Submitted:
19 November 2024
Posted:
21 November 2024
You are already at the latest version
Abstract
Keywords:
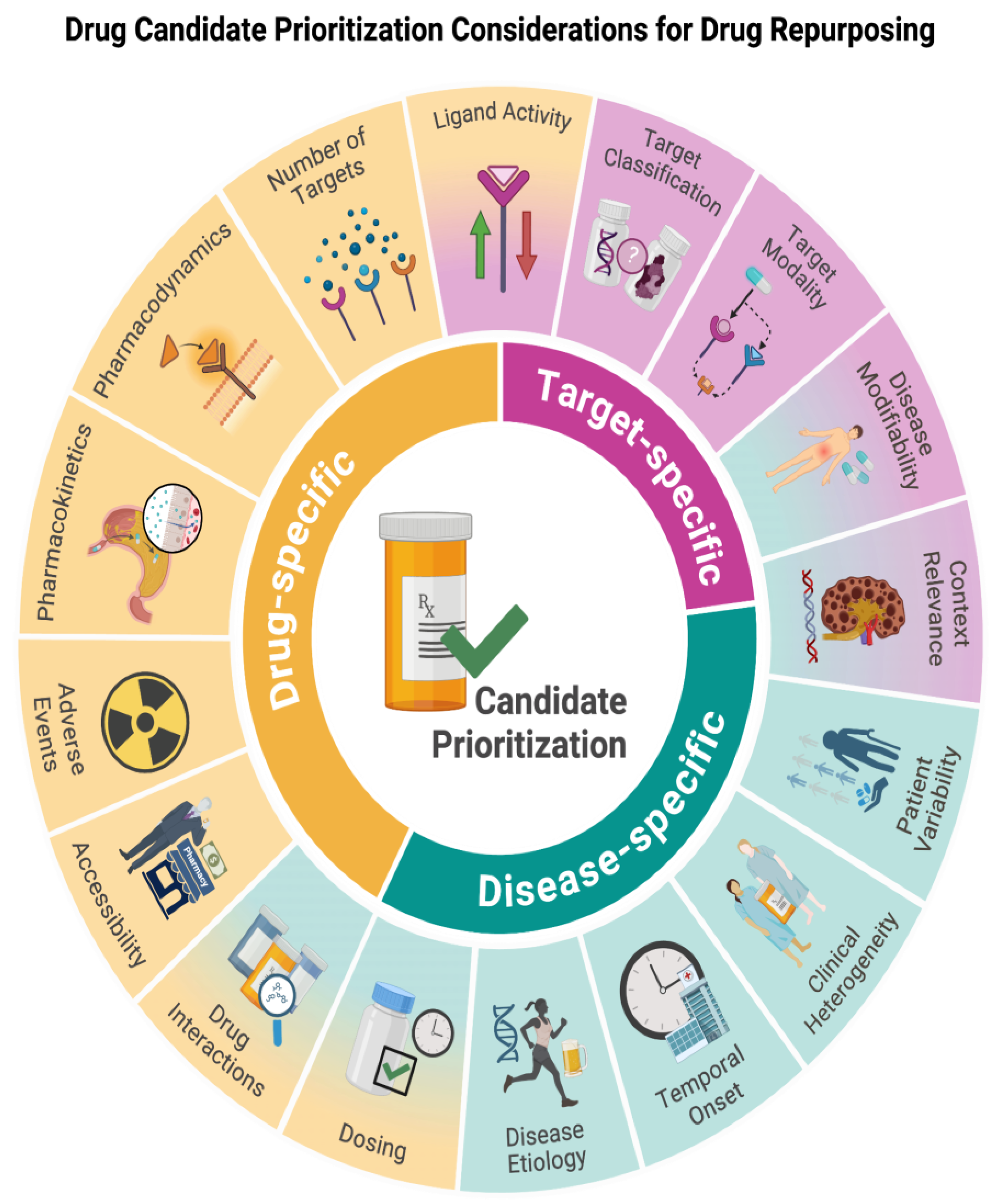
Introduction
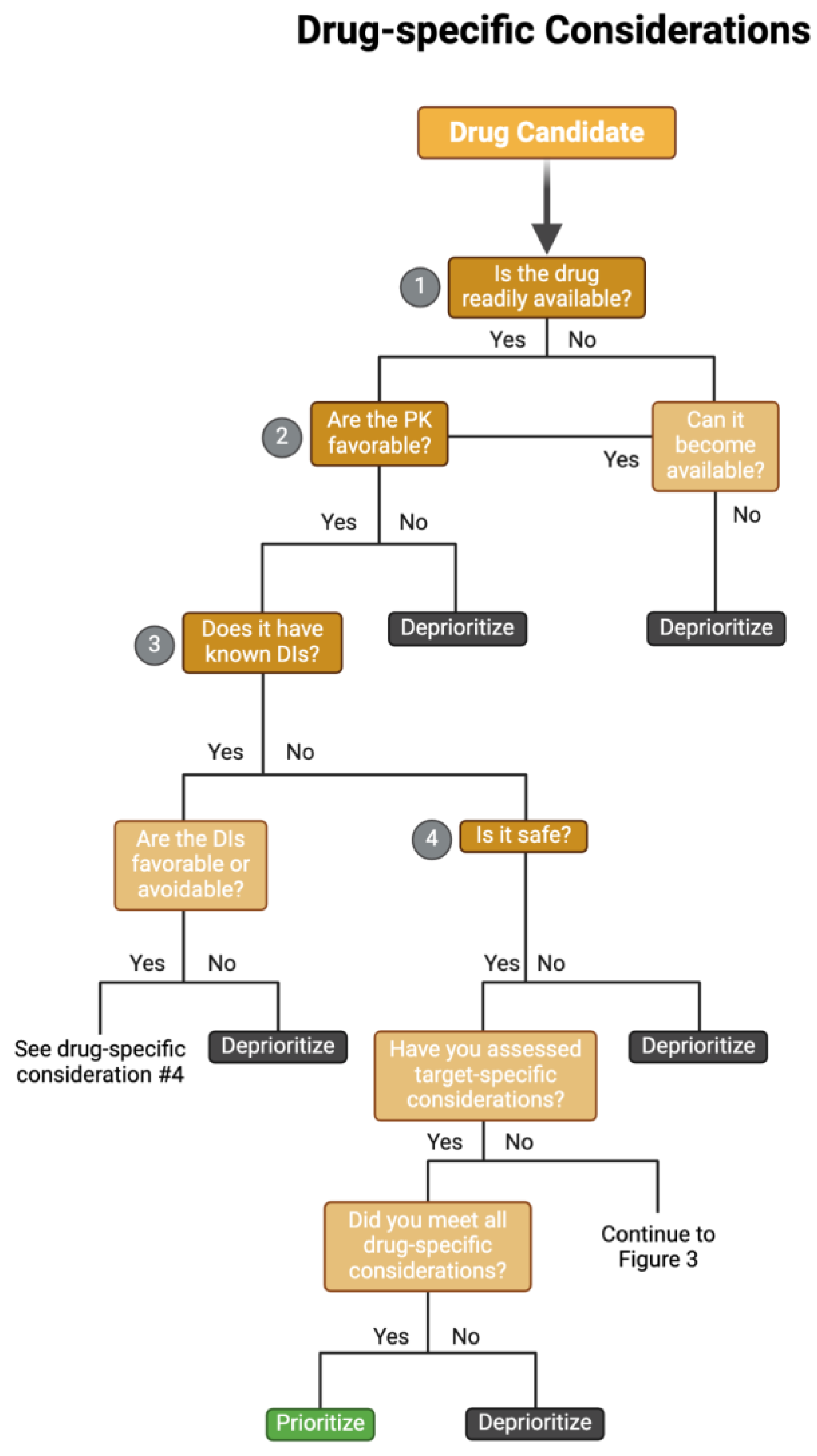
Drug-Specific Considerations
Evaluating Drug Safety
Candidate Drug Accessibility
Determining Drug Targets
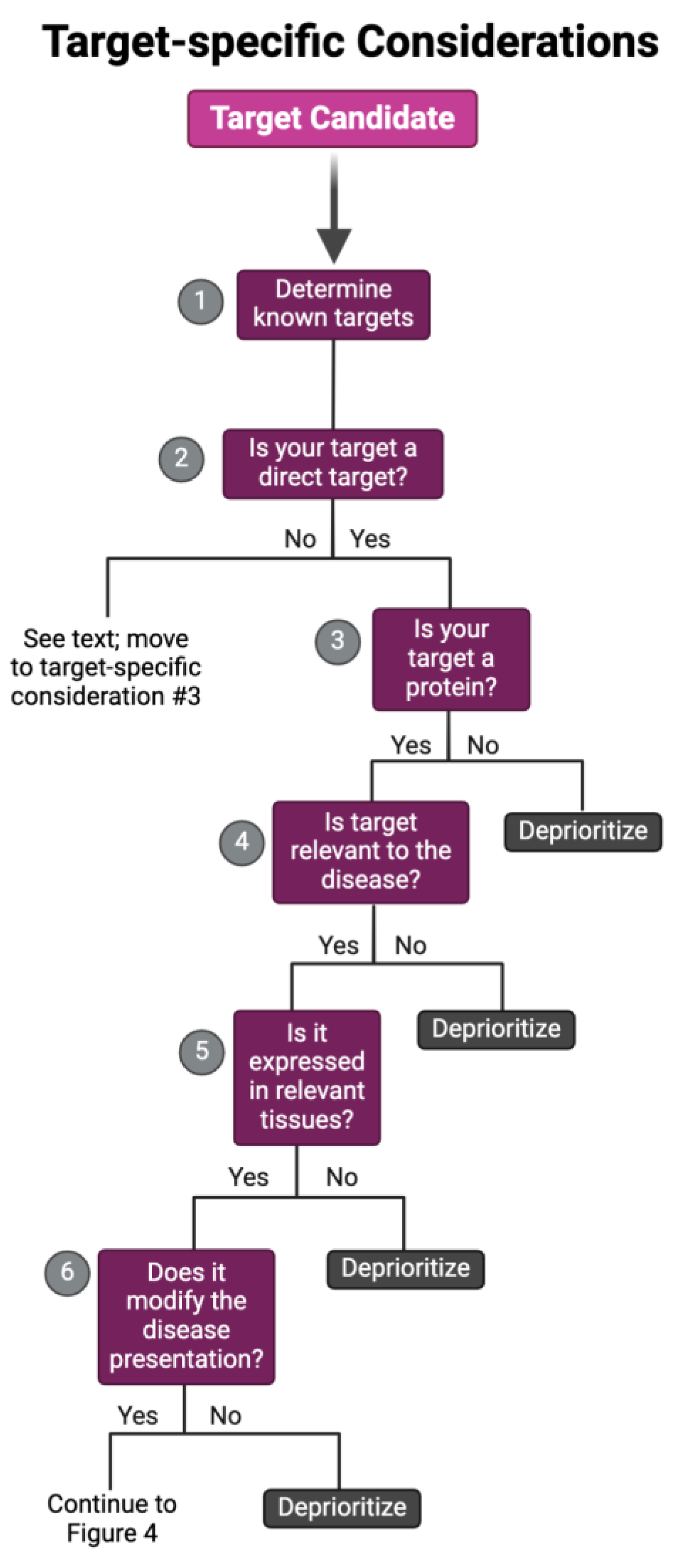
Target-Specific Considerations
Target Classification
Target Modality
Target- and Disease-Specific Considerations
Context Relevance
Disease-Modifying Effect
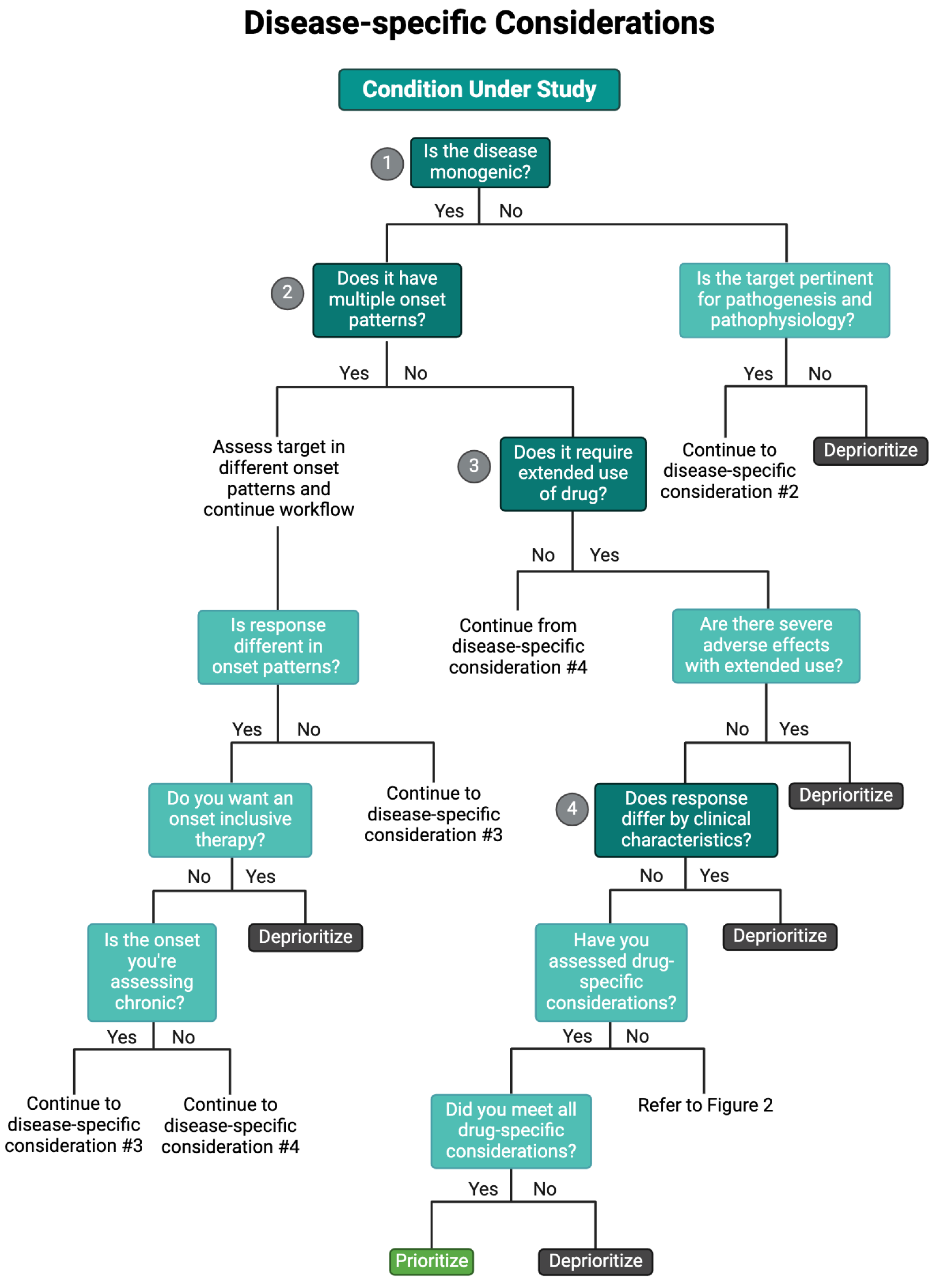
Disease-Specific Considerations
Disease Etiology
Temporal Onset
Clinical Heterogeneity
Patient Variability
Polypharmacy
Dosing
Discussion
Acknowledgments
Conflicts of Interest
References
- Aittokallio, T. What are the current challenges for machine learning in drug discovery and repurposing? Expert Opin Drug Discov 2022, 17, 423–425. [Google Scholar] [CrossRef] [PubMed]
- Alavi, S.E.; Shahmabadi, H.E. Anthelmintics for drug repurposing: Opportunities and challenges. Saudi Pharm. J. 2021, 29, 434–445. [Google Scholar] [CrossRef] [PubMed]
- Al-Bari, M.A.A. Chloroquine analogues in drug discovery: new directions of uses, mechanisms of actions and toxic manifestations from malaria to multifarious diseases. J Antimicrob Chemother 2015, 70, 1608–1621. [Google Scholar] [CrossRef] [PubMed]
- Albishri, J.; MBCh, B. NSAIDs and hypertension. 2024. Available online: https://applications.emro.who.int/imemrf/Anaesth_Pain_Intensive_Care/Anaesth_Pain_Intensive_Care_2013_17_2_171_173.pdf (accessed on 5 November 2024).
- Andreana, I.; Bincoletto, V.; Milla, P.; Dosio, F.; Stella, B.; Arpicco, S. Nanotechnological approaches for pentamidine delivery. Drug Deliv. Transl. Res. 2022, 12, 1911–1927. [Google Scholar] [CrossRef]
- Andreana, I.; Repellin, M.; Carton, F.; Kryza, D.; Briançon, S.; Chazaud, B.; Mounier, R.; Arpicco, S.; Malatesta, M.; Stella, B.; et al. Nanomedicine for Gene Delivery and Drug Repurposing in the Treatment of Muscular Dystrophies. Pharmaceutics 2021, 13, 278. [Google Scholar] [CrossRef]
- Astashkina, A.; Mann, B.; Grainger, D.W. A critical evaluation of in vitro cell culture models for high-throughput drug screening and toxicity. Pharmacol. Ther. 2012, 134, 82–106. [Google Scholar] [CrossRef]
- Ayoub, N.M. Editorial: Novel Combination Therapies for the Treatment of Solid Cancers. Front. Oncol. 2021, 11. [Google Scholar] [CrossRef]
- Baek, S.; Kwon, S.H.; Jeon, J.Y.; Lee, G.Y.; Ju, H.S.; Yun, H.J.; Cho, D.J.; Lee, K.P.; Nam, M.H. Radotinib attenuates TGFβ -mediated pulmonary fibrosis in vitro and in vivo: exploring the potential of drug repurposing. BMC Pharmacol. Toxicol. 2022, 23, 1–9. [Google Scholar] [CrossRef]
- Bai, Y.-F.; Zeng, C.; Jia, M.; Xiao, B. Molecular mechanisms of topiramate and its clinical value in epilepsy. 2022, 98, 51–56. [Google Scholar] [CrossRef]
- Bakheet, T.M.; Doig, A.J. Properties and identification of human protein drug targets. Bioinformatics 2009, 25, 451–457. [Google Scholar] [CrossRef]
- Bartolucci, D.; Pession, A.; Hrelia, P.; Tonelli, R. Precision Anti-Cancer Medicines by Oligonucleotide Therapeutics in Clinical Research Targeting Undruggable Proteins and Non-Coding RNAs. Pharmaceutics 2022, 14, 1453. [Google Scholar] [CrossRef] [PubMed]
- Begley, C.G.; Ashton, M.; Baell, J.; Bettess, M.; Brown, M.P.; Carter, B.; Charman, W.N.; Davis, C.; Fisher, S.; Frazer, I.; et al. Drug repurposing: Misconceptions, challenges, and opportunities for academic researchers. Sci. Transl. Med. 2021, 13, eabd5524. [Google Scholar] [CrossRef] [PubMed]
- Behroozi, R.; Dehghanian, E. Drug repurposing study of levofloxacin: Structural properties, lipophilicity along with experimental and computational DNA binding. J. Mol. Liq. 2024, 407. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z. The Protein Data Bank. Nucleic Acids Res 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Bhanot, S.; Sharma, A. App Review Series: Epocrates. J. Digit. Imaging 2017, 30, 534–536. [Google Scholar] [CrossRef]
- Boolell, M.; Allen, M.J.; Ballard, S.A.; Gepi-Attee, S.; Muirhead, G.J.; Naylor, A.M.; Osterloh, I.H.; Gingell, C. Sildenafil: An orally active type 5 cyclic GMP-specific phosphodiesterase inhibitor for the treatment of penile erectile dysfunction. Int. J. Impot. Res. 1996, 8, 47–52. [Google Scholar]
- Breckenridge, A.; Jacob, R. Overcoming the legal and regulatory barriers to drug repurposing. Nat. Rev. Drug Discov. 2018, 18, 1–2. [Google Scholar] [CrossRef]
- Buddolla, A.L.; Kim, S. Recent insights into the development of nucleic acid-based nanoparticles for tumor-targeted drug delivery. Colloids Surfaces B: Biointerfaces 2018, 172, 315–322. [Google Scholar] [CrossRef]
- Cabrerizo, S.; Cuadras, D.; Gomez-Busto, F.; Artaza-Artabe, I.; Marín-Ciancas, F.; Malafarina, V. Serum albumin and health in older people: Review and meta analysis. Maturitas 2015, 81, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Weinstein, J.N.; Collisson, E.A.; Mills, G.B.; Shaw, K.R.M.; Ozenberger, B.A.; Ellrott, K.; Shmulevich, I.; Sander, C.; Stuart, J.M.; The Cancer Genome Atlas Research Network. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 2013, 45, 1113–1120. [Google Scholar] [CrossRef] [PubMed]
- Castiglione, V.; Aimo, A.; Vergaro, G.; Saccaro, L.; Passino, C.; Emdin, M. Biomarkers for the diagnosis and management of heart failure. Hear. Fail. Rev. 2022, 27, 625–643. [Google Scholar] [CrossRef]
- Chandak, P.; Huang, K.; Zitnik, M. Building a knowledge graph to enable precision medicine. Sci. Data 2023, 10, 1–16. [Google Scholar] [CrossRef]
- Chan, J.; Wang, X.; A Turner, J.; E Baldwin, N.; Gu, J. Breaking the paradigm: Dr Insight empowers signature-free, enhanced drug repurposing. Bioinformatics 2019, 35, 2818–2826. [Google Scholar] [CrossRef]
- Chartier, M.; Morency, L.-P.; Zylber, M.I.; Najmanovich, R.J. Large-scale detection of drug off-targets: hypotheses for drug repurposing and understanding side-effects. BMC Pharmacol. Toxicol. 2017, 18, 1–16. [Google Scholar] [CrossRef]
- Cha Y, Erez T, Reynolds IJ, et al (2018) Drug repurposing from the perspective of pharmaceutical companies: Drug repurposing in pharmaceutical companies. Br J Pharmacol 175:168–180.
- Cheng, F.; Kovacs, I.A.; Barabási, A.-L. Network-based prediction of drug combinations. Nat. Commun. 2019, 10, 1197. [Google Scholar] [CrossRef]
- Chen, Y.-A.; Yogo, E.; Kurihara, N.; Ohno, T.; Higuchi, C.; Rokushima, M.; Mizuguchi, K. Assessing drug target suitability using TargetMine. F1000Research 2019, 8, 233. [Google Scholar] [CrossRef]
- Cheung CK, Barratt J (2016) Is IgA Nephropathy a Single Disease? In: Pathogenesis and Treatment in IgA Nephropathy. Springer Japan, Tokyo, pp 3–17.
- Chong CR, Sullivan DJ Jr (2007) New uses for old drugs. Nature 448:645–646.
- Chou, T. Mathematical definitions of “additive effect of two (or more) drugs” and their synergism and/or antagonism based on mass-action law (MAL) algorithms for pharmacodynamics (PD), biodynamics (BD) and bioinformatics (BI) simulations. FASEB J. 2022, 36. [Google Scholar] [CrossRef]
- Cichonska, A.; Rousu, J.; Aittokallio, T.; Msc; D, P. Identification of drug candidates and repurposing opportunities through compound–target interaction networks. Expert Opin. Drug Discov. 2015, 10, 1333–1345. [Google Scholar] [CrossRef]
- Cnossen, W.R.; Drenth, J.P.H. Polycystic liver disease: an overview of pathogenesis, clinical manifestations and management. Orphanet J. Rare Dis. 2014, 9, 69–69. [Google Scholar] [CrossRef]
- Correia, A.S.; Gärtner, F.; Vale, N. Drug combination and repurposing for cancer therapy: the example of breast cancer. Heliyon 2021, 7, e05948. [Google Scholar] [CrossRef] [PubMed]
- Corsello, S.; Bittker, J.A.; Liu, Z.; Gould, J.; McCarren, P.; Hirschman, J.E.; Johnston, S.E.; Vrcic, A.; Wong, B.; Khan, M.; et al. The Drug Repurposing Hub: a next-generation drug library and information resource. Nat. Med. 2017, 23, 405–408. [Google Scholar] [CrossRef]
- Corsello SM, Nagari RT, Spangler RD, et al (2020) Discovering the anti-cancer potential of non-oncology drugs by systematic viability profiling. Nat Cancer 1:235–248.
- Crockett, S.D.; Schectman, R.; Stürmer, T.; Kappelman, M.D. Topiramate Use Does Not Reduce Flares of Inflammatory Bowel Disease. Dig. Dis. Sci. 2014, 59, 1535–1543. [Google Scholar] [CrossRef]
- Crooke, S.T. A call to arms against ultra-rare diseases. Nat. Biotechnol. 2021, 39, 671–677. [Google Scholar] [CrossRef]
- Ctortecka, C.; Palve, V.; Kuenzi, B.M.; Fang, B.; Sumi, N.J.; Izumi, V.; Novakova, S.; Kinose, F.; Rix, L.L.R.; Haura, E.B.; et al. Functional Proteomics and Deep Network Interrogation Reveal a Complex Mechanism of Action of Midostaurin in Lung Cancer Cells. Mol. Cell. Proteom. 2018, 17, 2434–2447. [Google Scholar] [CrossRef]
- CZI Single-Cell Biology Program, Abdulla S, Aevermann B, et al (2023) CZ CELL×GENE Discover: A single-cell data platform for scalable exploration, analysis and modeling of aggregated data. bioRxiv 2023.10.30.563174.
- Debisschop, A.; Bogaert, B.; Muntean, C.; De Smedt, S.C.; Raemdonck, K. Beyond chloroquine: Cationic amphiphilic drugs as endosomal escape enhancers for nucleic acid therapeutics. Curr. Opin. Chem. Biol. 2024, 83, 102531. [Google Scholar] [CrossRef]
- Del Hoyo, D.; Salinas, M.; Lomas, A.; Ulzurrun, E.; Campillo, N.E.; Sorzano, C.O. Scipion-Chem: An Open Platform for Virtual Drug Screening. J. Chem. Inf. Model. 2023, 63, 7873–7885. [Google Scholar] [CrossRef]
- Doshi, S.; Chepuri, S.P. A computational approach to drug repurposing using graph neural networks. Comput. Biol. Med. 2022, 150, 105992–105992. [Google Scholar] [CrossRef]
- Drews, J. Drug Discovery: A Historical Perspective. Science 2000, 287, 1960–1964. [Google Scholar] [CrossRef]
- Duan, Q.-Q.; Wang, H.; Su, W.-M.; Gu, X.-J.; Shen, X.-F.; Jiang, Z.; Ren, Y.-L.; Cao, B.; Li, G.-B.; Wang, Y.; et al. TBK1, a prioritized drug repurposing target for amyotrophic lateral sclerosis: evidence from druggable genome Mendelian randomization and pharmacological verification in vitro. BMC Med. 2024, 22, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Evans, D.S.; A Miller, R.; Schork, N.J.; Cummings, S.R.; Girke, T. signatureSearch: environment for gene expression signature searching and functional interpretation. Nucleic Acids Res. 2020, 48, e124–e124. [Google Scholar] [CrossRef] [PubMed]
- Dudley, J.T.; Sirota, M.; Shenoy, M.; Pai, R.K.; Roedder, S.; Chiang, A.P.; Morgan, A.A.; Sarwal, M.M.; Pasricha, P.J.; Butte, A.J. Computational Repositioning of the Anticonvulsant Topiramate for Inflammatory Bowel Disease. Sci. Transl. Med. 2011, 3, 96ra76. [Google Scholar] [CrossRef] [PubMed]
- Emmerich, C.H.; Gamboa, L.M.; Hofmann, M.C.J.; Bonin-Andresen, M.; Arbach, O.; Schendel, P.; Gerlach, B.; Hempel, K.; Bespalov, A.; Dirnagl, U.; et al. Improving target assessment in biomedical research: the GOT-IT recommendations. Nat. Rev. Drug Discov. 2020, 20, 64–81. [Google Scholar] [CrossRef] [PubMed]
- Evans, E.F.; Shyr, Z.A.; Traynor, B.J.; Zheng, W. Therapeutic development approaches to treat haploinsufficiency diseases: restoring protein levels. Drug Discov. Today 2024, 104201. [Google Scholar] [CrossRef]
- Evans, J.M.M.; Donnelly, L.A.; Emslie-Smith, A.M.; Alessi, D.R.; Morris, A.D. Metformin and reduced risk of cancer in diabetic patients. Br. Med. J. 2005, 330, 1304–1305. [Google Scholar] [CrossRef]
- Feng, Y.; Wang, Q.; Wang, T. Drug Target Protein-Protein Interaction Networks: A Systematic Perspective. BioMed Res. Int. 2017, 2017, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Fisher, J.L.; Clark, A.D.; Jones, E.F.; Lasseigne, B.N. Sex-biased gene expression and gene-regulatory networks of sex-biased adverse event drug targets and drug metabolism genes. BMC Pharmacol. Toxicol. 2024, 25, 1–19. [Google Scholar] [CrossRef]
- Fisher, J.L.; Jones, E.F.; Flanary, V.L.; Williams, A.S.; Ramsey, E.J.; Lasseigne, B.N. Considerations and challenges for sex-aware drug repurposing. Biol. Sex Differ. 2022, 13, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Fisher, J.L.; Wilk, E.J.; Oza, V.H.; Gary, S.E.; Howton, T.C.; Flanary, V.L.; Clark, A.D.; Hjelmeland, A.B.; Lasseigne, B.N. Signature reversion of three disease-associated gene signatures prioritizes cancer drug repurposing candidates. FEBS Open Bio 2024, 14, 803–830. [Google Scholar] [CrossRef]
- Flanary, V.L.; Fisher, J.L.; Wilk, E.J.; Howton, T.C.; Lasseigne, B.N. Computational Advancements in Cancer Combination Therapy Prediction. JCO Precis. Oncol. 2023, 7, e2300261. [Google Scholar] [CrossRef]
- Floris, M.; Olla, S.; Schlessinger, D.; Cucca, F. Genetic-Driven Druggable Target Identification and Validation. Trends Genet. 2018, 34, 558–570. [Google Scholar] [CrossRef]
- Foretz, M.; Guigas, B.; Viollet, B. Metformin: update on mechanisms of action and repurposing potential. Nat. Rev. Endocrinol. 2023, 19, 460–476. [Google Scholar] [CrossRef] [PubMed]
- Foretz, M.; Guigas, B.; Viollet, B. Understanding the glucoregulatory mechanisms of metformin in type 2 diabetes mellitus. Nat. Rev. Endocrinol. 2019, 15, 569–589. [Google Scholar] [CrossRef] [PubMed]
- Fournier, J.-P.; Sommet, A.; Bourrel, R.; Oustric, S.; Pathak, A.; Lapeyre-Mestre, M.; Montastruc, J.-L. Non-steroidal anti-inflammatory drugs (NSAIDs) and hypertension treatment intensification: a population-based cohort study. Eur. J. Clin. Pharmacol. 2012, 68, 1533–1540. [Google Scholar] [CrossRef]
- Frankish A, Diekhans M, Ferreira A-M, et al (2019) GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res 47:D766–D773.
- Gaber, D.M.; Ibrahim, S.S.; Awaad, A.K.; Shahine, Y.M.; Elmallah, S.; Barakat, H.S.; Khamis, N.I. A drug repurposing approach of Atorvastatin calcium for its antiproliferative activity for effective treatment of breast cancer: In vitro and in vivo assessment. Int. J. Pharm. X 2024, 7, 100249. [Google Scholar] [CrossRef]
- Gaedigk, A.; Casey, S.T.; Whirl-Carrillo, M.; Miller, N.A.; Klein, T.E. Pharmacogene Variation Consortium: A Global Resource and Repository for Pharmacogene Variation. Clin. Pharmacol. Ther. 2021, 110, 542–545. [Google Scholar] [CrossRef]
- Gaulton A, Hersey A, Nowotka M, et al (2017) The ChEMBL database in 2017. Nucleic Acids Res 45:D945–D954.
- Gimeno A, Ojeda-Montes MJ, Tomás-Hernández S, et al (2019) The light and dark sides of virtual screening: What is there to know? Int J Mol Sci 20:1375.
- Gonzalez-Cavazos, A.C.; Tanska, A.; Mayers, M.; Carvalho-Silva, D.; Sridharan, B.; Rewers, P.A.; Sankarlal, U.; Jagannathan, L.; Su, A.I. DrugMechDB: A Curated Database of Drug Mechanisms. Sci. Data 2023, 10, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Gregori-Puigjané, E.; Setola, V.; Hert, J.; Crews, B.A.; Irwin, J.J.; Lounkine, E.; Marnett, L.; Roth, B.L.; Shoichet, B.K. Identifying mechanism-of-action targets for drugs and probes. Proc. Natl. Acad. Sci. 2012, 109, 11178–11183. [Google Scholar] [CrossRef]
- Gronde T van der, Uyl-de Groot CA, Pieters T (2017) Addressing the challenge of high-priced prescription drugs in the era of precision medicine: A systematic review of drug life cycles, therapeutic drug markets and regulatory frameworks. PLoS One 12:e0182613.
- Grzegorzewski, J.; Brandhorst, J.; Green, K.; Eleftheriadou, D.; Duport, Y.; Barthorscht, F.; Köller, A.; Ke, D.Y.J.; De Angelis, S.; König, M. PK-DB: pharmacokinetics database for individualized and stratified computational modeling. Nucleic Acids Res. 2020, 49, D1358–D1364. [Google Scholar] [CrossRef] [PubMed]
- GTEx Consortium (2013) The Genotype-Tissue Expression (GTEx) project. Nat Genet 45:580–585.
- Hammond SM (2015) An overview of microRNAs. Adv Drug Deliv Rev 87:3–14.
- Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100:57–7073. Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100:57–70.
- Harris MA, Clark J, Ireland A, et al (2004) The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res 32:D258–61.
- Harrison PW, Amode MR, Austine-Orimoloye O, et al (2024) Ensembl 2024. Nucleic Acids Res 52:D891–D899.
- He, B.; Xiao, Y.; Liang, H.; Huang, Q.; Du, Y.; Li, Y.; Garmire, D.; Sun, D.; Garmire, L.X. ASGARD is A Single-cell Guided Pipeline to Aid Repurposing of Drugs. Nat. Commun. 2023, 14, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Hicks, S.M.; Frias, J.A.; Mishra, S.K.; Scotti, M.; Muscato, D.R.; Valero, M.C.; Adams, L.M.; Cleary, J.D.; Nakamori, M.; Wang, E.; et al. Alternative splicing dysregulation across tissue and therapeutic approaches in a mouse model of myotonic dystrophy type 1. Mol. Ther. - Nucleic Acids 2024, 35, 102338. [Google Scholar] [CrossRef] [PubMed]
- Hogg, R.J.; Bay, R.C.; Jennette, J.C.; Sibley, R.; Kumar, S.; Fervenza, F.C.; Appel, G.; Cattran, D.; Fischer, D.; Hurley, R.M.; et al. Randomized Controlled Trial of Mycophenolate Mofetil in Children, Adolescents, and Adults With IgA Nephropathy. Am. J. Kidney Dis. 2015, 66, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Hou FF, Xie D, Wang J, et al (2023) Effectiveness of mycophenolate mofetil among patients with progressive IgA nephropathy: A randomized clinical trial. JAMA Netw Open 6:e2254054.
- Huang, K.; Chandak, P.; Wang, Q.; Havaldar, S.; Vaid, A.; Leskovec, J.; Nadkarni, G.N.; Glicksberg, B.S.; Gehlenborg, N.; Zitnik, M. A foundation model for clinician-centered drug repurposing. Nat. Med. 2024, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Hughes JP, Rees S, Kalindjian SB, Philpott KL (2011) Principles of early drug discovery: Principles of early drug discovery. Br J Pharmacol 162:1239–1249.
- Ingle, R.G.; Fang, W.-J. An Overview of the Stability and Delivery Challenges of Commercial Nucleic Acid Therapeutics. Pharmaceutics 2023, 15, 1158. [Google Scholar] [CrossRef]
- Ishiguro, C.; Fujita, T.; Omori, T.; Fujii, Y.; Mayama, T.; Sato, T. Assessing the Effects of Non-steroidal Anti-inflammatory Drugs on Antihypertensive Drug Therapy Using Post-Marketing Surveillance Database. J. Epidemiology 2008, 18, 119–124. [Google Scholar] [CrossRef]
- Issa, N.T.; Stathias, V.; Schürer, S.; Dakshanamurthy, S. Machine and deep learning approaches for cancer drug repurposing. Semin. Cancer Biol. 2020, 68, 132–142. [Google Scholar] [CrossRef]
- Jackson, A.L.; Linsley, P.S. Recognizing and avoiding siRNA off-target effects for target identification and therapeutic application. Nat. Rev. Drug Discov. 2010, 9, 57–67. [Google Scholar] [CrossRef]
- Jenquin JR, Coonrod LA, Silverglate QA, et al (2018) Furamidine rescues myotonic dystrophy type I associated mis-splicing through multiple mechanisms. ACS Chem Biol 13:2708–2718.
- Jenquin JR, Yang H, Huigens RW 3rd, et al (2019) Combination treatment of erythromycin and furamidine provides additive and synergistic rescue of mis-splicing in myotonic dystrophy type 1 models. ACS Pharmacol Transl Sci 2:247–263.
- Jéquier, E.; Constant, F. Water as an essential nutrient: the physiological basis of hydration. Eur. J. Clin. Nutr. 2010, 64, 115–123. [Google Scholar] [CrossRef]
- Jia, J.; Zhu, F.; Ma, X.; Cao, Z.W.; Li, Y.X.; Chen, Y.Z. Mechanisms of drug combinations: interaction and network perspectives. Nat. Rev. Drug Discov. 2009, 8, 111–128. [Google Scholar] [CrossRef]
- Jiang, W.; Hu, J.-W.; He, X.-R.; Jin, W.-L.; He, X.-Y. Statins: a repurposed drug to fight cancer. J. Exp. Clin. Cancer Res. 2021, 40, 241. [Google Scholar] [CrossRef] [PubMed]
- Jia, Z.; Liu, Y.; Guan, N.; Bo, X.; Luo, Z.; Barnes, M.R. Cogena, a novel tool for co-expressed gene-set enrichment analysis, applied to drug repositioning and drug mode of action discovery. BMC Genom. 2016, 17, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Jin, G.; Wong, S.T. Toward better drug repositioning: prioritizing and integrating existing methods into efficient pipelines. Drug Discov. Today 2014, 19, 637–644. [Google Scholar] [CrossRef] [PubMed]
- Jones EF, Howton TC, Soelter TM, et al (2024) Cell-type-specific alternative splicing in the cerebral cortex of a Schinzel-Giedion Syndrome patient variant mouse model. Genomics.
- Jonker, A.H.; O’connor, D.; Cavaller-Bellaubi, M.; Fetro, C.; Gogou, M.; Hoen, P.A.C. .; de Kort, M.; Stone, H.; Valentine, N.; Pasmooij, A.M.G. Drug repurposing for rare: progress and opportunities for the rare disease community. Front. Med. 2024, 11, 1352803. [Google Scholar] [CrossRef] [PubMed]
- Kandasamy, T.; Sarkar, S.; Ghosh, S.S. Harnessing Drug Repurposing to Combat Breast Cancer by Targeting Altered Metabolism and Epithelial-to-Mesenchymal Transition Pathways. ACS Pharmacol. Transl. Sci. 2024. [Google Scholar] [CrossRef]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alfoldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 2020, 581, 434–443. [Google Scholar] [CrossRef]
- Keijsers CJPW, Segers WS, de Wildt DJ, et al (2015) Implementation of the WHO-6-step method in the medical curriculum to improve pharmacology knowledge and pharmacotherapy skills: Effect of WHO-6-step implemented in a medical curriculum. Br J Clin Pharmacol 79:896–906.
- Kopanos, C.; Tsiolkas, V.; Kouris, A.; Chapple, C.E.; Aguilera, M.A.; Meyer, R.; Massouras, A. VarSome: The human genomic variant search engine. Bioinformatics 2019, 35, 1978–1980. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Krishnamurthy, N.; Grimshaw, A.A.; Axson, S.A.; Choe, S.H.; Miller, J.E. Drug repurposing: a systematic review on root causes, barriers and facilitators. BMC Heal. Serv. Res. 2022, 22, 1–17. [Google Scholar] [CrossRef]
- Kuan, V.; Denaxas, S.; Patalay, P.; Nitsch, D.; Mathur, R.; Gonzalez-Izquierdo, A.; Sofat, R.; Partridge, L.; Roberts, A.; Wong, I.C.K.; et al. Identifying and visualising multimorbidity and comorbidity patterns in patients in the English National Health Service: a population-based study. Lancet Digit Health 2023, 5, e16–e27. [Google Scholar] [CrossRef]
- Kuenzi, B.M.; Rix, L.L.R.; Kinose, F.; Kroeger, J.L.; Lancet, J.E.; Padron, E.; Rix, U. Off-target based drug repurposing opportunities for tivantinib in acute myeloid leukemia. Sci. Rep. 2019, 9, 1–12. [Google Scholar] [CrossRef]
- Kulkarni, J.A.; Witzigmann, D.; Thomson, S.B.; Chen, S.; Leavitt, B.R.; Cullis, P.R.; van der Meel, R. The current landscape of nucleic acid therapeutics. Nat. Nanotechnol. 2021, 16, 630–643. [Google Scholar] [CrossRef]
- Kwon, O.-S.; Kim, W.; Cha, H.-J.; Lee, H. In silico drug repositioning: from large-scale transcriptome data to therapeutics. Arch. Pharmacal Res. 2019, 42, 879–889. [Google Scholar] [CrossRef] [PubMed]
- Lago, S.G.; Bahn, S. The druggable schizophrenia genome: from repurposing opportunities to unexplored drug targets. npj Genom. Med. 2022, 7, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Lamb, J.; Crawford, E.D.; Peck, D.; Modell, J.W.; Blat, I.C.; Wrobel, M.J.; Lerner, J.; Brunet, J.-P.; Subramanian, A.; Ross, K.N.; et al. The Connectivity Map: Using Gene-Expression Signatures to Connect Small Molecules, Genes, and Disease. Science 2006, 313, 1929–1935. [Google Scholar] [CrossRef] [PubMed]
- Landrum, M.J.; Lee, J.M.; Riley, G.R.; Jang, W.; Rubinstein, W.S.; Church, D.M.; Maglott, D.R. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014, 42, D980–D985. [Google Scholar] [CrossRef]
- Lauffer, M.C.; van Roon-Mom, W.; Aartsma-Rus, A. Possibilities and limitations of antisense oligonucleotide therapies for the treatment of monogenic disorders. Commun. Med. 2024, 4, 1–11. [Google Scholar] [CrossRef]
- Surdo, P.L.; Iannuccelli, M.; Contino, S.; Castagnoli, L.; Licata, L.; Cesareni, G.; Perfetto, L. SIGNOR 3.0, the SIGnaling network open resource 3.0: 2022 update. Nucleic Acids Res. 2022, 51, D631–D637. [Google Scholar] [CrossRef]
- Maes, B.D.; Oyen, R.; Claes, K.; Evenepoel, P.; Kuypers, D.; Vanwalleghem, J.; Van Damme, B.; Vanrenterghem, Y.F.C. Mycophenolate mofetil in IgA nephropathy: Results of a 3-year prospective placebo-controlled randomized study. Kidney Int. 2004, 65, 1842–1849. [Google Scholar] [CrossRef]
- Ma, J.; Wang, J.; Ghoraie, L.S.; Men, X.; Liu, L.; Dai, P. Network-based method for drug target discovery at the isoform level. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Manirajan, P.; Sivanandy, P. A Pharmacist Perspective on Optimizing Pharmacotherapy among Older Adults – A Review. F1000Research 2024, 13, 748. [Google Scholar] [CrossRef]
- Masnoon, N.; Shakib, S.; Kalisch-Ellett, L.; Caughey, G.E. What is polypharmacy? A systematic review of definitions. BMC Geriatr. 2017, 17, 230. [Google Scholar] [CrossRef]
- McDowall, S.; Aung-Htut, M.; Wilton, S.; Li, D.; McDowall, \. Antisense oligonucleotides and their applications in rare neurological diseases. Front. Neurosci. 2024, 18, 1414658. [Google Scholar] [CrossRef]
- McKusick, V.A. Mendelian Inheritance in Man and Its Online Version, OMIM. Am. J. Hum. Genet. 2007, 80, 588–604. [Google Scholar] [CrossRef] [PubMed]
- Meier, S.M.; Deckert, J. Genetics of Anxiety Disorders. Curr. Psychiatry Rep. 2019, 21, 16. [Google Scholar] [CrossRef] [PubMed]
- Meng, Z.; Lu, M. RNA Interference-Induced Innate Immunity, Off-Target Effect, or Immune Adjuvant? Front. Immunol. 2017, 8, 331. [Google Scholar] [CrossRef]
- Messa, L.; Testa, C.; Carelli, S.; Rey, F.; Jacchetti, E.; Cereda, C.; Raimondi, M.T.; Ceri, S.; Pinoli, P. Non-Negative Matrix Tri-Factorization for Representation Learning in Multi-Omics Datasets with Applications to Drug Repurposing and Selection. Int. J. Mol. Sci. 2024, 25, 9576. [Google Scholar] [CrossRef]
- Miller, J.A.; Ding, S.-L.; Sunkin, S.M.; Smith, K.A.; Ng, L.; Szafer, A.; Ebbert, A.; Riley, Z.L.; Royall, J.J.; Aiona, K.; et al. Transcriptional landscape of the prenatal human brain. Nature 2014, 508, 199–206. [Google Scholar] [CrossRef]
- Miranda, A.M.A.; Janbandhu, V.; Maatz, H.; Kanemaru, K.; Cranley, J.; Teichmann, S.A.; Hübner, N.; Schneider, M.D.; Harvey, R.P.; Noseda, M. Single-cell transcriptomics for the assessment of cardiac disease. Nat. Rev. Cardiol. 2022, 20, 289–308. [Google Scholar] [CrossRef]
- Morris, R.; Ali, R.; Cheng, F. Drug Repurposing Using FDA Adverse Event Reporting System (FAERS) Database. Curr. Drug Targets 2024, 25, 454–464. [Google Scholar] [CrossRef]
- Ng, M.S.Y.; Kaur, G.; Francis, R.S.; Hawley, C.M.; Johnson, D.W. Drug repurposing for glomerular diseases: an underutilized resource. Nat. Rev. Nephrol. 2024, 20, 707–721. [Google Scholar] [CrossRef]
- Nosengo N (2016) Can you teach old drugs new tricks? Nature 534:314–316.
- Novak, J.; Goldberg, A.; Dharmarajan, K.; Amini, A.; Maggiore, R.J.; Presley, C.J.; Nightingale, G. Polypharmacy in older adults with cancer undergoing radiotherapy: A review. J. Geriatr. Oncol. 2022, 13, 778–783. [Google Scholar] [CrossRef]
- Oliva, M.; Muñoz-Aguirre, M.; Kim-Hellmuth, S.; Wucher, V.; Gewirtz, A.D.H.; Cotter, D.J.; Parsana, P.; Kasela, S.; Balliu, B.; Viñuela, A.; et al. The impact of sex on gene expression across human tissues. Science 2020, 369, eaba3066. [Google Scholar] [CrossRef] [PubMed]
- Oprea, T.I.; Bauman, J.E.; Bologa, C.G.; Buranda, T.; Chigaev, A.; Edwards, B.S.; Jarvik, J.W.; Gresham, H.D.; Haynes, M.K.; Hjelle, B.; et al. Drug repurposing from an academic perspective. Drug Discov. Today: Ther. Strat. 2012, 8, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Oprea, T.I.; Mestres, J. Drug Repurposing: Far Beyond New Targets for Old Drugs. AAPS J. 2012, 14, 759–763. [Google Scholar] [CrossRef]
- Palve, V.; Liao, Y.; Rix, L.L.R.; Rix, U. Turning liabilities into opportunities: Off-target based drug repurposing in cancer. Semin. Cancer Biol. 2020, 68, 209–229. [Google Scholar] [CrossRef]
- Pantziarka, P.; Verbaanderd, C.; Huys, I.; Bouche, G.; Meheus, L. Repurposing drugs in oncology: From candidate selection to clinical adoption. Semin. Cancer Biol. 2021, 68, 186–191. [Google Scholar] [CrossRef]
- Pantziarka, P.; Verbaanderd, C.; Capistrano, R.; Crispino, S.; Gyawali, B.; Rooman, I.; Van Nuffel, A.M.; Meheus, L.; Sukhatme, V.P.; Bouche, G. ReDO_DB: the repurposing drugs in oncology database. ecancermedicalscience 2018, 12, 886. [Google Scholar] [CrossRef]
- Park, K. A review of computational drug repurposing. Transl. Clin. Pharmacol. 2019, 27, 59–63. [Google Scholar] [CrossRef]
- Parvathaneni V, Kulkarni NS, Muth A, Gupta V (2019) Drug repurposing: a promising tool to accelerate the drug discovery process. Drug Discov Today 24:2076–2085.
- Pascual-Gilabert, M.; Artero, R.; López-Castel, A. The myotonic dystrophy type 1 drug development pipeline: 2022 edition. Drug Discov. Today 2023, 28, 103489. [Google Scholar] [CrossRef] [PubMed]
- Paysan-Lafosse T, Blum M, Chuguransky S, et al (2023) InterPro in 2022. Nucleic Acids Res 51:D418–D427.
- Perez, D.R.; Sklar, L.A.; Chigaev, A.; Matlawska-Wasowska, K. Drug repurposing for targeting cyclic nucleotide transporters in acute leukemias - A missed opportunity. Semin. Cancer Biol. 2020, 68, 199–208. [Google Scholar] [CrossRef]
- Pinzi, L.; Bisi, N.; Rastelli, G. How drug repurposing can advance drug discovery: challenges and opportunities. Front. Drug Discov. 2024, 4, 1460100. [Google Scholar] [CrossRef]
- Polamreddy P, Gattu N (2019) The drug repurposing landscape from 2012 to 2017: evolution, challenges, and possible solutions. Drug Discov Today 24:789–795. [CrossRef]
- Pollard, J.A.; Furutani, E.; Liu, S.; Esrick, E.B.; Cohen, L.E.; Bledsoe, J.R.; Liu, C.-W.; Lu, K.; de Haro, M.J.R.; Surrallés, J.; et al. Metformin for treatment of cytopenias in children and young adults with Fanconi anemia. Blood Adv. 2022, 6, 3803–3811. [Google Scholar] [CrossRef]
- Preissner, S.C.; Hoffmann, M.F.; Preissner, R.; Dunkel, M.; Gewiess, A. Polymorphic Cytochrome P450 Enzymes (CYPs) and Their Role in Personalized Therapy. PLOS ONE 2013, 8, e82562. [Google Scholar] [CrossRef] [PubMed]
- Pushpakom S, Iorio F, Eyers PA, et al (2019) Drug repurposing: progress, challenges and recommendations. Nat Rev Drug Discov 18:41–58.
- Qiu, S.; Cai, Y.; Yao, H.; Lin, C.; Xie, Y.; Tang, S.; Zhang, A. Small molecule metabolites: discovery of biomarkers and therapeutic targets. Signal Transduct. Target. Ther. 2023, 8, 132. [Google Scholar] [CrossRef]
- Qi, X.; Zhao, L.; Tian, C.; Li, Y.; Chen, Z.-L.; Huo, P.; Chen, R.; Liu, X.; Wan, B.; Yang, S.; et al. Predicting transcriptional responses to novel chemical perturbations using deep generative model for drug discovery. Nat. Commun. 2024, 15, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Rafikova, E.; Nemirovich-Danchenko, N.; Ogmen, A.; Parfenenkova, A.; Velikanova, A.; Tikhonov, S.; Peshkin, L.; Rafikov, K.; Spiridonova, O.; Belova, Y.; et al. Open Genes—a new comprehensive database of human genes associated with aging and longevity. Nucleic Acids Res. 2023, 52, D950–D962. [Google Scholar] [CrossRef]
- Rodchenkov, I.; Babur, O.; Luna, A.; Aksoy, B.A.; Wong, J.V.; Fong, D.; Franz, M.; Siper, M.C.; Cheung, M.; Wrana, M.; et al. Pathway Commons 2019 Update: integration, analysis and exploration of pathway data. Nucleic Acids Res. 2020, 48, D489–D497. [Google Scholar] [CrossRef]
- Roessler HI, Knoers NVAM, van Haelst MM, van Haaften G (2021) Drug repurposing for rare diseases. Trends Pharmacol Sci 42:255–267144. Roessler HI, Knoers NVAM, van Haelst MM, van Haaften G (2021) Drug repurposing for rare diseases. Trends Pharmacol Sci 42:255–267.
- Saiyed, M.M.; Ong, P.S.; Chew, L. Off-label drug use in oncology: a systematic review of literature. J. Clin. Pharm. Ther. 2017, 42, 251–258. [Google Scholar] [CrossRef] [PubMed]
- Sangkuhl, K.; Whirl-Carrillo, M.; Whaley, R.M.; Woon, M.; Lavertu, A.; Altman, R.B.; Carter, L.; Verma, A.; Ritchie, M.D.; Klein, T.E. Pharmacogenomics Clinical Annotation Tool (PharmCAT). Clin. Pharmacol. Ther. 2019, 107, 203–210. [Google Scholar] [CrossRef]
- Scannell, J.W.; Blanckley, A.; Boldon, H.; Warrington, B. Diagnosing the decline in pharmaceutical R&D efficiency. Nat. Rev. Drug Discov. 2012, 11, 191–200. [Google Scholar] [CrossRef]
- Schenone, M.; Dančík, V.; Wagner, B.K.; Clemons, P.A. Target identification and mechanism of action in chemical biology and drug discovery. Nat. Chem. Biol. 2013, 9, 232–240. [Google Scholar] [CrossRef]
- Setoain, J.; Franch, M.; Martínez, M.; Tabas-Madrid, D.; Sorzano, C.O.S.; Bakker, A.; Gonzalez-Couto, E.; Elvira, J.; Pascual-Montano, A. NFFinder: an online bioinformatics tool for searching similar transcriptomics experiments in the context of drug repositioning. Nucleic Acids Res. 2015, 43, W193–W199. [Google Scholar] [CrossRef] [PubMed]
- Shao, R.; Li, Z.; Xin, H.; Jiang, S.; Zhu, Y.; Liu, J.; Huang, R.; Xu, K.; Shi, X. Biomarkers as targets for CAR-T/NK cell therapy in AML. Biomark. Res. 2023, 11, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Sheils, T.K.; Mathias, S.L.; Kelleher, K.J.; Siramshetty, V.B.; Nguyen, D.-T.; Bologa, C.G.; Jensen, L.J.; Vidović, D.; Koleti, A.; Schürer, S.C.; et al. TCRD and Pharos 2021: mining the human proteome for disease biology. Nucleic Acids Res. 2020, 49, D1334–D1346. [Google Scholar] [CrossRef] [PubMed]
- Shin, C.; Kim, Y.-K. Ketamine in Major Depressive Disorder: Mechanisms and Future Perspectives. Psychiatry Investig. 2020, 17, 181–192. [Google Scholar] [CrossRef] [PubMed]
- Silva, I.; Mendes, P.; Guerra, S.; Pinto, R.; Mateus, V. Anti-Inflammatory Effect of Topiramate in a Chronic Model of TNBS-Induced Colitis. Int. J. Mol. Sci. 2022, 23, 9127. [Google Scholar] [CrossRef]
- Sirtori, C.R. The pharmacology of statins. Pharmacol. Res. 2014, 88, 3–11. [Google Scholar] [CrossRef]
- Sleire, L.; Førde, H.E.; Netland, I.A.; Leiss, L.; Skeie, B.S.; Enger, P.O. Drug repurposing in cancer. Pharmacol. Res. 2017, 124, 74–91. [Google Scholar] [CrossRef]
- Sollis, E.; Mosaku, A.; Abid, A.; Buniello, A.; Cerezo, M.; Gil, L.; Groza, T.; Güneş, O.; Hall, P.; Hayhurst, J.; et al. The NHGRI-EBI GWAS Catalog: knowledgebase and deposition resource. Nucleic Acids Res. 2022, 51, D977–D985. [Google Scholar] [CrossRef]
- Sun, W.; Sanderson, P.E.; Zheng, W. Drug combination therapy increases successful drug repositioning. Drug Discov. Today 2016, 21, 1189–1195. [Google Scholar] [CrossRef]
- Takimoto, Chris H.. 2001. Basic Pharmacokinetics and Pharmacodynamic Principles. 106: 85–101. [CrossRef]
- Talevi, A.; Bellera, C.L. Challenges and opportunities with drug repurposing: finding strategies to find alternative uses of therapeutics. Expert Opin. Drug Discov. 2019, 15, 397–401. [Google Scholar] [CrossRef]
- Tanaka Y, Chen HY, Belloni P, et al. 2024. OnSIDES (ON-label SIDE effectS resource) database : Extracting adverse drug events from drug labels using Natural Language Processing models. medRxiv 2024.03.22.24304724.
- Tan, G.S.; Sloan, E.K.; Lambert, P.; Kirkpatrick, C.M.; Ilomäki, J. Drug repurposing using real-world data. Drug Discov. Today 2022, 28, 10–13. [Google Scholar] [CrossRef] [PubMed]
- Tan, X.; Jia, F.; Wang, P.; Zhang, K. Nucleic acid-based drug delivery strategies. J. Control. Release 2020, 323, 240–252. [Google Scholar] [CrossRef] [PubMed]
- The White House. 2023. FACT SHEET: President Biden’s Cap on the Cost of Insulin Could Benefit Millions of Americans in All 50 States. In: The White House. https://www.whitehouse.gov/briefing-room/statements-releases/2023/03/02/fact-sheet-president-bidens-cap-on-the-cost-of-insulin-could-benefit-millions-of-americans-in-all-50-states/. 2024.
- Thorman, A.W.; Reigle, J.; Chutipongtanate, S.; Yang, J.; Shamsaei, B.; Pilarczyk, M.; Fazel-Najafabadi, M.; Adamczak, R.; Kouril, M.; Bhatnagar, S.; et al. Accelerating drug discovery and repurposing by combining transcriptional signature connectivity with docking. Sci. Adv. 2024, 10, eadj3010. [Google Scholar] [CrossRef] [PubMed]
- Tseng, C.-W.; Masuda, C.; Chen, R.; Hartung, D.M. Impact of Higher Insulin Prices on Out-of-Pocket Costs in Medicare Part D. Diabetes Care 2020, 43, E50–E51. [Google Scholar] [CrossRef] [PubMed]
- Uhlén, M.; Fagerberg, L.; Hallström, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, Å.; Kampf, C.; Sjöstedt, E.; Asplund, A.; et al. Proteomics. Tissue-Based Map of the Human Proteome. Science 2015, 347, 1260419. [Google Scholar] [CrossRef]
- Unger, J.M.; Vaidya, R.; Albain, K.S.; LeBlanc, M.; Minasian, L.M.; Gotay, C.C.; Henry, N.L.; Fisch, M.J.; Lee, S.M.; Blanke, C.D.; et al. Sex Differences in Risk of Severe Adverse Events in Patients Receiving Immunotherapy, Targeted Therapy, or Chemotherapy in Cancer Clinical Trials. J. Clin. Oncol. 2022, 40, 1474–1486. [Google Scholar] [CrossRef]
- UniProt Consortium (2023) UniProt: The universal protein knowledgebase in 2023. Nucleic Acids Res 51:D523–D531. [CrossRef]
- van Dam P-J, Daelemans S, Ross E, et al (2018) Histopathological growth patterns as a candidate biomarker for immunomodulatory therapy. Semin Cancer Biol 52:86–93.
- Varey, A.H.R.; Rennel, E.S.; Qiu, Y.; Bevan, H.S.; Perrin, R.M.; Raffy, S.; Dixon, A.R.; Paraskeva, C.; Zaccheo, O.; Hassan, A.B.; et al. VEGF165b, an antiangiogenic VEGF-A isoform, binds and inhibits bevacizumab treatment in experimental colorectal carcinoma: balance of pro- and antiangiogenic VEGF-A isoforms has implications for therapy. Br. J. Cancer 2008, 98, 1366–1379. [Google Scholar] [CrossRef]
- Volpi E, Nazemi R, Fujita S (2004) Muscle tissue changes with aging. Curr Opin Clin Nutr Metab Care 7:405–410.
- Wang, W.; He, S.; Dong, G.; Sheng, C. Nucleic-Acid-Based Targeted Degradation in Drug Discovery. J. Med. Chem. 2022, 65, 10217–10232. [Google Scholar] [CrossRef]
- Wang, Z.; Clark, N.R.; Ma’ayan, A. Drug-induced adverse events prediction with the LINCS L1000 data. Bioinformatics 2016, 32, 2338–2345. [Google Scholar] [CrossRef]
- Waring, M.J.; Arrowsmith, J.; Leach, A.R.; Leeson, P.D.; Mandrell, S.; Owen, R.M.; Pairaudeau, G.; Pennie, W.D.; Pickett, S.D.; Wang, J.; et al. An analysis of the attrition of drug candidates from four major pharmaceutical companies. Nat. Rev. Drug Discov. 2015, 14, 475–486. [Google Scholar] [CrossRef]
- Watson, S.; Caster, O.; A Rochon, P.; Ruijter, H.D. Reported adverse drug reactions in women and men: Aggregated evidence from globally collected individual case reports during half a century. eClinicalMedicine 2019, 17, 100188. [Google Scholar] [CrossRef]
- Weth, F.R.; Hoggarth, G.B.; Weth, A.F.; Paterson, E.; White, M.P.J.; Tan, S.T.; Peng, L.; Gray, C. Unlocking hidden potential: advancements, approaches, and obstacles in repurposing drugs for cancer therapy. Br. J. Cancer 2023, 130, 703–715. [Google Scholar] [CrossRef] [PubMed]
- Whirl-Carrillo, M.; Huddart, R.; Gong, L.; Sangkuhl, K.; Thorn, C.F.; Whaley, R.; Klein, T.E. An Evidence-Based Framework for Evaluating Pharmacogenomics Knowledge for Personalized Medicine. Clin. Pharmacol. Ther. 2021, 110, 563–572. [Google Scholar] [CrossRef] [PubMed]
- Whitlock JH, Soelter TM, Howton TC, et al (2023) Cell-type-specific gene expression and regulation in the cerebral cortex and kidney of atypical Setbp1S858R Schinzel Giedion Syndrome mice. J Cell Mol Med 27:3565–3577.
- Wilk, E.J.; Howton, T.C.; Fisher, J.L.; Oza, V.H.; Brownlee, R.T.; McPherson, K.C.; Cleary, H.L.; Yoder, B.K.; George, J.F.; Mrug, M.; et al. Prioritized polycystic kidney disease drug targets and repurposing candidates from pre-cystic and cystic mouse Pkd2 model gene expression reversion. Mol. Med. 2023, 29, 1–21. [Google Scholar] [CrossRef]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; et al. DrugBank 5.0: A Major Update to the DrugBank Database for 2018. Nucleic Acids Res. 2018, 46, D1074–D1082. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.; Feng, Q.; Kerchberger, V.E.; Nelson, S.D.; Chen, Q.; Li, B.; Edwards, T.L.; Cox, N.J.; Phillips, E.J.; Stein, C.M.; et al. Integrating gene expression and clinical data to identify drug repurposing candidates for hyperlipidemia and hypertension. Nat. Commun. 2022, 13, 1–12. [Google Scholar] [CrossRef]
- Xia, Y.; Sun, M.; Huang, H.; Jin, W.-L. Drug repurposing for cancer therapy. Signal Transduct. Target. Ther. 2024, 9, 1–33. [Google Scholar] [CrossRef]
- Xu B, Forthman KL, Kuplicki R, et al (2024) Genetic correlates of treatment-resistant depression: Insights from polygenic scores across cognitive, temperamental, and sleep traits in the all of US cohort. medRxiv 2024.07.03.24309914.
- Yan, C.; Grabowska, M.E.; Dickson, A.L.; Li, B.; Wen, Z.; Roden, D.M.; Stein, C.M.; Embí, P.J.; Peterson, J.F.; Feng, Q.; et al. Leveraging generative AI to prioritize drug repurposing candidates for Alzheimer’s disease with real-world clinical validation. npj Digit. Med. 2024, 7, 1–6. [Google Scholar] [CrossRef]
- Yao, Z.; van Velthoven, C.T.J.; Kunst, M.; Zhang, M.; McMillen, D.; Lee, C.; Jung, W.; Goldy, J.; Abdelhak, A.; Aitken, M.; et al. A high-resolution transcriptomic and spatial atlas of cell types in the whole mouse brain. Nature 2023, 624, 317–332. [Google Scholar] [CrossRef]
- Yu, A.-M.; Choi, Y.H.; Tu, M.-J. RNA Drugs and RNA Targets for Small Molecules: Principles, Progress, and Challenges. Pharmacol. Rev. 2020, 72, 862–898. [Google Scholar] [CrossRef]
- Yuhan J, Zhu L, Zhu L, et al (2022) Cell-specific aptamers as potential drugs in therapeutic applications: A review of current progress. J Control Release 346:405–420.
- Yu, H.-H.; Chiang, B.-L. Diagnosis and classification of IgA nephropathy. Autoimmun. Rev. 2014, 13, 556–559. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Chen, J.; Li, D.; Wang, L.; Wang, W.; Liu, H. Systematic Analysis of Adverse Event Reports for Sex Differences in Adverse Drug Events. Sci. Rep. 2016, 6, 24955–24955. [Google Scholar] [CrossRef] [PubMed]
- Zahoránszky-Kőhalmi, G.; Sheils, T.; Oprea, T.I. SmartGraph: a network pharmacology investigation platform. J. Chemin- 2020, 12, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Wang, F.; Hu, J.; Sorrentino, R. Exploring the relationship between drug side-effects and therapeutic indications. . 2013, 2013, 1568–77. [Google Scholar] [PubMed]
- Zhang, S.-D.; Gant, T.W. sscMap: An extensible Java application for connecting small-molecule drugs using gene-expression signatures. BMC Bioinform. 2009, 10, 236–236. [Google Scholar] [CrossRef]
- Zhao Z, Ukidve A, Kim J, Mitragotri S (2020) Targeting strategies for tissue-specific drug delivery. Cell 181:151–167.
- Zhou, S.-S.; Jin, J.-P.; Wang, J.-Q.; Zhang, Z.-G.; Freedman, J.H.; Zheng, Y.; Cai, L. miRNAS in cardiovascular diseases: potential biomarkers, therapeutic targets and challenges. Acta Pharmacol. Sin. 2018, 39, 1073–1084. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhang, Y.; Lian, X.; Li, F.; Wang, C.; Zhu, F.; Qiu, Y.; Chen, Y. Therapeutic target database update 2022: facilitating drug discovery with enriched comparative data of targeted agents. Nucleic Acids Res. 2021, 50, D1398–D1407. [Google Scholar] [CrossRef]
| Specificity Type | Name of consideration | Resources |
|---|---|---|
| Drug |
Accessibility | ClinCalc (https://clincalc.com/) Epocrates (https://www.epocrates.com/)* Medi-Span (https://www.wolterskluwer.com/en/solutions/medi-span)* ReDO (Pantziarka et al. 2018) |
| Drug |
Pharmacokinetics |
Medi-Span (https://www.wolterskluwer.com/en/solutions/medi-span)* PHAROS (Sheils et al. 2021) DrugBank (Wishart et al. 2018) ChEMBL (Gaulton et al. 2017)PharmGKB (Whirl-Carrillo et al. 2021) PK-DB (Grzegorzewski et al. 2021) |
| Drug | Pharmacodynamics | Medi-Span (https://www.wolterskluwer.com/en/solutions/medi-span)* PHAROS (Sheils et al. 2021) DrugBank (Wishart et al. 2018) ChEMBL (Gaulton et al. 2017) DrugMechDB (Gonzalez-Cavazos et al. 2023) |
| Drug; disease | Drug interactions; patient variability - polypharmacy |
Medi-Span (https://www.wolterskluwer.com/en/solutions/medi-span)* Epocrates (https://www.epocrates.com/)* UpToDate (https://www.uptodate.com/contents/search)* DrugBank (Wishart et al. 2018) |
| Drug |
Adverse effects | ClinicalTrials.gov (https://www.clinicaltrials.gov/) Medi-Span (https://www.wolterskluwer.com/en/solutions/medi-span)* FAERS (https://www.fda.gov/drugs/fdas-adverse-event-reporting-system-faers/fda-adverse-event-reporting-system-faers-public-dashboard) DrugBank (Wishart et al. 2018) |
| Drug |
Number of targets | DrugBank (Wishart et al. 2018) PHAROS (Sheils et al. 2021) SmartGraph (Zahoránszky-Kőhalmi et al. 2020) Therapeutic Target Database (Zhou et al. 2022) Drug Repurposing Hub (Corsello et al. 2017) |
| Drug; target | Ligand activity | PHAROS (Sheils et al. 2021) |
| Target |
Target classification |
PHAROS (Sheils et al. 2021) Therapeutic Target Database (Zhou et al. 2022) |
| Target; disease |
Context relevance - protein family | Uniprot (UniProt Consortium 2023) Interpro (Paysan-Lafosse et al. 2023) The Protein Data Bank (Berman et al. 2000) |
| Target; disease |
Context relevance - biological pathways | Pathway Commons (Rodchenkov et al. 2020) Gene Ontologies (Harris et al. 2004) |
| Target; disease |
Context relevance - genome-wide associations | GTEx (GTEx Consortium 2013) GWAS Catalog (Sollis et al. 2023) |
| Target; disease | Context relevance - other | MedlinePlus (https://medlineplus.gov) Gene Ontologies (Harris et al. 2004) ReDO (Pantziarka et al. 2018) |
| Target; disease |
Context relevance and disease modifiable effect - gene expression | GTEx (GTEx Consortium 2013) Human Protein Atlas (Uhlén et al. 2015) Allen Brain Map (Yao et al. 2023) TCGA (Cancer Genome Atlas Research Network et al. 2013) CellxGene (CZI Single-Cell Biology Program et al. 2023) |
| Target; disease | Context relevance - protein expression properties | Human Protein Atlas (Uhlén et al. 2015) |
| Target; disease |
Context relevance - alternatively spliced transcripts/isoforms | Ensembl (Harrison et al. 2024) GENCODE (Frankish et al. 2019) |
| Target; disease | Disease modifiable effect - clinical information | MedlinePlus (https://medlineplus.gov) ClinicalTrials.gov (https://www.clinicaltrials.gov/) Retrospective clinical data |
| Disease | Disease etiology | OMIM (McKusick 2007) Retrospective clinical data MedlinePlus (https://medlineplus.gov) |
| Disease |
Temporal onset | MedlinePlus (https://medlineplus.gov) ClinicalTrials.gov (https://www.clinicaltrials.gov/) Retrospective clinical data |
| Disease |
Clinical specificity - phenotypic properties | Retrospective clinical data MedlinePlus (https://medlineplus.gov) |
| Disease |
Clinical specificity - clinical variants | Varsome (Kopanos et al. 2019) GnomAD (Karczewski et al. 2020) ClinVar (Landrum et al. 2014) PharmVar (Gaedigk et al. 2021) PharmGKB (Whirl-Carrillo et al. 2021) PharmCAT (Sangkuhl et al. 2020) |
| Disease | Patient variability - patient characteristics | ClinicalTrials.gov (https://www.clinicaltrials.gov/) Electronic health records MedlinePlus (https://medlineplus.gov)TCGA (Cancer Genome Atlas Research Network et al. 2013) ReDO (Pantziarka et al. 2018)PharmGKB (Whirl-Carrillo et al. 2021) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




