Submitted:
26 November 2024
Posted:
28 November 2024
You are already at the latest version
Abstract
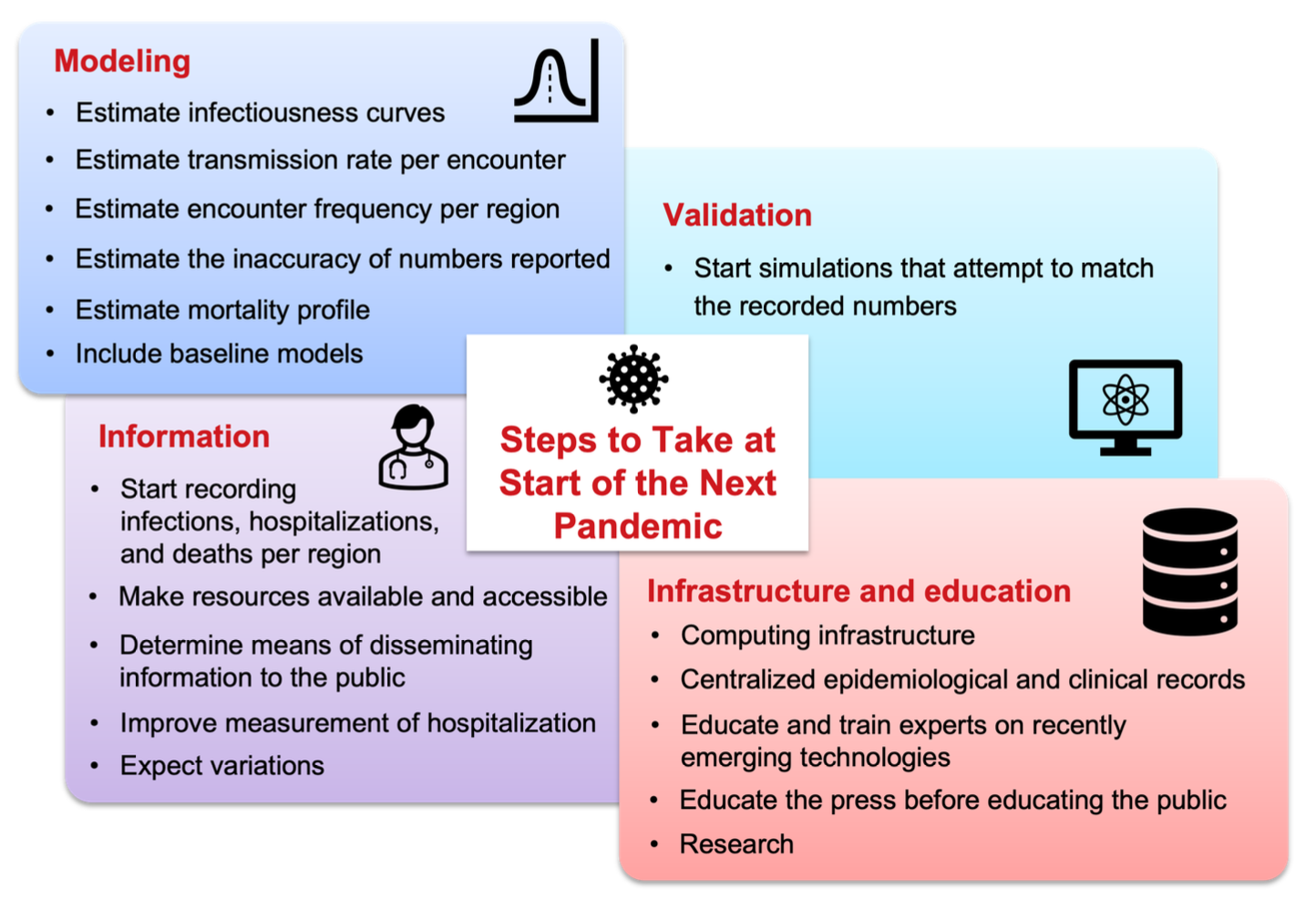
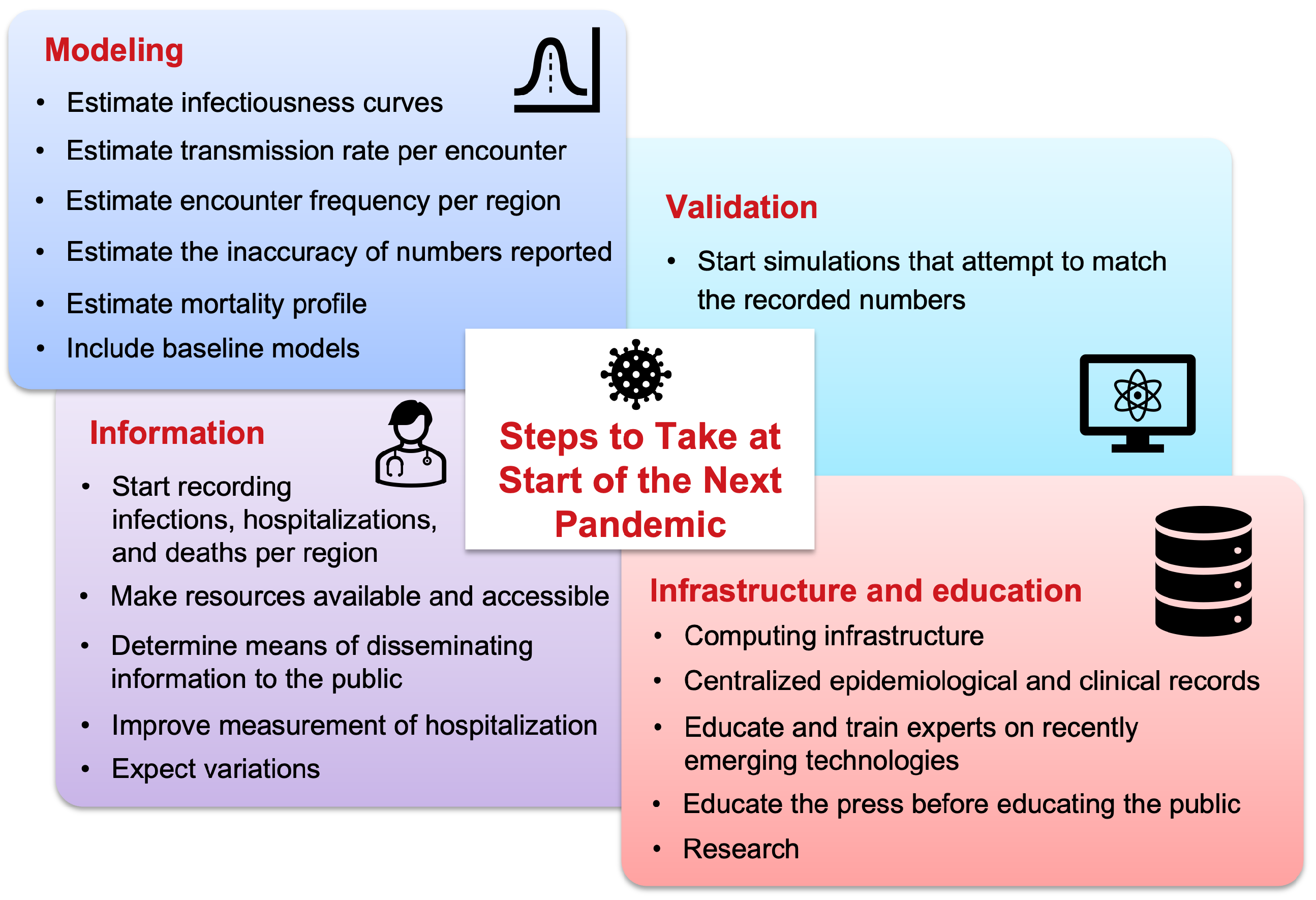
The COVID-19 pandemic spurred many computational modeling efforts. Many mistakes were made and many lessons were learned. This study attempts to list the key lessons learned from a modeling perspective, highlighting both the successes and shortcomings observed during the pandemic. Additionally, this work attempts to compile a set of critical steps and best practices that the authors believe would prove helpful and should be implemented before the start of the next pandemic to avoid inaccuracies in modeling pandemic scenarios. This will help to improve preparedness and ensure that computational models can more effectively guide decision-making in future pandemics.
Keywords:
Introduction
1. Record Infections, Hospitalizations, and Deaths per Region
2. Estimate Infectiousness Curves
3. Estimate Transmission Rate per Encounter
4. Estimate Encounter Frequency per Region
5. Estimate Mortality Profile
6. Estimate the Inaccuracy of Reported Numbers
7. Start Simulations that Will Attempt to Match the Recorded Numbers
8. Make Resources Available and Accessible
9. Improve Measurement of Hospitalization
10. Include Baseline Models
10.1. For Government Officials
10.2. For Researchers
11. Expect Variations
12. Determine Means of Disseminating Information to the Public
12.1. Computing Infrastructure
12.2. Centralized Epidemiological and Clinical Records
12.3. Educate and Train Experts on Recently Emerging Technologies
12.4. Educate the Press Before Educating the Public
12.5. Research
Contributions
Conflict of Interests Statement
Transparency
Acknowledgments
References
- Chharia, A. et al. Accuracy of US CDC COVID-19 forecasting models. Frontiers in Public Health.
- The COVID Tracking Project at the Atlantic. https://covidtracking.com/. Accessed: 2024-9-6.
- Varkila, M. R. et al. Use of wastewater metrics to track COVID-19 in the US. JAMA Network Open. [CrossRef]
- Böttcher, L. , Xia, M. & Chou, T. Why case fatality ratios can be misleading: individual-and population-based mortality estimates and factors influencing them. Physical biology 17, 065003. [Google Scholar] [CrossRef]
- Schreiber, M. CDC coding error led to overcount of 72,000 Covid deaths. https://www.theguardian.com/world/2022/mar/24/cdc-coding-error-overcount-covid-deaths (2022).
- U.S. Department of Homeland Security. Master Question List for COVID-19 (caused by SARS-CoV-2) Weekly Report . https://www.dhs.gov/sites/default/files/publications/mql_sars-cov-2_-_cleared_for_public_release_20200526.pdf. 26 May.
- Ke, R. , Zitzmann, C., Ribeiro, R. M. & Perelson, A. S. Kinetics of SARS-CoV-2 infection in the human upper and lower respiratory tracts and their relationship with infectiousness. MedRxiv. [CrossRef]
- Hart, W. S. , Maini, P. K. & Thompson, R. N. High infectiousness immediately before COVID-19 symptom onset highlights the importance of continued contact tracing. Elife. [CrossRef]
- Del Valle, S. Y. , Hyman, J. M., Hethcote, H. W. & Eubank, S. G. Mixing patterns between age groups in social networks. Social Networks. [CrossRef]
- Edmunds, W. J. , O’callaghan, C. & Nokes, D. Who mixes with whom? A method to determine the contact patterns of adults that may lead to the spread of airborne infections. Proceedings of the Royal Society of London. Series B: Biological Sciences. [CrossRef]
- Mobility trends reports. https://covid19.apple.com/mobility. Accessed: 2020-11-7.
- United States Census Bureau: Explore Census Data. https://data.census.gov (2020).
- Forgoston, E. & Thorne, M. A. Strategies for controlling the spread of COVID-19. MedRxiv. [CrossRef]
- Forgoston, E. Stay at home orders by state for COVID-19: Spreadsheets containing start dates and relaxation dates for stay-at-home and closure orders by state in the USA. https://github.com/eric-forgoston/Stay-at-home-orders-by-state-for-COVID-19.
- Assisting businesses with COVID-19 orders and helping them effectively continue operations. https://www.huschblackwell.com/state-by-state-covid-19-guidance. Accessed: 2024-3-27.
- Bialek, S. 12 February 2019; 16. [CrossRef]
- Castiglione, F. , Deb, D., Srivastava, A. P., Liò, P. & Liso, A. From infection to immunity: understanding the response to SARS-CoV2 through in-silico modeling. Frontiers in Immunology. [CrossRef]
- Barhak, J. COVID-19 mortality model by Filippo Castiglione et. al. https://github.com/Jacob-Barhak/COVID19Models/tree/main/COVID19_Mortality_Castiglione (2021).
- Barhak, J. SimTK: The Reference Model for disease progression. https://simtk.org/projects/therefmodel.
- Barhak, J. The Reference Model: An Initial Use Case for COVID-19. Online: https://www.cureus.com/articles/36677-the-reference-model-an-initial-use-case-for-covid-19, Interactive Results: https://jacob-barhak.netlify. 2020. [Google Scholar] [CrossRef]
- Hart, W. S. et al. Generation time of the alpha and delta SARS-CoV-2 variants: an epidemiological analysis. The Lancet Infectious Diseases. [CrossRef]
- Nandakishore, P. et al. Deviations in predicted covid-19 cases in the us during early months of 2021 relate to rise in B.1.526 and its family of variants. 1.526 and its family of variants. medRxiv 2021–12. [CrossRef]
- Rufino, J. et al. Using survey data to estimate the impact of the omicron variant on vaccine efficacy against COVID-19 infection. Scientific Reports. [CrossRef]
- Natraj, S. et al. COVID-19 activity risk calculator as a gamified public health intervention tool. Scientific Reports. [CrossRef]
- Barhak, J. The Reference Model for COVID-19 attempts to explain USA data. MODSIM World 2023 -23 booth, Presentation: https://www.clinicalunitmapping.com/show/COVID19_Ensemble_Latest.html (2023). 22 May.
- Barhak, J. MIST: Micro-Simulation Tool to Support Disease Modeling. SciPy, Bioinformatics track, https://github.com/scipy/scipy2013_talks/tree/master/talks/jacob_barhak, Presentation also available at: https://jacob-barhak.github.io/old/SciPy2013_MIST_Presented_2013_06_26.pptx Video of Talk: https://www.youtube.com/watch?v=AD896WakR94 (2013).
- Kerr, C. Python vs. the pandemic: writing high-performance models in a jiffy. https://youtu.be/tG3K4Oq2W6M?si=VuZLFjnPYKYEOye0 (2021).
- Institute for Disease Modeling. COVID-19 Agent-based Simulator (Covasim): a model for exploring coronavirus dynamics and interventions. https://github.com/InstituteforDiseaseModeling/covasim (2024).
- Institute of Health Metrics and Evaluation (IHME). Vivarium: A python microsimulation framework. https://github.com/ihmeuw/vivarium.
- Deason, A. Small Potatoes: Microsimulation With Vivarium. PyTexas https://youtu.be/A7yWl4oeLhQ?si=Psceb1u_FlO-8sTm (2017).
- Barhak, J. Reference model for disease progression. https://ppubs.uspto.gov/dirsearch-public/print/downloadPdf/9858390 (2018). U.S. Patent US9858390B2.
- Barhak, J. Analysis and Verification of Models Derived from Clinical Trials Data Extracted from a Database. https://ppubs.uspto.gov/dirsearch-public/print/downloadPdf/10923234 (2021). U.S. Patent US20170286627A1.
- Mathieu, E. et al. Coronavirus Pandemic (COVID-19). Our World in Data.
- Hadfield, J. et al. Nextstrain: real-time tracking of pathogen evolution. Bioinformatics. [CrossRef]
- Kermack, W. O. & McKendrick, A. G. A contribution to the mathematical theory of epidemics. Proceedings of the royal society of london. Series A, Containing papers of a mathematical and physical character. [CrossRef]
- Lam, S. K. , Pitrou, A. & Seibert, 1–6 (2015)., S. Numba: A llvm-based python jit compiler. In Proceedings of the Second Workshop on the LLVM Compiler Infrastructure in HPC.
- Seibert, S. Evening of Python Coding 2021-09-21: Numba Demo. https://youtu.be/dl8JnpO7vBY?si=rf6jVXrzppAlKzJu, https://numba.pydata.org/ (2021).
- Nickolls, J. , Buck, I., Garland, M. & Skadron, K. Scalable parallel programming with cuda: Is cuda the parallel programming model that application developers have been waiting for? Queue, 2008. [Google Scholar]
- Barhak, J. The Reference Model: A Decade of Healthcare Predictive Analytics with Python. PyTexas 2017, Nov 18-19, 2017, Galvanize, Austin TX. Video: https://youtu.be/Pj_N4izLmsI Presentation: http://sites.google.com/site/jacobbarhak/home/PyTexas2017_Upload_2017_11_18.pptx (2017).
| a | Rescale High-Performance Computing https://rescale.com/
|
| b | The Microsoft Cloud https://www.microsoft.com/en-us/microsoft-cloud
|
| c | Amazon AWS https://aws.amazon.com/
|
| d | MIDAS Network https://midasnetwork.us/
|
| e | COVID-19 Observatory https://covid19obs.fbk.eu/#/
|
| f | Genomic epidemiology of SARS-CoV-2 with subsampling focused globally over the past 6 months: https://nextstrain.org/ncov/gisaid/global/6m
|
| g | |
| h | MIDAS network COVID papers https://midasnetwork.us/covid-19/
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



