Submitted:
09 December 2024
Posted:
10 December 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Patient Characteristics
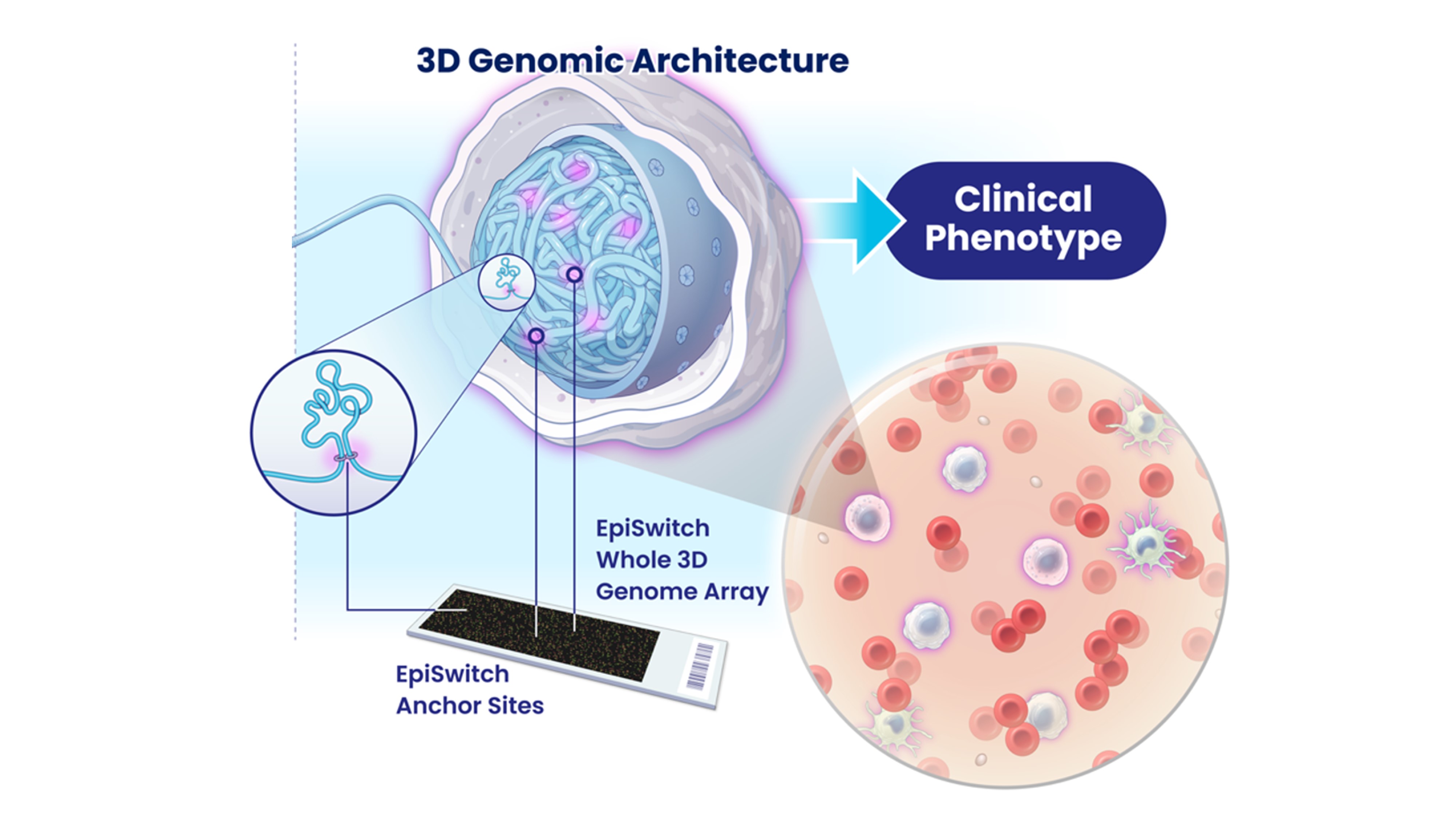
Preparation of 3D Genomic Templates
Custom Microarray Design
Microarray Statistical Analysis
Translation of Array-Based 3D Genomic Markers to PCR Readouts
EpiSwitch® PCR
PCR Statistical Analysis
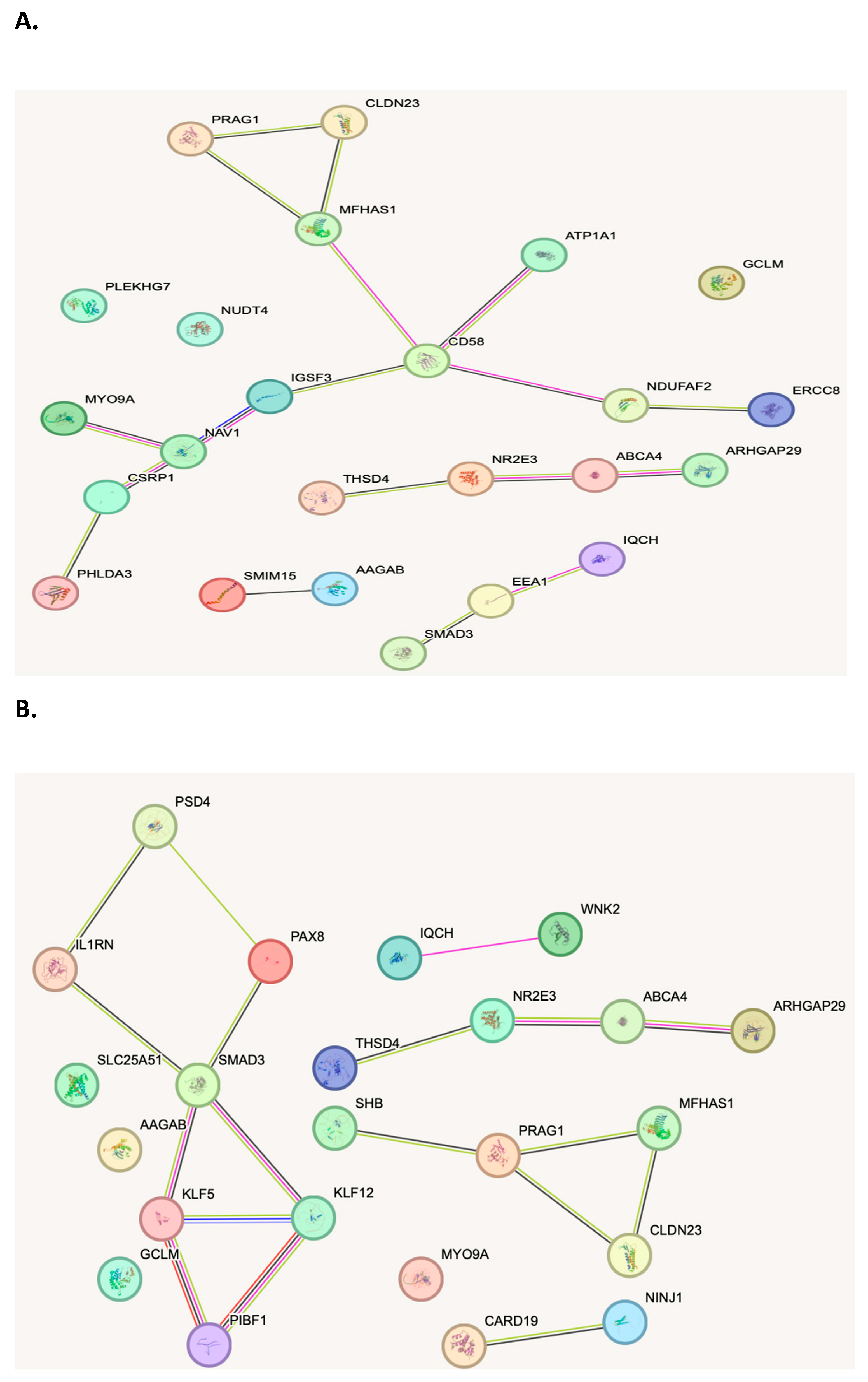
Biological Network/Pathway Analysis
Results
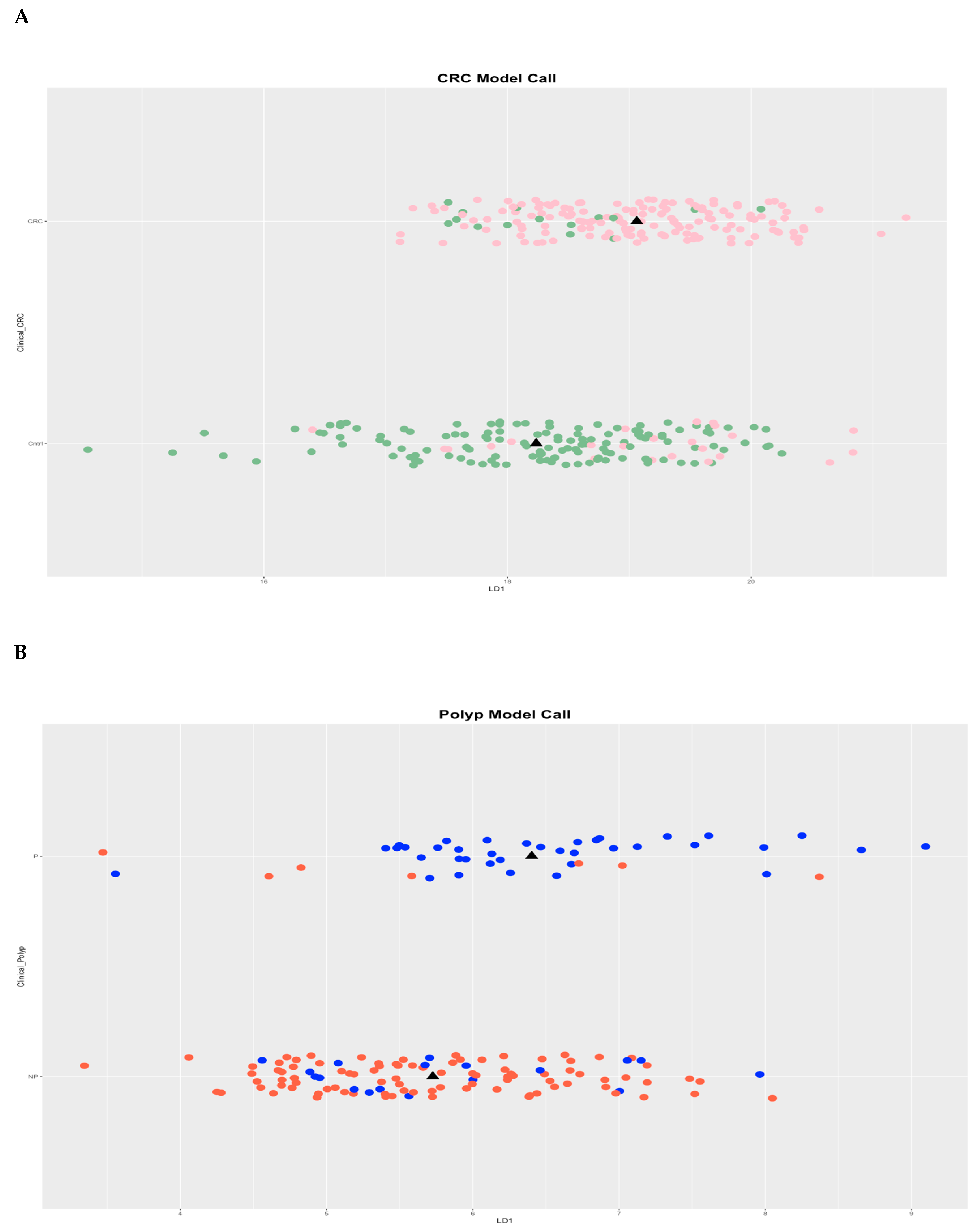
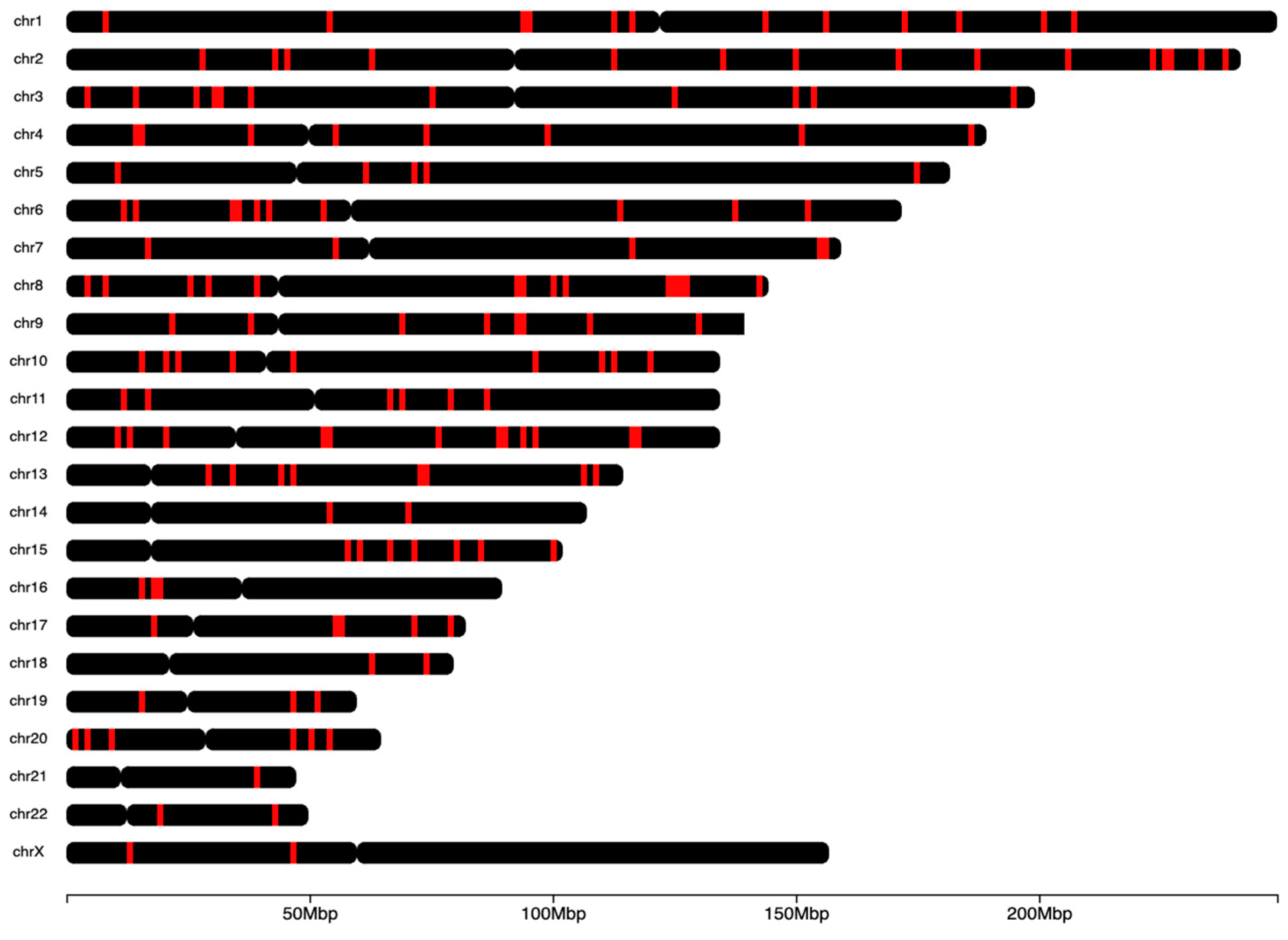
Microarrays
qPCR Validation of Biomarkers
Discussion
Conclusions
Supplementary Materials
Author Contributions
Funding
Consent for publication
Ethical consent and guidelines
Acknowledgments
References
- Morgan E, Arnold M, Gini A, Lorenzoni V, Cabasag CJ, Laversanne M, Morgan E, Arnold M, Gini A, Lorenzoni V, Cabasag CJ, Laversanne M, Vignat J, Ferlay J, Murphy N, and Abou-Kheir W.Bray F. Global burden of colorectal cancer in 2020 and 2040: incidence and mortality estimates from GLOBOCAN. Gut. 2023 Feb;72(2):338-44. [CrossRef] [PubMed]
- Nguyen LH, Goel A, Chung DC. Pathways of Colorectal Carcinogenesis. Gastroenterology. 2020 Jan;158(2):291-302. [CrossRef] [PubMed] [PubMed Central]
- Sullivan BA, Lieberman DA. Colon Polyp Surveillance: Separating the Wheat From the Chaff. Gastroenterology. 2024 May;166(5):743-57. [CrossRef] [PubMed]
- Vyas DA, Eisenstein LG, Jones DS. Hidden in Plain Sight - Reconsidering the Use of Race Correction in Clinical Algorithms. N Engl J Med. 2020 Aug 27;383(9):874-82. [PubMed]
- Fadlallah H, El Masri J, Fakhereddine H, Youssef J, Chemaly C, Doughan S, and Abou-Kheir W. Colorectal cancer: Recent advances in management and treatment. World journal of clinical oncology. 2024 Sep 24;15(9):1136-56. [CrossRef] [PubMed]
- Gharib E, Robichaud GA. From Crypts to Cancer: A Holistic Perspective on Colorectal Carcinogenesis and Therapeutic Strategies. International journal of molecular sciences. 2024 Aug 30;25(17). [CrossRef] [PubMed] [PubMed Central]
- Force USPST, Bibbins-Domingo K, Grossman DC, Curry SJ, Davidson KW, Epling JW, Jr. arcía FAR, Gillman MW, Harper DM, Kemper AR, et al. Screening for Colorectal Cancer: US Preventive Services Task Force Recommendation Statement. Jama. 2016 Jun 21;315(23):2564-75. PubMed PMID: 27304597FHe J, Efron JE. Screening for colorectal cancer. Advances in surgery. 2011;45:31-44. [PubMed]
- He J, Efron JE. Screening for colorectal cancer. Advances in surgery. 2011;45:31-44.
- Seager A, Sharp L, Neilson LJ, Brand A, Hampton JS, Lee TJW, Evans R, Vale L, Whelpton J, Bestwick N et al. Polyp detection with colonoscopy assisted by the GI Genius artificial intelligence endoscopy module compared with standard colonoscopy in routine colonoscopy practice (COLO-DETECT): a multicentre, open-label, parallel-arm, pragmatic randomised controlled trial. The Lancet Gastroenterology & hepatology. 2024 Oct;9(10):911-23. [CrossRef]
- Li SJ, Sharples LD, Benton SC, Blyuss O, Mathews C, Sasieni P, and Duffy SW. Faecal immunochemical testing in bowel cancer screening: Estimating outcomes for different diagnostic policies. Journal of medical screening. 2021 Sep;28(3):277-85. [CrossRef] [PubMed]
- Wieszczy P, Kaminski MF, Loberg M, Bugajski M, Bretthauer M, Kalager M. Estimation of overdiagnosis in colorectal cancer screening with sigmoidoscopy and faecal occult blood testing: comparison of simulation models. BMJ open. 2021 Apr 14;11(4):e042158. [CrossRef] [PubMed] [PubMed Central]
- Fatemi N, Tierling S, Es HA, Varkiani M, Mojarad EN, Aghdaei HA, Walter J, and Totonchi M. DNA methylation biomarkers in colorectal cancer: Clinical applications for precision medicine. Int J Cancer. 2022 Dec 15;151(12):2068-81. [CrossRef] [PubMed]
- Qin J, Wen B, Liang Y, Yu W, Li H. Histone Modifications and their Role in Colorectal Cancer (Review). Pathology oncology research : POR. 2020 Oct;26(4):2023-33. [CrossRef] [PubMed] [PubMed Central]
- Mukhopadhyay S, Ramadass AS, Akoulitchev A, Gordon S. Formation of distinct chromatin conformation signatures epigenetically regulate macrophage activation. International immunopharmacology. 2014 Jan;18(1):7-11. [CrossRef] [PubMed]
- Seaman L, Chen H, Brown M, Wangsa D, Patterson G, Camps J, Omenn G.S.,Ried T., and Rajapakse I. Nucleome Analysis Reveals Structure-Function Relationships for Colon Cancer. Molecular cancer research : MCR. 2017 Jul;15(7):821-30. PubMed PMID: 28258094. Pubmed Central PMCID: 7934948. [CrossRef] [PubMed] [PubMed Central]
- Dekker J, Rippe K, Dekker M, Kleckner N. Capturing chromosome conformation. Science. 2002 Feb 15;295(5558):1306-11. [CrossRef] [PubMed]
- Bastonini E, Jeznach M, Field M, Juszczyk K, Corfield E, Dezfouli M, Ahmat N, Smith A, Womersley H, Jordan P, et al. Chromatin barcodes as biomarkers for melanoma. Pigment cell & melanoma research. 2014 Sep;27(5):788-800. [CrossRef] [PubMed]
- Jakub JW, Grotz TE, Jordan P, Hunter E, Pittelkow M, Ramadass A, Akoulitchev A, and Markovic S. A pilot study of chromosomal aberrations and epigenetic changes in peripheral blood samples to identify patients with melanoma. Melanoma research. 2015 Oct;25(5):406-11. [CrossRef] [PubMed]
- Alshaker H, Mills R, Hunter E, Salter M, Ramadass A, Skinner BM, Westra W, Green J, Akoulitchev A, Winkler M et al. Chromatin conformation changes in peripheral blood can detect prostate cancer and stratify disease risk groups. Journal of translational medicine. 2021 Jan 28;19(1):46. [CrossRef] [PubMed] [PubMed Central]
- Alshaker H, Hunter E, Salter M, Ramadass A, Westra W, Winkler M, Green J, Akoulitchev A, and Pchejetski D. Monocytes acquire prostate cancer-specific chromatin conformations upon indirect co-culture with prostate cancer cells. Front Oncol. 2022;12:990842. [CrossRef] [PubMed] [PubMed Central]
- Pchejetski D, Hunter E, Dezfouli M, Salter M, Powell R, Green J, Naithani T, Koutsothanasi C, Alshaker H, Jaipuria J, et al. Circulating Chromosome Conformation Signatures Significantly Enhance PSA Positive Predicting Value and Overall Accuracy for Prostate Cancer Detection. Cancers. 2023 Jan 29;15(3). [CrossRef] [PubMed] [PubMed Central]
- Hunter E, Salter M, Powell R, Dring A, Naithani T, Chatziioannou ME, Gebregzabhar A, Issa M, Green J, Ng S et al. Development and Validation of Blood-Based Predictive Biomarkers for Response to PD-1/PD-L1 Checkpoint Inhibitors: Evidence of a Universal Systemic Core of 3D Immunogenetic Profiling across Multiple Oncological Indications. Cancers. 2023 May 10;15(10). [PubMed] [PubMed Central]
- Salter M, Corfield E, Ramadass A, Grand F, Green J, Westra J, Lim CR, Farrimond L, Feneberg E, Scaber J, et al. Initial Identification of a Blood-Based Chromosome Conformation Signature for Aiding in the Diagnosis of Amyotrophic Lateral Sclerosis. EBioMedicine. 2018 Jul;33:169-84. [CrossRef] [PubMed] [PubMed Central]
- Carini C, Hunter E, Scottish Early Rheumatoid Arthritis Inception cohort I, Ramadass AS, Green J, Akoulitchev A, et al. Chromosome conformation signatures define predictive markers of inadequate response to methotrexate in early rheumatoid arthritis. Journal of translational medicine. 2018 Jan 29;16(1):18. [CrossRef] [PubMed] [PubMed Central]
- Hunter E, McCord R, Ramadass AS, Green J, Westra JW, Mundt K, and Akoulitchev A. Comparative molecular cell-of-origin classification of diffuse large B-cell lymphoma based on liquid and tissue biopsies. Translational Medicine Communications. 2020 2020/03/24;5(1):5. [CrossRef]
- Yan H, Hunter E, Akoulitchev A, Park P, Winchester DJ, Moo-Young TA, and Prinz RA. Epigenetic chromatin conformation changes in peripheral blood can detect thyroid cancer. Surgery. 2019 Jan;165(1):44-9. [CrossRef] [PubMed]
- Hunter E, Koutsothanasi C, Wilson A, Santos FC, Salter M, Powell R, et al. 3D genomic capture of regulatory immuno-genetic profiles in COVID-19 patients for prognosis of severe COVID disease outcome. bioRxiv. 2021:2021.03.14.435295.
- Tsourkas A, Behlke MA, Xu Y, Bao G. Spectroscopic features of dual fluorescence/luminescence resonance energy-transfer molecular beacons. Anal Chem. 2003 Aug 1;75(15):3697-703. [CrossRef] [PubMed]
- Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences of the United States of America. 2005 Oct 25;102(43):15545-50. PubMed PMID: 16199517. [CrossRef] [PubMed]
- Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, et al. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017 Jan 4;45(D1):D362-D8. Epub 20161018. [PubMed]
- Jain S, Maque J, Galoosian A, Osuna-Garcia A, May FP. Optimal Strategies for Colorectal Cancer Screening. Current treatment options in oncology. 2022 Apr;23(4):474-93. [CrossRef] [PubMed] [PubMed Central]
- Knudsen AB, Zauber AG, Rutter CM, Naber SK, Doria-Rose VP, Pabiniak C, et al. Estimation of Benefits, Burden, and Harms of Colorectal Cancer Screening Strategies: Modeling Study for the US Preventive Services Task Force. Jama. 2016 Jun 21;315(23):2595-609. [PubMed] [PubMed Central]
- Lin JS, Perdue LA, Henrikson NB, Bean SI, Blasi PR. Screening for Colorectal Cancer: Updated Evidence Report and Systematic Review for the US Preventive Services Task Force. Jama. 2021 May 18;325(19):1978-98. [PubMed]
- Imperiale TF, Porter K, Zella J, Gagrat ZD, Olson MC, Statz S, et al. Next-Generation Multitarget Stool DNA Test for Colorectal Cancer Screening. N Engl J Med. 2024 Mar 14;390(11):984-93. [CrossRef] [PubMed]
- Chung DC, Gray DM, 2nd, Singh H, Issaka RB, Raymond VM, Eagle C, et al. A Cell-free DNA Blood-Based Test for Colorectal Cancer Screening. N Engl J Med. 2024 Mar 14;390(11):973-83. [PubMed]
- Dupont C, Armant DR, Brenner CA. Epigenetics: definition, mechanisms and clinical perspective. Seminars in reproductive medicine. 2009 Sep;27(5):351-7. [CrossRef] [PubMed] [PubMed Central]
- Feinberg AP. The Key Role of Epigenetics in Human Disease Prevention and Mitigation. N Engl J Med. 2018 Apr 5;378(14):1323-34. [CrossRef] [PubMed]
- Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA, Jr., Kinzler KW. Cancer genome landscapes. Science. 2013 Mar 29;339(6127):1546-58. [PubMed] [PubMed Central]
- Frazer KA, Murray SS, Schork NJ, Topol EJ. Human genetic variation and its contribution to complex traits. Nature reviews Genetics. 2009 Apr;10(4):241-51. [CrossRef] [PubMed]
- Yan W, Guo M. Epigenetics of colorectal cancer. Methods Mol Biol. 2015;1238:405-24. [PubMed]
- Beckedorff FC, Amaral MS, Deocesano-Pereira C, Verjovski-Almeida S. Long non-coding RNAs and their implications in cancer epigenetics. Bioscience reports. 2013;33(4). Epub 2013/07/24. eng. [CrossRef] [PubMed] [PubMed Central]
- Lai F, Orom UA, Cesaroni M, Beringer M, Taatjes DJ, Blobel GA, et al. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature. 2013 Feb 28;494(7438):497-501. [CrossRef] [PubMed]
- Kosaka N, Ochiya T. Unraveling the Mystery of Cancer by Secretory microRNA: Horizontal microRNA Transfer between Living Cells. Frontiers in genetics. 2011;2:97. Epub 2012/02/04. eng. [CrossRef] [PubMed] [PubMed Central]
- Vinson KE, George DC, Fender AW, Bertrand FE, Sigounas G. The Notch pathway in colorectal cancer. Int J Cancer. 2016 Apr 15;138(8):1835-42. [CrossRef] [PubMed]
- Fleming NI, Jorissen RN, Mouradov D, Christie M, Sakthianandeswaren A, Palmieri M, et al. SMAD2, SMAD3 and SMAD4 mutations in colorectal cancer. Cancer Res. 2013 Jan 15;73(2):725-35. [PubMed]
- Xu S, Wen Z, Jiang Q, Zhu L, Feng S, Zhao Y, et al. CD58, a novel surface marker, promotes self-renewal of tumor-initiating cells in colorectal cancer. Oncogene. 2015 Mar 19;34(12):1520-31. [CrossRef] [PubMed]
- Ding N, Luo H, Zhang T, Peng T, Yao Y, He Y. Correlation between SMADs and Colorectal Cancer Expression, Prognosis, and Immune Infiltrates. International journal of analytical chemistry. 2023;2023:8414040. [CrossRef] [PubMed] [PubMed Central]
- Wu B, Zhan X, Jiang M. CD58 defines regulatory macrophages within the tumor microenvironment. Communications biology. 2024 Aug 21;7(1):1025. 10.1038/s42003-024-06712-6. [PubMed] [PubMed Central]
- Koncina E, Nurmik M, Pozdeev VI, Gilson C, Tsenkova M, Begaj R, et al. IL1R1(+) cancer-associated fibroblasts drive tumor development and immunosuppression in colorectal cancer. Nature communications. 2023 Jul 17;14(1):4251. [PubMed] [PubMed Central]
| Cohort | N (total) | Prospective | Retrospective | Male | Female | Age (mean) |
|---|---|---|---|---|---|---|
| Control | 110 | 42 | 68 | 56 | 54 | 61 |
|
Polyp CRC |
44 171 |
44 14 |
0 157 |
29 89 |
15 82 |
63 64 |
| qPCR markers | Array marker | Probe sequence | P.Value | adj.P.Val | FC | Gene | GeneDist |
|---|---|---|---|---|---|---|---|
| obd156_q1177_q1179 | ORF1_1_116481182_116484855_116627241_116630872_RF | TTGACATAGGACCTCAGCAGAGAGCAGCTCGAGATCCACCCACGTTGTTGCATGTATCAA | 0.0263014 | 1 | -1.295889575 | RP5-1086K13.1;CD58;NAP1L4P1;MIR548AC;IGSF3;AL355794.1;RP4-655J12.4;MIR320B1 | 0;0;0;0;0;0;2341;40878 |
| obd156_q1313_q1315 | ORF1_5_61009121_61015983_61116919_61125541_FR | GAGGCAGGCAGATCACAAGGTCAAGAGATCGATAAGTACATGAGAAATAAACAAAATTCA | 2.33E-07 | 8.13E-05 | -1.389502347 | NDUFAF2;CTC-436P18.4;ERCC8;CTC-436P18.5 | 0;0;64049;20306 |
| obd156_q1301_q1303 | ORF1_12_93013996_93019448_93102345_93106201_FR | TGATGGACTTATGGACTCATTCACTGCATCGATATGGCTCATGCCATTTTATGTGCTATC | 2.02E-08 | 2.64E-05 | 1.445482624 | RP11-511B23.1;RP11-511B23.2;Y_RNA;RP11-511B23.4;RPL41P5;RP11-202G11.2;AC138123.1;RNU6-1329P;NACAP3 | 0;0;0;0;0;0;0;61252;17863 |
| obd156_q1185_q1187 | ORF1_1_201477609_201480715_201569360_201570965_RF | ACAAAGCTATCTCATTTCCTGAGCTTCATCGAGGTGAGGAGATCATGGATGAGTTTTTTA | 0.0247499 | 1 | 1.538252027 | CSRP1;RP11-134G8.7;RP11-134G8.5;RP11-134G8.6;PHLDA3;NAV1 | 0;0;0;0;8373;51921 |
| obd156_q1245_q1247 | ORF1_8_8307248_8309141_8529093_8530943_RF | CAATAATTCATTCTTCTTCATCAGTCCTTCGAACTCCTGACTCAGGAGATCTATCCACCT | 0.0161572 | 1 | -1.342913514 | SGK223;CTA-398F10.1;CTA-398F10.2;FAM86B3P;CTD-3023L14.3 | 0;0;0;62384;24796 |
| obd156_q1217_q1219 | ORF1_1_94060570_94064104_94081020_94084795_RF | TCTTGCCGGGAGTACTCTTCAAACTCCTTCGACATGATGGAGAAGCTGTCCAGGAACCAG | 0.0000011 | 0.000163 | 1.535480941 | ABCA4;RP5-837O21.2;RP11-78O9.1 | 0;125327;60317 |
| obd156_q1297_q1299 | ORF1_15_71449255_71457687_71567140_71571578_RR | GTACTGAATAATAGTGTATGTGTTTATGTCGACTGTACTGGCGGACCCTATAAGAGGCAG | 6.85E-06 | 0.000421 | 1.462480028 | THSD4;RP11-1123I8.1;RP11-592N21.2;AC104938.1 | 0;0;100785;201033 |
| obd156_q1225_q1227 | ORF1_15_67079527_67081854_67195948_67198335_RF | ATCTGTCCCAATCCTTTATCCTTCTAGCTCGAGTCAGCAGTGTTGACTGTTAGCAAATCA | 1.8E-07 | 7.03E-05 | 1.652685053 | SMAD3;RP11-342M21.2;RP11-798K3.2;AAGAB | 0;0;20275;2699 |
| Test | Present | n | Absent | n | Total |
|---|---|---|---|---|---|
| Yes | True positive | 125 | False positive | 25 | 150 |
| No | False negative | 14 | True negative | 89 | 103 |
| Total | 139 | 114 | |||
| Statistic | Value (%) | 95% Cl | |||
| Sensitivity | 89.93 | 83.68 to 94.38 | |||
| Specificity | 79.46 | 70.80 to 86.51 | |||
| Positive Likelihood Ratio | 4.38 | 3.03 to 6.33 | |||
| Negative Likelihood Ratio | 0.13 | 0.08 to 0.21 | |||
| Disease prevalence | 55.38 | 49.00 to 61.63 | |||
| Positive Predictive Value | 84.46 | 78.99 to 88.71 | |||
| Negative Predictive value | 86.41 | 79.31 to 91.33 | |||
| Accuracy | 85.26 | 80.26 to 89.40 | |||
| Test | Present | n | Absent | n | Total |
|---|---|---|---|---|---|
| Yes | True positive | 31 | False positive | 23 | 54 |
| No | False negative | 6 | True negative | 89 | 95 |
| Total | 37 | 112 | |||
| Statistic | Value (%) | 95% Cl | |||
| Sensitivity | 83.78 | 67.99 to 93.81 | |||
| Specificity | 79.46 | 70.80 to 86.51 | |||
| Positive Likelihood Ratio | 4.08 | 2.76 to 6.03 | |||
| Negative Likelihood Ratio | 0.2 | 0.10 to 0.43 | |||
| Disease prevalence | 24.83 | 18.13 to 32.57 | |||
| Positive Predictive Value | 57.41 | 47.69 to 66.58 | |||
| Negative Predictive value | 93.68 | 87.64 to 96.88 | |||
| Accuracy | 80.54 | 73.26 to 86.56 | |||
| qPCR markers | Array marker | Probe sequence | P.Value | adj.P.Val | FC | Gene | GeneDist |
|---|---|---|---|---|---|---|---|
| obd156_q1205_q1207 | ORF1_13_73435053_73437099_73484222_73486544_RF | ACACACAGTAGGTAATTAATACGGTGGATCGAAGTACGCTCTAGTTATACGAGGCTTGTT | 4.43E-08 | 3.46E-05 | 1.424866474 | LINC00393;MARK2P12;LINC00392 | 0;26702;77701 |
| obd156_q1213_q1215 | ORF1_9_37919925_37923489_38002100_38004773_FR | CCGAGGTCCCGAGACTATCTGCCAATCCTCGATTCTCTGGTTTTCCAGTTTGTCTATCTT | 2.49E-07 | 8.27E-05 | -1.373971322 | RP11-613M10.9;SHB;RNU7-124P;SLC25A51;AL161448.1 | 0;0;0;15573;141722 |
| obd156_q1273_q1275 | ORF1_2_113209902_113215780_113275966_113277494_FR | CCAACACCACCCCAAATGCCGGGGCACGTCGAGCGTCCCCGGTTATTGGGAAGGGTGCGC | 0.0178566 | 1 | -1.456428166 | PAX8-AS1;PAX8;RP11-65I12.1;PSD4;IGKV1OR2-108 | 0;0;0;507;128903 |
| obd156_q1293_q1295 | ORF1_9_93218632_93223726_93274460_93278066_RF | TTTATATAACAATGTTTTTTTCAAGGCTTCGAGCAGACATTTCCCCGTCAGGAAGTAACA | 1.12E-07 | 5.53E-05 | -1.438635096 | WNK2;RP11-370F5.4;C9orf129 | 0;70077;40134 |
| obd156_q1245_q1247 | ORF1_8_8307248_8309141_8529093_8530943_RF | CAATAATTCATTCTTCTTCATCAGTCCTTCGAACTCCTGACTCAGGAGATCTATCCACCT | 0.0161572 | 1 | -1.342913514 | SGK223;CTA-398F10.1;CTA-398F10.2;FAM86B3P;CTD-3023L14.3 | 0;0;0;62384;24796 |
| obd156_q1217_q1219 | ORF1_1_94060570_94064104_94081020_94084795_RF | TCTTGCCGGGAGTACTCTTCAAACTCCTTCGACATGATGGAGAAGCTGTCCAGGAACCAG | 0.0000011 | 0.000163 | 1.535480941 | ABCA4;RP5-837O21.2;RP11-78O9.1 | 0;125327;60317 |
| obd156_q1297_q1299 | ORF1_15_71449255_71457687_71567140_71571578_RR | GTACTGAATAATAGTGTATGTGTTTATGTCGACTGTACTGGCGGACCCTATAAGAGGCAG | 6.85E-06 | 0.000421 | 1.462480028 | THSD4;RP11-1123I8.1;RP11-592N21.2;AC104938.1 | 0;0;100785;201033 |
| obd156_q1225_q1227 | ORF1_15_67079527_67081854_67195948_67198335_RF | ATCTGTCCCAATCCTTTATCCTTCTAGCTCGAGTCAGCAGTGTTGACTGTTAGCAAATCA | 1.8E-07 | 7.03E-05 | 1.652685053 | SMAD3;RP11-342M21.2;RP11-798K3.2;AAGAB | 0;0;20275;2699 |
| Test | Present | n | Absent | n | Total |
|---|---|---|---|---|---|
| Yes | True positive | 27 | False positive | 18 | 45 |
| No | False negative | 7 | True negative | 90 | 97 |
| Total | 34 | 108 | |||
| Statistic | Value (%) | 95% Cl | |||
| Sensitivity | 79.41 | 62.10 to 91.30 | |||
| Specificity | 83.33 | 74.94 to 89.81 | |||
| Positive Likelihood Ratio | 4.76 | 3.02 to 7.51 | |||
| Negative Likelihood Ratio | 0.25 | 0.13 to 0.48 | |||
| Disease prevalence | 23.94 | 17.19 to 31.82 | |||
| Positive Predictive Value | 60 | 48.76 to 70.28 | |||
| Negative Predictive value | 92.78 | 86.86 to 96.16 | |||
| Accuracy | 82.39 | 75.12 to 88.27 | |||
| Detection of precancerous lesions (polyps) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| EpiSwitch® NST | Cologuard | FIT | Freenome PREEMPT CRC® | Guardant Shield® | Colonoscopy | ||||
| Sensitivity | 79% | 43% | 23% | 13% | 13% | 75% | |||
| Specificity | 83% | 91% | 95% | 92% | 90% | 89% | |||
| PPV | 60% | 36% | 35% | 39% | 17% | 40% | |||
| NPV | 93% | 93% | 91% | 73% | 86% | 99% | |||
| Accuracy | 82% | 86% | 87% | 70% | 77% | 86% | |||
| Detection of early-stage (I/II) CRC | |||||||||
| EpiSwitch® NST | Cologuard | FIT | Freenome PREEMPT CRC® | Guardant Shield® | Colonoscopy | ||||
| Sensitivity | 84% | 90% | 60% | 79% | 65% | 75% | |||
| Specificity | 79% | 91% | 95% | 92% | 90% | 89% | |||
| PPV | 57% | 2% | 3% | 3% | 1% | 80% | |||
| NPV | 94% | 100% | 100% | 100% | 100% | 86% | |||
| Accuracy | 81% | 91% | 95% | 92% | 90% | 80% | |||
| Detection of CRC (stages I - IV) | |||||||||
| EpiSwitch® NST | Cologuard | FIT | Freenome PREEMPT CRC® | Guardant Shield® | Colonoscopy | ||||
| Sensitivity | 90% | 97% | 71% | 82% | 83% | 75% | |||
| Specificity | 79% | 91% | 95% | 92% | 90% | 89% | |||
| PPV | 84% | 2% | 3% | 3% | 2% | 80% | |||
| NPV | 86% | 100% | 100% | 100% | 99% | 86% | |||
| Accuracy | 85% | 91% | 95% | 92% | 90% | 80% | |||
| Abbreviations: positive predictive value, PPV ; negative predictive value, NPV | |||||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




