Submitted:
05 December 2024
Posted:
06 December 2024
You are already at the latest version
Abstract
(1) Background: Hepatoblastoma and medulloblastoma are two types of pediatric tumors with embryonic origins. Both tumor types can exhibit genetic alterations that affect beta-catenin and Wnt pathway; (2) Materials and Methods: This study used bioinformatics and integrative analysis of multi-omics data at both the tumor and single-cell levels to investigate these two distinct pediatric tumors: medulloblastoma and hepatoblastoma; (3) Results: Cross-transcriptome analysis revealed a commonly regulated expression signature between hepatoblastoma and medulloblastoma tumors. Among the commonly upregulated genes, the transcription factor LEF1 was significantly expressed in both tumor types. In medulloblastoma, LEF1 upregulation is associated with the WNT-subtype. Analysis of LEF1 genome binding occupancy in H1 embryonic stem cells identified 141 LEF1 proximal targets activated in WNT-medulloblastoma, 13 of which are involved in Wnt pathway regulation: RNF43, LEF1, NKD1, AXIN2, DKK4, DKK1, LGR6, FGFR2, NXN, TCF7L1, STK3, YAP1, and NFATC4. An expression score based on these 13 WNT-LEF1 targets accurately predicted the WNT-subtype in two independent medulloblastoma transcriptome cohorts. At the single-cell level, the WNT-LEF1 expression score was exclusively positive in WNT-medulloblastoma tumor cells. This WNT-LEF1-dependent signature was also confirmed as activated in the hepatoblastoma tumor transcriptome. At the single-cell level, the WNT-LEF1 expression score was higher in tumor cells from both human hepatoblastoma samples and a hepatoblastoma patient-derived xenotransplant model; (4) Discussion: This study uncovered a shared transcriptional activation of a LEF1-dependent embryonic program, which orchestrates the regulation of the Wnt signaling pathway in tumor cells from both hepatoblastoma and medulloblastoma.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Public Transcriptome Datasets
2.1.1. MB-PBTA (MB, OpenPBTA)
2.1.2. GSE37418 (MB)
2.1.3. GSE44971 (Normal Cerebellum)
2.1.4. GSE104766 (HB and Normal Liver)
2.1.5. GSE131329 (HB)
2.2. Public Single Cell Transcriptome Datasets
2.2.1. scRNA-seq Medulloblastoma
2.2.2. scRNA-seq Hepatoblastoma:
2.3. ChIP-Sequencing in Human Pluripotent H1 Cell Line
- GSM1579343: LEF1 untreated; Homo sapiens with SRA alias SRR1745491,
- GSM1579344: LEF1 Wnt3a; Homo sapiens with SRA alias SRR1745492,
- GSM1693959: Input H1; Homo sapiens with SRA alias SRR2037029,
- GSM1579348: H3K27ac Wnt3a; Homo sapiens with SRA alias SRR1745496,
- GSM1579350: H3K27me3 Wnt3a; Homo sapiens with SRA alias SRR1745498.
2.4. Transcriptome Cross Normalization for Common Signature Between Hepatoblastoma and Medulloblastoma Tumors
2.5. ChIP-Sequencing Analyses
2.6. WNT-LEF1 Expression Score
2.7. ElasticNet Modeling on WNT-LEF1 Expression Signature
3. Results
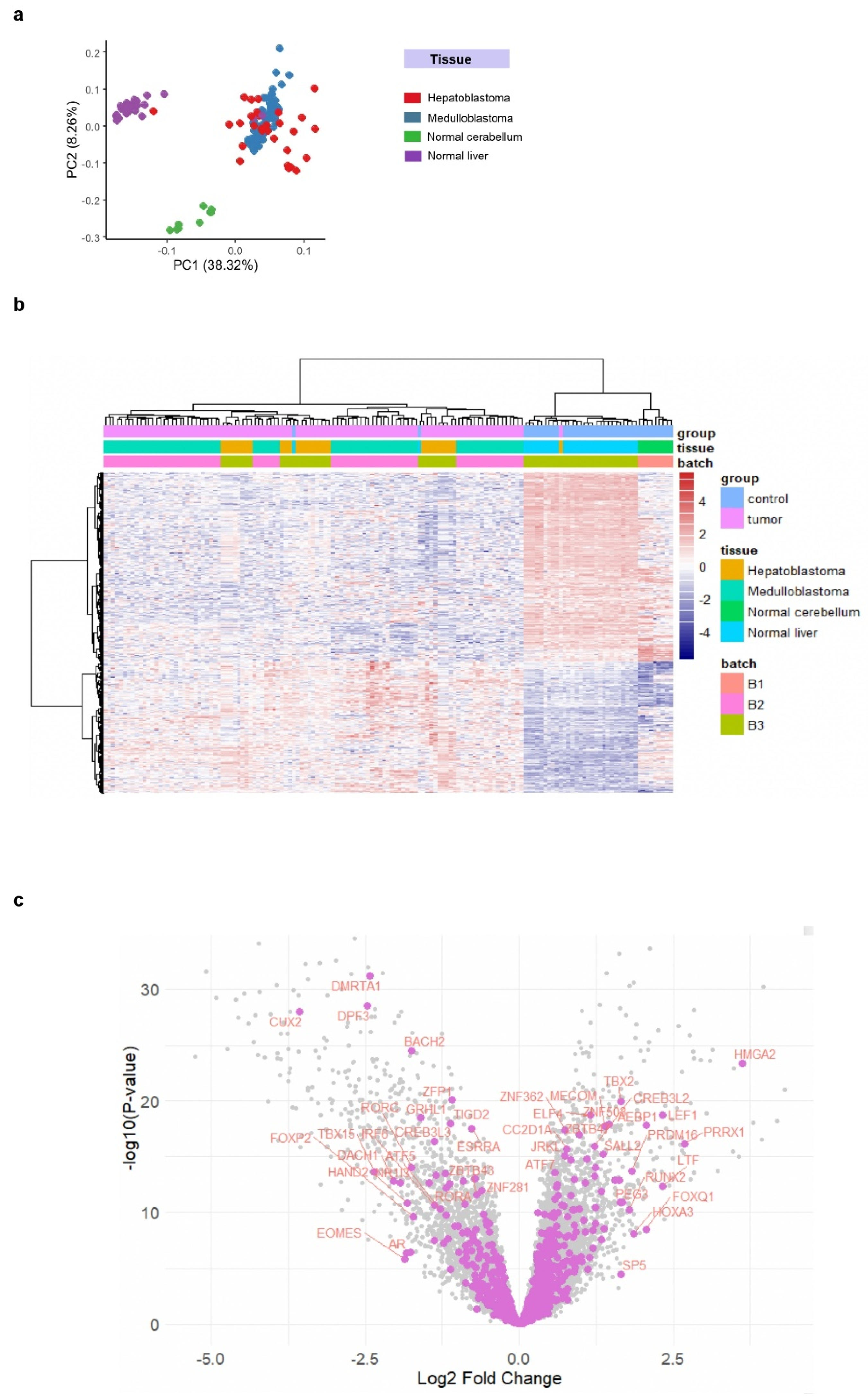
3.1. LEF1 Upregulation as a Shared Molecular Feature in Hepatoblastoma and Medulloblastoma
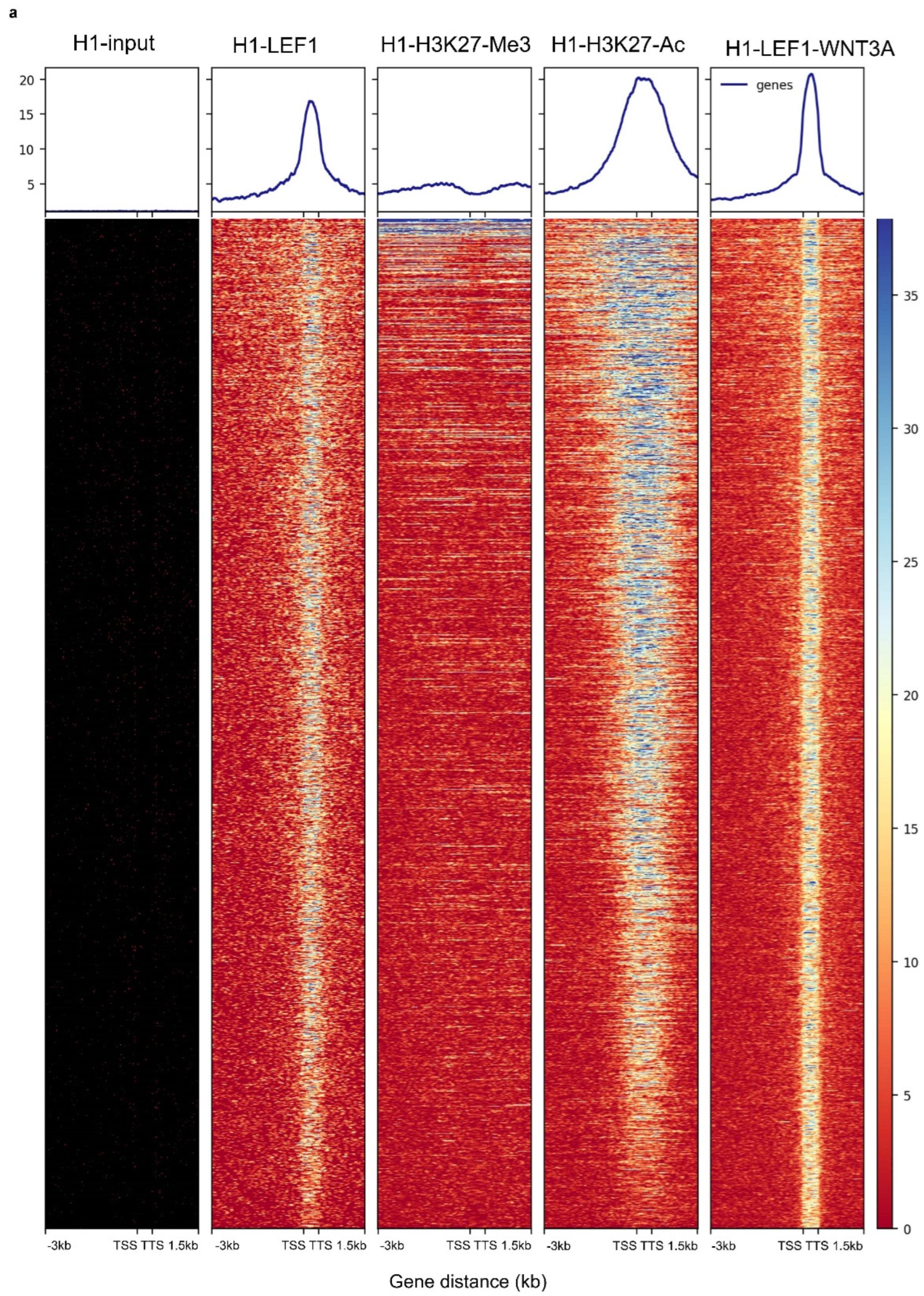
3.2. WNT3A Stimulation Activates LEF1 Chromatin Binding Program with H3K27ac Co-Occupancy in H1 Embryonic Cells
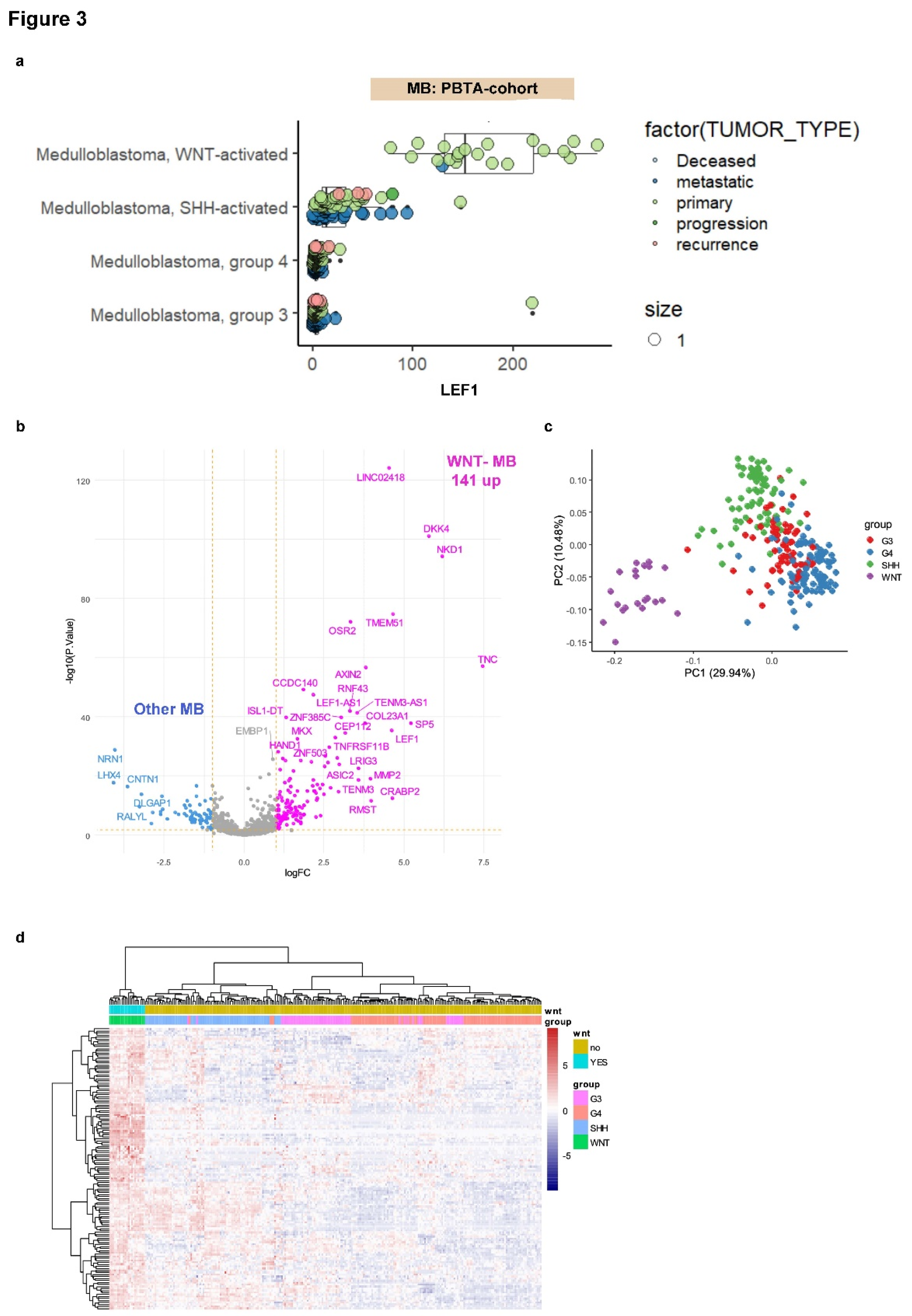
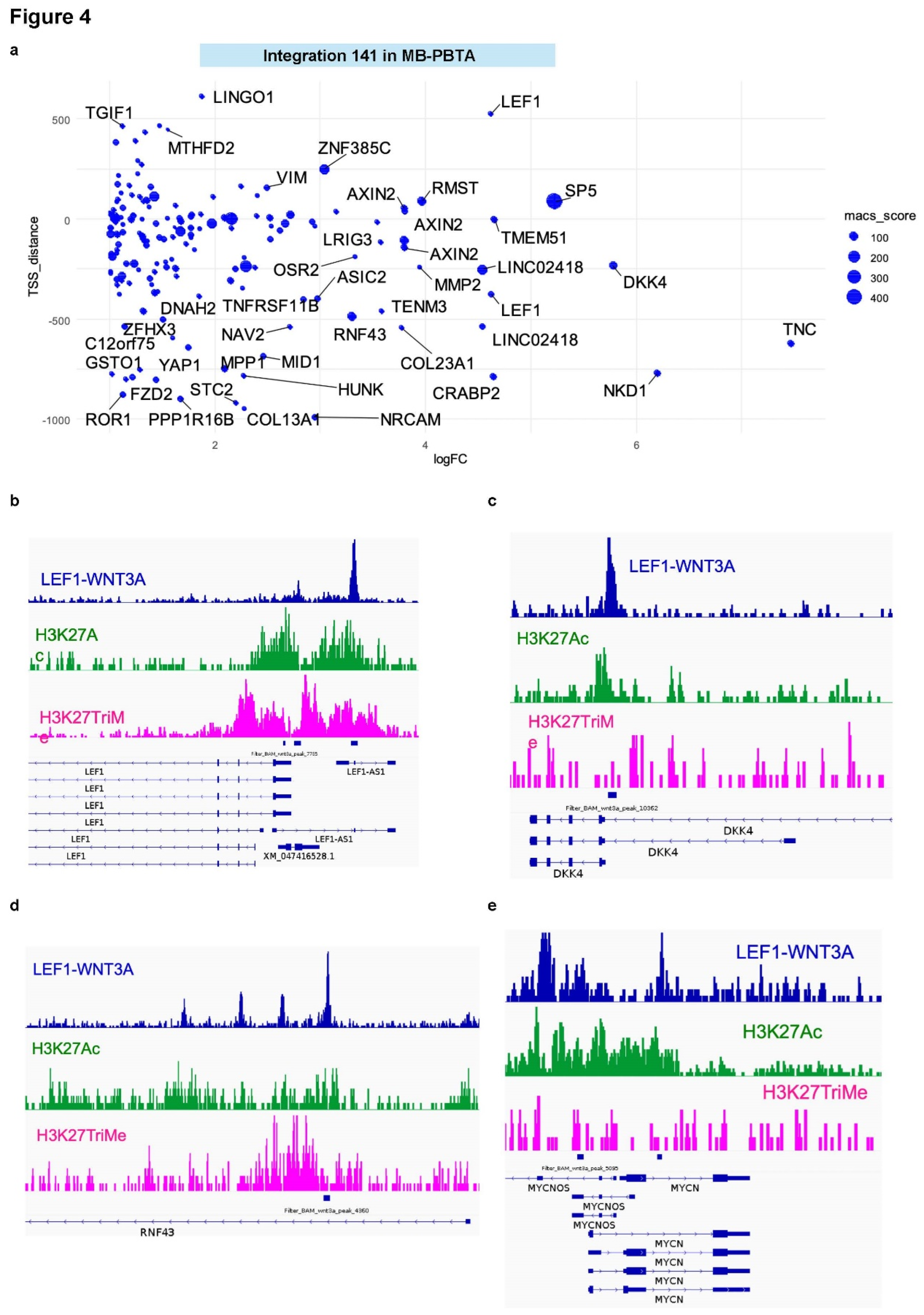
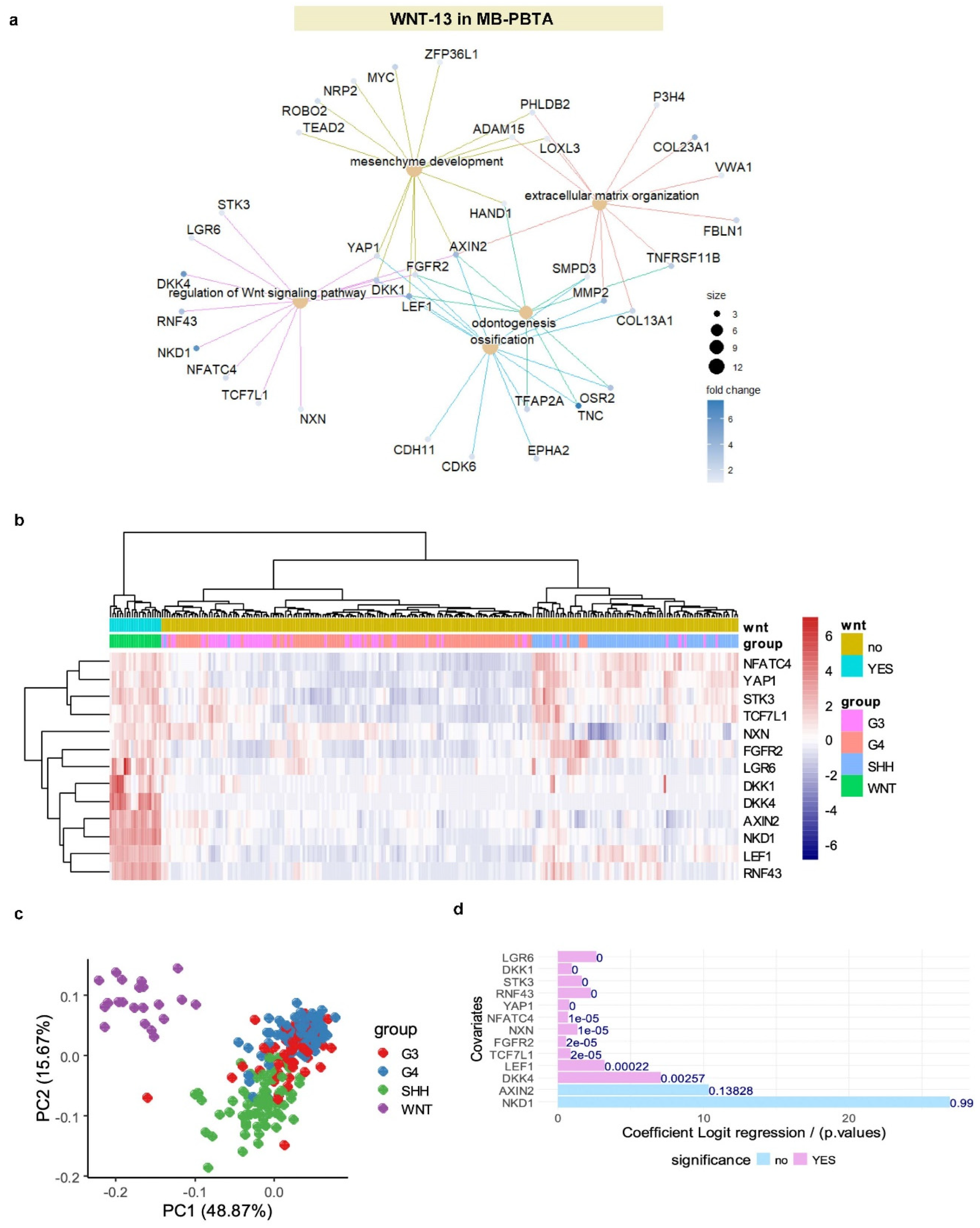
3.3. LEF1 Embryonic Transcriptional Network Is a Hallmark of WNT-Subtype Medulloblastoma
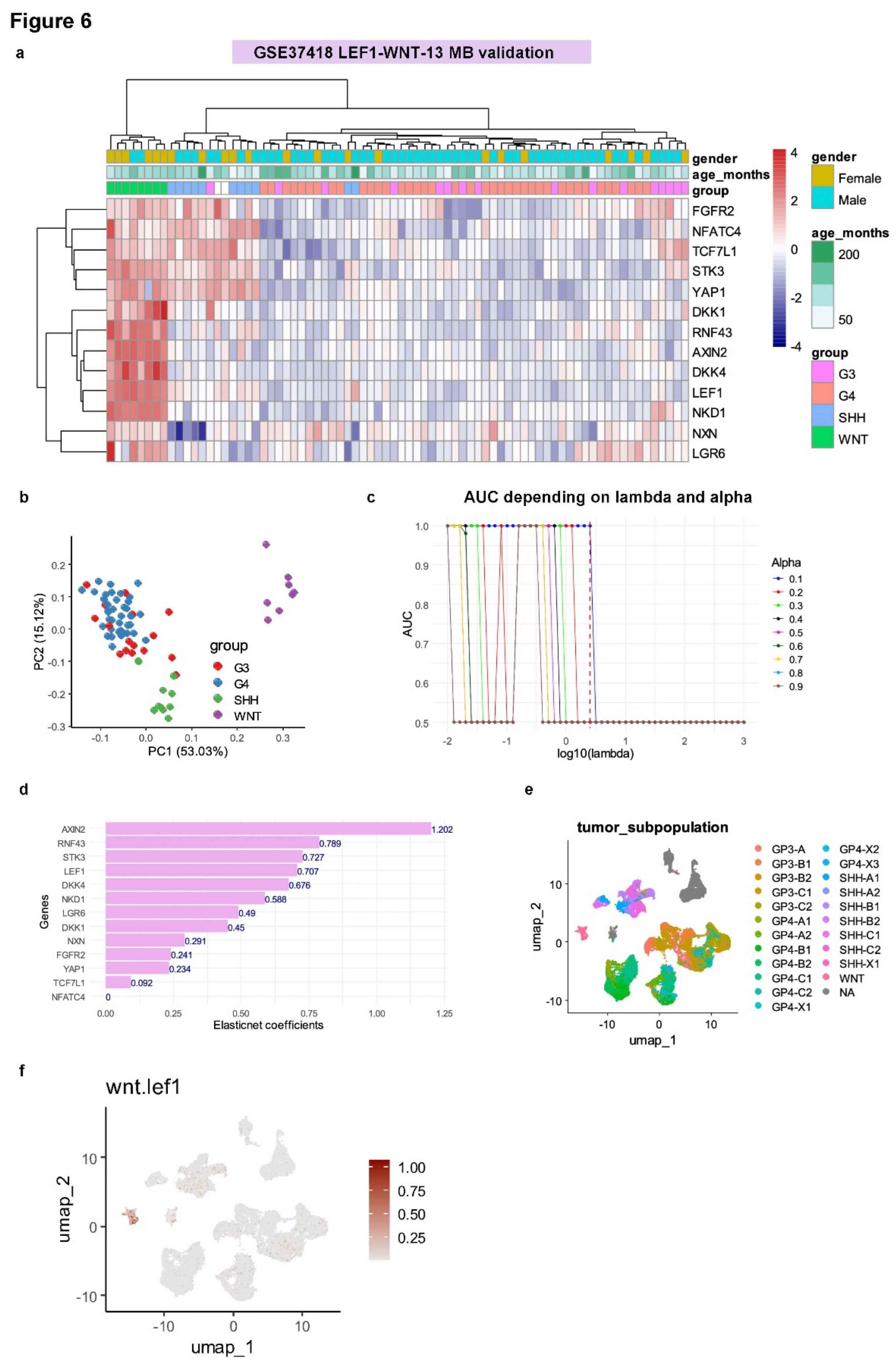
3.4. WNT-LEF1 Gene Signature Accurately Predicts WNT Subtype in Medulloblastoma Transcriptomes
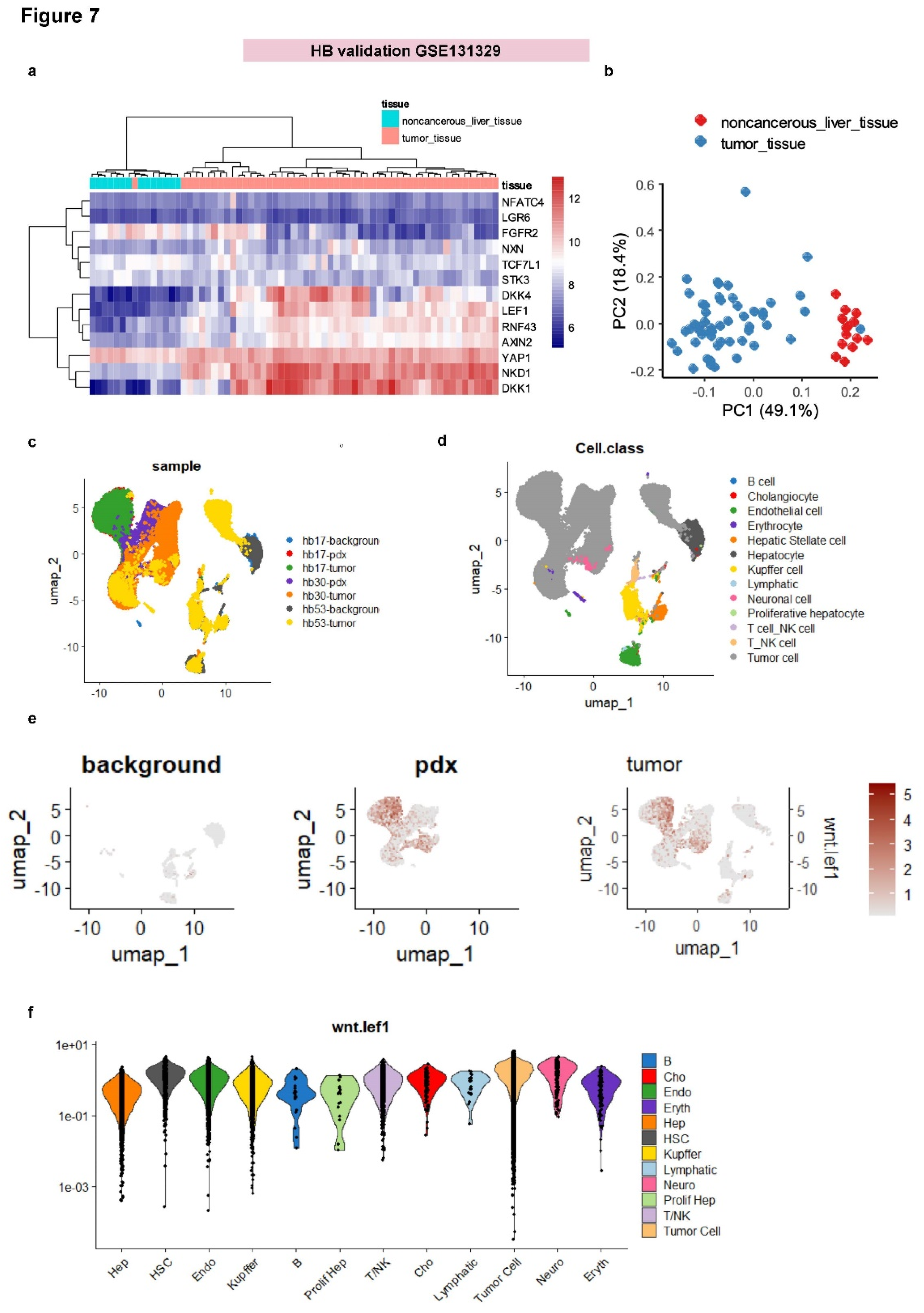
3.5. Single-Cell Analysis Validates WNT-LEF1 Signature Activation in Hepatoblastoma Tumor Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sharma D, Subbarao G, Saxena R. Hepatoblastoma. Semin Diagn Pathol. 2017 Mar;34(2):192-200. Epub 2016 Dec 23. [CrossRef] [PubMed]
- Cotter JA, Hawkins C. Medulloblastoma: WHO 2021 and Beyond. Pediatr Dev Pathol. 2022 Jan-Feb;25(1):23-33. [CrossRef] [PubMed]
- Liu J, Xiao Q, Xiao J, Niu C, Li Y, Zhang X, Zhou Z, Shu G, Yin G. Wnt/β-catenin signalling: function, biological mechanisms, and therapeutic opportunities. Signal Transduct Target Ther. 2022 Jan 3;7(1):3. [CrossRef] [PubMed] [PubMed Central]
- He S, Tang S. WNT/β-catenin signaling in the development of liver cancers. Biomed Pharmacother. 2020 Dec;132:110851. Epub 2020 Oct 17. [CrossRef] [PubMed]
- Krishnamurthy N, Kurzrock R. Targeting the Wnt/beta-catenin pathway in cancer: Update on effectors and inhibitors. Cancer Treat Rev. 2018 Jan;62:50-60. Epub 2017 Nov 13. [CrossRef] [PubMed] [PubMed Central]
- Crippa S, Ancey PB, Vazquez J, Angelino P, Rougemont AL, Guettier C, Zoete V, Delorenzi M, Michielin O, Meylan E. Mutant CTNNB1 and histological heterogeneity define metabolic subtypes of hepatoblastoma. EMBO Mol Med. 2017 Nov;9(11):1589-1604. [CrossRef] [PubMed] [PubMed Central]
- Chiang J, Moreira DC, Pytel NJ, Liu YC, Blackburn PR, Shi Z, Cardenas M, Wheeler DA, Furtado LV. A CTNNB1-altered medulloblastoma shows the immunophenotypic, DNA methylation and transcriptomic profiles of SHH-activated, and not WNT-activated, medulloblastoma. Neuropathol Appl Neurobiol. 2022 Aug;48(5):e12815. Epub 2022 Mar 30. [CrossRef] [PubMed] [PubMed Central]
- Staal FJ, Clevers H. Tcf/Lef transcription factors during T-cell development: unique and overlapping functions. Hematol J. 2000;1(1):3-6. [CrossRef] [PubMed]
- Gao L, Cui W, Kelting S, Woodroof J, Li H, Li L, Zhang D. Canonical and non-canonical Wnt signal pathway in classic Hodgkin lymphoma and the prognostic significance of LEF1, β-catenin, FZD6 and Wnt5a/b. Am J Blood Res. 2022 Aug 15;12(4):136-143. PMCID: PMC9490108. [PubMed]
- Katoh, M. Multi layered prevention and treatment of chronic inflammation, organ fibrosis and cancer associated with canonical WNT/β catenin signaling activation (Review). Int J Mol Med. 2018 Aug;42(2):713-725. Epub 2018 May 17. [CrossRef] [PubMed] [PubMed Central]
- Fakhr E, Zare F, Azadmanesh K, Teimoori-Toolabi L. LEF1 silencing sensitizes colorectal cancer cells to oxaliplatin, 5-FU, and irinotecan. Biomed Pharmacother. 2021 Nov;143:112091. Epub 2021 Aug 30. [CrossRef] [PubMed]
- Blazquez R, Rietkötter E, Wenske B, Wlochowitz D, Sparrer D, Vollmer E, Müller G, Seegerer J, Sun X, Dettmer K, Barrantes-Freer A, Stange L, Utpatel K, Bleckmann A, Treiber H, Bohnenberger H, Lenz C, Schulz M, Reimelt C, Hackl C, Grade M, Büyüktas D, Siam L, Balkenhol M, Stadelmann C, Kube D, Krahn MP, Proescholdt MA, Riemenschneider MJ, Evert M, Oefner PJ, Klein CA, Hanisch UK, Binder C, Pukrop T. LEF1 supports metastatic brain colonization by regulating glutathione metabolism and increasing ROS resistance in breast cancer. Int J Cancer. 2020 Jun 1;146(11):3170-3183. Epub 2019 Nov 11. [CrossRef] [PubMed]
- Robinson, G., Parker, M., Kranenburg, T. A., Lu, C., Chen, X., Ding, L., Phoenix, T. N., Hedlund, E., Wei, L., Zhu, X., Chalhoub, N., Baker, S. J., Huether, R., Kriwacki, R., Curley, N., Thiruvenkatam, R., Wang, J., Wu, G., Rusch, M., Hong, X., Becksfort, J., Gupta, P., Ma, J., Easton, J., Vadodaria, B., Onar-Thomas, A., Lin, T., Li, S., Pounds, S., Paugh, S., Zhao, D., Kawauchi, D., Roussel, M. F., Finkelstein, D., Ellison, D. W., Lau, C. C., Bouffet, E., Hassall, T., Gururangan, S., Cohn, R., Fulton, R. S., Fulton, L. L., Dooling, D. J., Ochoa, K., Gajjar, A., Mardis, E. R., Wilson, R. K., Downing, J. R., Zhang, J., and Gilbertson, R. J. (2012) Novel mutations target distinct subgroups of medulloblastoma. Nature 488, 43–48.
- Barrett, T., Wilhite, S. E., Ledoux, P., Evangelista, C., Kim, I. F., Tomashevsky, M., Marshall, K. A., Phillippy, K. H., Sherman, P. M., Holko, M., Yefanov, A., Lee, H., Zhang, N., Robertson, C. L., Serova, N., Davis, S., and Soboleva, A. (2013) NCBI GEO: archive for functional genomics data sets--update. Nucleic Acids Res. 41, D991-995.
- Davis, S. and Meltzer, P. S. (2007) GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23, 1846–1847.
- Lambert, S. R., Witt, H., Hovestadt, V., Zucknick, M., Kool, M., Pearson, D. M., Korshunov, A., Ryzhova, M., Ichimura, K., Jabado, N., Fontebasso, A. M., Lichter, P., Pfister, S. M., Collins, V. P., and Jones, D. T. W. (2013) Differential expression and methylation of brain developmental genes define location-specific subsets of pilocytic astrocytoma. Acta Neuropathol 126, 291–301.
- Riemondy, K. A., Venkataraman, S., Willard, N., Nellan, A., Sanford, B., Griesinger, A. M., Amani, V., Mitra, S., Hankinson, T. C., Handler, M. H., Sill, M., Ocasio, J., Weir, S. J., Malawsky, D. S., Gershon, T. R., Garancher, A., Wechsler-Reya, R. J., Hesselberth, J. R., Foreman, N. K., Donson, A. M., and Vibhakar, R. (2022) Neoplastic and immune single-cell transcriptomics define subgroup-specific intra-tumoral heterogeneity of childhood medulloblastoma. Neuro Oncol 24, 273–286.
- Butler, A., Hoffman, P., Smibert, P., Papalexi, E., and Satija, R. (2018) Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420.
- Bondoc, A., Glaser, K., Jin, K., Lake, C., Cairo, S., Geller, J., Tiao, G., and Aronow, B. (2021) Identification of distinct tumor cell populations and key genetic mechanisms through single cell sequencing in hepatoblastoma. Commun Biol 4, 1049.
- Zappia, L., Lun, A., Kamm, J., and Cannoodt, R. (2024) zellkonverter: Conversion Between scRNA-seq Objects.
- Harikumar, A. and Meshorer, E. (2015) Chromatin remodeling and bivalent histone modifications in embryonic stem cells. EMBO Rep 16, 1609–1619.
- Leinonen, R., Sugawara, H., Shumway, M., and on behalf of the International Nucleotide Sequence Database Collaboration. (2011) The Sequence Read Archive. Nucleic Acids Research 39, D19–D21.
- Shapiro, J. A., Gaonkar, K. S., Spielman, S. J., Savonen, C. L., Bethell, C. J., Jin, R., Rathi, K. S., Zhu, Y., Egolf, L. E., Farrow, B. K., Miller, D. P., Yang, Y., Koganti, T., Noureen, N., Koptyra, M. P., Duong, N., Santi, M., Kim, J., Robins, S., Storm, P. B., Mack, S. C., Lilly, J. V., Xie, H. M., Jain, P., Raman, P., Rood, B. R., Lulla, R. R., Nazarian, J., Kraya, A. A., Vaksman, Z., Heath, A. P., Kline, C., Scolaro, L., Viaene, A. N., Huang, X., Way, G. P., Foltz, S. M., Zhang, B., Poetsch, A. R., Mueller, S., Ennis, B. M., Prados, M., Diskin, S. J., Zheng, S., Guo, Y., Kannan, S., Waanders, A. J., Margol, A. S., Kim, M. C., Hanson, D., Van Kuren, N., Wong, J., Kaufman, R. S., Coleman, N., Blackden, C., Cole, K. A., Mason, J. L., Madsen, P. J., Koschmann, C. J., Stewart, D. R., Wafula, E., Brown, M. A., Resnick, A. C., Greene, C. S., Rokita, J. L., Taroni, J. N., Children’s Brain Tumor Network, and Pacific Pediatric Neuro-Oncology Consortium. (2023) OpenPBTA: The Open Pediatric Brain Tumor Atlas. Cell Genom 3, 100340.
- Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E., and Storey, J. D. (2012) The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883.
- Zhao, Y., Wong, L., and Goh, W. W. B. (2020) How to do quantile normalization correctly for gene expression data analyses. Sci Rep 10, 15534.
- Ritchie, M. E., Phipson, B., Wu, D., Hu, Y., Law, C. W., Shi, W., and Smyth, G. K. (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47.
- Wu, T., Hu, E., Xu, S., Chen, M., Guo, P., Dai, Z., Feng, T., Zhou, L., Tang, W., Zhan, L., Fu, X., Liu, S., Bo, X., and Yu, G. (2021) clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. The Innovation 2, 100141.
- Xu, S., Hu, E., Cai, Y., Xie, Z., Luo, X., Zhan, L., Tang, W., Wang, Q., Liu, B., Wang, R., Xie, W., Wu, T., Xie, L., and Yu, G. (2024) Using clusterProfiler to characterize multiomics data. Nat Protoc 19, 3292–3320.
- Ashburner, M., Ball, C. A., Blake, J. A., Botstein, D., Butler, H., Cherry, J. M., Davis, A. P., Dolinski, K., Dwight, S. S., Eppig, J. T., Harris, M. A., Hill, D. P., Issel-Tarver, L., Kasarskis, A., Lewis, S., Matese, J. C., Richardson, J. E., Ringwald, M., Rubin, G. M., and Sherlock, G. (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25, 25–29.
- Landt, S. G., Marinov, G. K., Kundaje, A., Kheradpour, P., Pauli, F., Batzoglou, S., Bernstein, B. E., Bickel, P., Brown, J. B., Cayting, P., Chen, Y., DeSalvo, G., Epstein, C., Fisher-Aylor, K. I., Euskirchen, G., Gerstein, M., Gertz, J., Hartemink, A. J., Hoffman, M. M., Iyer, V. R., Jung, Y. L., Karmakar, S., Kellis, M., Kharchenko, P. V., Li, Q., Liu, T., Liu, X. S., Ma, L., Milosavljevic, A., Myers, R. M., Park, P. J., Pazin, M. J., Perry, M. D., Raha, D., Reddy, T. E., Rozowsky, J., Shoresh, N., Sidow, A., Slattery, M., Stamatoyannopoulos, J. A., Tolstorukov, M. Y., White, K. P., Xi, S., Farnham, P. J., Lieb, J. D., Wold, B. J., and Snyder, M. (2012) ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813–1831.
- Bolger, A. M., Lohse, M., and Usadel, B. (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120.
- Langmead, B. and Salzberg, S. L. (2012) Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359.
- Li, H., Handsaker, B., Wysoker, A., Fennell, T., Ruan, J., Homer, N., Marth, G., Abecasis, G., Durbin, R., and 1000 Genome Project Data Processing Subgroup. (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079.
- Barnett, D. W., Garrison, E. K., Quinlan, A. R., Strömberg, M. P., and Marth, G. T. (2011) BamTools: a C++ API and toolkit for analyzing and managing BAM files. Bioinformatics 27, 1691–1692.
- Zhang, Y., Liu, T., Meyer, C. A., Eeckhoute, J., Johnson, D. S., Bernstein, B. E., Nusbaum, C., Myers, R. M., Brown, M., Li, W., and Liu, X. S. (2008) Model-based analysis of ChIP-Seq (MACS). Genome Biol 9, R137.
- Feng, J., Liu, T., Qin, B., Zhang, Y., and Liu, X. S. (2012) Identifying ChIP-seq enrichment using MACS. Nat Protoc 7, 1728–1740.
- Liang, K. and Keleş, S. (2012) Normalization of ChIP-seq data with control. BMC Bioinformatics 13, 199.
- Heinz, S., Benner, C., Spann, N., Bertolino, E., Lin, Y. C., Laslo, P., Cheng, J. X., Murre, C., Singh, H., and Glass, C. K. (2010) Simple Combinations of Lineage-Determining Transcription Factors Prime cis-Regulatory Elements Required for Macrophage and B Cell Identities. Molecular Cell 38, 576–589.
- Frankish, A., Diekhans, M., Jungreis, I., Lagarde, J., Loveland, J. E., Mudge, J. M., Sisu, C., Wright, J. C., Armstrong, J., Barnes, I., Berry, A., Bignell, A., Boix, C., Carbonell Sala, S., Cunningham, F., Di Domenico, T., Donaldson, S., Fiddes, I. T., García Girón, C., Gonzalez, J. M., Grego, T., Hardy, M., Hourlier, T., Howe, K. L., Hunt, T., Izuogu, O. G., Johnson, R., Martin, F. J., Martínez, L., Mohanan, S., Muir, P., Navarro, F. C. P., Parker, A., Pei, B., Pozo, F., Riera, F. C., Ruffier, M., Schmitt, B. M., Stapleton, E., Suner, M.-M., Sycheva, I., Uszczynska-Ratajczak, B., Wolf, M. Y., Xu, J., Yang, Y. T., Yates, A., Zerbino, D., Zhang, Y., Choudhary, J. S., Gerstein, M., Guigó, R., Hubbard, T. J. P., Kellis, M., Paten, B., Tress, M. L., and Flicek, P. (2021) GENCODE 2021. Nucleic Acids Res 49, D916–D923.
- Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A., and Manke, T. (2014) deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187-191.
- Thiele, C. and Hirschfeld, G. (2021) cutpointr: Improved Estimation and Validation of Optimal Cutpoints in R. J. Stat. Soft. 98.
- Kuhn, M. (2008) Building Predictive Models in R Using the caret Package. J. Stat. Soft. 28.
- Tay, J. K., Narasimhan, B., and Hastie, T. (2023) Elastic Net Regularization Paths for All Generalized Linear Models. J. Stat. Soft. 106.
- Lambert, S. A., Jolma, A., Campitelli, L. F., Das, P. K., Yin, Y., Albu, M., Chen, X., Taipale, J., Hughes, T. R., and Weirauch, M. T. (2018) The Human Transcription Factors. Cell 172, 650–665.
- Estarás, C., Benner, C., and Jones, K. A. (2015) SMADs and YAP compete to control elongation of β-catenin:LEF-1-recruited RNAPII during hESC differentiation. Mol Cell 58, 780–793.
- Geng, Z., Wafula, E., Corbett, R. J., Zhang, Y., Jin, R., Gaonkar, K. S., Shukla, S., Rathi, K. S., Hill, D., Lahiri, A., Miller, D. P., Sickler, A., Keith, K., Blackden, C., Chroni, A., Brown, M. A., Kraya, A. A., Koschmann, C. J., Aldape, K., Huang, X., Rood, B. R., Mason, J. L., Trooskin, G. R., Abdullaev, Z., Wang, P., Zhu, Y., Farrow, B. K., Farrel, A., Dybas, J. M., Zhong, C., Van Kuren, N., Zhang, B., Santi, M., Phul, S., Chinwalla, A. T., Resnick, A. C., Diskin, S. J., Tasian, S., Stefankiewicz, S., Maris, J. M., Ennis, B. M., Lueder, M. R., Naqvi, A. S., Coleman, N., Ma, W., Taylor, D., and Rokita, J. L. (2024) The Open Pediatric Cancer Project.
- Cairo, S., Armengol, C., De Reyniès, A., Wei, Y., Thomas, E., Renard, C.-A., Goga, A., Balakrishnan, A., Semeraro, M., Gresh, L., Pontoglio, M., Strick-Marchand, H., Levillayer, F., Nouet, Y., Rickman, D., Gauthier, F., Branchereau, S., Brugières, L., Laithier, V., Bouvier, R., Boman, F., Basso, G., Michiels, J.-F., Hofman, P., Arbez-Gindre, F., Jouan, H., Rousselet-Chapeau, M.-C., Berrebi, D., Marcellin, L., Plenat, F., Zachar, D., Joubert, M., Selves, J., Pasquier, D., Bioulac-Sage, P., Grotzer, M., Childs, M., Fabre, M., and Buendia, M.-A. (2008) Hepatic stem-like phenotype and interplay of Wnt/beta-catenin and Myc signaling in aggressive childhood liver cancer. Cancer Cell 14, 471–484.
- Ramaswamy, V., Remke, M., Bouffet, E., Bailey, S., Clifford, S. C., Doz, F., Kool, M., Dufour, C., Vassal, G., Milde, T., Witt, O., von Hoff, K., Pietsch, T., Northcott, P. A., Gajjar, A., Robinson, G. W., Padovani, L., André, N., Massimino, M., Pizer, B., Packer, R., Rutkowski, S., Pfister, S. M., Taylor, M. D., and Pomeroy, S. L. (2016) Risk stratification of childhood medulloblastoma in the molecular era: the current consensus. Acta Neuropathol 131, 821–831.
- Wang, D., Gong, J., Zhang, H., Liu, Y., Sun, N., Hao, X., and Mu, K. (2022) Immunohistochemical staining of LEF-1 is a useful marker for distinguishing WNT-activated medulloblastomas. Diagn Pathol 17, 69.
- Aboubakr, O., Métais, A., Doz, F., Saffroy, R., Masliah-Planchon, J., Hasty, L., Beccaria, K., Ayrault, O., Dufour, C., Varlet, P., and Tauziède-Espariat, A. (2024) LEF-1 immunohistochemistry, a better diagnostic biomarker than β-catenin for medulloblastoma, WNT-activated subtyping. J Neuropathol Exp Neurol 83, 136–138.
- Ye, S., Zhang, T., Tong, C., Zhou, X., He, K., Ban, Q., Liu, D., and Ying, Q.-L. (2017) Depletion of Tcf3 and Lef1 maintains mouse embryonic stem cell self-renewal. Biol Open 6, 511–517.
- Kobayashi, W. and Ozawa, M. (2013) The transcription factor LEF-1 induces an epithelial–mesenchymal transition in MDCK cells independent of β-catenin. Biochemical and Biophysical Research Communications 442, 133–138.
- Zirkel, A., Lederer, M., Stöhr, N., Pazaitis, N., and Hüttelmaier, S. (2013) IGF2BP1 promotes mesenchymal cell properties and migration of tumor-derived cells by enhancing the expression of LEF1 and SNAI2 (SLUG). Nucleic Acids Research 41, 6618–6636.
- Liang, J., Li, X., Li, Y., Wei, J., Daniels, G., Zhong, X., Wang, J., Sfanos, K., Melamed, J., Zhao, J., and Lee, P. (2015) LEF1 targeting EMT in prostate cancer invasion is mediated by miR-181a. Am J Cancer Res 5, 1124–1132.
- Eberherr, C., Beck, A., Vokuhl, C., Becker, K., Häberle, B., Von Schweinitz, D., and Kappler, R. (2019) Targeting excessive MYCN expression using MLN8237 and JQ1 impairs the growth of hepatoblastoma cells. Int J Oncol 54, 1853–1863.
- Bourdeaut, F., Grison, C., Maurage, C.-A., Laquerriere, A., Vasiljevic, A., Delisle, M.-B., Michalak, S., Figarella-Branger, D., Doz, F., Richer, W., Pierron, G., Miquel, C., Delattre, O., and Couturier, J. (2013) MYC and MYCN amplification can be reliably assessed by aCGH in medulloblastoma. Cancer Genet 206, 124–129.
- de Lau, W., Peng, W. C., Gros, P., and Clevers, H. (2014) The R-spondin/Lgr5/Rnf43 module: regulator of Wnt signal strength. Genes Dev 28, 305–316.
- Bell, I., Khan, H., Stutt, N., Horn, M., Hydzik, T., Lum, W., Rea, V., Clapham, E., Hoeg, L., and Van Raay, T. J. (2024) Nkd1 functions downstream of Axin2 to attenuate Wnt signaling. Mol Biol Cell 35, ar93.
- Fatima, S., Lee, N. P., Tsang, F. H., Kolligs, F. T., Ng, I. O. L., Poon, R. T. P., Fan, S. T., and Luk, J. M. (2012) Dickkopf 4 (DKK4) acts on Wnt/β-catenin pathway by influencing β-catenin in hepatocellular carcinoma. Oncogene 31, 4233–4244.
- Li, Y., Lu, W., King, T. D., Liu, C.-C., Bijur, G. N., and Bu, G. (2010) Dkk1 stabilizes Wnt co-receptor LRP6: implication for Wnt ligand-induced LRP6 down-regulation. PLoS One 5, e11014.
- Feng, Q., Li, S., Ma, H.-M., Yang, W.-T., and Zheng, P.-S. (2021) LGR6 activates the Wnt/β-catenin signaling pathway and forms a β-catenin/TCF7L2/LGR6 feedback loop in LGR6high cervical cancer stem cells. Oncogene 40, 6103–6114.
- Bourdeaut, F., Miquel, C., Di Rocco, F., Grison, C., Richer, W., Brugieres, L., Pierron, G., James, S., Baujat, G., Delattre, O., and Collet, C. (2013) Germline mutations in FGF receptors and medulloblastomas. American J of Med Genetics Pt A 161, 382–385.
- Valek, L. and Tegeder, I. (2021) Nucleoredoxin Knockdown in SH-SY5Y Cells Promotes Cell Renewal. Antioxidants (Basel) 10, 449.
- Athanasouli, P., Balli, M., De Jaime-Soguero, A., Boel, A., Papanikolaou, S., van der Veer, B. K., Janiszewski, A., Vanhessche, T., Francis, A., El Laithy, Y., Nigro, A. L., Aulicino, F., Koh, K. P., Pasque, V., Cosma, M. P., Verfaillie, C., Zwijsen, A., Heindryckx, B., Nikolaou, C., and Lluis, F. (2023) The Wnt/TCF7L1 transcriptional repressor axis drives primitive endoderm formation by antagonizing naive and formative pluripotency. Nat Commun 14, 1210.
- Li, P., Chen, Y., Mak, K. K., Wong, C. K., Wang, C. C., and Yuan, P. (2013) Functional role of Mst1/Mst2 in embryonic stem cell differentiation. PLoS One 8, e79867.
- Kim, W., Khan, S. K., Gvozdenovic-Jeremic, J., Kim, Y., Dahlman, J., Kim, H., Park, O., Ishitani, T., Jho, E.-H., Gao, B., and Yang, Y. (2017) Hippo signaling interactions with Wnt/β-catenin and Notch signaling repress liver tumorigenesis. J Clin Invest 127, 137–152.
- Park, H. W., Kim, Y. C., Yu, B., Moroishi, T., Mo, J.-S., Plouffe, S. W., Meng, Z., Lin, K. C., Yu, F.-X., Alexander, C. M., Wang, C.-Y., and Guan, K.-L. (2015) Alternative Wnt Signaling Activates YAP/TAZ. Cell 162, 780–794.
- Azzolin, L., Panciera, T., Soligo, S., Enzo, E., Bicciato, S., Dupont, S., Bresolin, S., Frasson, C., Basso, G., Guzzardo, V., Fassina, A., Cordenonsi, M., and Piccolo, S. (2014) YAP/TAZ incorporation in the β-catenin destruction complex orchestrates the Wnt response. Cell 158, 157–170.
- Morfouace, M., Cheepala, S., Jackson, S., Fukuda, Y., Patel, Y. T., Fatima, S., Kawauchi, D., Shelat, A. A., Stewart, C. F., Sorrentino, B. P., Schuetz, J. D., and Roussel, M. F. (2015) ABCG2 Transporter Expression Impacts Group 3 Medulloblastoma Response to Chemotherapy. Cancer Res 75, 3879–3889.
| Variable | Level | HIGH (n= 21) | low (n=233) | Total (n=254) | p-value |
|---|---|---|---|---|---|
| SEX | Male | 13 (61.9) | 142 (60.9) | 155 (61.0) | |
| Female | 8 (38.1) | 91 (39.1) | 99 (39.0) | 1.00000 | |
| TUMOR TYPE | primary | 20 (95.2) | 135 (57.9) | 155 (61.0) | |
| progression | 0 (0.0) | 3 (1.3) | 3 (1.2) | ||
| metastatic | 1 (4.8) | 83 (35.6) | 84 (33.1) | ||
| recurrence | 0 (0.0) | 11 (4.7) | 11 (4.3) | ||
| Deceased | 0 (0.0) | 1 (0.4) | 1 (0.4) | 0.02348 | |
| group | WNT | 21 (100.0) | 0 (0.0) | 21 (8.3) | |
| G3 | 0 (0.0) | 60 (25.8) | 60 (23.6) | ||
| G4 | 0 (0.0) | 99 (42.5) | 99 (39.0) | ||
| SHH | 0 (0.0) | 74 (31.8) | 74 (29.1) | < 1e-04 | |
| CNS_REGION | Ventricles | 5 (23.8) | 12 (5.2) | 17 (6.7) | |
| Mixed | 3 (14.3) | 63 (27.2) | 66 (26.1) | ||
| Posterior fossa | 13 (61.9) | 152 (65.5) | 165 (65.2) | ||
| Spine | 0 (0.0) | 1 (0.4) | 1 (0.4) | ||
| Hemispheric | 0 (0.0) | 3 (1.3) | 3 (1.2) | ||
| Other | 0 (0.0) | 1 (0.4) | 1 (0.4) | 0.03963 | |
| missing | 0 | 1 | 1 | ||
| EFS_STATUS | 1:Recurrence | 1 (4.8) | 36 (15.5) | 37 (14.6) | |
| 0:No Event | 19 (90.5) | 121 (51.9) | 140 (55.1) | ||
| 1:Progressive - Metastatic | 1 (4.8) | 33 (14.2) | 34 (13.4) | ||
| 1:Deceased - due to disease | 0 (0.0) | 7 (3.0) | 7 (2.8) | ||
| 1:Recurrence - Metastatic | 0 (0.0) | 23 (9.9) | 23 (9.1) | ||
| 1:Second Malignancy | 0 (0.0) | 6 (2.6) | 6 (2.4) | ||
| 1:Progressive | 0 (0.0) | 7 (3.0) | 7 (2.8) | 0.06538 |
| Variable | Level | Low (n=66) | Hight (n= 8) | Total (n=74) | p-value |
|---|---|---|---|---|---|
| gender | Male | 51 (77.3) | 2 (25.0) | 53 (71.6) | |
| Female | 15 (22.7) | 6 (75.0) | 21 (28.4) | 0.00732 | |
| age_months | mean (sd) | 98.4 (38.7) | 104.5 (17) | 99.1 (37) | 0.66172 |
| anapath | CL | 43 (65.2) | 8 (100.0) | 51 (68.9) | |
| DN | 6 (9.1) | 0 (0.0) | 6 (8.1) | ||
| AN | 17 (25.8) | 0 (0.0) | 17 (23.0) | 0.13231 | |
| ethnie | White | 38 (57.6) | 4 (50.0) | 42 (56.8) | |
| Black | 5 (7.6) | 0 (0.0) | 5 (6.8) | ||
| Asian | 5 (7.6) | 1 (12.5) | 6 (8.1) | ||
| other | 2 (3.0) | 1 (12.5) | 3 (4.1) | ||
| Hispanic | 12 (18.2) | 1 (12.5) | 13 (17.6) | ||
| Asian Indian | 4 (6.1) | 0 (0.0) | 4 (5.4) | ||
| Pacific Islander | 0 (0.0) | 1 (12.5) | 1 (1.4) | 0.07854 | |
| group | G4 | 39 (59.1) | 0 (0.0) | 39 (52.7) | |
| WNT | 0 (0.0) | 8 (100.0) | 8 (10.8) | ||
| SHH | 11 (16.7) | 0 (0.0) | 11 (14.9) | ||
| G3 | 16 (24.2) | 0 (0.0) | 16 (21.6) | < 1e-04 |
| Variable | Level | tumor_tissue (n=53) | Noncancerous liver tissue (n=14) | Total (n=67) | p-value |
|---|---|---|---|---|---|
| gender | Female | 25 (47.2) | 8 (57.1) | 33 (49.3) | |
| Male | 28 (52.8) | 6 (42.9) | 34 (50.7) | 0.716363 | |
| Histological type | Well_differentiated | 30 (56.6) | 14 (100.0) | 44 (65.7) | |
| Other | 2 (3.8) | 0 (0.0) | 2 (3.0) | ||
| Poorly_differentiated | 21 (39.6) | 0 (0.0) | 21 (31.3) | 0.009797 | |
| age_months | mean (sd) | 27.2 (24.1) | 27.6 (26.4) | 27.3 (24.4) | 0.964822 |
| pretext_stage | P3 | 18 (34.0) | 0 (0.0) | 18 (34.0) | |
| P2 | 15 (28.3) | 0 (0.0) | 15 (28.3) | ||
| P4 | 11 (20.8) | 0 (0.0) | 11 (20.8) | ||
| P1 | 9 (17.0) | 0 (0.0) | 9 (17.0) | NA | |
| chic_risk stratification | Standard | 31 (58.5) | 0 (0.0) | 31 (58.5) | |
| High | 14 (26.4) | 0 (0.0) | 14 (26.4) | ||
| Intermediate | 8 (15.1) | 0 (0.0) | 8 (15.1) | NA | |
| ctnnbi_gene alteration | Deletion | 23 (43.4) | 0 (0.0) | 23 (43.4) | |
| Wild_Type | 14 (26.4) | 0 (0.0) | 14 (26.4) | ||
| Mutation (exon3) | 16 (30.2) | 0 (0.0) | 16 (30.2) | NA | |
| clinical_course | Alive | 38 (71.7) | 0 (0.0) | 38 (71.7) | |
| Dead | 15 (28.3) | 0 (0.0) | 15 (28.3) | NA |
| Comparison | Mean Groupe1 | Mean Groupe2 | p_value |
|---|---|---|---|
| Hepatocyte vs Tumor cell | -1.088 | 0.450 | 0.00E+00 |
| Hepatic Stellate cell vs Tumor cell | 0.168 | 0.450 | 2.93E-07 |
| Endothelial cell vs Tumor cell | -0.264 | 0.450 | 0.00E+00 |
| Kupffer cell vs Tumor cell | -0.686 | 0.450 | 0.00E+00 |
| B cell vs Tumor cell | -0.628 | 0.450 | 1.24E+05 |
| Proliferative hepatocyte vs Tumor cell | -1.270 | 0.450 | 2.40E-24 |
| T cell_NK cell vs Tumor cell | -0.652 | 0.450 | 9.86E-91 |
| Cholangiocyte vs Tumor cell | -0.208 | 0.450 | 2.26E-07 |
| Lymphatic vs Tumor cell | -0.416 | 0.450 | 7.62E+07 |
| T_NK cell vs Tumor cell | -0.267 | 0.450 | 1.25E-59 |
| Neuronal cell vs Tumor cell | 0.375 | 0.450 | 4.13E-01 |
| Erythrocyte vs Tumor cells | -0.619 | 0.450 | 4.20E-50 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




