Submitted:
22 December 2024
Posted:
23 December 2024
You are already at the latest version
Abstract
Keywords:
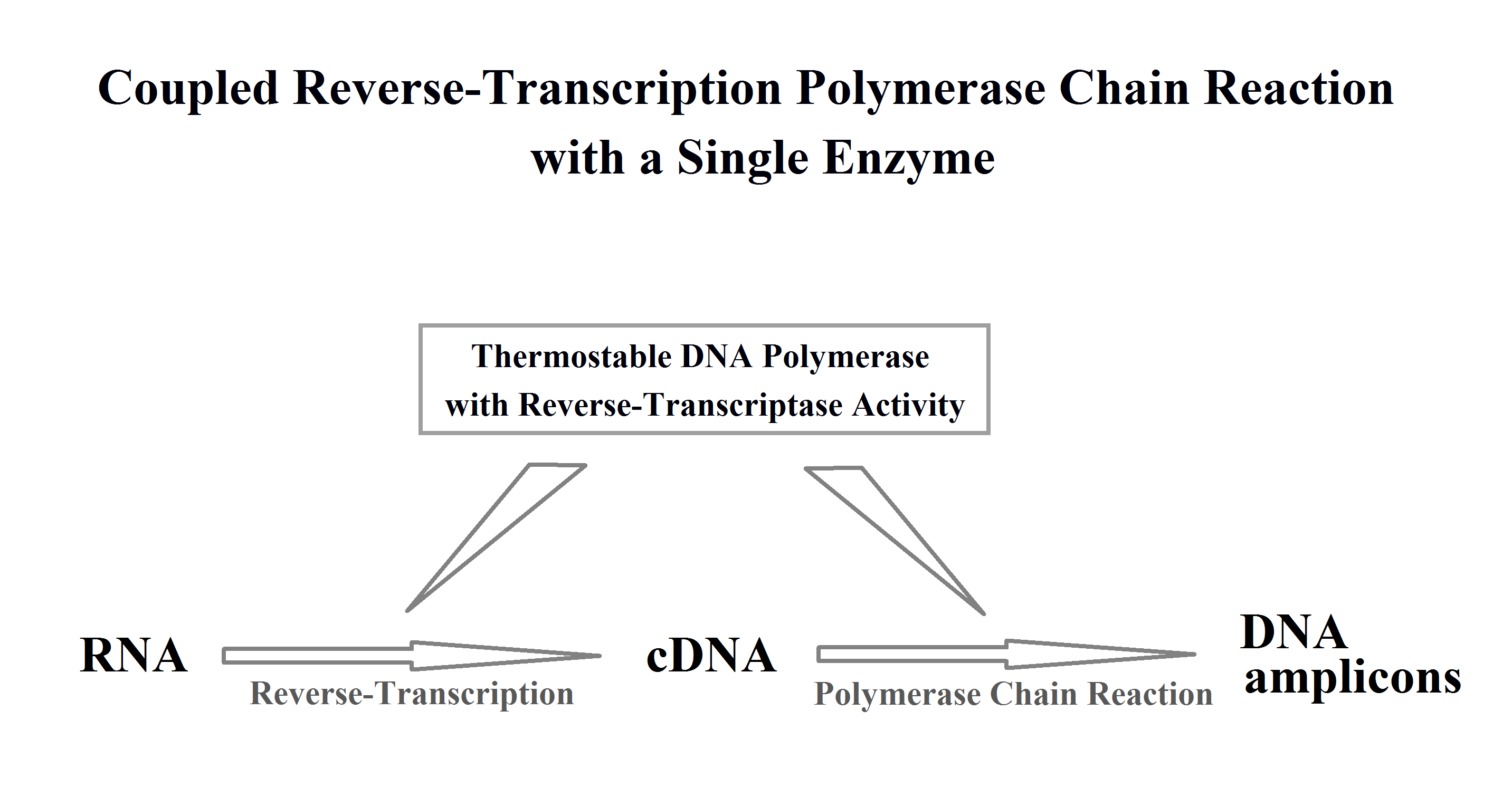
1. Introduction
2. Materials and Methods
2.1. Enzymes and Reagents
2.2. Coupled End-Point RT-PCR
2.3. Coupled Real-Time RT-qPCR with an Intercalating Dye
2.4. Coupled Real-Time RT-qPCR with a TaqMan Probe
2.5. Coupled Real-Time RT-qPCR with a TaqMan Probe
3. Results
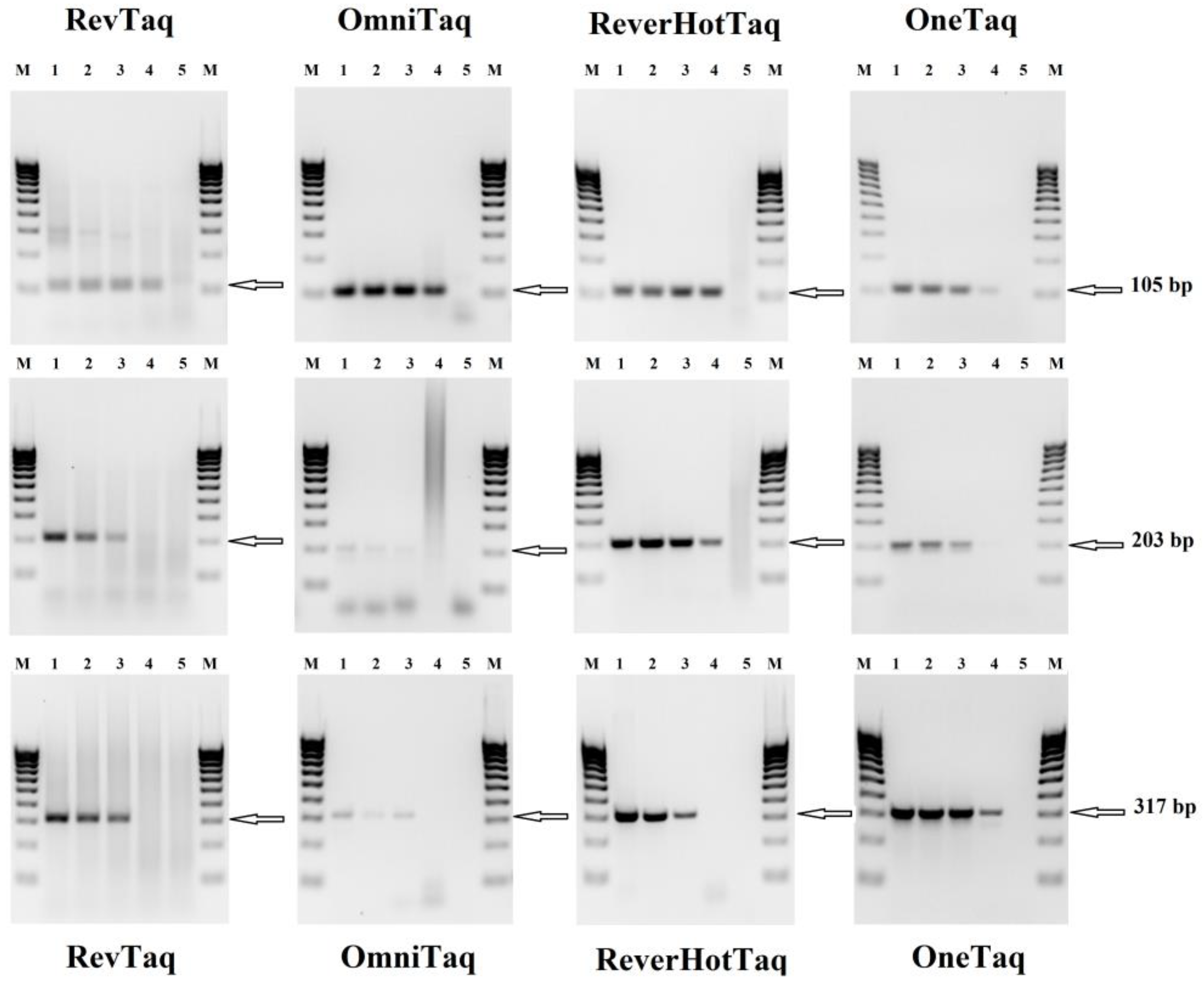
3.1. End-point RT-PCR
3.2. Real-Time RT-qPCR with an Intercalating Dye
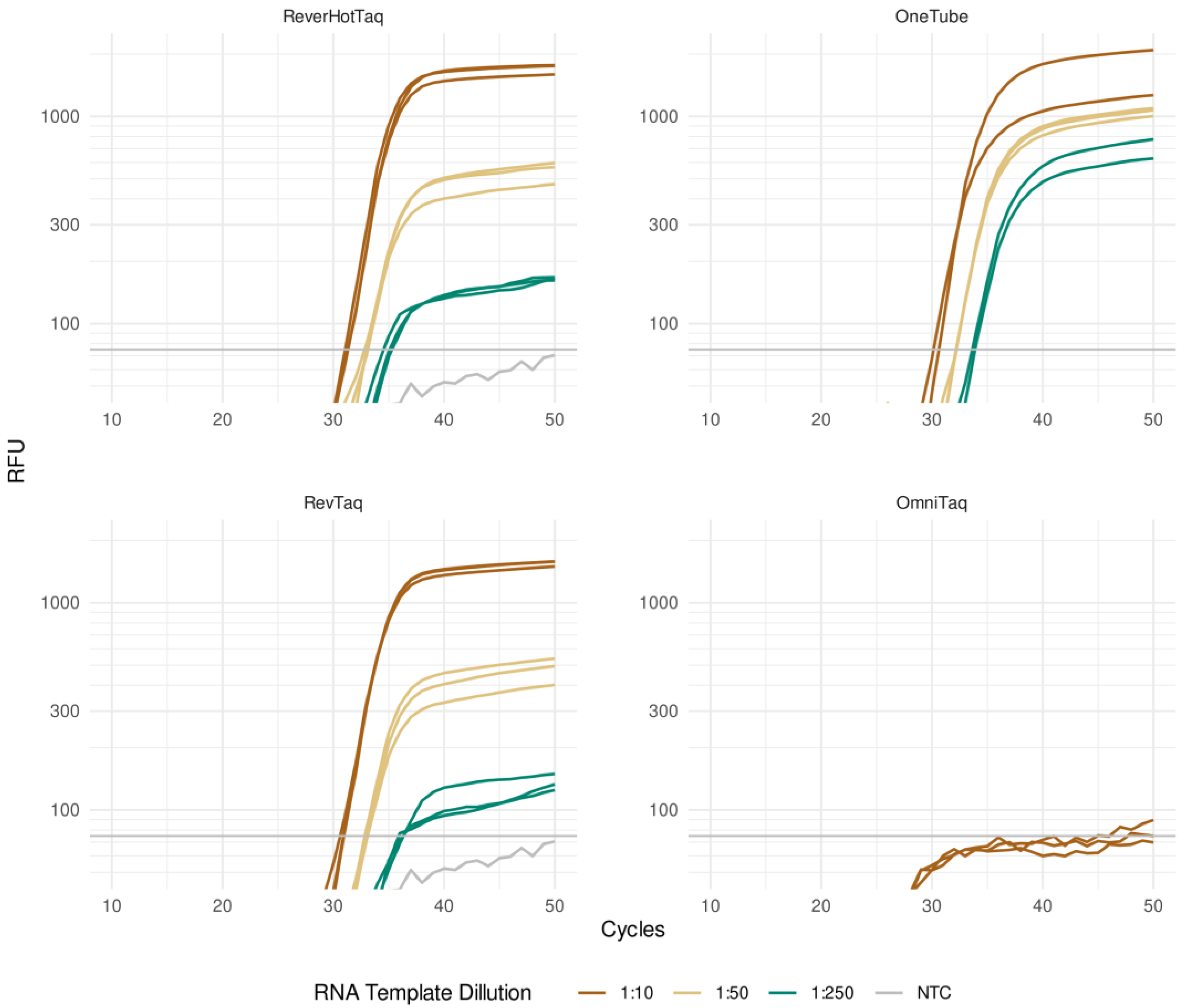
3.3. Real-Time RT-qPCR with a TaqMan Probe
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Varlamov, D.A.; Blagodatskikh, K.A.; Smirnova, E.V.; Kramarov, V.M.; Ignatov, K.B. Combinations of PCR and Isothermal Amplification Techniques Are Suitable for Fast and Sensitive Detection of SARS-CoV-2 Viral RNA. Front. Bioeng. Biotechnol. 2020, 8, 604793. [CrossRef]
- Garafutdinov, R.R.; Mavzyutov, A.R.; Alekseev, Ya.I.; Vorobev, A.A.; Nikonorov, Yu.M.; Chubukova, O.V.; Matniyazov, R.T.; Baymiev, An.Kh.; Maksimov, I.V.; Kuluev, B.R.; et al. Human Betacoronaviruses and Their Highly Sensitive Detection by PCR and Other Amplification Methods. Bmcs 2020, 12, 121–179. [CrossRef]
- Udugama, B.; Kadhiresan, P.; Kozlowski, H.N.; Malekjahani, A.; Osborne, M.; Li, V.Y.C.; Chen, H.; Mubareka, S.; Gubbay, J.B.; Chan, W.C.W. Diagnosing COVID-19: The Disease and Tools for Detection. ACS Nano 2020, 14, 3822–3835. [CrossRef]
- Ignatov, K.B.; Barsova, E.V.; Fradkov, A.F.; Blagodatskikh, K.A.; Kramarova, T.V.; Kramarov, V.M. A Strong Strand Displacement Activity of Thermostable DNA Polymerase Markedly Improves the Results of DNA Amplification. BioTechniques 2014, 57, 81–87. [CrossRef]
- Sellner, L.N.; Coelen, R.J.; Mackenzie, J.S. Reverse Transcriptase Inhibits Taq Polymerase Activity. Nucl Acids Res 1992, 20, 1487–1490. [CrossRef]
- Fehlmann, C.; Krapf, R.; Solioz, M. Reverse Transcriptase Can Block Polymerase Chain Reaction. Clinical Chemistry 1993, 39, 368–369. [CrossRef]
- Chumakov, K.M. Reverse Transcriptase Can Inhibit PCR and Stimulate Primer-Dimer Formation. Genome Research 1994, 4, 62–64. [CrossRef]
- Suslov, O. PCR Inhibition by Reverse Transcriptase Leads to an Overestimation of Amplification Efficiency. Nucleic Acids Research 2005, 33, e181–e181. [CrossRef]
- Tse, W.T.; Forget, B.G. Reverse Transcription and Direct Amplification of Cellular RNA Transcripts by Taq Polymerase. Gene 1990, 88, 293–296. [CrossRef]
- Myers, T.W.; Gelfand, D.H. Reverse Transcription and DNA Amplification by a Thermus Thermophilus DNA Polymerase. Biochemistry 1991, 30, 7661–7666. [CrossRef]
- Grebennikova, T.V.; Glukhov, A.I.; Chistiakova, L.G.; Kiselev, V.I.; Severin, E.S. [Use of thermostable DNA polymerase from Thermus thermophilus KTP in a combined reverse transcription and amplification reaction of detecting interleukin 2alpha RNA and determining expression of the multidrug resistance gene (MDR-1)]. Mol Biol (Mosk) 1995, 29, 930–941.
- Grabko, V.I.; Chistyakova, L.G.; Lyapustin, V.N.; Korobko, V.G.; Miroshnikov, A.I. Reverse Transcription, Amplification and Sequencing of Poliovirus RNA by Taq DNA Polymerase. FEBS Letters 1996, 387, 189–192. [CrossRef]
- Ong, J.L.; Loakes, D.; Jaroslawski, S.; Too, K.; Holliger, P. Directed Evolution of DNA Polymerase, RNA Polymerase and Reverse Transcriptase Activity in a Single Polypeptide. Journal of Molecular Biology 2006, 361, 537–550. [CrossRef]
- Vichier-Guerre, S.; Ferris, S.; Auberger, N.; Mahiddine, K.; Jestin, J. A Population of Thermostable Reverse Transcriptases Evolved from Thermus Aquaticus DNA Polymerase I by Phage Display. Angew Chem Int Ed 2006, 45, 6133–6137. [CrossRef]
- Sauter, K.B.M.; Marx, A. Evolving Thermostable Reverse Transcriptase Activity in a DNA Polymerase Scaffold. Angew Chem Int Ed 2006, 45, 7633–7635. [CrossRef]
- Kranaster, R.; Drum, M.; Engel, N.; Weidmann, M.; Hufert, F.T.; Marx, A. One-step RNA Pathogen Detection with Reverse Transcriptase Activity of a Mutated Thermostable Thermus Aquaticus DNA Polymerase. Biotechnology Journal 2010, 5, 224–231. [CrossRef]
- Blatter, N.; Bergen, K.; Nolte, O.; Welte, W.; Diederichs, K.; Mayer, J.; Wieland, M.; Marx, A. Structure and Function of an RNA-Reading Thermostable DNA Polymerase. Angew Chem Int Ed 2013, 52, 11935–11939. [CrossRef]
- Kuiper, J.W.P.; Baade, T.; Kremer, M.; Kranaster, R.; Irmisch, L.; Schuchmann, M.; Zander, J.; Marx, A.; Hauck, C.R. Detection of SARS-CoV-2 from Raw Patient Samples by Coupled High Temperature Reverse Transcription and Amplification. PLoS ONE 2020, 15, e0241740. [CrossRef]
- Babler, K.M.; Sharkey, M.E.; Abelson, S.; Amirali, A.; Benitez, A.; Cosculluela, G.A.; Grills, G.S.; Kumar, N.; Laine, J.; Lamar, W.; et al. Degradation Rates Influence the Ability of Composite Samples to Represent 24-Hourly Means of SARS-CoV-2 and Other Microbiological Target Measures in Wastewater. Science of The Total Environment 2023, 867, 161423. [CrossRef]
- Heller, R.C.; Chung, S.; Crissy, K.; Dumas, K.; Schuster, D.; Schoenfeld, T.W. Engineering of a Thermostable Viral Polymerase Using Metagenome-Derived Diversity for Highly Sensitive and Specific RT-PCR. Nucleic Acids Research 2019, 47, 3619–3630. [CrossRef]
- Barnes, W.M.; Zhang, Z.; Kermekchiev, M.B. A Single Amino Acid Change to Taq DNA Polymerase Enables Faster PCR, Reverse Transcription and Strand-Displacement. Front. Bioeng. Biotechnol. 2021, 8, 553474. [CrossRef]
- Whiting, S.H.; Champoux, J.J. Strand Displacement Synthesis Capability of Moloney Murine Leukemia Virus Reverse Transcriptase. J Virol 1994, 68, 4747–4758. [CrossRef]
- Luo, Z.; Xue, Y.; Chen, X.; Zhang, J.; Lu, C. Manganese-Independent Reverse Transcriptase Activity of Tth DNAPolymerase with Two Amino Acid Substitutions. PPL 2023, 30, 193–200. [CrossRef]
- .
- Rödiger, S.; Burdukiewicz, M.; Spiess, A.-N.; Blagodatskikh, K. Enabling Reproducible Real-Time Quantitative PCR Research: The RDML Package. Bioinformatics 2017, 33, 4012–4014. [CrossRef]
- Rödiger, S.; Burdukiewicz, M.; Schierack, P. chipPCR: An R Package to Pre-Process Raw Data of Amplification Curves. Bioinformatics 2015, 31, 2900–2902. [CrossRef]
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Use R!; 2nd ed. 2016.; Springer International Publishing : Imprint: Springer: Cham, 2016; ISBN 978-3-319-24277-4.
- Balaji, S.; Vanniarajan, A. Implication of Pseudo Reference Genes in Normalization of Data from Reverse Transcription-Quantitative PCR. Gene 2020, 757, 144948. [CrossRef]
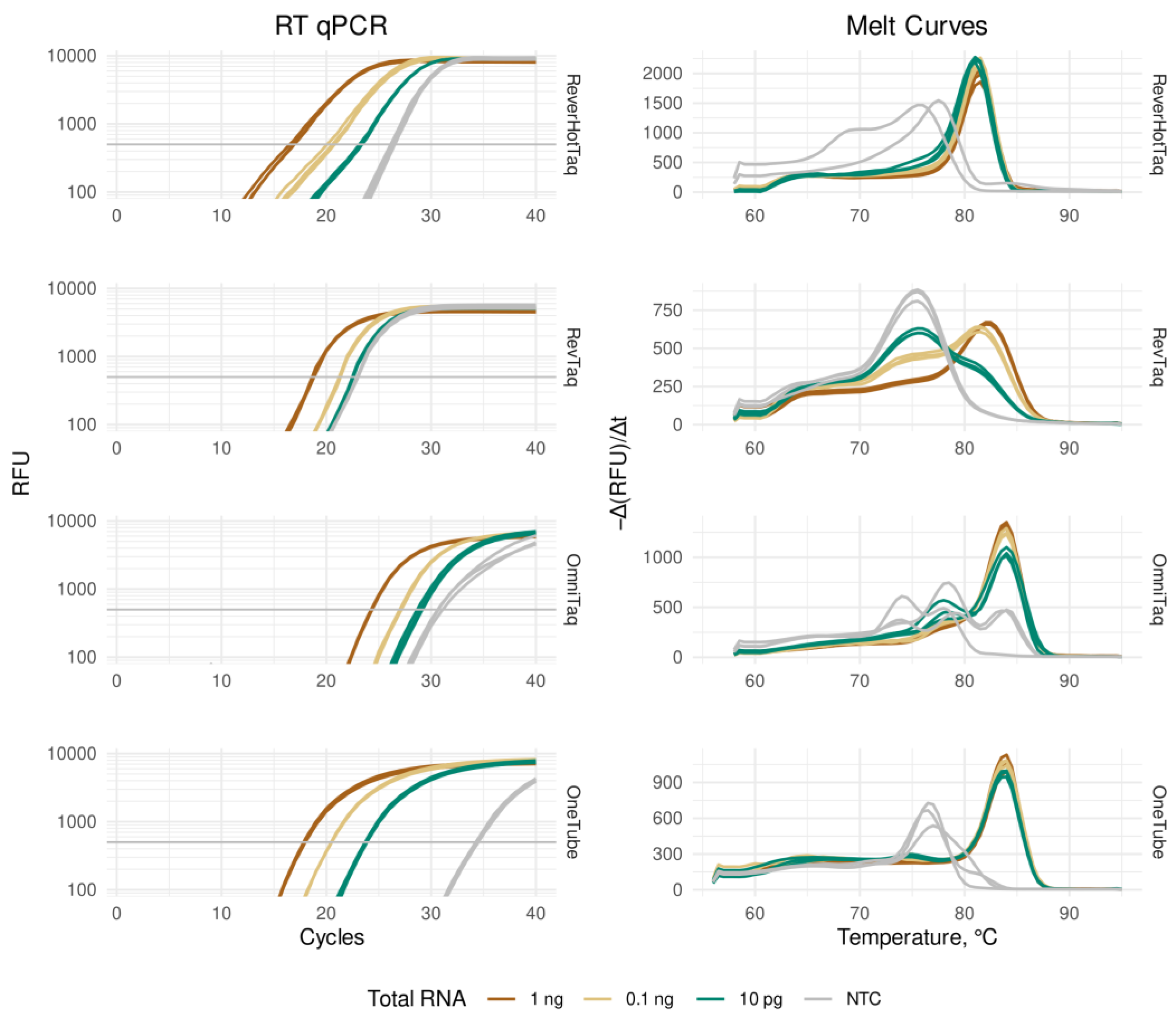
| (A) RT-qPCR of GAPDH mRNA with EvaGreen intercalating dye | ||||
| Human total RNA per reaction | 1 ng | 0.1 ng | 0.01 ng | NTC |
| ReverHotTaq | Cq 16.52 ± 0.25 ∆Cq ‒1.12 |
Cq 20.22 ± 0.32 ∆Cq 0.04 |
Cq 22.98 ± 0.09 ∆Cq −0.46 |
Cq 25.96 ± 0.15 ∆Cq −8.14 |
| RevTaq | Cq 18.34 ± 0.07 ∆Cq 0.69 |
Cq 20.89 ± 0.05 ∆Cq 0.71 |
Cq 22.12 ± 0.05 ∆Cq −1.33 |
Cq 22.53 ± 0.17 ∆Cq −11.58 |
| OmniTaq2 | Cq 24.01 ± 0.04 ∆Cq 6.36 |
Cq 26.81 ± 0.06 ∆Cq 6.63 |
Cq 28.65 ± 0.28 ∆Cq 5.19 |
Cq 30.61 ± 0.30 ∆Cq −3.51 |
| OneTube RT-PCRMix | Cq 17.65 ± 0.11 ∆Cq 0 |
Cq 20.18 ± 0.03 ∆Cq 0 |
Cq 23.45 ± 0.11 ∆Cq 0 |
Cq 34.11 ± 0.12 ∆Cq 0 |
| (B) RT-qPCR of SARS-CoV-2 viral RNA with TaqMan probe | ||||
| RNA template dilution | 1:10 | 1:50 | 1:250 | NTC |
| ReverHotTaq | Cq 31.51 ± 0.12 ∆Cq 1.01 |
Cq 32.89 ± 0.14 ∆Cq 0.65 |
Cq 34.94 ± 0.44 ∆Cq 1.24 |
N/A1 |
| RevTaq | Cq 31.15 ± 0.15 ∆Cq 0.65 |
Cq 32.83 ± 0.06 ∆Cq 0.59 |
Cq 36.72 ± 0.45 ∆Cq 3.02 |
N/A1 |
| OmniTaq2 | N/A | N/A | N/A | N/A1 |
| OneTube RT-PCRMix | Cq 30.5 ± 0.49 ∆Cq 0 |
Cq 32.24 ± 0.06 ∆Cq 0 |
Cq 33.7 ± 0.17 ∆Cq 0 |
N/A1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




