Submitted:
03 December 2024
Posted:
04 December 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Pre-Processing
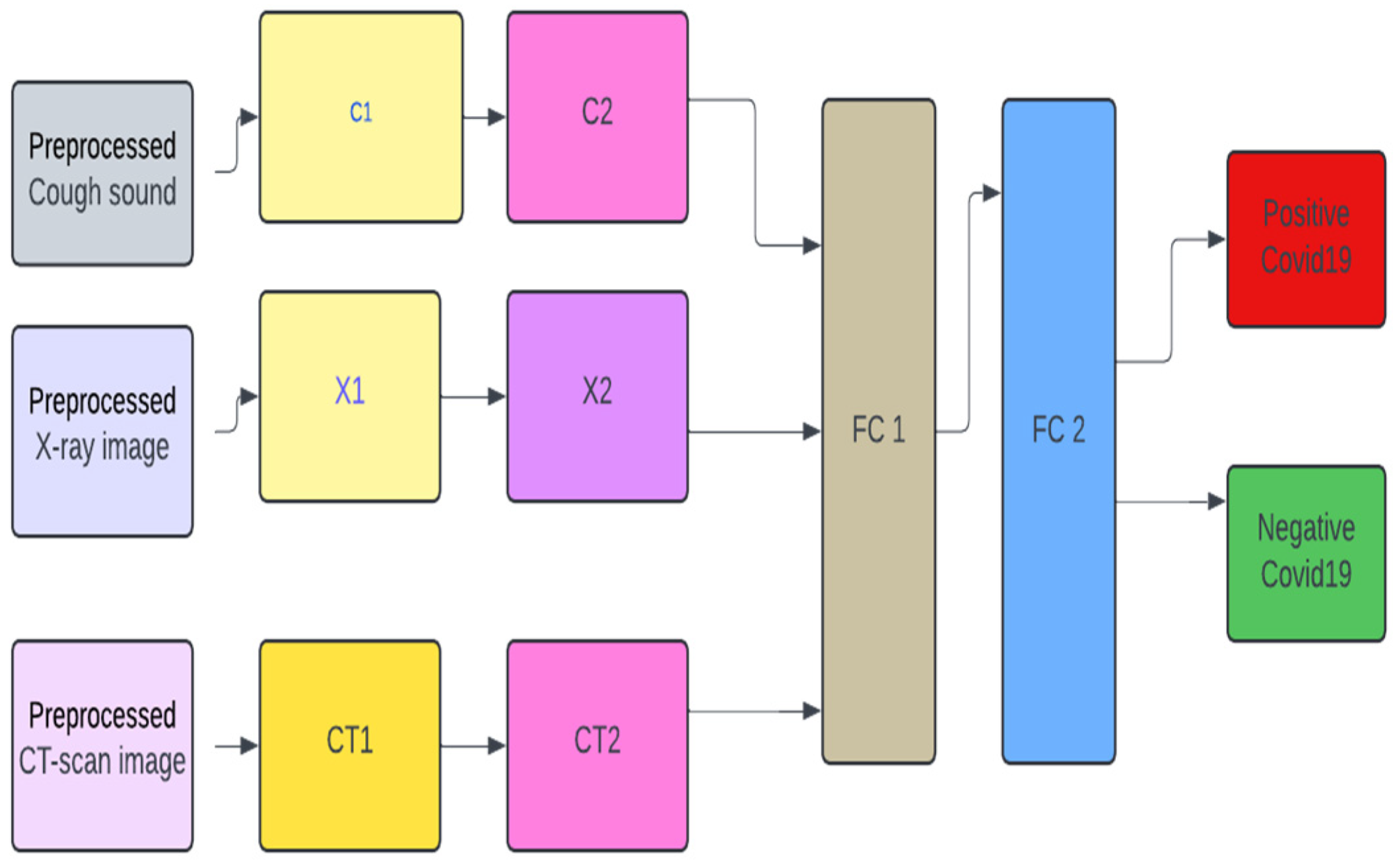
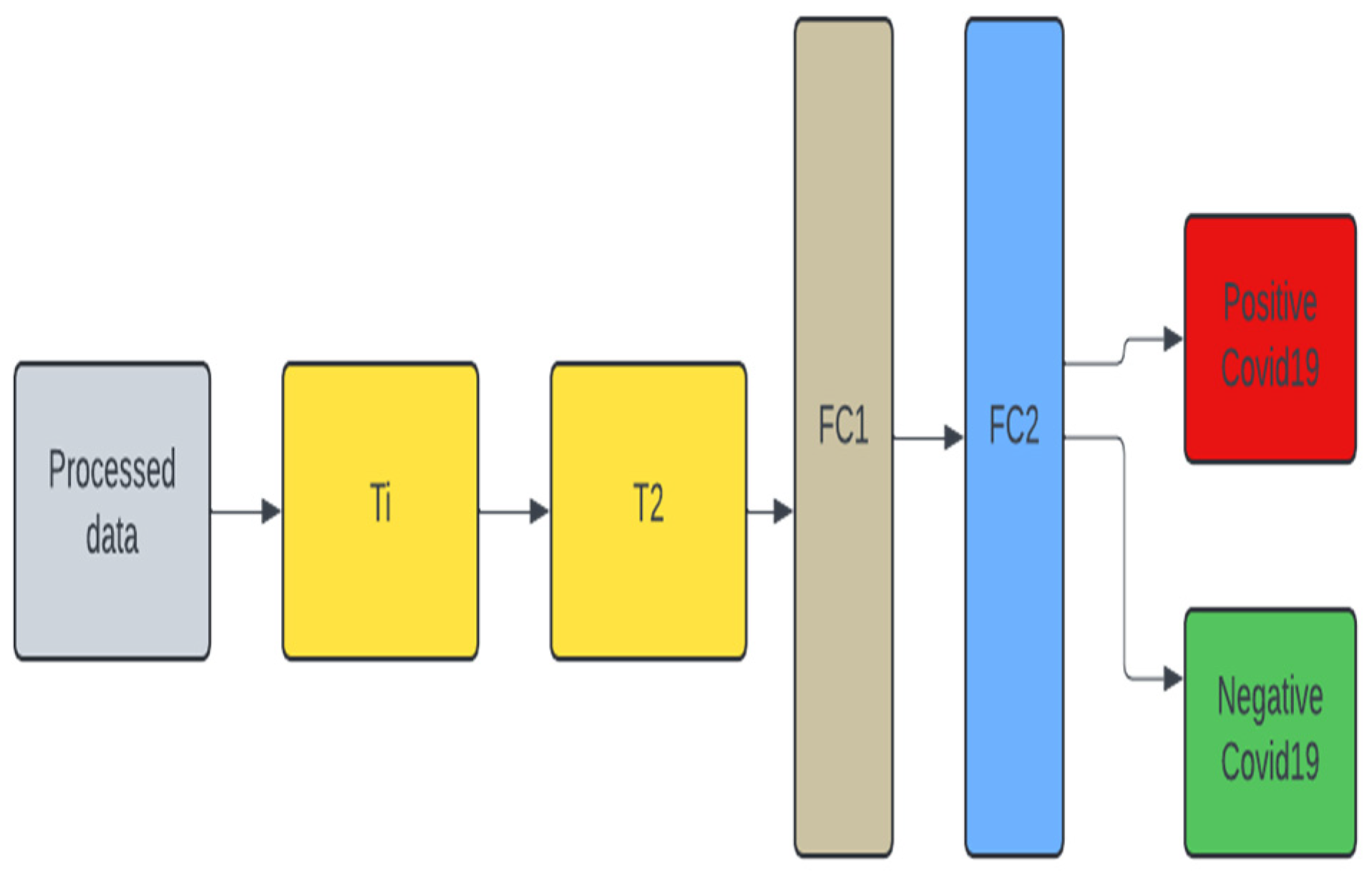
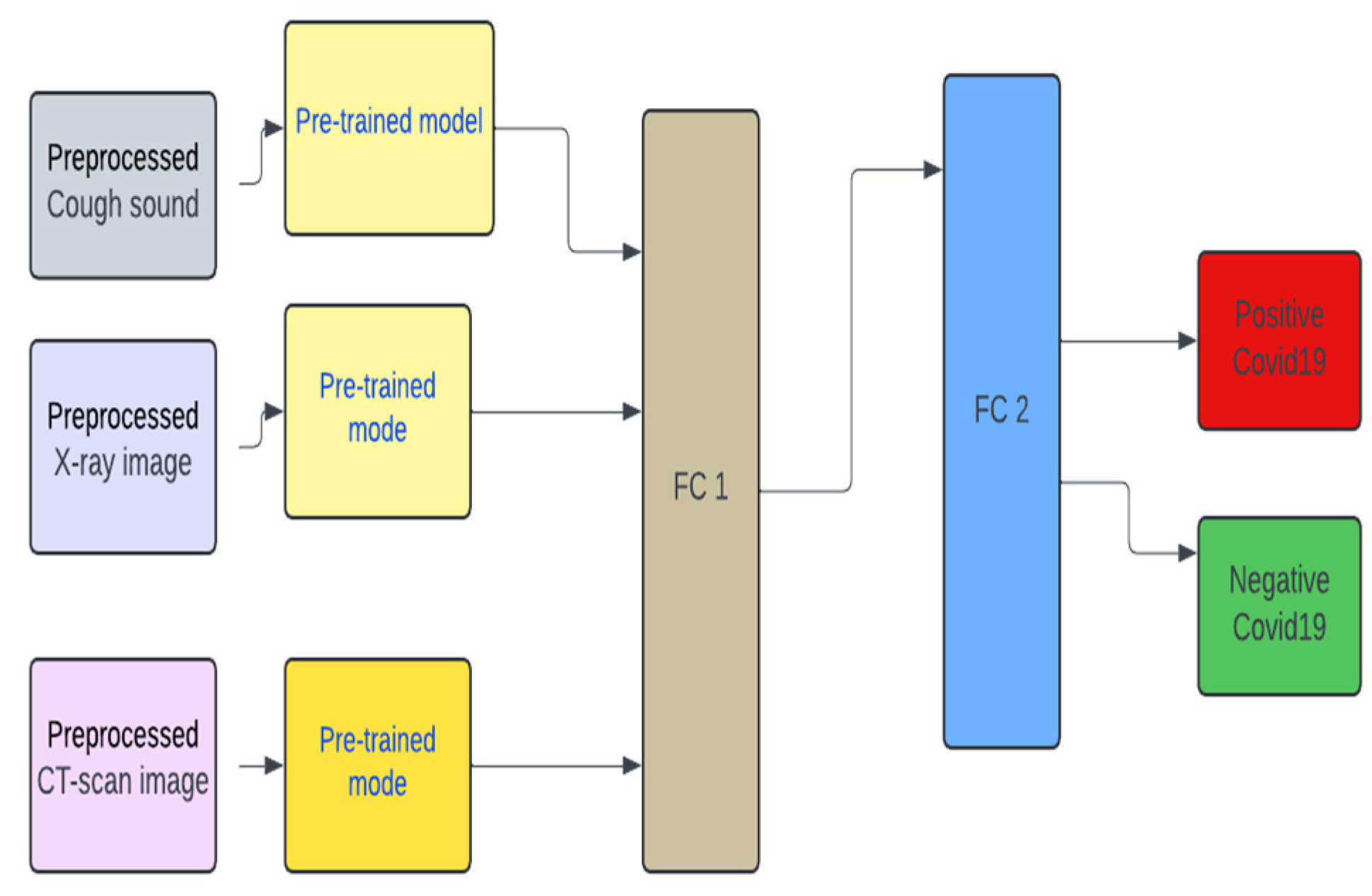
2.2. Model Design
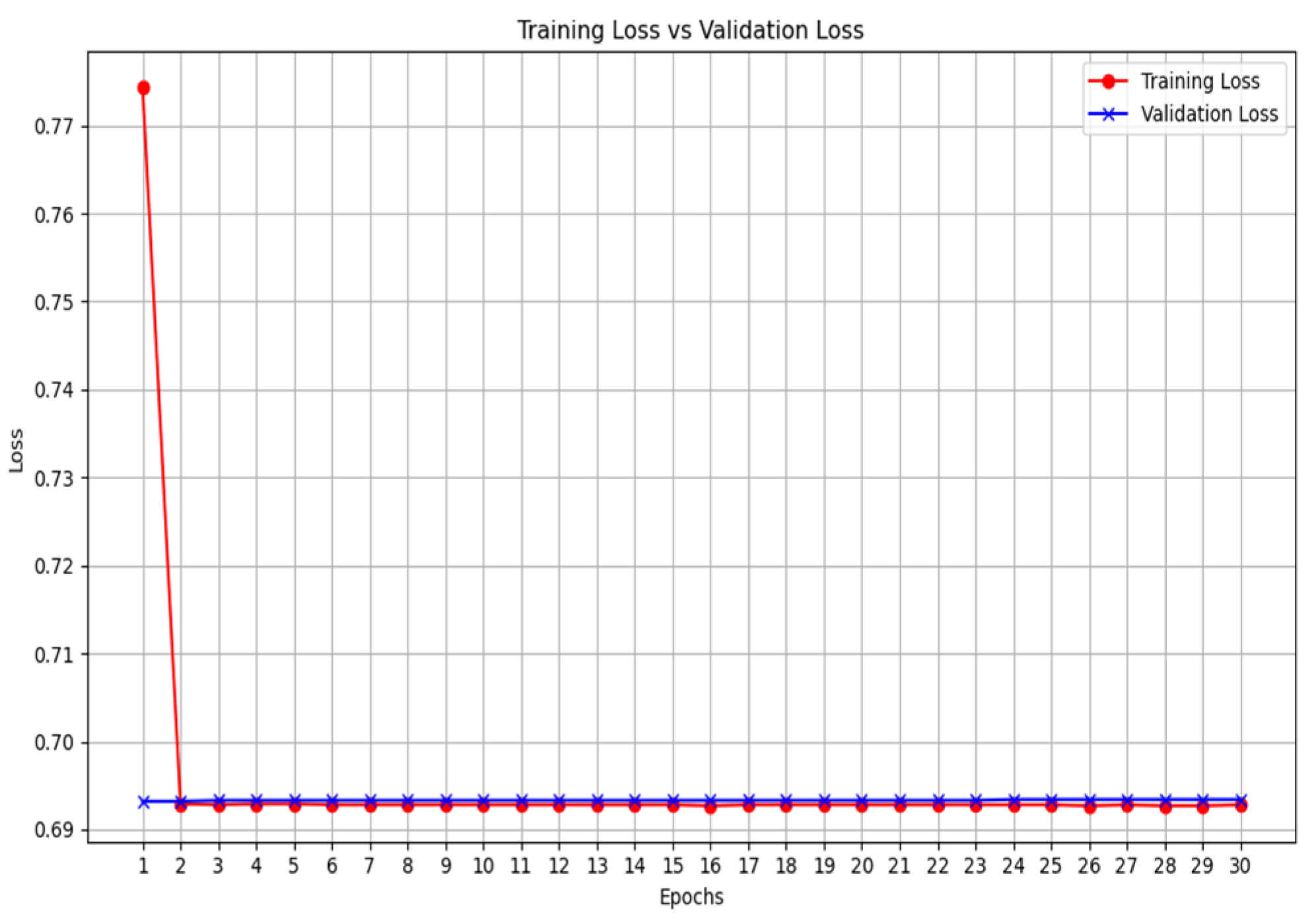
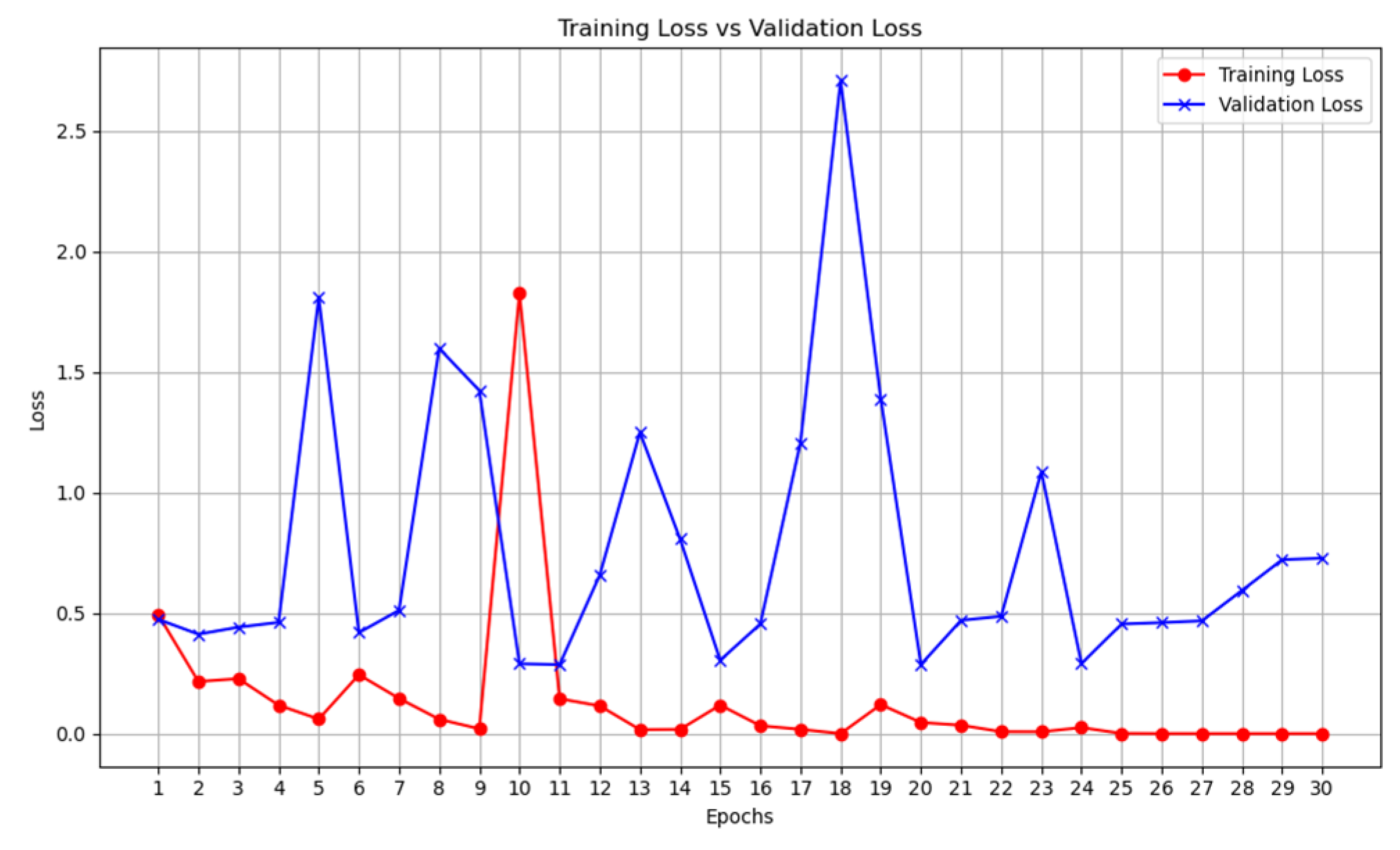
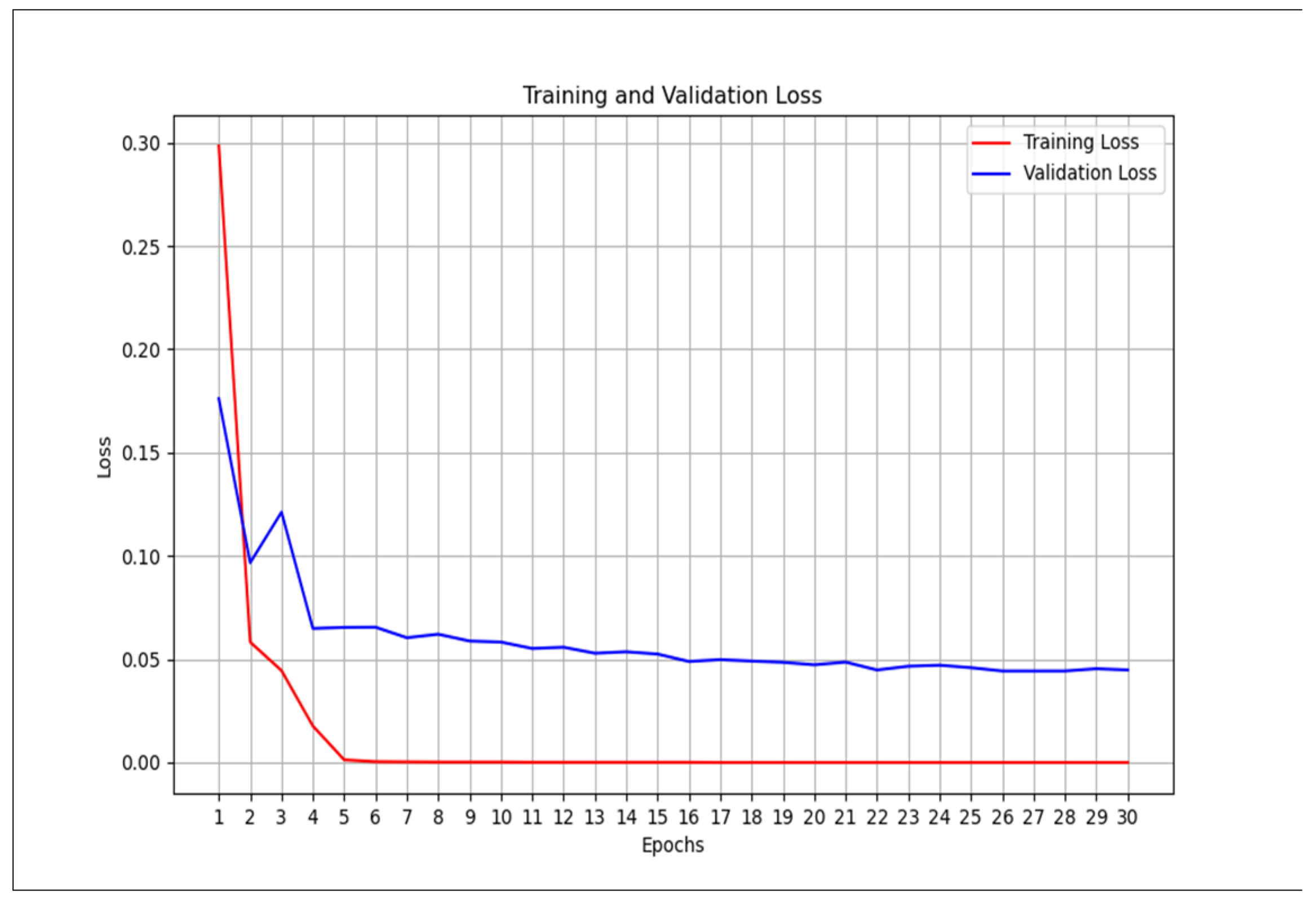
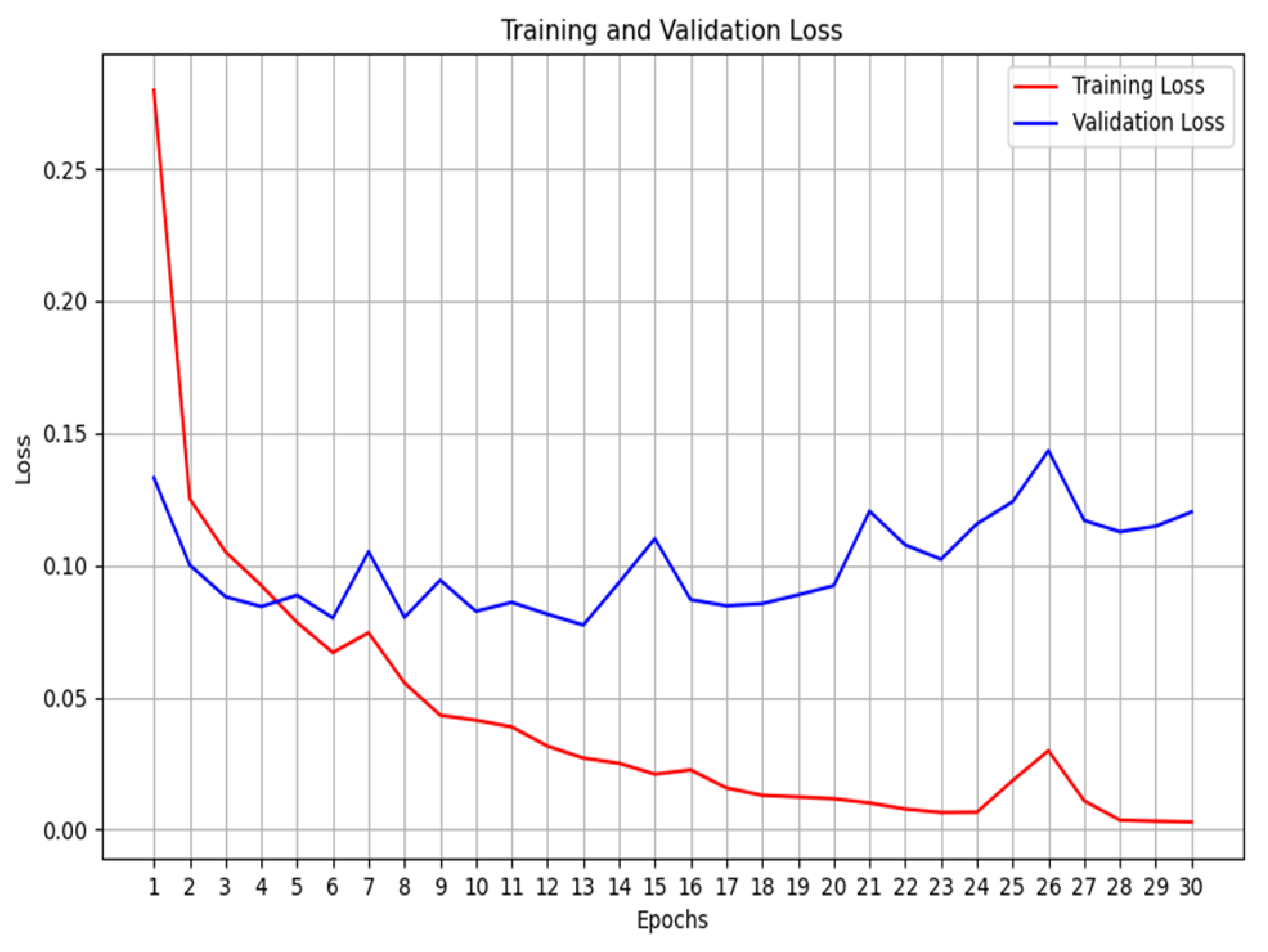
2.3. Training
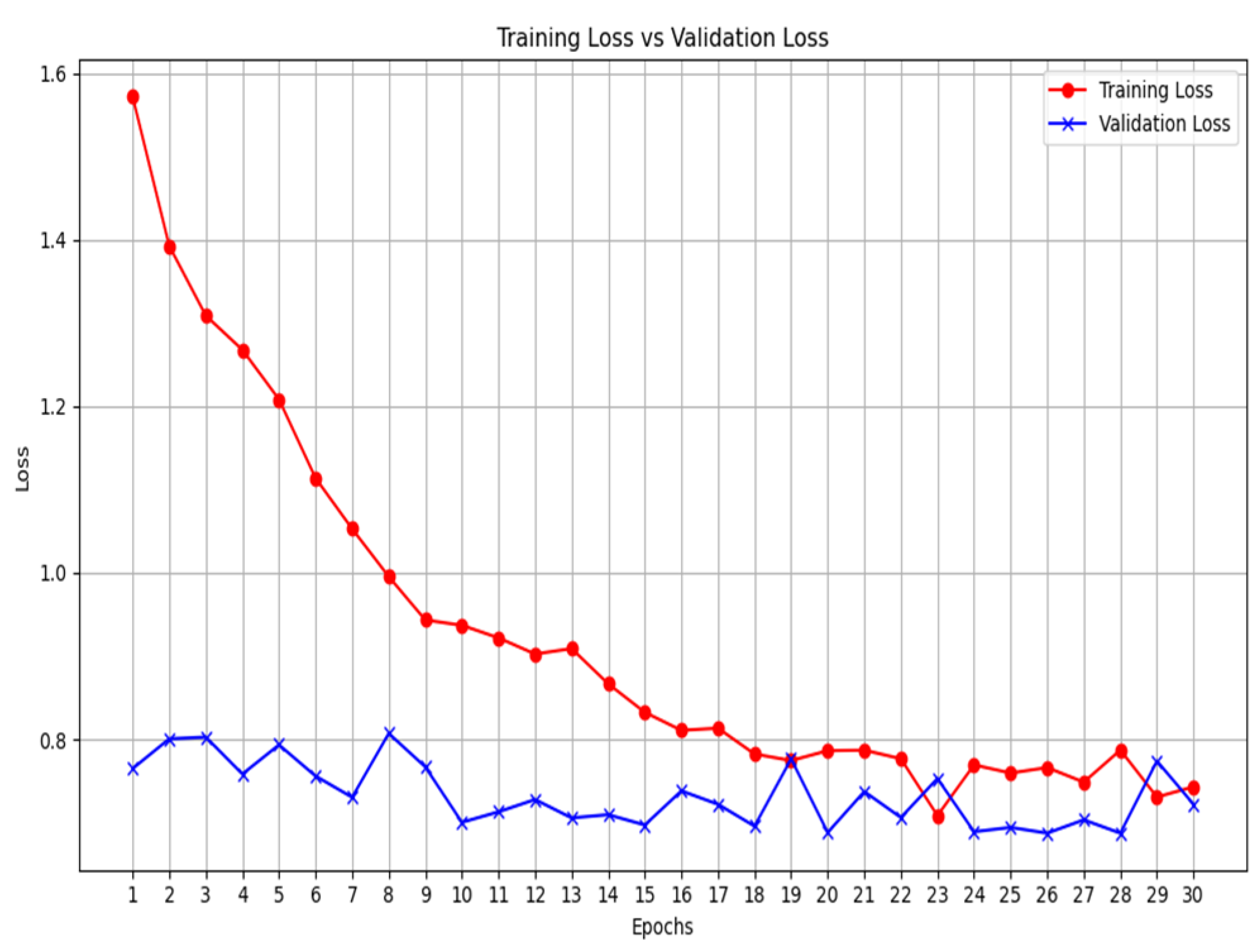
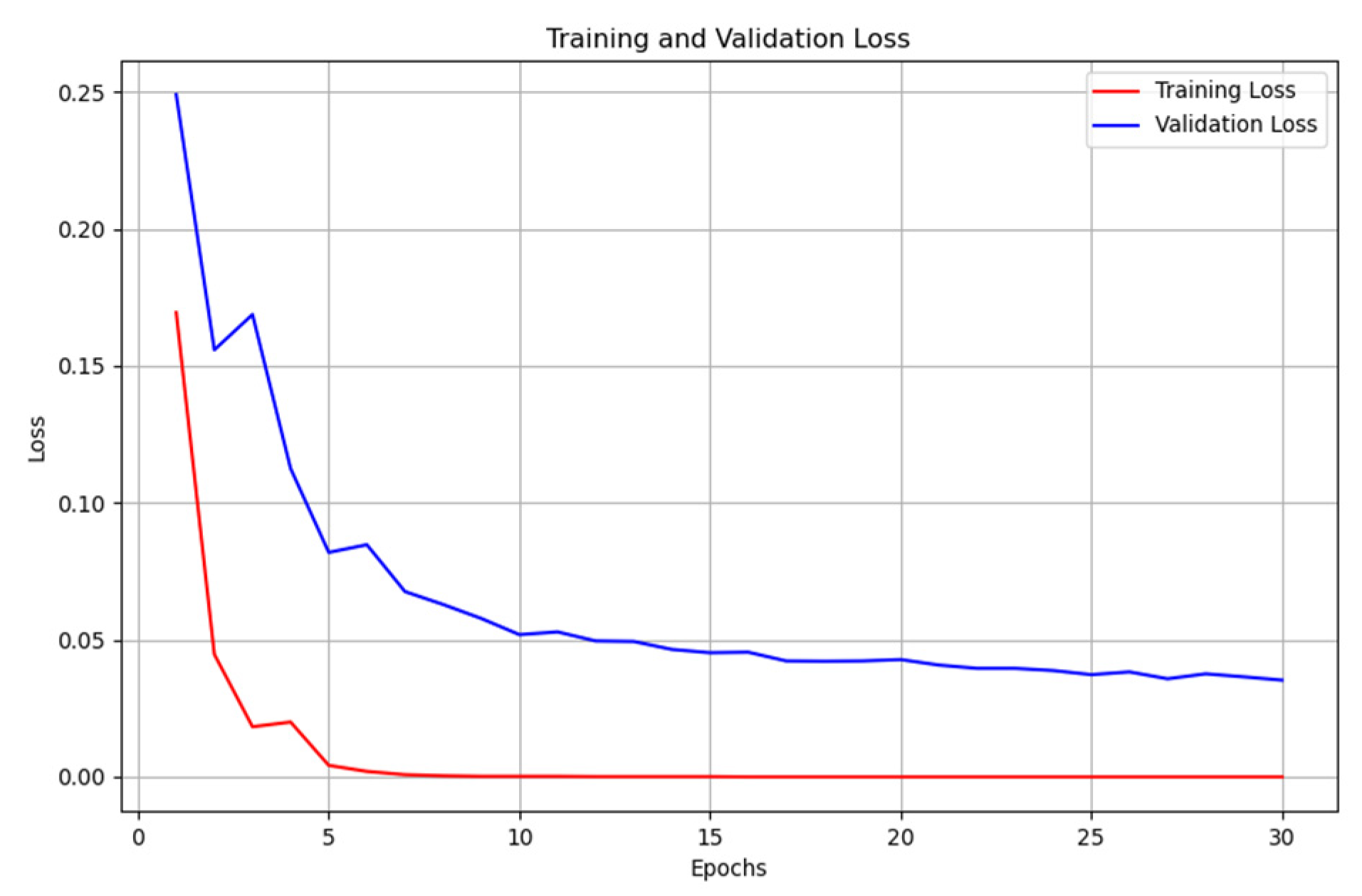
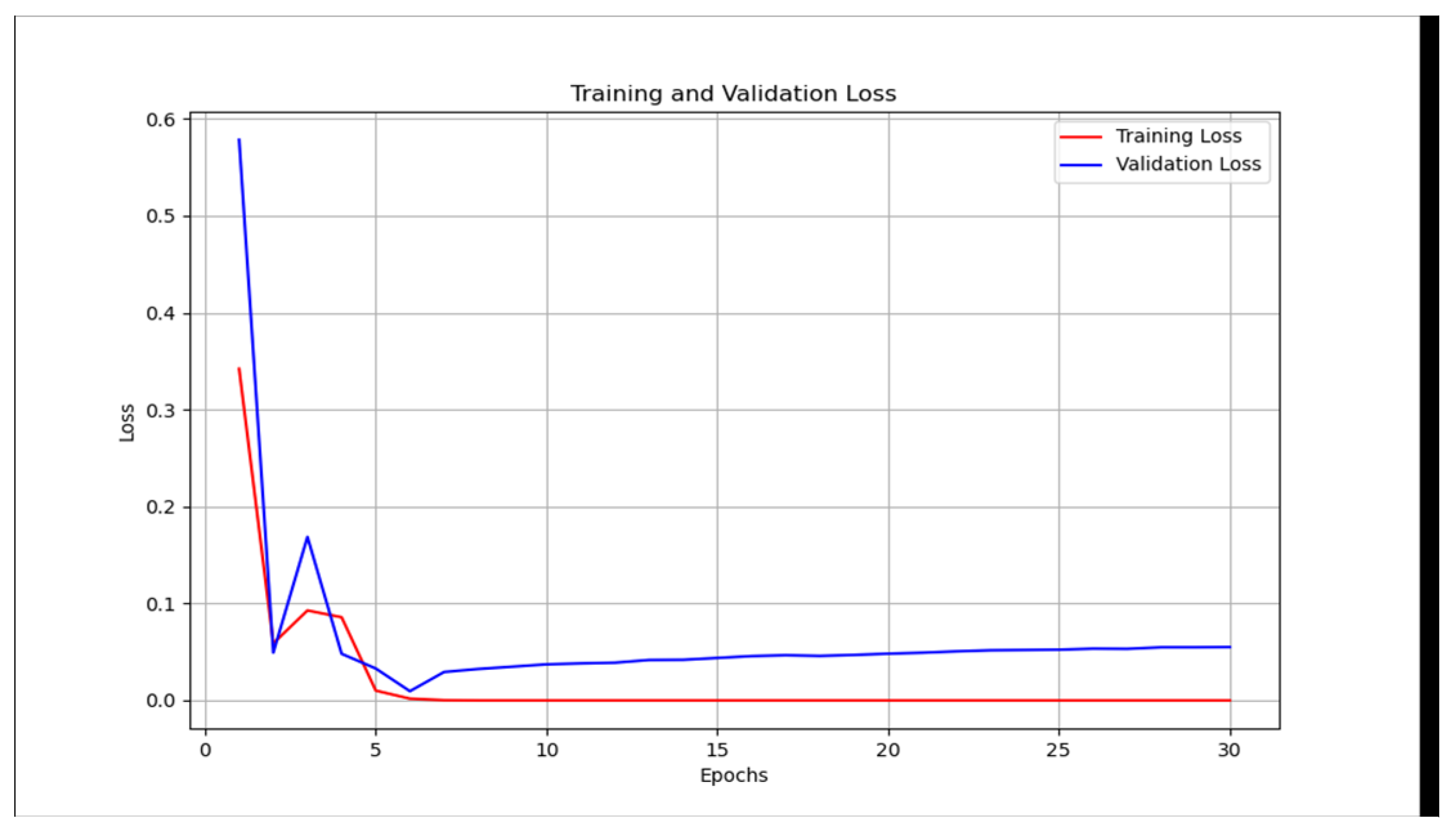
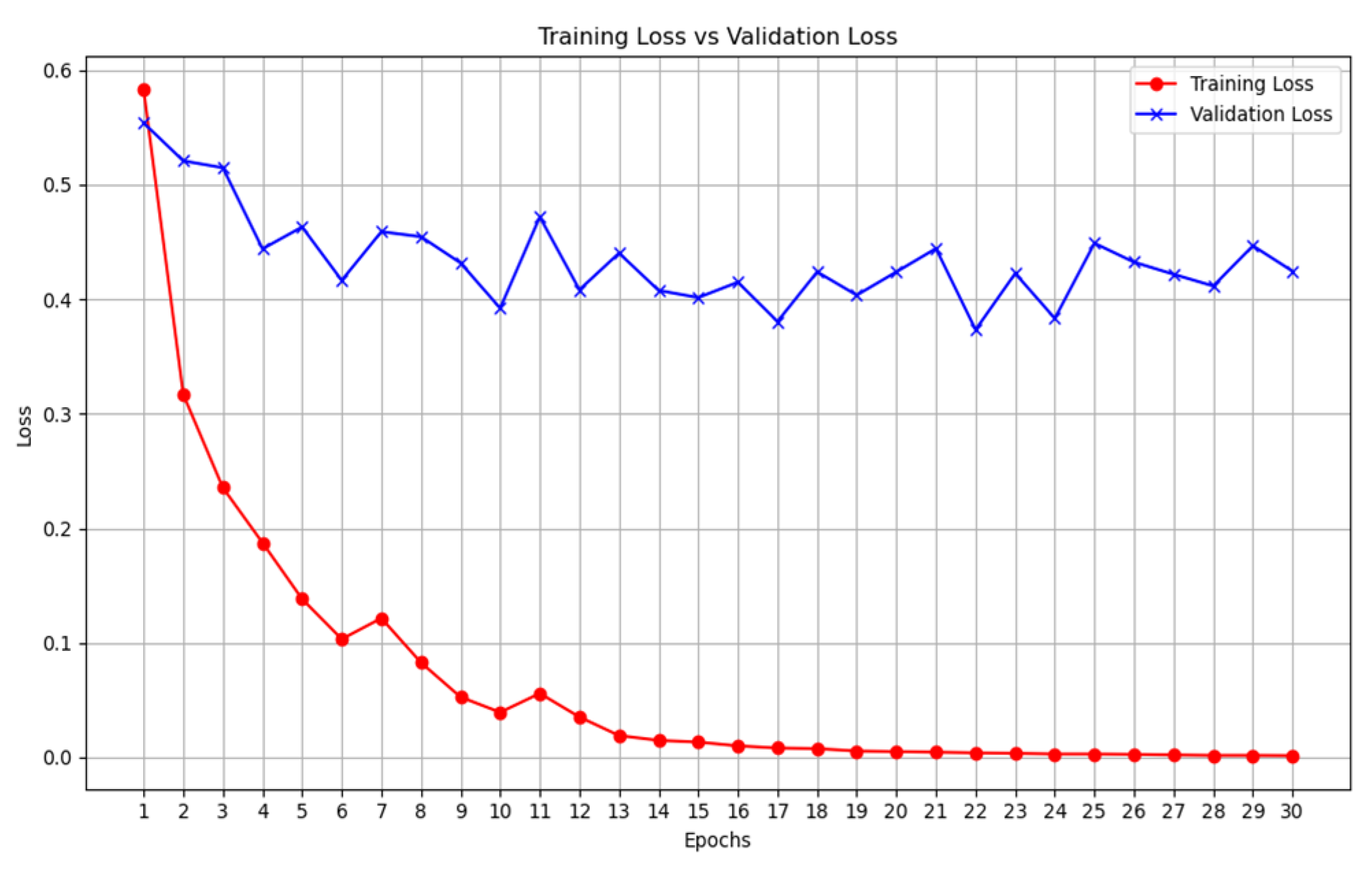
2.4. Validation
- Initialize model’s best loss
- Forward pass
- Compute loss
- Compute accuracy
- We then calculate the total loss on the validation dataset for each epoch.
2.5. Unimodal Systems
2.6. Multimodal Pre-Trained Systems
3. Evaluation
4. Results
5. Discussion
6. Conclusion
Author Contributions
Funding
Informed Consent Statement
Data-Availability-Statement
Acknowledgments
Conflicts of Interest
References
- B. Hu, H. Guo, P. Zhou, and Z.-L. Shi,, "Characteristics of SARS-CoV-2 and COVID-19," Nature Reviews Microbiology, vol. 19, no. 3, pp. 141-154, 2021.
- Y. J. Chen, W. H. Jian, Z. Y. Liang, W. J. Guan, W. H. Liang, R. C. Chen, C. L. Tang, T. Wang, H. R. Liang, Y. M. Li, X. Q. Liu, L. Sang, L. L. Cheng, F. Ye, S. Y. Li, N. F. Zhang, Z. Zhang, Y. Fang, J. X. He, N. S. Zhong, and J. P. Zheng, "Earlier diagnosis improves COVID-19 prognosis: a nationwide retrospective cohort analysis,," Ann. Transl. Med, vol. 9, no. 11, p. 941, 2021. [CrossRef]
- L. J. Layfield, S. Camp, K. Bowers, and D. C. Miller,, "SARS-CoV-2 detection by reverse transcriptase polymerase chain reaction testing: Analysis of false positive results and recommendations for quality control measures," Pathology - Research and Practice, vol. 225, p. 153579, 2021.
- M. Teymouri, S. Mollazadeh, H. Mortazavi, Z. N. Ghale-Noie, V. Keyvani, F. Aghababaei, M. R. Hamblin, G. Abbaszadeh-Goudarzi, H. Pourghadamyari, S. M. R. Hashemian, and H. Mirzaei, , "Recent advances and challenges of RT-PCR tests for the diagnosis of COVID19," athology - Research and Practice, vol. 221, p. 153443, 2021. [CrossRef]
- ". E. M. F. El Houby, "COVID-19 detection from chest X-ray images using transfer learning," Scientific Reports, , vol. 14, p. 11639, 2024. [CrossRef]
- R. Fusco, R. Grassi, V. Granata, S. V. Setola, F. Grassi, D. Cozzi, B. Pecori, F. Izzo, and A. Petrillo, , "Artificial Intelligence and COVID-19 Using Chest CT Scan and Chest X-ray Images: Machine Learning and Deep Learning Approaches for Diagnosis and Treatment," J. Pers. Med, vol. 11, no. 10, p. 993, 2021. [CrossRef]
- M. Melek, "Diagnosis of COVID-19 and non-COVID-19 patients by classifying only a single cough sound,," Neural Comput. Appl., vol. 33, no. 24, p. 17621–17632, 2021. [CrossRef]
- K. Nguyen-Trong and K. Nguyen-Hoang, "Multi-modal Approach for COVID-19 Detection Using Coughs and Self-reported Symptoms," Journal of Intelligent & Fuzzy Systems, vol. 44, no. 3, p. 3501 – 3513, 2023. [CrossRef]
- M. Constantinou, T. Exarchos, A. G. Vrahatis, and P. Vlamos, "COVID-19 classification on chest X-ray images using deep learning methods," Int. J. Environ. Res. Public Health, vol. 20, no. 3, p. 2035, 2023. [CrossRef]
- Z. Li et al.,, "COVID19-ResCapsNet: A novel residual capsule network for COVID-19 detection from chest X-ray scans images," IEEE Access,, vol. 11, pp. 52923-52937, 2023. [CrossRef]
- M. Nahiduzzaman et al, " A novel method for multivariant pneumonia classification based on hybrid CNN-PCA based feature extraction using extreme learning machine with CXR images," IEEE Access, vol. 9, pp. 147512-147526, 2021. [CrossRef]
- U. Haruna, R. Ali, and M. Man, , "A new modification CNN using VGG19 and ResNet50V2 for classification of COVID-19 from X-ray radiograph images," Indonesian Journal of Electrical Engineering and Computer Science, vol. 31, no. 1, pp. 369-377, 2023. [CrossRef]
- K. K. Mohbey, S. Sharma, S. Kumar, and M. Sharma,, "COVID-19 identification and analysis using CT scan images: Deep transfer learning-based approach," Blockchain Applications for Healthcare Informatics, p. 447–470, 2022.
- P. Angelov and E. Soares, , "Towards explainable deep neural networks (xDNN)," Neural Networks, , vol. 130, p. 185–194, 2020. [CrossRef]
- W. Zhao, W. Jiang, and X. Qiu, "Deep learning for COVID-19 detection based on CT images," Scientific Reports, vol. 11, no. 1, p. 14353, 2021. [CrossRef]
- R. Han, L. Huang, H. Jiang, J. Dong, H. Peng, and D. Zhang,, " Early clinical and CT manifestations of coronavirus disease 2019 (COVID-19) pneumonia," American Journal of Roentgenology, vol. 215, no. 2, pp. 338-343, 2020. [CrossRef]
- W. H. Organization, "Clinical management of severe acute respiratory infection (SARI) when COVID-19 disease is suspected: interim guidance," Tech. Rep, 2020.
- J. Han, C. Brown, J. Chauhan, A. Grammenos, A. Hasthanasombat, D. Spathis, T. Xia, P. Cicuta, and C. Mascolo, , "Exploring automatic COVID-19 diagnosis via voice and symptoms from crowdsourced data," in EEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), 2021, pp. 8328-8332.
- I. A. Imran, I. Posokhova, H. N. Qureshi, U. Masood, M. S. Riaz, K. Ali, C. N. John, M. I. Hussain, and M. Nabeel,, " AI4COVID-19: AI enabled preliminary diagnosis for COVID-19 from cough samples via an app.," Informatics in Medicine Unlocked, vol. 20, no. 10, p. 100378, 2020. [CrossRef]
- N. Nasir, A. Kansal, F. Barneih, O. Al-Shaltone, T. Bonny, M. Al-Shabi, and A. Al Shammaa, "Multi-modal image classification of COVID-19 cases using computed tomography and X-rays scans," Intelligent Systems with Applications, vol. 15, p. 200160, 2023. [CrossRef]
- N. Hilmizen, A. Bustamam, and D. Sarwinda, "The Multimodal Deep Learning for Diagnosing COVID-19 Pneumonia from Chest CT-Scan and X-Ray Images," in 3rd International Seminar on Research of Information Technology and Intelligent Systems (ISRITI),, Yogyakarta, Indonesia,, 2020.
- S. Kumar, M. K. Chaube, S. H. Alsamhi, S. K. Gupta, M. Guizani, R. Gravina, and G. Fortino, , "A novel multimodal fusion framework for early diagnosis and accurate classification of COVID-19 patients using X-ray images and speech signal processing techniques," Comput. Methods Programs Biomed, vol. 226, p. 107109, 2022. [CrossRef]
- S. E. Mukhi, R. T. Varshini, and S. E. F. Sherley,, " Diagnosis of COVID-19 from Multimodal Imaging Data Using Optimized Deep Learning Techniques," SN Comput. Sci.,, vol. 4, p. 212, 2023.
- E. Jangam, A. A. D. Barreto, and C. S. R. Annavarapu, "E. Jangam, A. A. D. Barreto, and C. S. R. Annavarapu, "Automatic detection of COVID-19 from chest Automatic detection of COVID-19 from chest CT scan and chest X-Rays images using deep learning, transfer learning and stacking," Appl. Intell.,, vol. 52, p. 2243–2259, 2022.
- C. Mahanty et al, "A Comprehensive Review on COVID-19 Detection Based on Cough Sounds, Symptoms, CXR, and CT Images," IEEE Access, vol. 12, pp. 75412-75425, 2024.
- A. M. V. D. Andrew, "COVID-19 Cough Audio Classification," Kaggle, 2020 . [Online]. Available: https://www.kaggle.com/datasets/andrewmvd/covid19-cough-audio-classification/discussion?sort=hotness. [Accessed 12 July 2024].
- D. Griffin and J. Lim, "Signal Estimation from Modified Short-Time Fourier Transform," IEEE Transactions on Acoustics, Speech, and Signal Processing,, vol. 32, no. 2, pp. 236-243,, 1984. [CrossRef]
- P. Prashant, "Chest X-ray Images (COVID-19, Pneumonia)," 2020. [Online]. Available: https://www.kaggle.com/datasets/prashant268/chest-xray-covid19-pneumonia. [Accessed 20 July 2024].
- J. P. Cohen, P. Morrison, L. Dao, K. Roth, T. Q. Duong, and M. Ghassemi, "COVID-19 Image Data Collection: Prospective Predictions Are the Future," 2020. [Online]. Available: https://github.com/ieee8023/covid-chestxray-dataset.. [Accessed 12 July 2024]. [CrossRef]
- A. G. Chung, "Figure 1 COVID Chest X-ray Dataset," 2020. [Online]. Available: https://github.com/agchung/Figure1-COVID-chestxray-dataset. [Accessed 12 July 2024].
- J. Tiptj, "Chest X-ray Pneumonia COVID-19 Tuberculosis Dataset," 2020. [Online]. Available: https://www.kaggle.com/datasets/jtiptj/chest-xray-pneumoniacovid19tuberculosis. [Accessed 12 July 2024].


| Evaluation metrics | Values |
|---|---|
| Accuracy | 50% |
| Sensitivity | 1 |
| Specificity | 0 |
| F1 Score | 0.6667 |
| Confusion matrix True Positives (TP) True Negatives (TN) False Positives (FP): False Negatives (FN): |
[[0 100] [0 100]] 100 0 100 0 |
| Evaluation metrics | Values |
|---|---|
| Accuracy | 57.50% |
| Sensitivity | 0.69 |
| Specificity | 0.46 |
| F1 Score | 0.6188 |
| Confusion matrix True Positives (TP) True Negatives (TN) False Positives (FP): False Negatives (FN): |
[[46 54] [32 69]] 69 46 54 32 |
| Evaluation metrics | Values |
|---|---|
| Accuracy | 98.00% |
| Sensitivity | 1 |
| Specificity | 0.960 |
| F1 Score | 0.9804 |
| Confusion matrix True Positives (TP) True Negatives (TN) False Positives (FP): False Negatives (FN): |
[[96 4] [0 100]] 100 96 4 0 |
| Evaluation metrics | Values |
|---|---|
| Accuracy | 99.00% |
| Sensitivity | 1 |
| Specificity | 0.98 |
| F1 Score | 0.9901 |
| Confusion matrix True Positives (TP) True Negatives (TN) False Positives (FP): False Negatives (FN): |
[[98 2] [0 100]] 100 98 2 0 |
| Evaluation metrics | Values |
|---|---|
| Accuracy | 91.00% |
| Sensitivity | 0.91 |
| Specificity | 0.91 |
| F1 Score | 0.91 |
| Confusion matrix True Positives (TP) True Negatives (TN) False Positives (FP): False Negatives (FN): |
[[91 9] [9 91]] 91 91 9 9 |
| Evaluation metrics | Values |
|---|---|
| Accuracy | 77.50% |
| Sensitivity | 0.55 |
| Specificity | 1 |
| F1 Score | 0.7097 |
| Confusion matrix True Positives (TP) True Negatives (TN) False Positives (FP): False Negatives (FN): |
[[100 0] [45 55]] 55 100 0 45 |
| Evaluation metrics | Values |
|---|---|
| Accuracy | 98.00% |
| Sensitivity | 1 |
| Specificity | 0.9600 |
| F1 Score | 0.9804 |
| Confusion matrix True Positives (TP) True Negatives (TN) False Positives (FP): False Negatives (FN): |
[[96 4] [0 100]] 100 96 4 0 |
| Evaluation metrics | Values |
|---|---|
| Accuracy | 98.00 % |
| Sensitivity | 0.98 |
| Specificity | 0.98 |
| F1 Score | 0.980 |
| Confusion matrix True Positives (TP) True Negatives (TN) False Positives (FP): False Negatives (FN): |
[[98 2] [2 98]] 98 98 2 2 |
| Evaluation metrics | Values |
|---|---|
| Accuracy | 50.00 % |
| Sensitivity | 1 |
| Specificity | 0 |
| F1 Score | 0.6667 |
| Confusion matrix True Positives (TP) True Negatives (TN) False Positives (FP): False Negatives (FN): |
[[0 100] [0 100]] 100 0 100 0 |
| Evaluation metrics | Values |
|---|---|
| Accuracy | 96 % |
| Sensitivity | 0.96 |
| Specificity | 0.96 |
| F1 Score | 0.96 |
| Confusion matrix True Positives (TP) True Negatives (TN) False Positives (FP): False Negatives (FN): |
[[96 4] [4 96]] 96 96 4 4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




