Submitted:
04 December 2024
Posted:
06 December 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
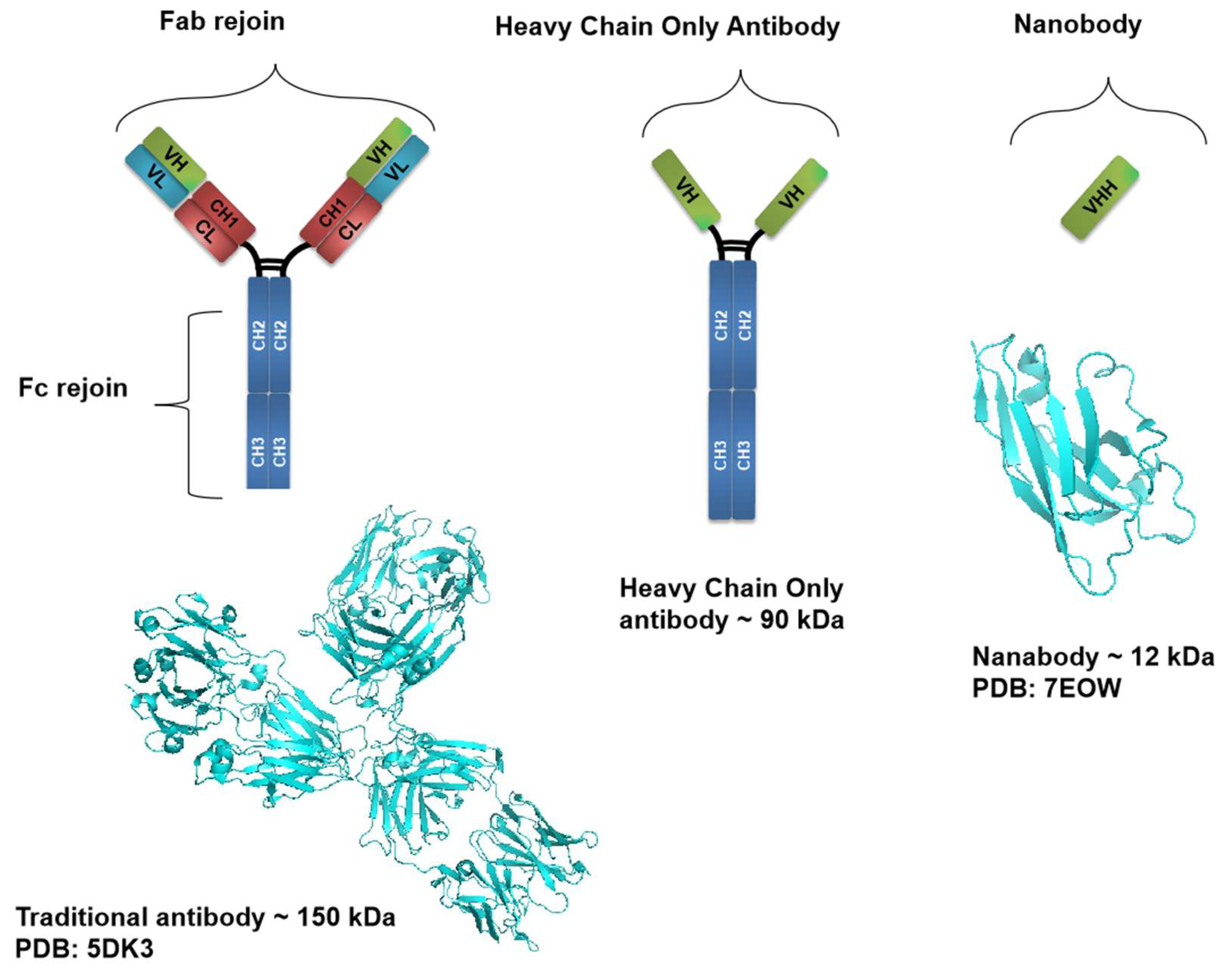
2. The Feature of Nanobodies
3. The Efficacy of Nanobodies Against Respiratory Viruses
| Sr.nu | Target Viral Protein | Disease | Mechanism of Action | Reference |
|---|---|---|---|---|
| 1 | Spike RBD | SARS-CoV-2 | Blocks receptor binding domain and prevents viral entry | [17] |
| 2 | Spike RBD | SARS-CoV-2 | Inhibits spike protein conformational change for fusion | [6] |
| 3 | Hemagglutinin | H5N1 | Prevents hemagglutinin from binding to sialic acid receptors | [8] |
| 4 | Hemagglutinin | H1N1 | Blocks hemagglutinin from facilitating viral entry | [9] |
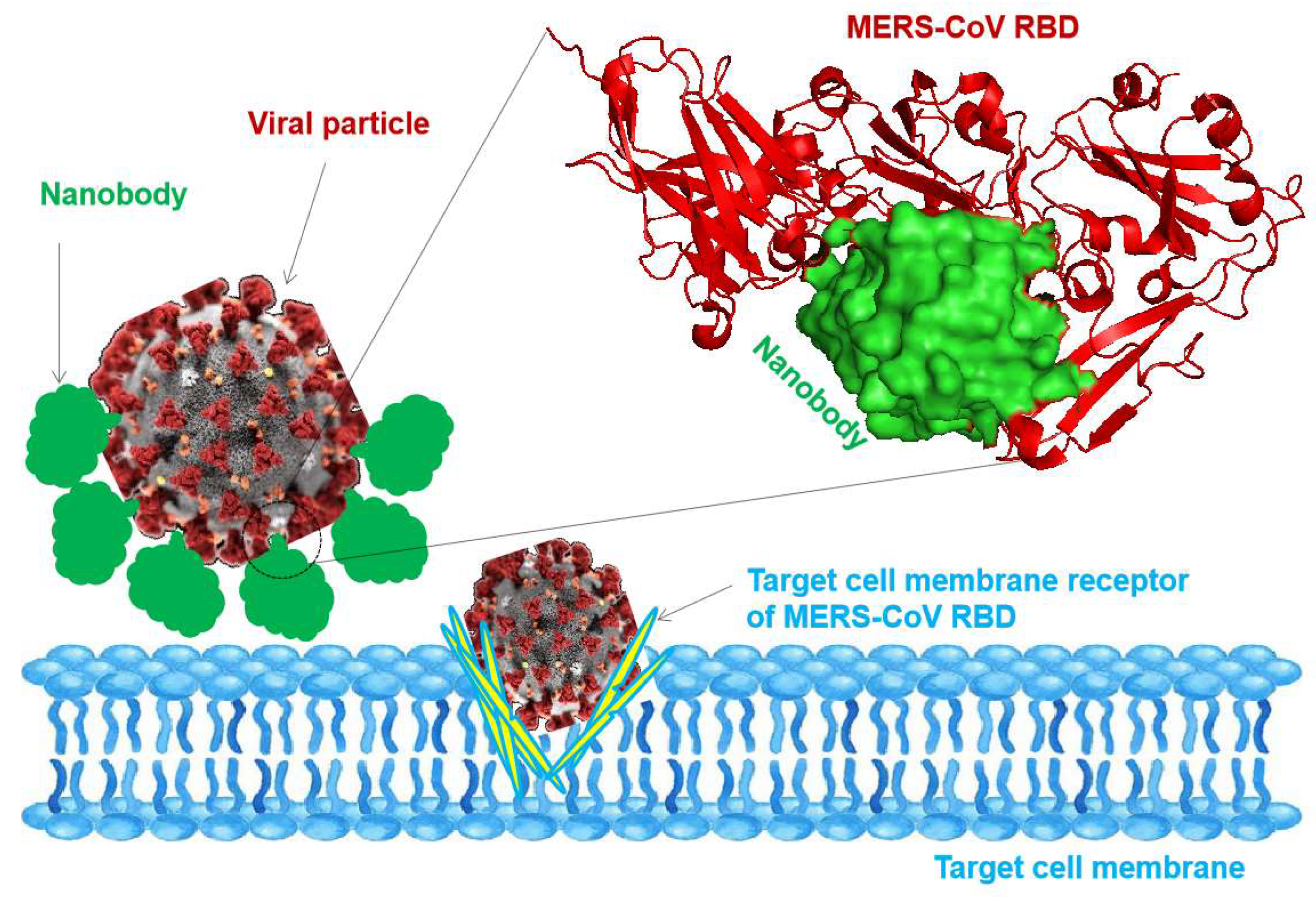
| 5 | Spike Protein | MERS-CoV | Inhibits interaction with DPP4 receptor, blocking entry | [4] |
| 6 | Spike Protein | SARS | Prevents viral entry by blocking spike protein interaction | [1] |
| 7 | F Protein | RSV | Stabilizes pre-fusion form, preventing membrane fusion | [11] |
| 8 | Spike RBD | SARS-CoV-2 | Binds to RBD, blocking ACE2 receptor interaction | [17] |
| 9 | Hemagglutinin | Influenza A | Inhibits receptor binding, preventing viral entry | [2] |
| 10 | Nucleocapsid | SARS-CoV-2 | Disrupts nucleocapsid structure, inhibiting replication | [10] |
| 11 | Hemagglutinin | H3N2 | Blocks hemagglutinin function, preventing viral entry | [14] |
| 12 | Spike S2 | SARS-CoV-2 | Inhibits spike-mediated membrane fusion | [15] |
| 13 | RBD | MERS-CoV | Blocks interaction with DPP4 receptor | [4] |
| 14 | Spike Protein | SARS-CoV-2 | Inhibits viral fusion with host cell membrane | [6] |
| 15 | G Protein | RSV | Blocks G protein interactions, inhibiting viral entry | [12] |
| 16 | ORF7a | SARS-CoV-2 | Modulates viral protein function, inhibiting replication | [13] |
4. Methodologies
4.1. Construction of Nanobody Libraries Using Phage Display Technology
- (1)
- Separation of lymphocytes: The first step is obtaining the lymphocytes from an immunized camelid or llama with the antigens of interest, typically this is done by drawing blood samples from the animals after the immunization period under ethical committee approval consent.
- (2)
- RNA isolation and cDNA synthesis: Using established protocols the total RNA will be extracted from the isolated lymphocytes, and using reverse transcriptase to convert the RNA into complementary DNA (cDNA).
- (3)
- Amplification of nanobody sequences: The purified cDNA containing the nanobody sequences will amplify using Polymerase Chain Reaction (PCR) and specific primers to the framework regions of the nanobodies will used to selectively amplify the variable regions of nanobodies.
- (4)
- Construction of phage display library: The amplified nanobody sequences are then cloned into a phagemid vector, which contains a gene encoding a coat protein of a bacteriophage, usually M13, phagemid vector confirms that the nanobodies are displayed on the surface of phage particles using molecular biology tool such as sequencing
- (5)
- Transformation and phage production: The phagemid vector containing the nanobody library will transform into Escherichia coli (E. coli) cells, and the transformed cells are then grown to a large number of phage particles displaying different nanobodies on the phage surface.
- (6)
- Phage selection: The nanobody phage library will be subjected to multiple rounds of affinity selection against the target antigen, usually (2-5 rounds). This involves incubating the library with the antigens of interest and washing away unbound phages several times to obtain a high affinity monoclonal nanobody phage.
- (7)
- Elution and amplification: The bound phages will be eluted from the antigen, and the selected nanobody genes will be amply using PCR to generate a new nanobody phage library enriched with nanobodies that have a high affinity for the antigens of interest.
4.2. Bio-Panning Screening of Nanobodies
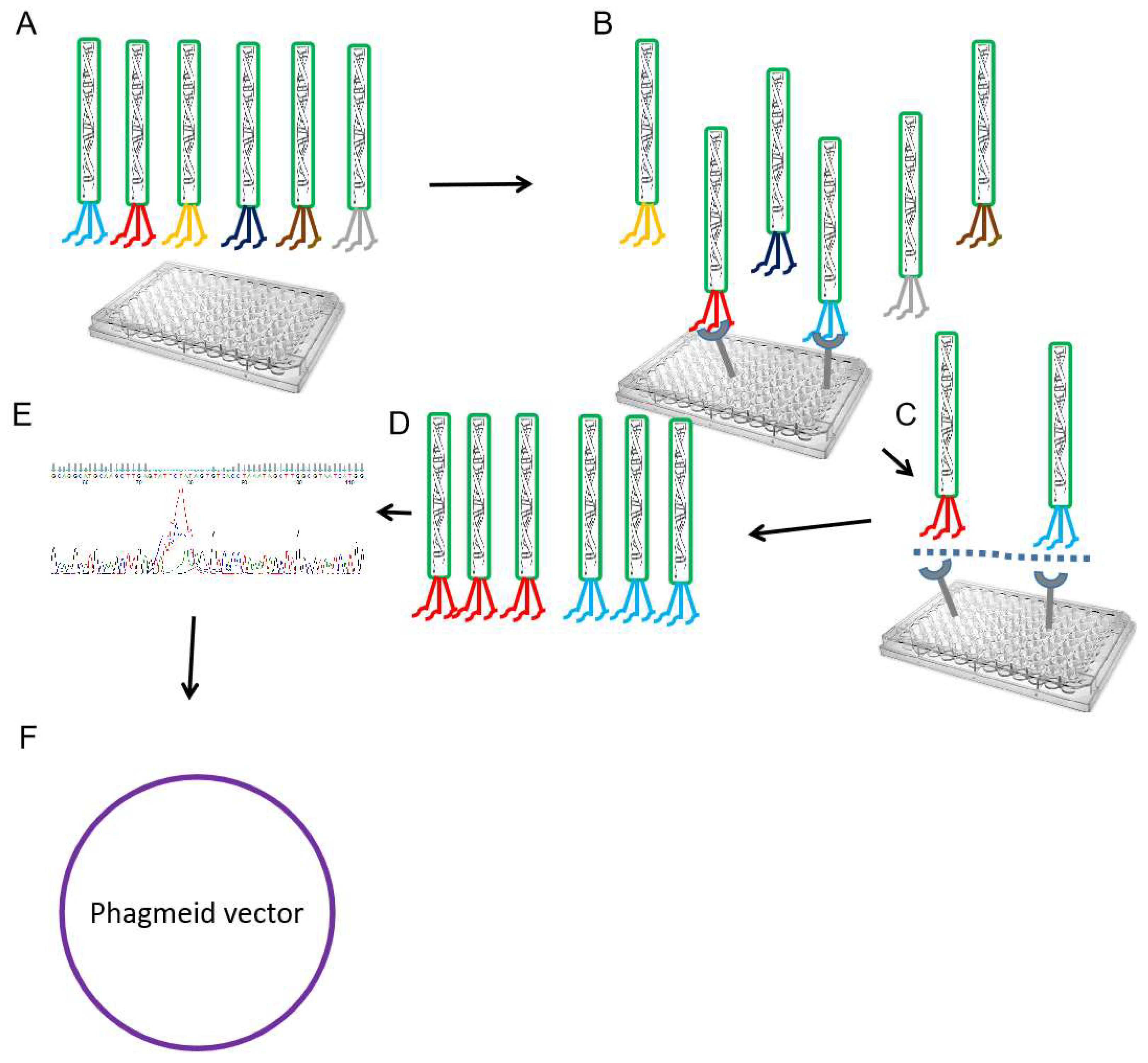
4.3. Cloning, Expression, and Purification of Selected Nanobodies Based on Viral Antigens
- Cloning of Nanobody Genes: The selected nanobody genes from the phage nanobodies display library are cloned into expression vectors suitable for protein production such as bacterial plasmid pET30a. These vectors often contain a promoter region to drive gene expression, as well as tags or fusion partners to aid in the purification and detection of the expressed nanobodies. Recently many scientists have engineered to clone the selected nanobody gene to the Fc region of a human IgG1 antibody to increase the affinity and efficacy of the nanobody-Fc antibody against the viral proteins.
- Expression of Nanobodies: After the cloning process is successful and confirmed with the PCR and sequencing, cloned expression vectors are transformed or transinfected into a suitable host system for protein expression. Generally used host systems such as bacteria (e.g., E. coli) or yeast (e.g., Saccharomyces cerevisiae), the host cells are then grown for three or four days under suitable conditions to induce the expression of the nanobodies protein.
- Purification of Nanobodies: When the nanobodies are expressed, they need to be purified from the host cell lysate or culture supernatant, affinity chromatography is often used, where the nanobodies are selectively bound to a column containing a ligand specific to the tag or fusion partner used during cloning such as Ni-column interacted with the polyhistidine at the vector, expressed with our target nanobodies protein. Other chromatographic techniques, such as ion exchange or size exclusion chromatography, can further purify the nanobodies and remove impurities.
4.4. Characterization and Validation of Nanobodies as Viral Antigens Inhibitors
- (1)
- Binding affinity determination: Nanobodies' binding affinity towards the viral antigen is evaluated using biophysical techniques such as surface plasmon resonance (SPR), isothermal titration calorimetry (ITC), size exclusion chromatography (SEC), or enzyme-linked immunosorbent assay (ELISA). These experiments provide information on the strength of the interaction between the nanobody and the viral antigen.
- (2)
- Epitope mapping: Epitope mapping studies aim to identify the specific region of the viral antigen recognized by the nanobody. This can be done using techniques such as peptide scanning arrays, alanine scanning mutagenesis, or X-ray crystallography. Understanding the binding site of the nanobody on the viral antigen can help elucidate its inhibitory mechanism.
- (3)
- Neutralization assays: To assess the inhibitory potential of nanobodies, neutralization assays are performed. These assays involve exposing the viral pathogen to the nanobody and evaluating its ability to prevent viral entry into host cells or block viral replication. Virus neutralization can be measured through infectivity assays, plaque reduction assays, or viral growth inhibition assays.
- (4)
- In vivo validation: Nanobodies showing promising inhibitory effects in vitro can be further validated in animal models. Animal studies, such as murine models or non-human primates, can provide insights into the nanobody's efficacy, safety, pharmacokinetics, and immunogenicity.
- (5)
- Structural characterization: Structural studies, such as X-ray crystallography or cryo-electron microscopy, can be conducted to determine the atomic level details of the nanobody viral antigen complex. These studies provide insights into the binding mode and conformational changes occurring upon complex formation.
- (6)
- Optimization and engineering: Nanobodies can be further optimized and engineered to enhance their binding affinity, stability, or half-life. Technologies like phage display based affinity maturation, site-directed mutagenesis, or antibody humanization can be applied to improve the nanobody's properties for therapeutic applications.
5. Conclusion and Future Directions
Author Contributions
Funding
Acknowledgements
Conflicts of Interest
References
- Sharma, A., et al., Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2): a global pandemic and treatment strategies. International journal of antimicrobial agents, 2020. 56(2): p. 106054.
- Meganck, R.M. and R.S. Baric, Developing therapeutic approaches for twenty-first-century emerging infectious viral diseases. Nature medicine, 2021. 27(3): p. 401-410. [CrossRef]
- Steeland, S., R.E. Vandenbroucke, and C. Libert, Nanobodies as therapeutics: big opportunities for small antibodies. Drug discovery today, 2016. 21(7): p. 1076-1113. [CrossRef]
- Sroga, P., D. Safronetz, and D.R. Stein, Nanobodies: a new approach for the diagnosis and treatment of viral infectious diseases. Future Virology, 2020. 15(3): p. 195-205. [CrossRef]
- Mohammed, A., et al., Generation, biochemical characterizations and validation of potent nanobodies derived from alpaca specific for human receptor of advanced glycation end product. Biochemical and Biophysical Research Communications, 2021. 581: p. 38-45. [CrossRef]
- Ma, H., et al., Potent neutralization of SARS-CoV-2 by hetero-bivalent alpaca nanobodies targeting the spike receptor-binding domain. Journal of virology, 2021. 95(10): p. 10.1128/jvi. 02438-20. doi:10.1128/JVI.
- Ma, H., et al., Hetero-bivalent nanobodies provide broad-spectrum protection against SARS-CoV-2 variants of concern including Omicron. Cell Research, 2022. 32(9): p. 831-842.
- Ibanez, L.I., et al., Nanobodies with in vitro neutralizing activity protect mice against H5N1 influenza virus infection. Journal of Infectious Diseases, 2011. 203(8): p. 1063-1072. [CrossRef]
- Skehel, J.J. and M.D. Waterfield, Studies on the primary structure of the influenza virus hemagglutinin. Proceedings of the National Academy of Sciences, 1975. 72(1): p. 93-97. [CrossRef]
- Kamat, S., M. Kumari, and C. Jayabaskaran, Nano-engineered tools in the diagnosis, therapeutics, prevention, and mitigation of SARS-CoV-2. Journal of Controlled Release, 2021. 338: p. 813-836. [CrossRef]
- Pantaleo, G., et al., Antibodies to combat viral infections: development strategies and progress. Nature Reviews Drug Discovery, 2022. 21(9): p. 676-696. [CrossRef]
- Alizadeh, M., et al., Designing a novel multi-epitope vaccine against Ebola virus using reverse vaccinology approach. Scientific reports, 2022. 12(1): p. 7757. [CrossRef]
- Bhattacharya, M., et al., Therapeutic applications of nanobodies against SARS-CoV-2 and other viral infections: Current update. International Journal of Biological Macromolecules, 2023. 229: p. 70-80.
- Gao, J., M. Gui, and Y. Xiang, Structural intermediates in the low pH-induced transition of influenza hemagglutinin. PLoS Pathogens, 2020. 16(11): p. e1009062.
- Chen, X., et al., A cell-free nanobody engineering platform rapidly generates SARS-CoV-2 neutralizing nanobodies. Nature communications, 2021. 12(1): p. 5506.
- Minatel, V.M., et al., Nanobodies: a promising approach to treatment of viral diseases. Frontiers in Immunology, 2024. 14: p. 1303353. [CrossRef]
- Huo, J., et al., Neutralizing nanobodies bind SARS-CoV-2 spike RBD and block interaction with ACE2. Nature structural & molecular biology, 2020. 27(9): p. 846-854.
- Esparza, T.J., et al., High affinity nanobodies block SARS-CoV-2 spike receptor binding domain interaction with human angiotensin converting enzyme. Scientific Reports, 2020. 10(1): p. 22370.
- Chen, Y.-L., et al., Research progresses and applications of fluorescent protein antibodies: a review focusing on nanobodies. International Journal of Molecular Sciences, 2023. 24(5): p. 4307. [CrossRef]
- Veugelen, S., et al., Screening and characterization strategies for nanobodies targeting membrane proteins. Methods in enzymology, 2017. 584: p. 59-97.
- Lu, Q., et al., Development of multivalent nanobodies blocking SARS-CoV-2 infection by targeting RBD of spike protein. Journal of nanobiotechnology, 2021. 19: p. 1-12.
- Li, J.-F., et al., Generation and characterization of a nanobody against SARS-CoV. Virologica Sinica, 2021. 36(6): p. 1484-1491.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




