Submitted:
07 November 2024
Posted:
08 November 2024
You are already at the latest version
Abstract

Keywords:
1. Background
2. Materials and Methods
2.1. Study Population and Sample Size Calculation
2.2. Method Validation
2.3. Selection of Stored Specimens
2.4. Extraction Based Protocol (Reference Method)
2.5. PCR Reagent Preparation (Master Mix Preparation)
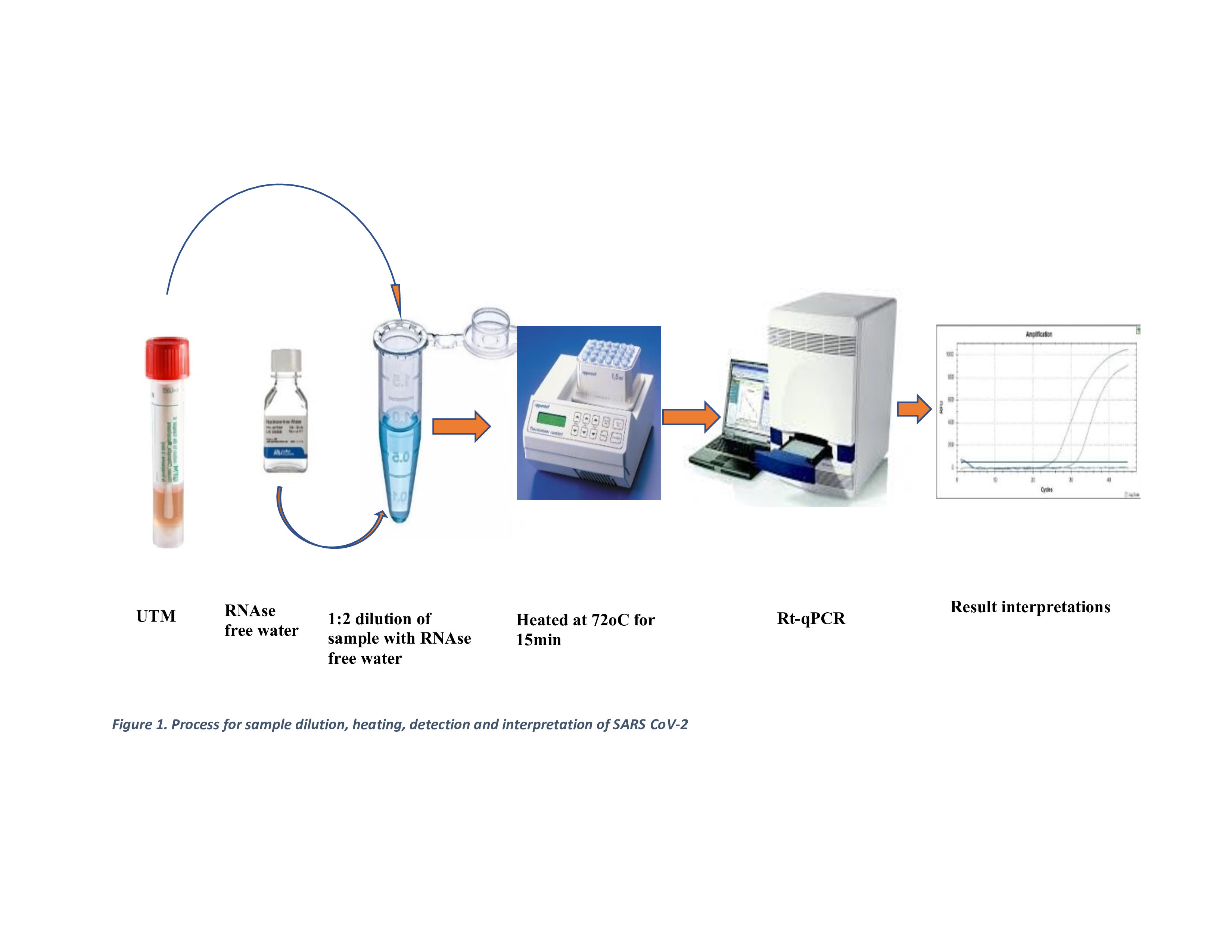
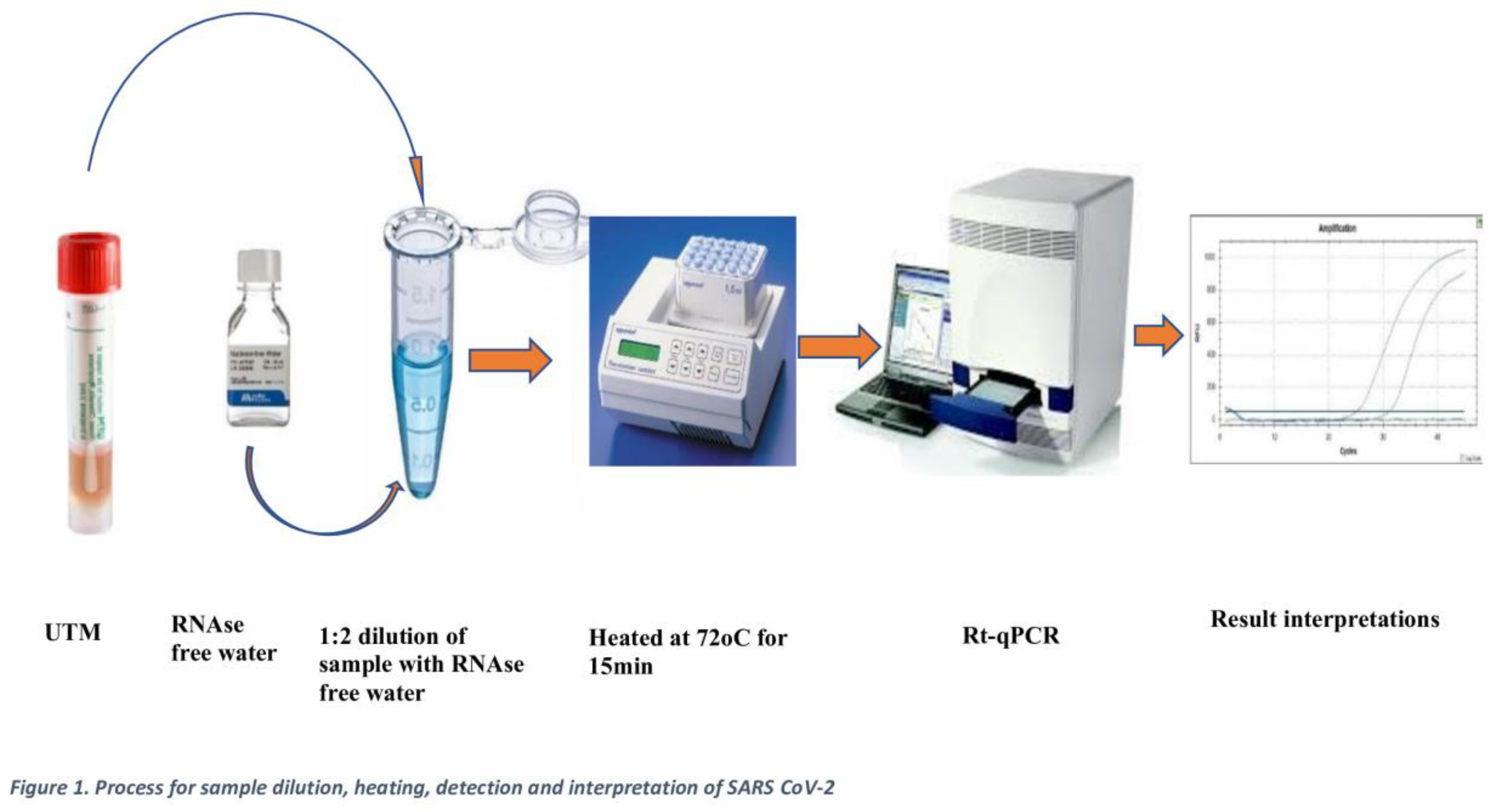
2.6. Extraction-Free (EFHD) Protocols (Index Method)
2.7. SARS-CoV-2 Detection (RT-qPCR)
2.8. Ethics Statement
2.9. Statistical Analysis
3. Results
3.1. Diagnostic Performance of Extraction-Free Method
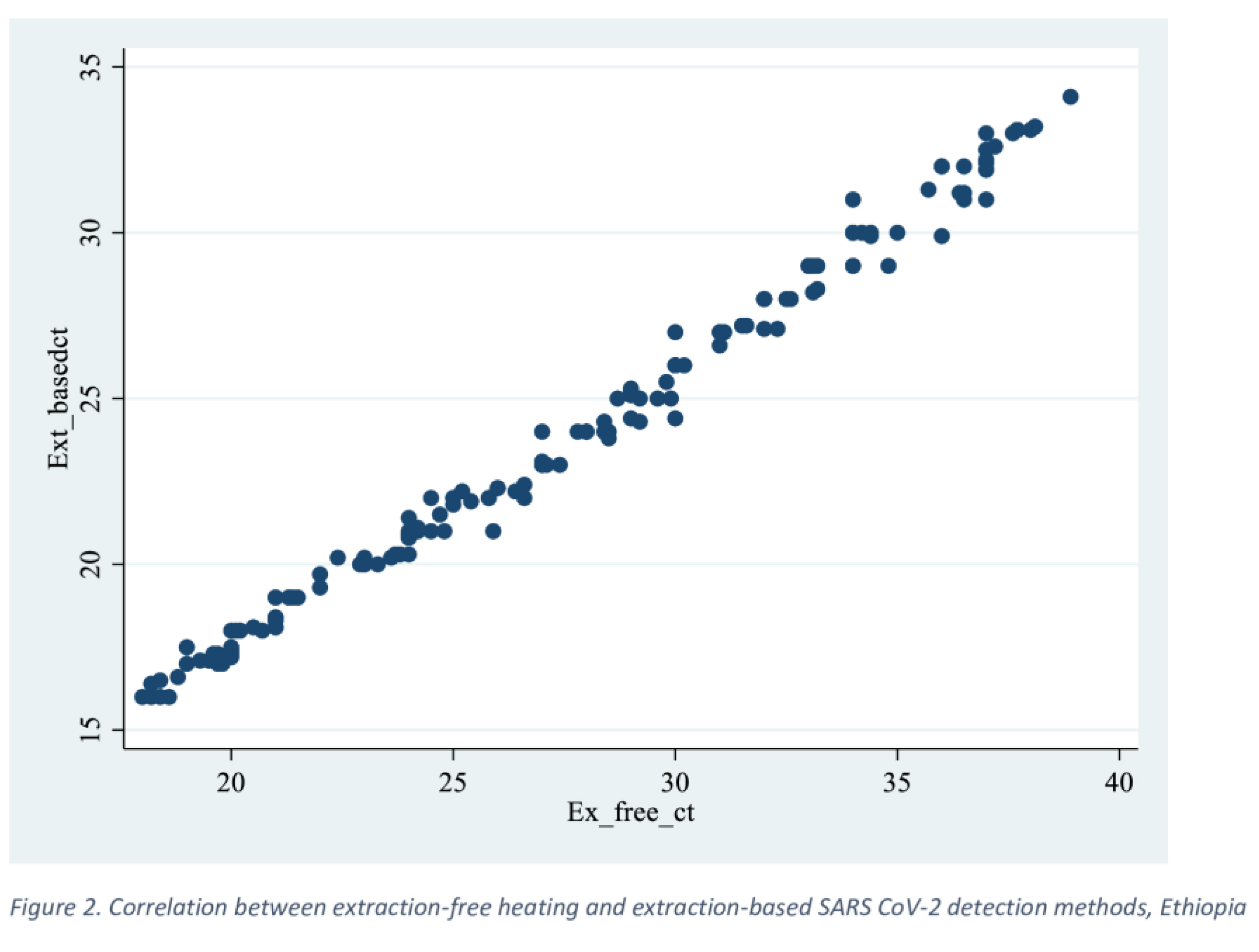
3.2. Comparison of Performance Characteristics in Terms of Ct-Value
4. Discussion
5. Conclusion
Author Contributions
Funding
Institutional Review Board Statatement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; et al. Genome Composition and Divergence of the Novel Coronavirus (2019-nCoV) Originating in China. Cell Host Microbe. 2020, 27, 325–328. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Yang, C.; Xu, X.F.; Xu, W.; Liu, S.W. Structural and functional properties of SARS-CoV-2 spike protein: Potential antivirus drug development for COVID-19. Acta Pharmacol Sin. 2020, 41, 1141–1149. [Google Scholar] [CrossRef] [PubMed]
- Harvey, W.T.; Carabelli, A.M.; Jackson, B.; Gupta, R.K.; Thomson, E.C.; Harrison, E.M.; et al. SARS-CoV-2 variants, spike mutations and immune escape. Nat Rev Microbiol. 2021, 19, 409–424. [Google Scholar] [CrossRef] [PubMed]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020, 25, 2000045. [Google Scholar] [CrossRef] [PubMed]
- Quantitatively assessing early detection strategies for mitigating COVID-19 and future pandemics | Nature Communications [Internet]. [cited 2024 May 2]. Available from: https://www.nature.com/articles/s41467-023-44199-7.
- Transmission risk of asymptomatic SARS-CoV-2 infection: A systematic review and meta-analysis - PMC [Internet]. [cited 2024 Jul 18]. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC9757919/.
- Joynt, G.M.; Wu, W.K. Understanding COVID-19: What does viral RNA load really mean? Lancet Infect Dis. 2020, 20, 635–636. [Google Scholar] [CrossRef]
- Sethuraman, N.; Jeremiah, S.S.; Ryo, A. Interpreting Diagnostic Tests for SARS-CoV-2. JAMA. 2020, 323, 2249–2251. [Google Scholar] [CrossRef]
- Loeffelholz, M.J.; Tang, Y.W. Laboratory diagnosis of emerging human coronavirus infections - the state of the art. Emerg Microbes Infect. 2020, 9, 747–756. [Google Scholar] [CrossRef]
- Cheng, V.C.C.; Lau, S.K.P.; Woo, P.C.Y.; Yuen, K.Y. Severe acute respiratory syndrome coronavirus as an agent of emerging and reemerging infection. Clin Microbiol Rev. 2007, 20, 660–694. [Google Scholar] [CrossRef]
- Ferrari, D.; Motta, A.; Strollo, M.; Banfi, G.; Locatelli, M. Routine blood tests as a potential diagnostic tool for COVID-19. Clin Chem Lab Med. 2020, 58, 1095–1099. [Google Scholar] [CrossRef]
- Clinical, laboratory and imaging features of COVID-19: A systematic review and meta-analysis - ScienceDirect [Internet]. [cited 2024 Jul 2]. Available from: https://www.sciencedirect.com/science/article/pii/S1477893920300910.
- Body Localization of ACE-2: On the Trail of the Keyhole of SARS-CoV-2 - PMC [Internet]. [cited 2024 Jul 20]. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7744810/.
- SARS-CoV-2: Structure, Biology, and Structure-Based Therapeutics Development - PMC [Internet]. [cited 2024 Jul 20]. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7723891/.
- How to Obtain a Nasopharyngeal Swab Specimen - PubMed [Internet]. [cited 2024 Jul 20]. Available from: https://pubmed.ncbi.nlm.nih.gov/32302471/.
- Böger, B.; Fachi, M.M.; Vilhena, R.O.; Cobre, A.F.; Tonin, F.S.; Pontarolo, R. Systematic review with meta-analysis of the accuracy of diagnostic tests for COVID-19. American Journal of Infection Control. 2021, 49, 21–29. [Google Scholar] [CrossRef]
- Smyrlaki, I.; Ekman, M.; Lentini, A.; Rufino De Sousa, N.; Papanicolaou, N.; Vondracek, M.; et al. Massive and rapid COVID-19 testing is feasible by extraction-free SARS-CoV-2 RT-PCR. Nat Commun. 2020, 11, 4812. [Google Scholar] [CrossRef] [PubMed]
- Bruce, E.A.; Huang, M.L.; Perchetti, G.A.; Tighe, S.; Laaguiby, P.; Hoffman, J.J.; et al. Direct RT-qPCR detection of SARS-CoV-2 RNA from patient nasopharyngeal swabs without an RNA extraction step. Lieberman RL, editor. PLoS Biol. 2020, 18, e3000896. [Google Scholar] [CrossRef] [PubMed]
- Alhamid, G.; Tombuloglu, H.; Rabaan, A.A.; Al-Suhaimi, E. SARS-CoV-2 detection methods: A comprehensive review. Saudi Journal of Biological Sciences 2022, 29. Available online: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC9512523/ (accessed on 20 July 2024). [CrossRef] [PubMed]
- RT-qPCR Testing of SARS-CoV-2: A Primer - PMC [Internet]. [cited 2024 Jul 20]. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7215906/.
- Aoki, M.N.; Coelho B de, O.; Góes, L.G.B.; Minoprio, P.; Durigon, E.L.; Morello, L.G.; et al. Colorimetric RT-LAMP SARS-CoV-2 diagnostic sensitivity relies on color interpretation and viral load. Scientific Reports 2021, 11. Available online: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8079700/ (accessed on 20 July 2024). [CrossRef] [PubMed]
- Tombuloglu, H.; Sabit, H.; Al-Suhaimi, E.; Al Jindan, R.; Alkharsah, K.R. Development of multiplex real-time RT-PCR assay for the detection of SARS-CoV-2. PLoS ONE. 2021, 16, e0250942. [Google Scholar] [CrossRef]
- SciELO - Brazil - Laboratory diagnosis for Covid-19: A mini-review Laboratory diagnosis for Covid-19: A mini-review [Internet]. [cited 2024 Jul 5]. Available from: https://www.scielo.br/j/rsbmt/a/P66HT68grCDFzSyshYgRqLf/.
- Lv, D.F.; Ying, Q.M.; Weng, Y.S.; Shen, C.B.; Chu, J.G.; Kong, J.P.; et al. Dynamic change process of target genes by RT-PCR testing of SARS-Cov-2 during the course of a Coronavirus Disease 2019 patient. Clin Chim Acta. 2020, 506, 172–175. [Google Scholar] [CrossRef]
- Tahmasebi, S.; Khosh, E.; Esmaeilzadeh, A. The outlook for diagnostic purposes of the 2019-novel coronavirus disease. J Cell Physiol. 2020, 235, 9211–9229. [Google Scholar] [CrossRef]
- Zhao, Y.; Cui, C.; Zhang, K.; Liu, J.; Xu, J.; Nisenbaum, E.; et al. COVID19: A Systematic Approach to Early Identification and Healthcare Worker Protection. Front Public Health. 2020, 8, 205. [Google Scholar] [CrossRef]
- Fomsgaard, A.S.; Rosenstierne, M.W. An alternative workflow for molecular detection of SARS-CoV-2 – escape from the NA extraction kit-shortage, Copenhagen, Denmark, March 2020. Eurosurveillance. 2020, 25, 2000398. [Google Scholar] [CrossRef]
- Villota, S.D.; Nipaz, V.E.; Carrazco-Montalvo, A.; Hernandez, S.; Waggoner, J.J.; Ponce, P.; et al. Alternative RNA extraction-free techniques for the real-time RT-PCR detection of SARS-CoV-2 in nasopharyngeal swab and sputum samples. Journal of Virological Methods. 2021, 298, 114302. [Google Scholar] [CrossRef]
- QIAGEN QIAamp® Viral RNA Mini Handbook, July 2020. - Google Search [Internet]. [cited 2023 Aug 21]. Available from: https://www.google.com/search?q=QIAGEN+QIAamp%C2%AE+Viral+RNA+Mini+Handbook%2C+July+2020.&rlz=1C1GCEU_enET1014ET1014&oq=QIAGEN+QIAamp%C2%AE+Viral+RNA+Mini+Handbook%2C+July+2020.&aqs=chrome..69i57.1026j0j4&sourceid=chrome&ie=UTF-8. 20 July.
- Jayaprakasam, M.; Aggarwal, S.; Mane, A.; Saxena, V.; Rao, A.; Bandopadhyay, B.; et al. RNA-extraction-free diagnostic method to detect SARS-CoV-2: An assessment from two states. India. Epidemiol Infect. 2021, 149, e245. [Google Scholar] [CrossRef]
- Download Citations | ACS Publications [Internet]. [cited 2024 May 2]. Available from: https://pubs.acs.org/action/showCitFormats?doi=10.1021/acs.analchem.0c03918&ref=pdf.
- Morecchiato, F.; Coppi, M.; Baccani, I.; Maggini, N.; Ciccone, N.; Antonelli, A.; et al. Evaluation of extraction-free RT-PCR methods for faster and cheaper detection of SARS-CoV-2 using two commercial systems. International Journal of Infectious Diseases. 2021, 112, 264–268. [Google Scholar] [CrossRef] [PubMed]
- Mahmoud, S.A.; Ganesan, S.; Ibrahim, E.; Thakre, B.; Teddy, J.G.; Raheja, P.; et al. Evaluation of RNA Extraction-Free Method for Detection of SARS-CoV-2 in Salivary Samples for Mass Screening for COVID-19. Gao SJ, editor. BioMed Research International. 2021, 2021, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Mokhtar, K.M. Improved COVID-19 testing by extraction free SARS-Cov-2 RT-PCR [Internet]. 2020 [cited 2024 May 2]. Available from: http://medrxiv.org/lookup/doi/10.1101/2020.08.10.20171512.
- Evaluation of extraction-free RT-qPCR methods for SARS-CoV-2 diagnostics - PubMed [Internet]. [cited 2024 Jun 24]. Available from: https://pubmed.ncbi.nlm.nih.gov/34302551/.
- Diagnostic Laboratory Tests for COVID-19 in US: Methodology and Performance - PMC [Internet]. [cited 2024 Jul 5]. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7386512/.
- Hasan, M.R.; Mirza, F.; Al-Hail, H.; Sundararaju, S.; Xaba, T.; Iqbal, M.; et al. Detection of SARS-CoV-2 RNA by direct RT-qPCR on nasopharyngeal specimens without extraction of viral RNA. Darlix JLE, editor. PLoS ONE. 2020, 15, e0236564. [Google Scholar]
- (PDF) Evaluation of extraction-free RT-qPCR methods for SARS-CoV-2 diagnostics [Internet]. [cited 2024 Jul 5]. Available from: https://www.researchgate.net/publication/353436476_Evaluation_of_extraction-free_RT-qPCR_methods_for_SARS-CoV-2_diagnostics.
| Institute | Gene Targets |
|---|---|
| China CDC, China | ORF 1ab and N |
| Institute Pasteur, France | Two targets in RdRP |
| US CDC, USA | Three targets in the N gene |
| National Institute of Infectious Disease, Japan | Pancrorona and multiple targets, Spike protein |
| Charité, Germany | RdRP, E, N |
| HKU, Hong Kong SAR | ORF 1b-nsp14, N |
| National Institute of Health, Thailand | N |
| Detection Based Method | |||
| Detection free Method (EFh+) | Positive | Negative | Total |
| Positive | 163 | 0 | 163 |
| Negative | 15 | 110 | 125 |
| Total | 178 | 110 | 288 |
| Statistic | Value | 95% CI |
|---|---|---|
| Apparent prevalence | 57% | 51% to 62% |
| True Prevalence | 62% | 56% to 67% |
| Sensitivity | 92% | 86% to 95% |
| Specificity | 100% | 85% to 100% |
| Negative Likelihood Ratio | 0.08 | 0.05 to 0.14 |
| Positive Predictive Value | 100% | 98% to 100% |
| Negative Predictive Value | 88% | 81% to 93% |
| Accuracy/Agreement/ | 95% | 92% to 97% |
| Extraction free | Extraction based | |||
| Positive | Negative | Total | ||
| Ct <20) | Positive | 38 | 0 | 38 |
| Negative | 0 | 0 | 0 | |
| Total | 38 | 0 | 38 | |
| Ct = 20-35 | Positive | 121 | 0 | 121 |
| Negative | 3 | 0 | 3 | |
| Total | 124 | 0 | 124 | |
| Ct > 35 | Positive | 0 | 0 | 0 |
| Negative | 9 | 110 | 119 | |
| Total | 9 | 110 | 119 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




