calling_Pipeline/.
- Patro, R.; Duggal, G.; Love, M.I.; Irizarry, R.A.; Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nature methods 2017, 14, 417–419. [CrossRef]
- Bioinformatics pipelines: mRNA-Seq analysis. https://docs.gdc.cancer.gov/Data/Bioinformatics_Pipelines/Expression_mRNA_Pipeline/.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 1996 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).



