Submitted:
26 December 2024
Posted:
29 December 2024
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
1.1. The Theories of Aging
1.2. Knowledge Gaps
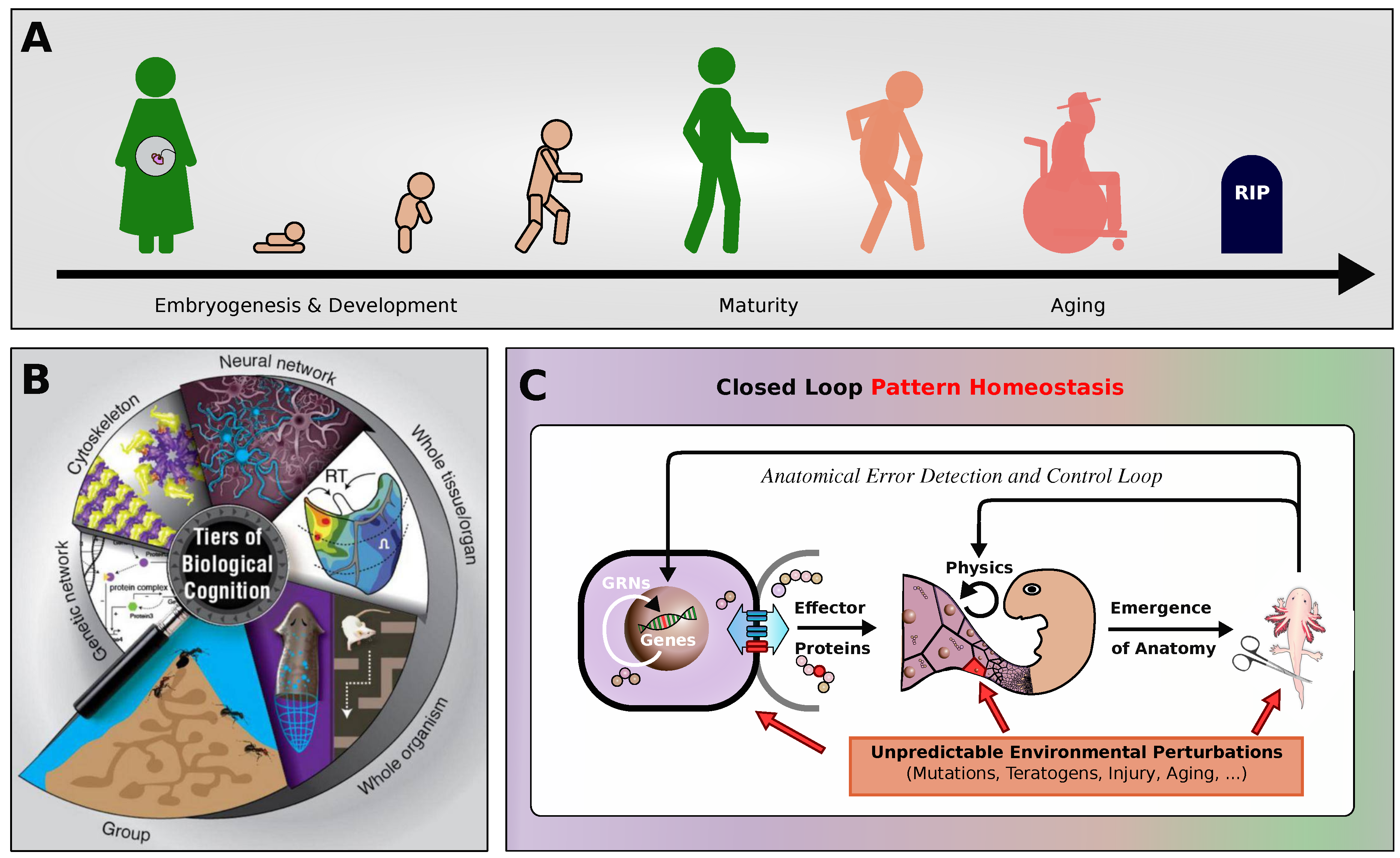
1.3. Biological Architecture of the Model
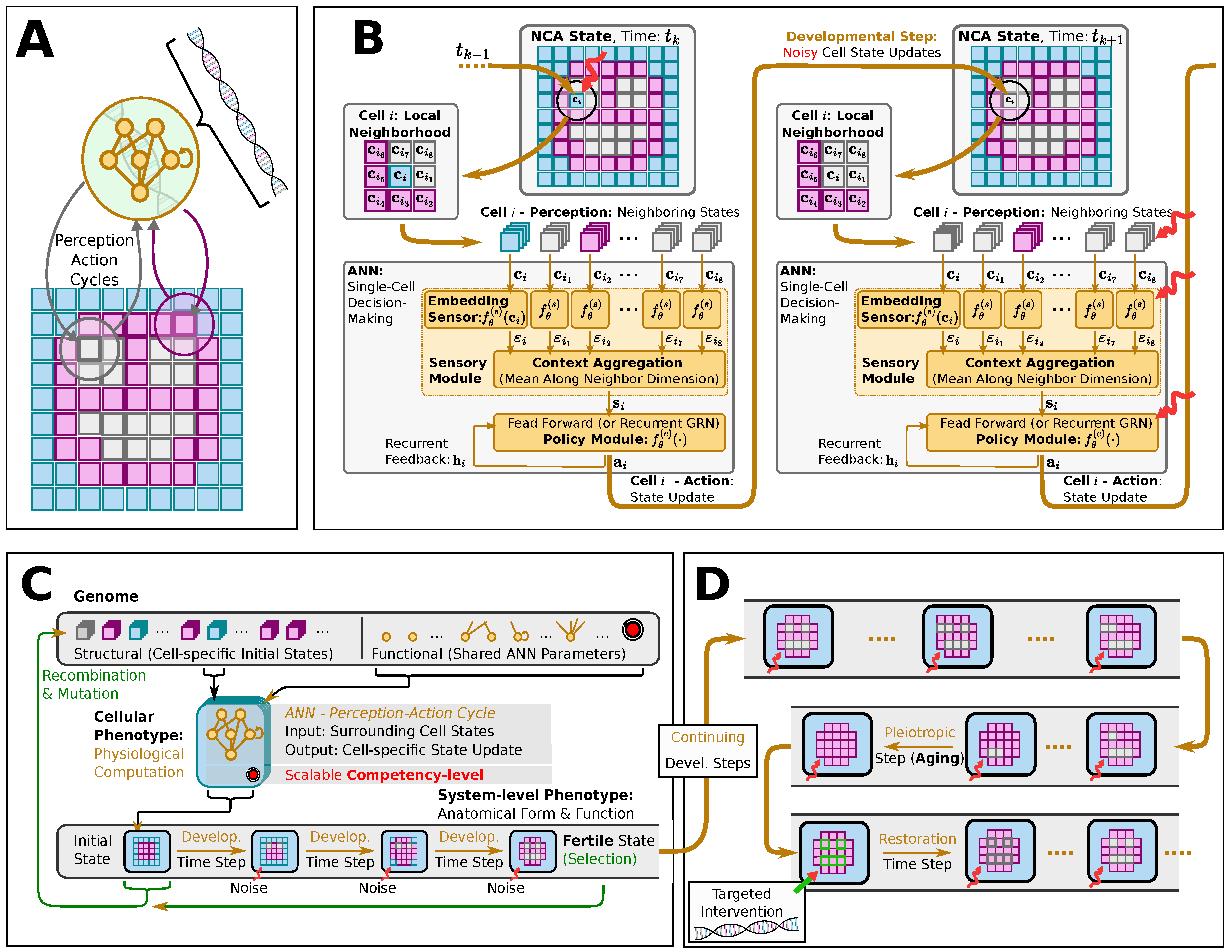
1.4. Our Computational Model
2. Material and Methods
2.1. Neural Cellular Automaton: A Multi-Agent Model for Morphogenesis, and Aging?
2.2. Neuroevolution of NCAs: an Evolutionary Algorithm Approach to Morphogenesis
2.3. Information-Theoretic Analysis: Active Information Storage, Transfer Entropy and Spatial Entropy on the NCA
- Active Information Storage (AIS) quantifies the amount of information from an agent’s past that is pertinent to predicting its future state. Specifically, AIS refers to the portion of stored information currently utilized for determining the agent’s next state [72]. Mathematically, the AIS of an agent Q is expressed as the local mutual information between its semi-infinite past as and its next state at time step :
- Transfer Entropy (TE) measures the information transferred from a source agent to a destination agent that is not contained in the past of the destination agent. We employed the local TE concept introduced by Lizier [72]. The local TE from a source agent Z to a destination agent Q is defined as the local mutual information between the previous state of the source and the next state of the destination agent , conditioned on the semi-infinite past of the destination (as ):
- Spatial Entropy (SE) measures the randomness or disorder within a spatial distribution of states. It provides insight into the complexity of spatial patterns in a system, such as an NCA with multiple states. In our study, we compute SE by evaluating the entropy of the distribution of cell states across the grid at each time step. Formally, the spatial entropy H at time step n is defined as:
3. System
4. Computational Results
4.1. Impact of Defects of Cellular Information Processing at Different Levels During Aging in a Multi-Scale Competency Architecture
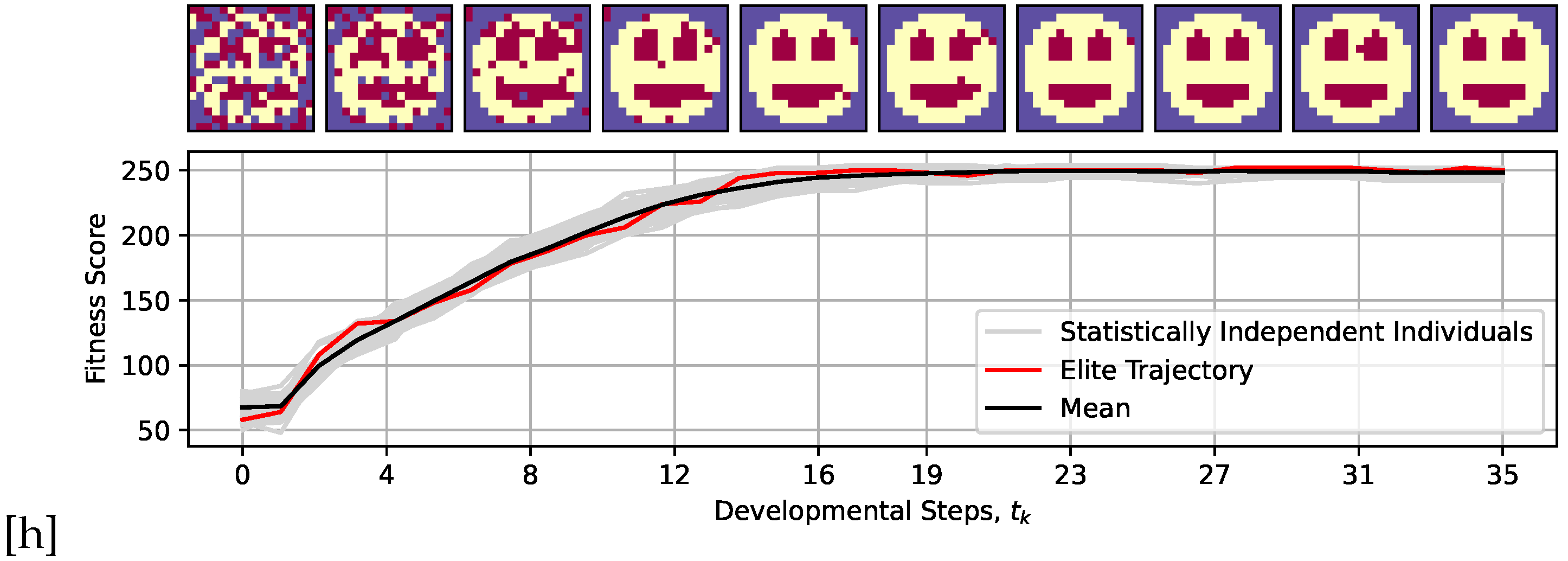
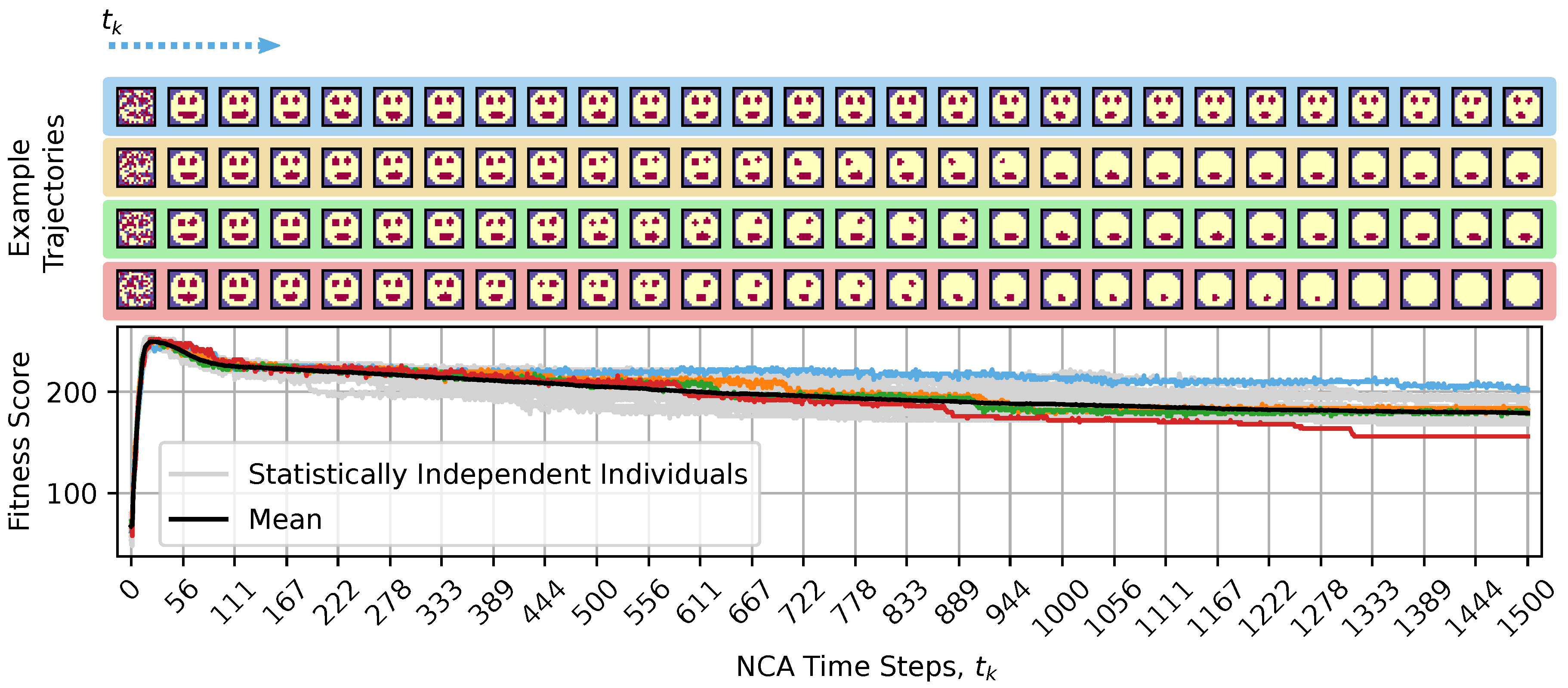
4.1.1. Aging as a Loss of Goal-Directedness: Organism Learned Development During Evolution, Not to Maintain Anatomical Homeostasis After Development
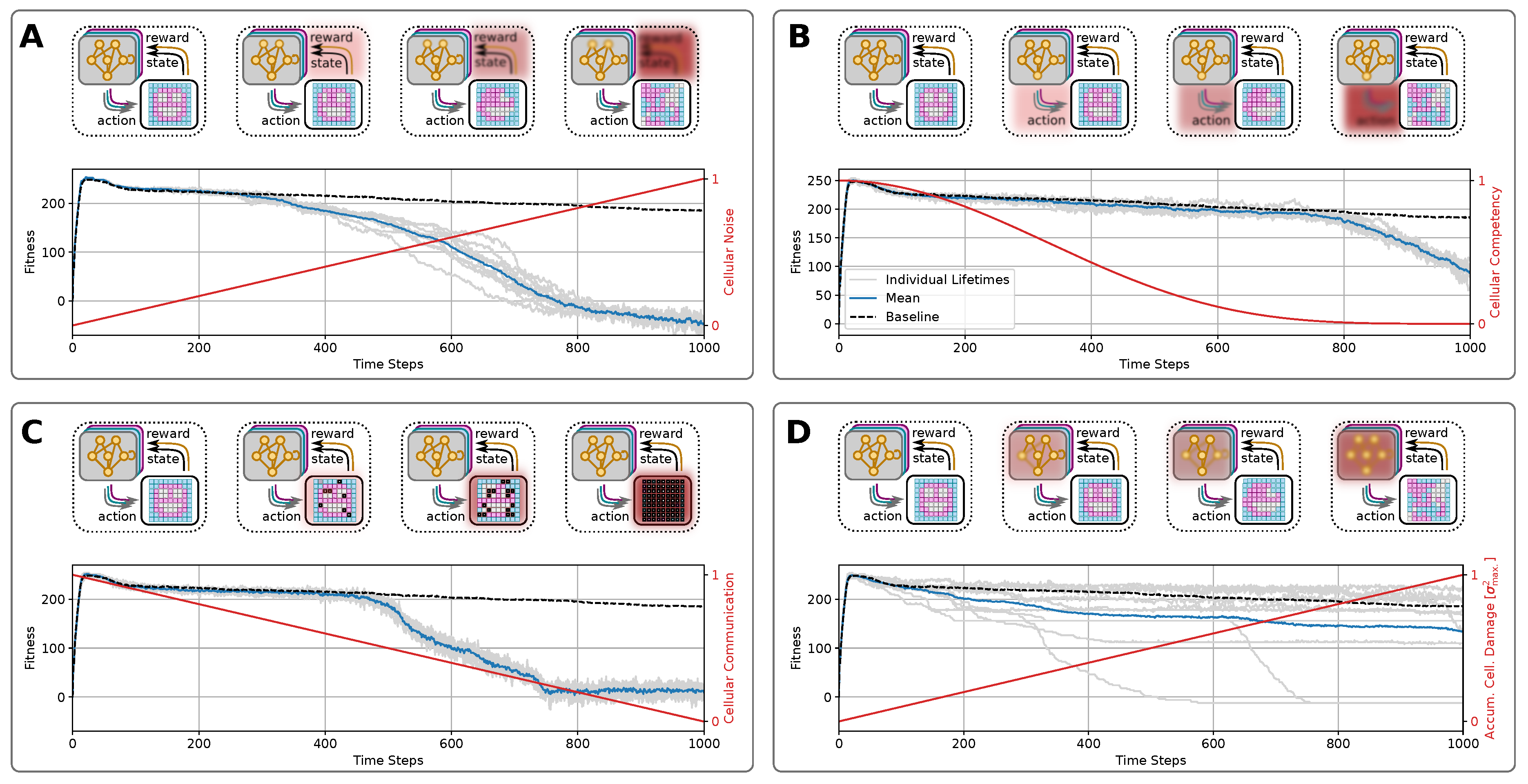
4.1.2. How Do Known Mechanisms of Aging (Defect of Cell-Cell Communication, Accumulation of Genetic Damage, Cellular Noise, Loss of Competency) Affect the Morphology in the Context of Competent Tissues
4.1.3. Cellular Noise
4.1.4. Cellular Competency
4.1.5. Cell-Cell Communication
4.1.6. Accumulation of Genetic Damage
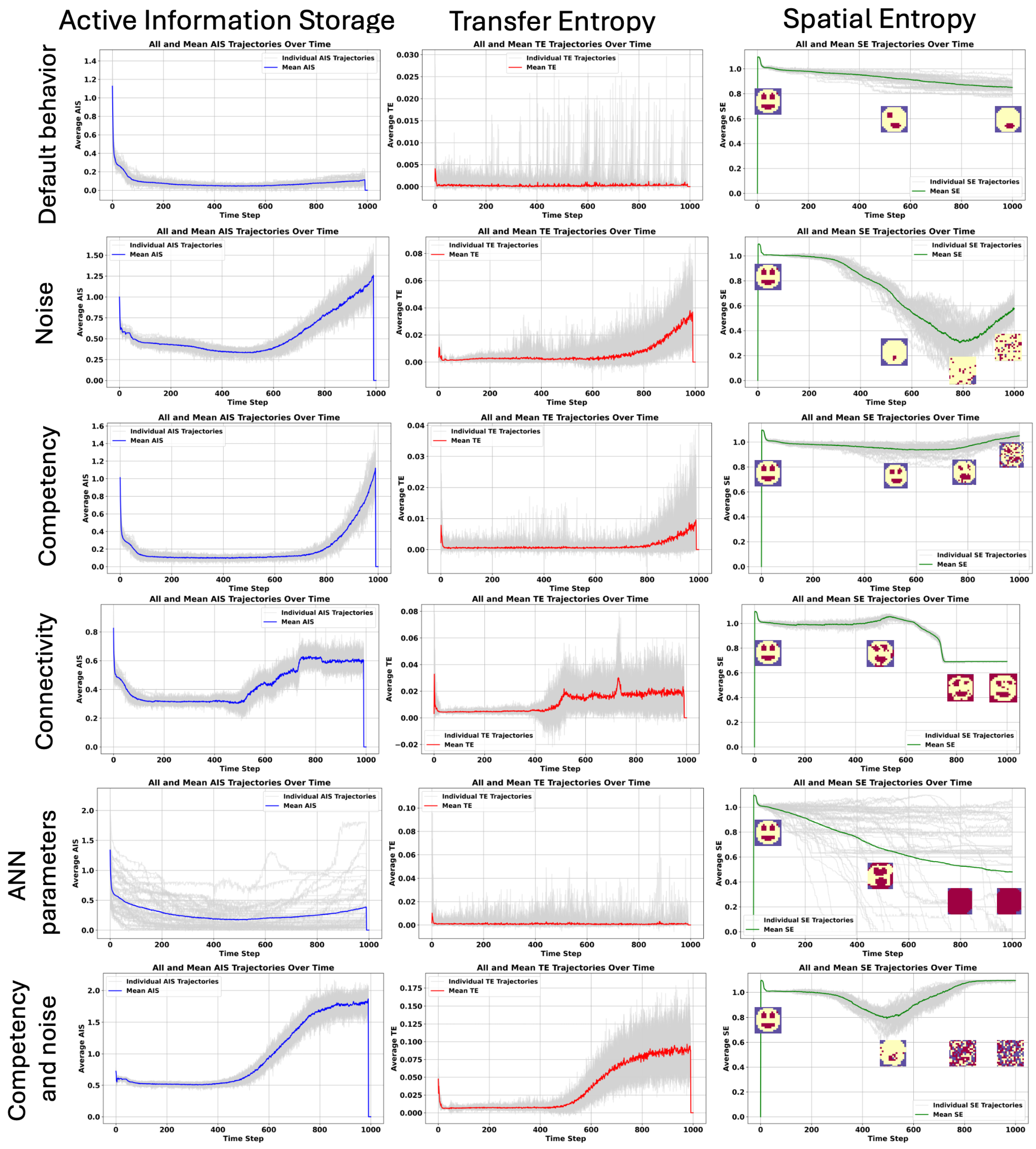
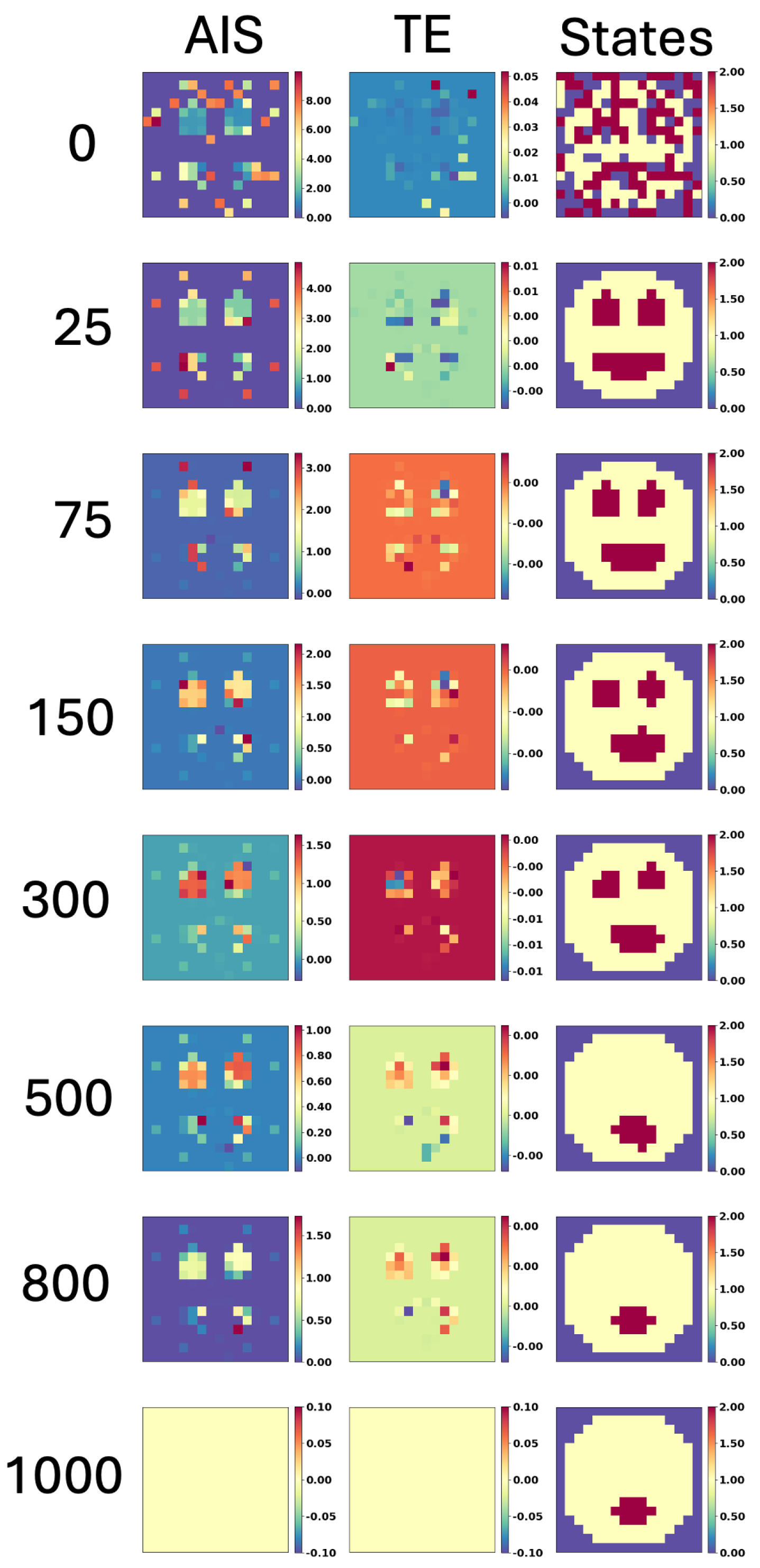
4.2. The Acceleration of Aging Is Linked to Increase in AIS and TE, While Spatial Entropy Revealed Two Different Kind of Aging: Loss of Structure and Proliferation, and Accumulation of Morphological Noise.
4.3. Regeneration as the Cure for Aging ?
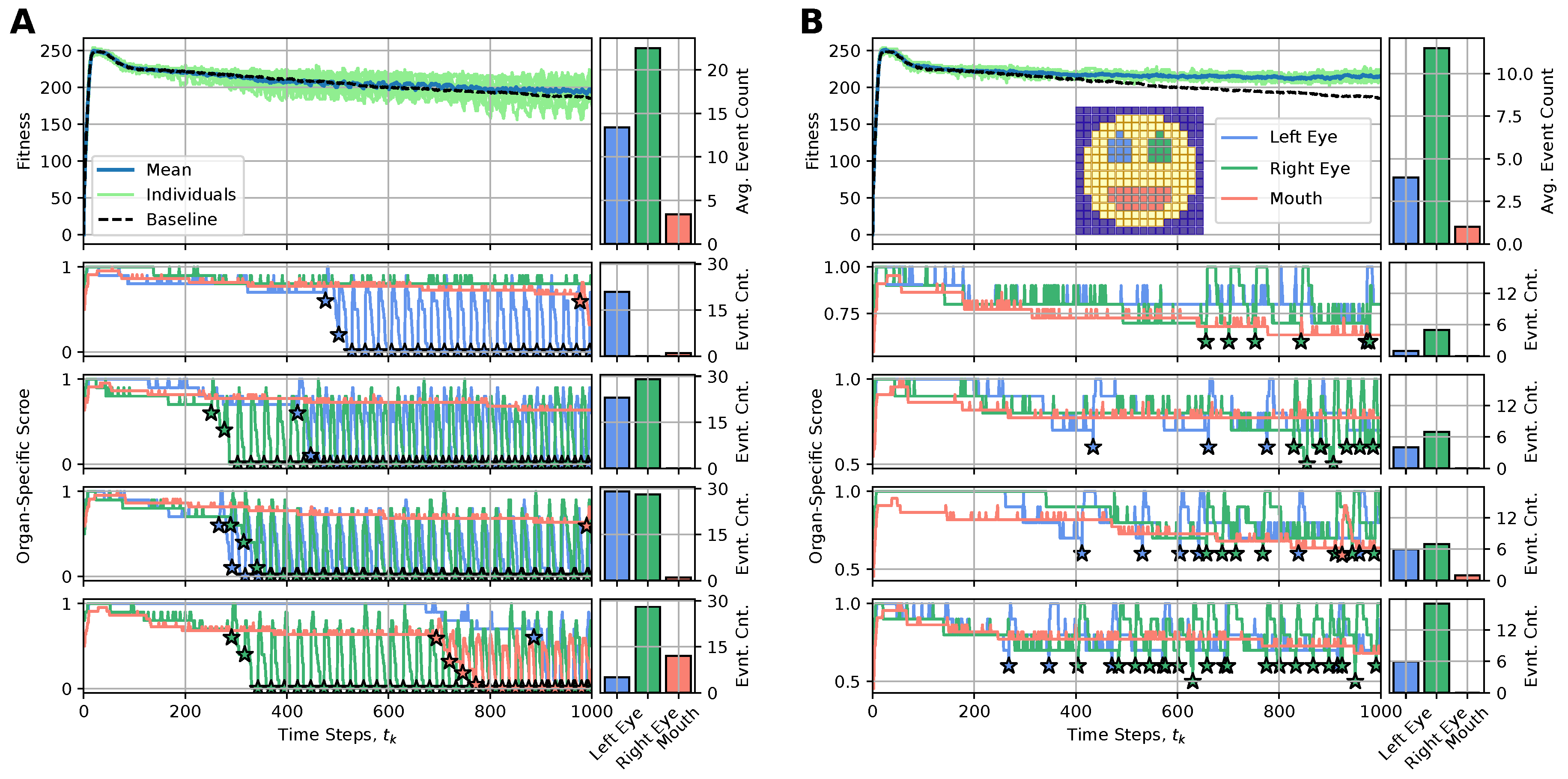
4.3.1. Loss of Organs Does Not Imply the Loss of Information About the Organ
4.3.2. Implication for Rejuvenation: A Simulated Experiment of Organ Restoration
4.3.3. Less Is More: Organ Restoration Induction by Injecting the Regenerative Information Only to Incorrectly-Positioned Cells
4.3.4. Boundaries Matter: Organ Restoration Is More Efficient with the Injection of a Differential Pattern Including the Organ Cell States and Neighboring Cell States
5. Discussion and Conclusion
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Levin, M. Darwin’s agential materials: evolutionary implications of multiscale competency in developmental biology. Cellular and Molecular Life Sciences 2023, 80, 142. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yeh, A.G.O. Neural-network-based cellular automata for simulating multiple land use changes using GIS. Int. J. Geogr. Inf. Sci. 2002, 16, 323–343. [Google Scholar] [CrossRef]
- Mordvintsev, A.; Randazzo, E.; Niklasson, E.; Levin, M. Growing Neural Cellular Automata. Distill 2020, 5. [Google Scholar] [CrossRef]
- Hartl, B.; Risi, S.; Levin, M. Evolutionary Implications of Self-Assembling Cybernetic Materials with Collective Problem-Solving Intelligence at Multiple Scales. Entropy 2024, 26. [Google Scholar] [CrossRef]
- Vandenberg, L.N.; Morrie, R.D.; Adams, D.S. V-ATPase-dependent ectodermal voltage and pH regionalization are required for craniofacial morphogenesis. Developmental Dynamics 2011, 240, 1889–1904. [Google Scholar] [CrossRef]
- de Magalhães, J.P. The biology of ageing: A primer. In Proceedings of the An Introduction to Gerontology; Stuart-Hamilton, I., Ed.; Cambridge University Press, 2011; pp. 21–47. [Google Scholar] [CrossRef]
- Partridge, L.; Deelen, J.; Slagboom, P.E. Facing up to the global challenges of ageing. Nature 2018, 561, 45–56. [Google Scholar] [CrossRef]
- Hayflick, L. Biological aging is no longer an unsolved problem. Annals of the New York academy of Sciences 2007, 1100, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Kirkwood, T.B.; Melov, S. On the programmed/non-programmed nature of ageing within the life history. Current Biology 2011, 21, R701–R707. [Google Scholar] [CrossRef]
- de Magalhães, J.P. Ageing as a software design flaw. Genome Biology 2023, 24, 1–20. [Google Scholar] [CrossRef]
- López-Otín, C.; Blasco, M.A.; Partridge, L.; Serrano, M.; Kroemer, G. Hallmarks of aging: An expanding universe. Cell 2023. [Google Scholar] [CrossRef] [PubMed]
- Gladyshev, V.N.; Kritchevsky, S.B.; Clarke, S.G.; Cuervo, A.M.; Fiehn, O.; de Magalhães, J.P.; Mau, T.; Maes, M.; Moritz, R.L.; Niedernhofer, L.J.; et al. Molecular damage in aging. Nature Aging 2021, 1, 1096–1106. [Google Scholar] [CrossRef]
- Kennedy, B.K.; Berger, S.L.; Brunet, A.; Campisi, J.; Cuervo, A.M.; Epel, E.S.; Franceschi, C.; Lithgow, G.J.; Morimoto, R.I.; Pessin, J.E.; et al. Geroscience: linking aging to chronic disease. Cell 2014, 159, 709–713. [Google Scholar] [CrossRef]
- de Magalhães, J.P.; Church, G.M. Genomes optimize reproduction: aging as a consequence of the developmental program. Physiology 2005, 20, 252–259. [Google Scholar] [CrossRef]
- Gems, D. The hyperfunction theory: an emerging paradigm for the biology of aging. Ageing research reviews 2022, 101557. [Google Scholar] [CrossRef]
- Skulachev, M.; Skulachev, V. New data on programmed aging—slow phenoptosis. Biochemistry (Moscow) 2014, 79, 977–993. [Google Scholar] [CrossRef]
- Austad, S.N.; Hoffman, J.M. Is antagonistic pleiotropy ubiquitous in aging biology? Evolution, medicine, and public health 2018, 2018, 287–294. [Google Scholar] [CrossRef]
- Blagosklonny, M.V. Answering the ultimate question “what is the proximal cause of aging?”. Aging (Albany NY) 2012, 4, 861. [Google Scholar] [CrossRef] [PubMed]
- Blagosklonny, M.V. The hyperfunction theory of aging: three common misconceptions. Oncoscience 2021, 8, 103. [Google Scholar] [CrossRef]
- Harris, A.K. The need for a concept of shape homeostasis. Biosystems 2018, 173, 65–72. [Google Scholar] [CrossRef]
- Pezzulo, G.; Levin, M. Top-down models in biology: explanation and control of complex living systems above the molecular level. Journal of The Royal Society Interface 2016, 13, 20160555. [Google Scholar] [CrossRef]
- McMillen, P.; Levin, M. Collective intelligence: A unifying concept for integrating biology across scales and substrates. Communications Biology 2024, 7, 378. [Google Scholar] [CrossRef] [PubMed]
- Levin, M. Collective Intelligence of Morphogenesis as a Teleonomic Process. In Evolution “On Purpose”: Teleonomy in Living Systems; The MIT Press, 2023; [https://direct.mit.edu/book/chapter-pdf/2155229/c011300_9780262376013.pdf]. [CrossRef]
- Pio-Lopez, L.; Levin, M. Aging as a loss of morphostatic information: a developmental bioelectricity perspective. Ageing Research Reviews 2024, 102310. [Google Scholar] [CrossRef]
- Yang, J.H.; Hayano, M.; Griffin, P.T.; Amorim, J.A.; Bonkowski, M.S.; Apostolides, J.K.; Salfati, E.L.; Blanchette, M.; Munding, E.M.; Bhakta, M.; et al. Loss of epigenetic information as a cause of mammalian aging. Cell 2023, 186, 305–326.e27. [Google Scholar] [CrossRef]
- Monzel, A.S.; Levin, M.; Picard, M. The energetics of cellular life transitions. Life metabolism 2024, 3, load051. [Google Scholar] [CrossRef] [PubMed]
- Mathews, J.; Chang, A.J.; Devlin, L.; Levin, M. Cellular signaling pathways as plastic, proto-cognitive systems: Implications for biomedicine. Patterns 2023, 4. [Google Scholar] [CrossRef]
- Levin, M. Endogenous bioelectrical networks store non-genetic patterning information during development and regeneration. The Journal of physiology 2014, 592, 2295–2305. [Google Scholar] [CrossRef]
- Noble, D. The role of stochasticity in biological communication processes. Progress in biophysics and molecular biology 2021, 162, 122–128. [Google Scholar] [CrossRef]
- Noble, D. A theory of biological relativity: no privileged level of causation. Interface focus 2012, 2, 55–64. [Google Scholar] [CrossRef] [PubMed]
- Moskalev, A.; Guvatova, Z.; Lopes, I.D.A.; Beckett, C.W.; Kennedy, B.K.; De Magalhaes, J.P.; Makarov, A.A. Targeting aging mechanisms: pharmacological perspectives. Trends in Endocrinology and Metabolism 2022, 33, 266–280. [Google Scholar] [CrossRef]
- Kenyon, C.J. The genetics of ageing. Nature 2010, 464, 504–512. [Google Scholar] [CrossRef] [PubMed]
- Pio-Lopez, L.; Levin, M. Morphoceuticals: Perspectives for discovery of drugs targeting anatomical control mechanisms in regenerative medicine, cancer and aging. Drug Discovery Today 2023, 28, 103585. [Google Scholar] [CrossRef] [PubMed]
- Levin, M. The wisdom of the body: future techniques and approaches to morphogenetic fields in regenerative medicine, developmental biology and cancer. Regenerative medicine 2011, 6, 667–673. [Google Scholar] [CrossRef] [PubMed]
- Pio-Lopez, L.; Bischof, J.; LaPalme, J.V.; Levin, M. The scaling of goals from cellular to anatomical homeostasis: an evolutionary simulation, experiment and analysis. Interface Focus 2023, 13, 20220072. [Google Scholar] [CrossRef] [PubMed]
- Grasso, C.; Bongard, J. Empowered neural cellular automata. In Proceedings of the Proceedings of the Genetic and Evolutionary Computation Conference Companion, 2022, pp. 108–111.
- Najarro, E.; Sudhakaran, S.; Glanois, C.; Risi, S. Hypernca: Growing developmental networks with neural cellular automata. arXiv 2022, arXiv:2204.11674 2022. [Google Scholar]
- Mordvintsev, A.; Randazzo, E.; Fouts, C. Growing isotropic neural cellular automata. In Proceedings of the Artificial Life Conference Proceedings 34. MIT Press One Rogers Street, Cambridge, MA 02142-1209, USA journals-info …, 2022, Vol. 2022, p. 65.
- McAuley, M.T.; Kenny, R.A.; Kirkwood, T.B.; Wilkinson, D.J.; Jones, J.J.; Miller, V.M. A mathematical model of aging-related and cortisol induced hippocampal dysfunction. BMC neuroscience 2009, 10, 1–14. [Google Scholar] [CrossRef]
- Viktorov, A.; Kholodnov, V.; Gladkikh, V.; Alekhnovich, A. Influence of environment on aging of living systems: a mathematical model. Advances in Gerontology 2013, 3, 255–260. [Google Scholar] [CrossRef]
- Xue, H.; Xian, B.; Dong, D.; Xia, K.; Zhu, S.; Zhang, Z.; Hou, L.; Zhang, Q.; Zhang, Y.; Han, J.D.J. A modular network model of aging. Molecular systems biology 2007, 3, 147. [Google Scholar] [CrossRef]
- Mc Auley, M.T.; Mooney, K.M. Computational systems biology for aging research. Interdiscip Top Gerontol 2015, 40, 35–48. [Google Scholar]
- Cavuoti, L.; Sacco, F.; Randazzo, E.; Levin, M. Adversarial takeover of neural cellular automata. In Proceedings of the ALIFE 2022: The 2022 Conference on Artificial Life. MIT Press, 2022.
- Levin, M. Technological Approach to Mind Everywhere: An Experimentally-Grounded Framework for Understanding Diverse Bodies and Minds. Frontiers in Systems Neuroscience 2022, 16. [Google Scholar] [CrossRef] [PubMed]
- Fields, C.; Levin, M. Competency in navigating arbitrary spaces as an invariant for analyzing cognition in diverse embodiments. Entropy 2022, 24, 819. [Google Scholar] [CrossRef]
- Pontes-Filho, S.; Walker, K.; Najarro, E.; Nichele, S.; Risi, S. A Unified Substrate for Body-Brain Co-evolution. In Proceedings of the From Cells to Societies: Collective Learning across Scales, 2022.
- Gawne, R.; McKenna, K.Z.; Levin, M. Competitive and coordinative interactions between body parts produce adaptive developmental outcomes. BioEssays 2020, 42, 1900245. [Google Scholar] [CrossRef] [PubMed]
- Noble, D. Genes and causation. Philosophical Transactions of the Royal Society A: Mathematical, Physical and Engineering Sciences 2008, 366, 3001–3015. [Google Scholar] [CrossRef]
- Ellis, G.F. On the nature of causation in complex systems. Transactions of the Royal Society of South Africa 2008, 63, 69–84. [Google Scholar] [CrossRef]
- Auletta, G.; Ellis, G.F.; Jaeger, L. Top-down causation by information control: from a philosophical problem to a scientific research programme. Journal of the Royal Society Interface 2008, 5, 1159–1172. [Google Scholar] [CrossRef] [PubMed]
- Walker, S.I.; Cisneros, L.; Davies, P.C. Evolutionary transitions and top-down causation. arXiv 2012, arXiv:1207.4808 2012. [Google Scholar]
- Scerri, E.R. Top-down causation regarding the chemistry–physics interface: a sceptical view. Interface Focus 2012, 2, 20–25. [Google Scholar] [CrossRef]
- Okasha, S. Emergence, hierarchy and top-down causation in evolutionary biology. Interface focus 2012, 2, 49–54. [Google Scholar] [CrossRef] [PubMed]
- Hall, D.G. Continuity and the persistence of objects: When the whole is greater than the sum of the parts. Cognitive Psychology 1998, 37, 28–59. [Google Scholar] [CrossRef]
- Lagasse, E.; Levin, M. Future medicine: from molecular pathways to the collective intelligence of the body. Trends in Molecular Medicine 2023. [Google Scholar] [CrossRef] [PubMed]
- Langton, C.G. Artificial life: An overview; Mit Press, 1997.
- Von Neumann, J.; Burks, A.W.; et al. Theory of self-reproducing automata. IEEE Transactions on Neural Networks 1966, 5, 3–14. [Google Scholar]
- Wolfram, S. A New Kind of Science; Wolfram Media, 2002.
- Cook, M. Universality in Elementary Cellular Automata. Complex Syst. 2004, 15. [Google Scholar] [CrossRef]
- Chan, B.W.C. Lenia: Biology of Artificial Life. Complex Systems 2019, 28, 251–286. [Google Scholar] [CrossRef]
- Games, M. The fantastic combinations of John Conway’s new solitaire game “life” by Martin Gardner. Sci. Am. 1970, 223, 120–123. [Google Scholar]
- Hartl, B.; Levin, M.; Zöttl, A. Neuroevolution of Decentralized Decision-Making in N-Bead Swimmers Leads to Scalable and Robust Collective Locomotion. ArXiv [in preparation] 2024. [Google Scholar]
- McMillen, P.; Levin, M. Collective intelligence: A unifying concept for integrating biology across scales and substrates. Communications Biology 2024, 7. [Google Scholar] [CrossRef]
- Mitchell, K.J.; Cheney, N. The Genomic Code: The genome instantiates a generative model of the organism. arXiv 2024, arXiv:2407.15908 2024. [Google Scholar]
- Katz, Y.; Fontana, W. Probabilistic inference with polymerizing biochemical circuits. Entropy 2022, 24, 629. [Google Scholar] [CrossRef]
- Gyurkó, D.M.; Veres, D.V.; Módos, D.; Lenti, K.; Korcsmáros, T.; Csermely, P. Adaptation and learning of molecular networks as a description of cancer development at the systems-level: potential use in anti-cancer therapies. In Proceedings of the Seminars in Cancer Biology. Elsevier, 2013, Vol. 23, pp. 262–269.
- Csermely, P.; Kunsic, N.; Mendik, P.; Kerestély, M.; Faragó, T.; Veres, D.V.; Tompa, P. Learning of signaling networks: molecular mechanisms. Trends in biochemical sciences 2020, 45, 284–294. [Google Scholar] [CrossRef] [PubMed]
- Herrera-Delgado, E.; Perez-Carrasco, R.; Briscoe, J.; Sollich, P. Memory functions reveal structural properties of gene regulatory networks. PLoS computational biology 2018, 14, e1006003. [Google Scholar] [CrossRef]
- Gabalda-Sagarra, M.; Carey, L.B.; Garcia-Ojalvo, J. Recurrence-based information processing in gene regulatory networks. Chaos: An Interdisciplinary Journal of Nonlinear Science 2018, 28. [Google Scholar] [CrossRef]
- Hansen, N.; Ostermeier, A. Completely Derandomized Self-Adaptation in Evolution Strategies. Evolutionary Computation 2001, 9, 159–195. [Google Scholar] [CrossRef] [PubMed]
- Vandenberg, L.N.; Morrie, R.D.; Adams, D.S. V-ATPase-dependent ectodermal voltage and ph regionalization are required for craniofacial morphogenesis. Developmental Dynamics 2011, 240, 1889–1904. [Google Scholar] [CrossRef]
- Lizier, J.T.; Prokopenko, M.; Zomaya, A.Y. Local measures of information storage in complex distributed computation. Information Sciences 2012, 208, 39–54. [Google Scholar] [CrossRef]
- Schreiber, T. Measuring information transfer. Physical review letters 2000, 85, 461. [Google Scholar] [CrossRef] [PubMed]
- Sutton, R.S.; Barto, A.G. Reinforcement Learning: An Introduction; The MIT Press, 2018.
- Tang, Y.; Ha, D. The Sensory Neuron as a Transformer: Permutation-Invariant Neural Networks for Reinforcement Learning. In Proceedings of the Advances in Neural Information Processing Systems; Beygelzimer, A.; Dauphin, Y.; Liang, P.; Vaughan, J.W., Eds., 2021.
- Rumelhart, D.E.; Hinton, G.E.; Williams, R.J., Learning internal representations by error propagation. In Parallel Distributed Processing: Explorations in the Microstructure of Cognition, Vol. 1: Foundations; MIT Press: Cambridge, MA, USA, 1986; p. 318–362.
- Hiscock, T.W. Adapting machine-learning algorithms to design gene circuits. BMC Bioinformatics 2019, 20, 214. [Google Scholar] [CrossRef]
- MacKay, D.J.; Mac Kay, D.J.; et al. Information theory, inference and learning algorithms; Cambridge university press, 2003.
- Aasen, T.; Hodgins, M.B.; Edward, M.; Graham, S.V. The relationship between connexins, gap junctions, tissue architecture and tumour invasion, as studied in a novel in vitro model of HPV-16-associated cervical cancer progression. Oncogene 2003, 22, 7969–7980. [Google Scholar] [CrossRef]
- Leithe, E.; Sirnes, S.; Omori, Y.; Rivedal, E. Downregulation of gap junctions in cancer cells. Critical Reviews™ in Oncogenesis 2006, 12. [Google Scholar] [CrossRef] [PubMed]
- Levin, M. Bioelectrical approaches to cancer as a problem of the scaling of the cellular self. Progress in biophysics and molecular biology 2021, 165, 102–113. [Google Scholar] [CrossRef]
- Turner, B.M. Epigenetic responses to environmental change and their evolutionary implications. Philosophical Transactions of the Royal Society B: Biological Sciences 2009, 364, 3403–3418. [Google Scholar] [CrossRef]
- Maden, M. Axolotl/newt. Molecular Embryology: Methods and Protocols 1999, pp. 415–428.
- Haluza, Y.; Zoller, J.A.; Lu, A.T.; Walters, H.E.; Lachnit, M.; Lowe, R.; Haghani, A.; Brooke, R.T.; Park, N.; Yun, M.H.; et al. Axolotl epigenetic clocks offer insights into the nature of negligible senescence. BioRxiv 2024, pp. 2024–09.
- Mašek, J.; Andersson, E.R. The developmental biology of genetic Notch disorders. Development 2017, 144, 1743–1763. [Google Scholar] [CrossRef] [PubMed]
- Pai, V.P.; Lemire, J.M.; Paré, J.F.; Lin, G.; Chen, Y.; Levin, M. Endogenous gradients of resting potential instructively pattern embryonic neural tissue via notch signaling and regulation of proliferation. Journal of Neuroscience 2015, 35, 4366–4385. [Google Scholar] [CrossRef]
- Pai, V.P.; Levin, M. HCN2 channel-induced rescue of brain, eye, heart and gut teratogenesis caused by nicotine, ethanol and aberrant notch signalling. Wound Repair and Regeneration 2022, 30, 681–706. [Google Scholar] [CrossRef]
- Levin, M.; Selberg, J.; Rolandi, M. Endogenous bioelectrics in development, cancer, and regeneration: drugs and bioelectronic devices as electroceuticals for regenerative medicine. Iscience 2019, 22, 519–533. [Google Scholar] [CrossRef]
- Whited, J.L.; Levin, M. Bioelectrical controls of morphogenesis: from ancient mechanisms of cell coordination to biomedical opportunities. Current opinion in genetics & development 2019, 57, 61–69. [Google Scholar]
- Chen, L.; Zaharia, M.; Zou, J. How is ChatGPT’s behavior changing over time? arXiv 2023, arXiv:2307.09009 2023. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



