Submitted:
28 October 2024
Posted:
30 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Works
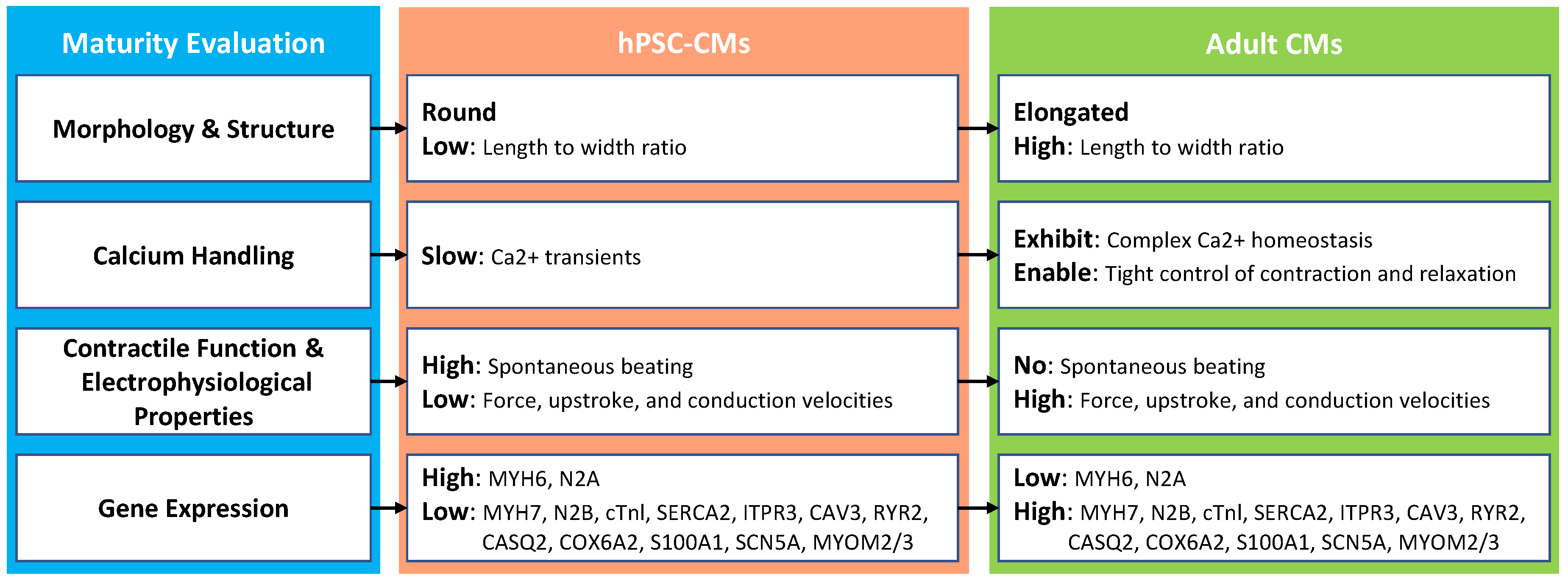
2.1. Morphology and Structure
2.2. Calcium Handling
2.3. Contractile Function and Electrophysiological Properties
2.4. Gene Expression
3. The hPSC-CM Maturity Evaluation Proxy
3.1. Cardiac-specific Gene Data Collection
3.2. Maturation Level v.s. Culture Duration
4. Data-Driven Maturity Quantification
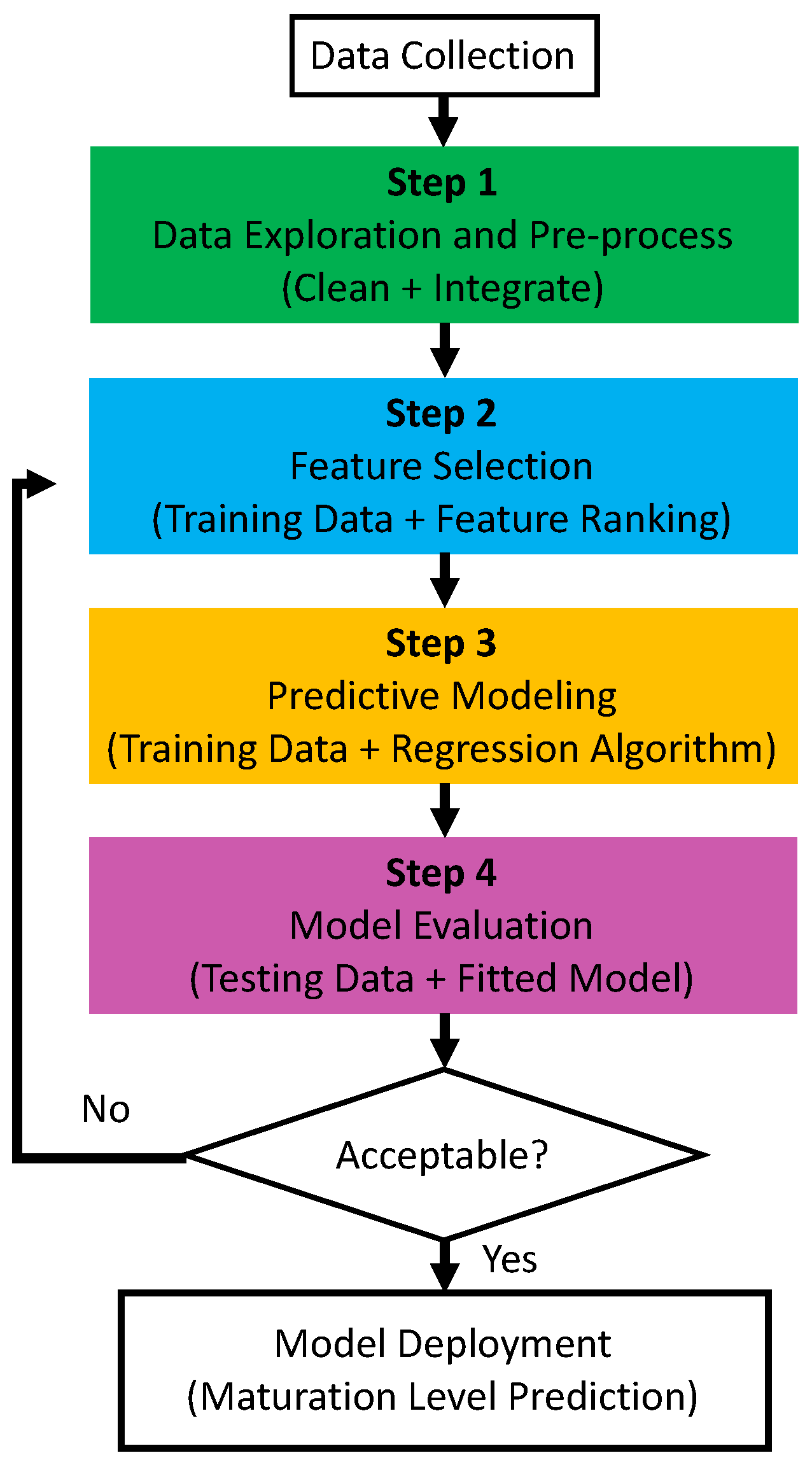
4.1. Data-Driven Maturity Quantification Pipeline
- Method 1 (): Filter method and linear regression
- Method 2 (): Wrapper method
- Method 3 (): Embedded method
- Method 4 (): Non-linear feature selection and non-linear regression
- Method 5 (): Non-linear feature selection and linear regression
4.2. — Filter Method (Pearson Correlation) + Linear Regression
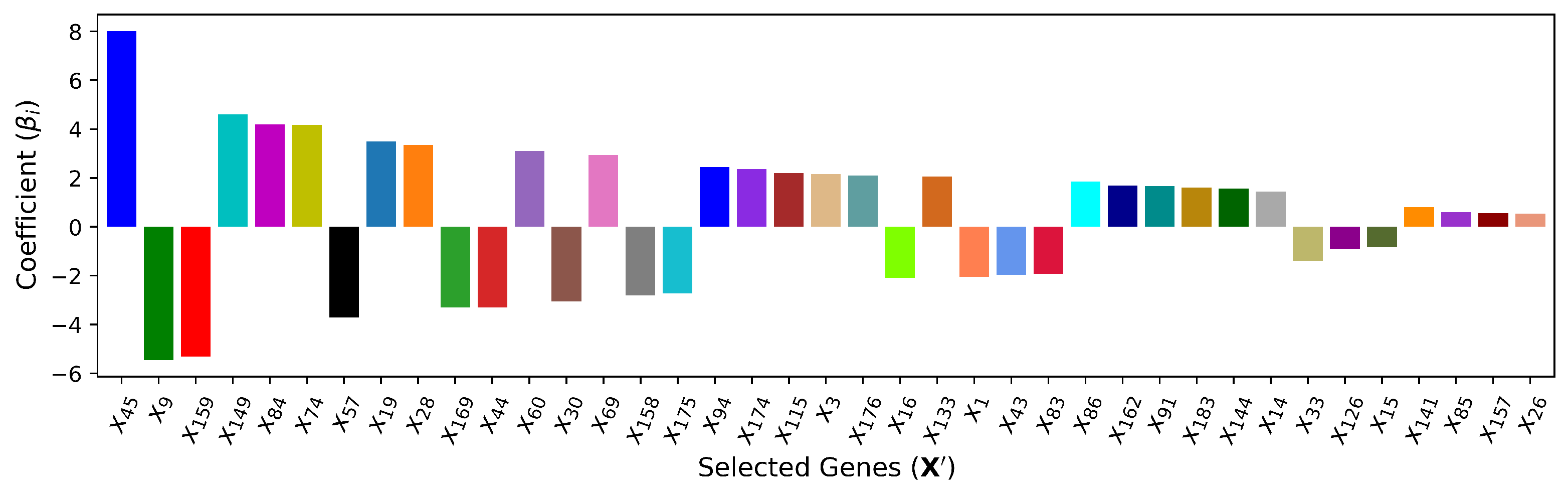
4.3. — Wrapper Method (Recursive Feature Elimination + Linear Regression)
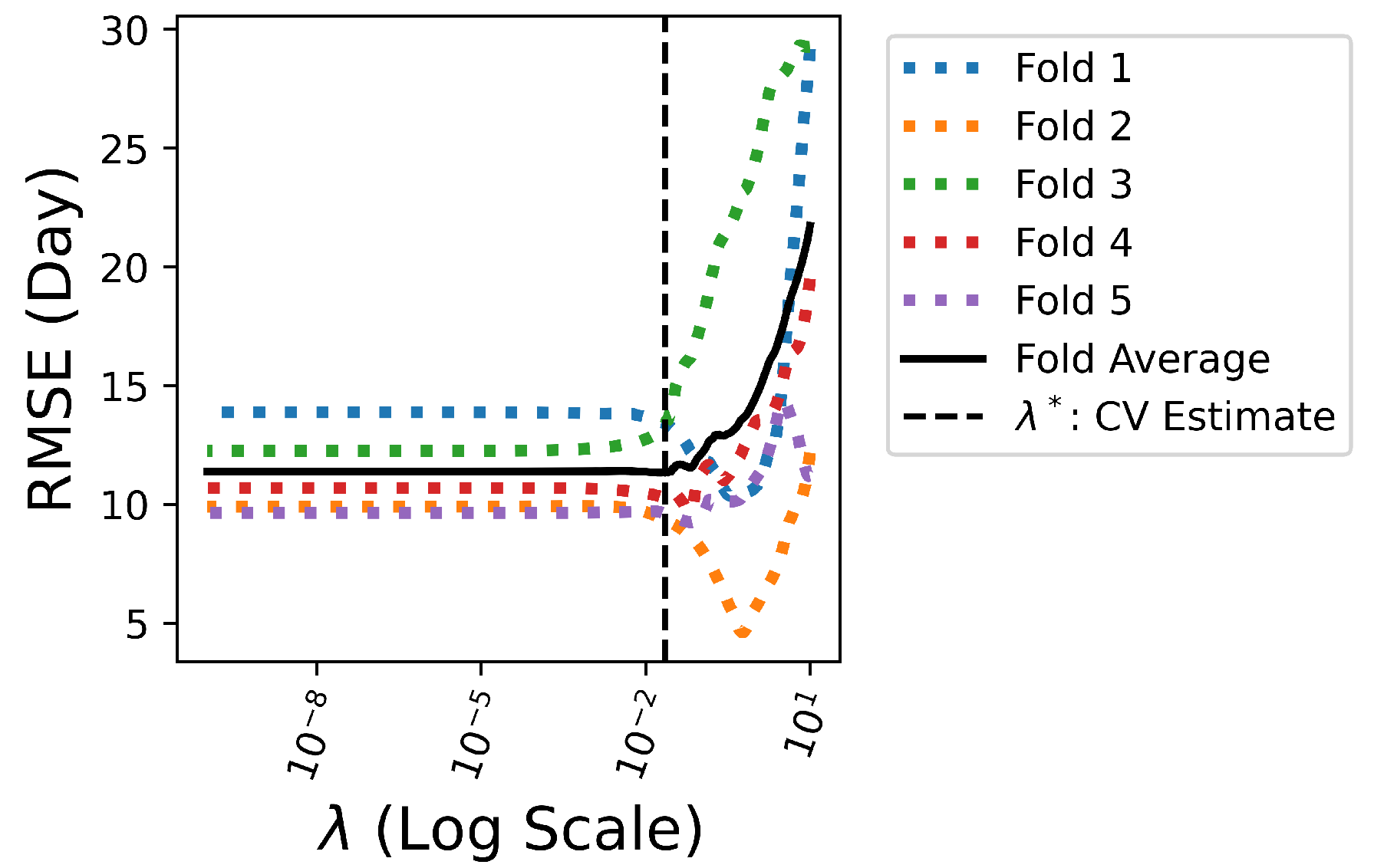
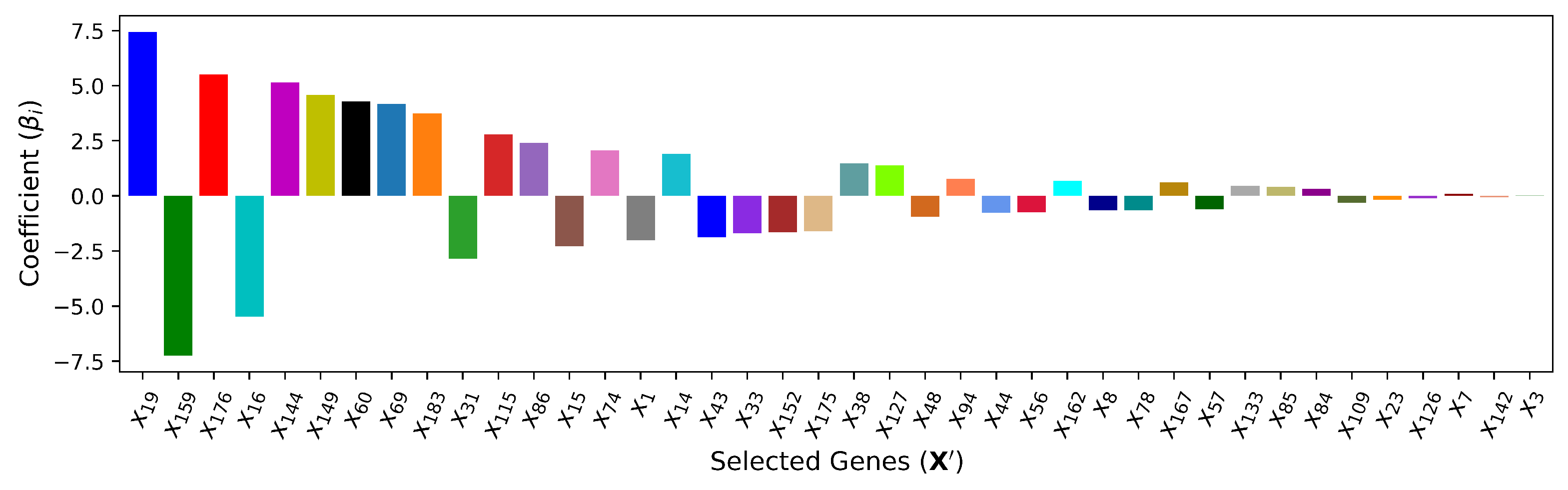
4.4. — Embedded Method (Lasso Regularization)
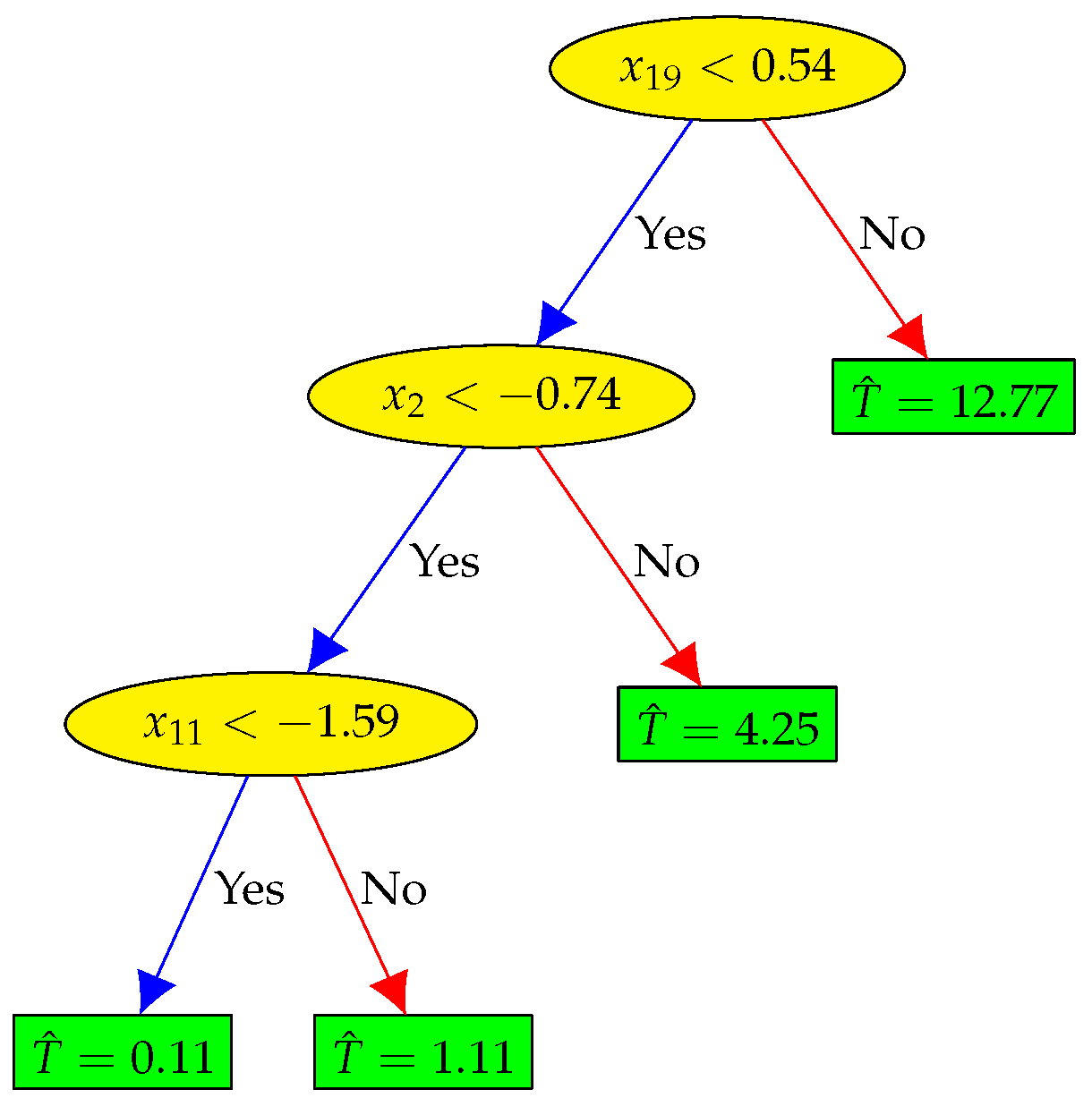
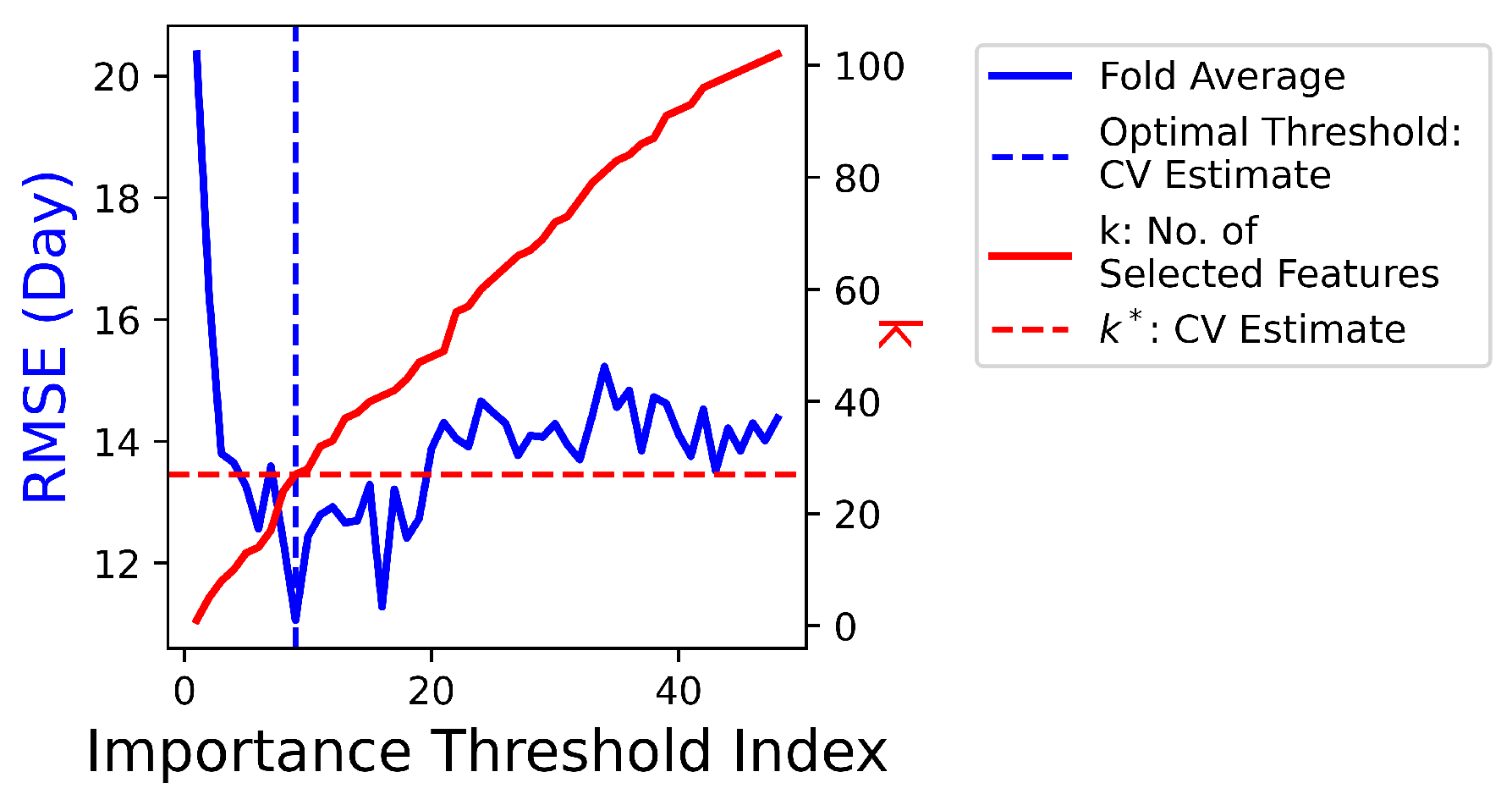
4.5. — Non-Linear Feature Selection and Regression (XGBoost Method)
4.6. — XGBoost Method + Linear Regression
5. Results and Discussions
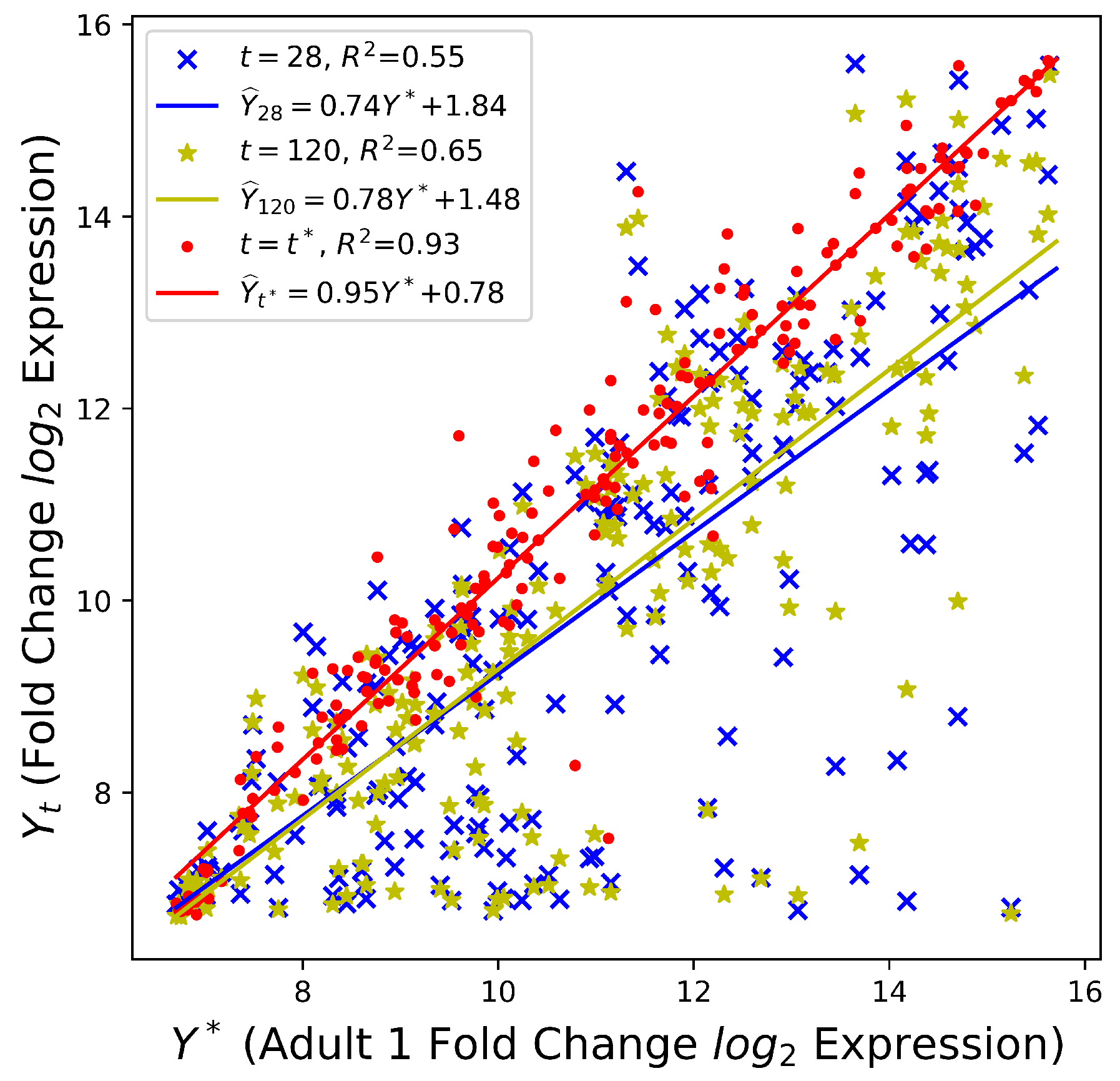
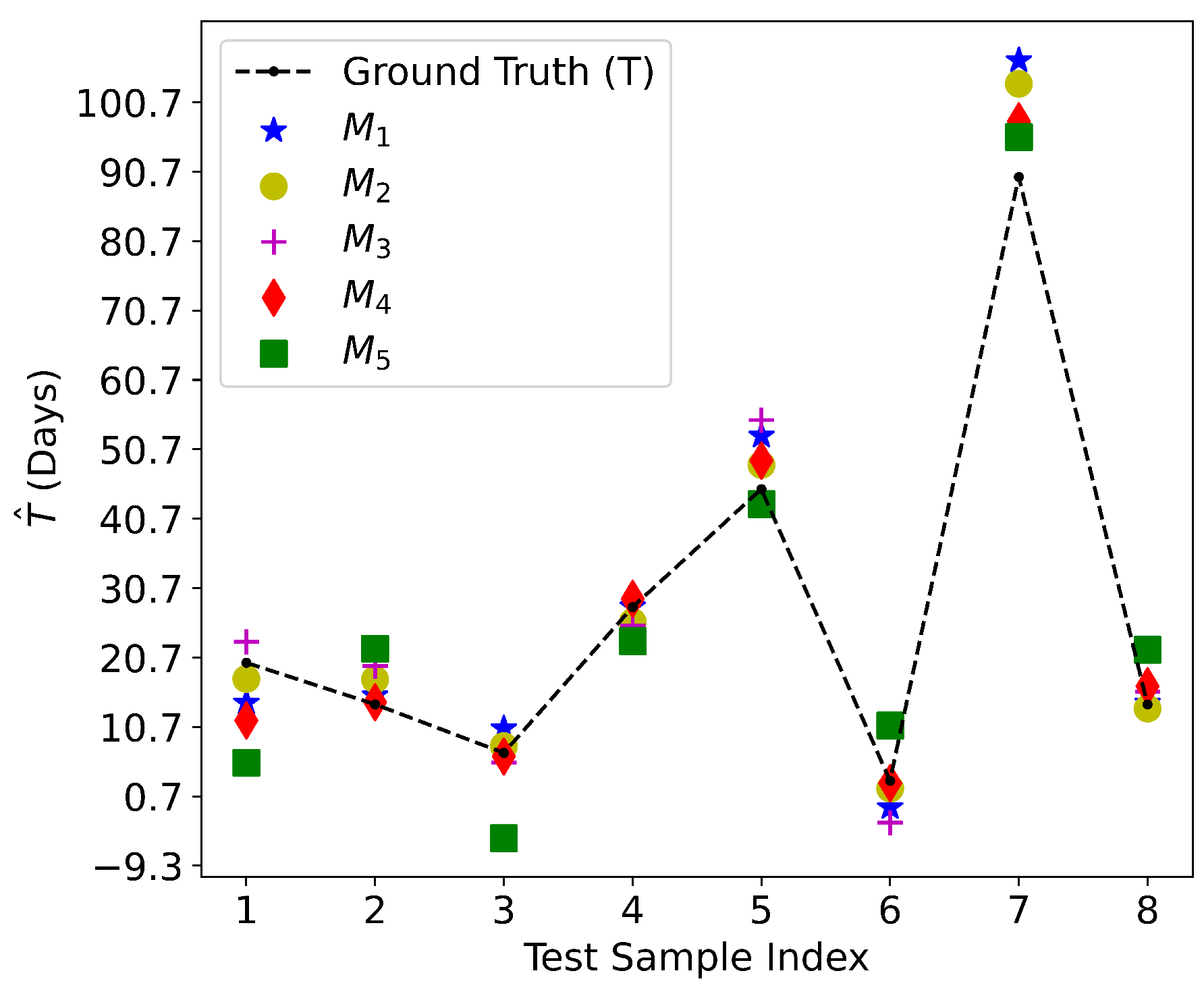
5.1. Culture Time Prediction
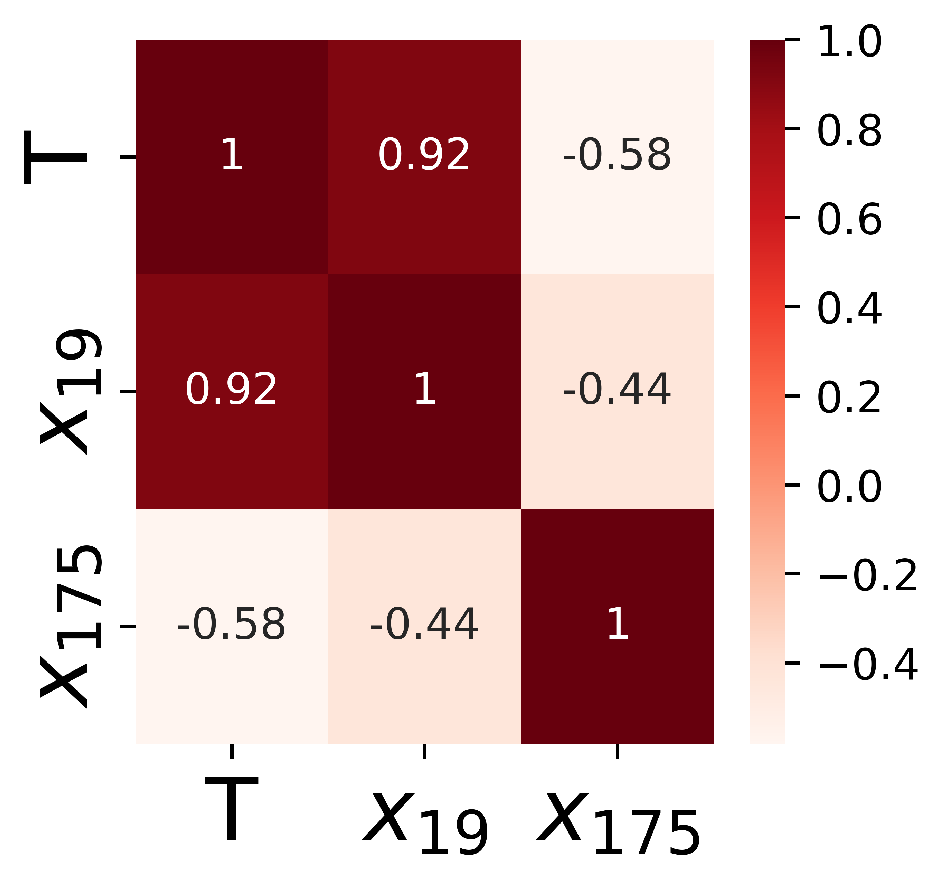
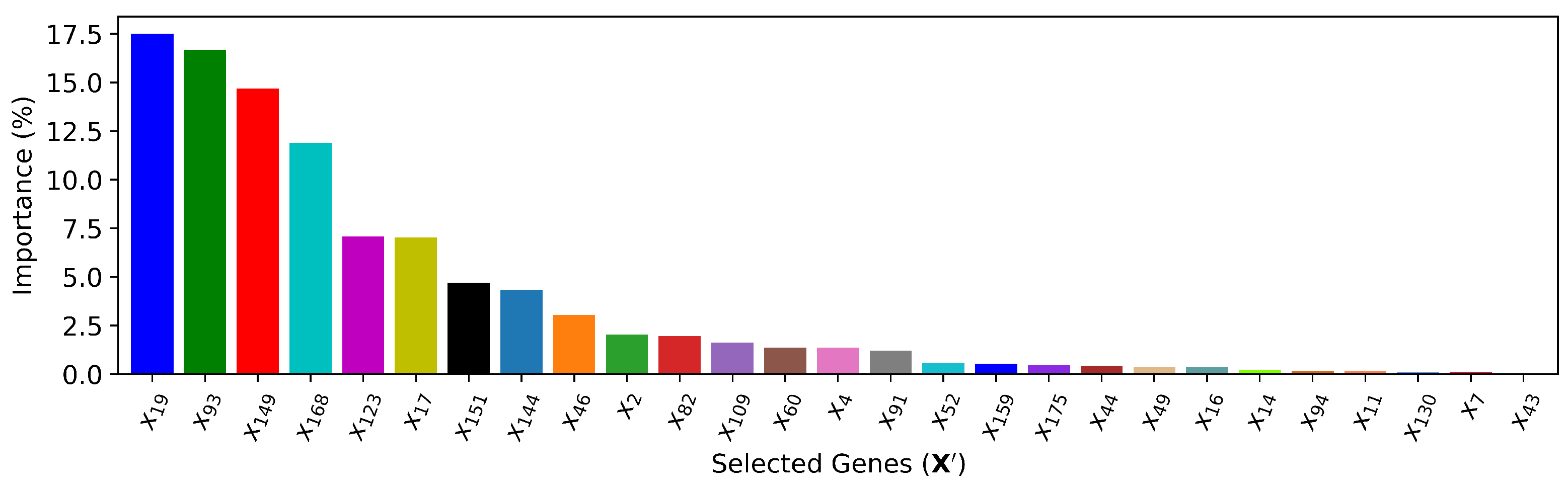
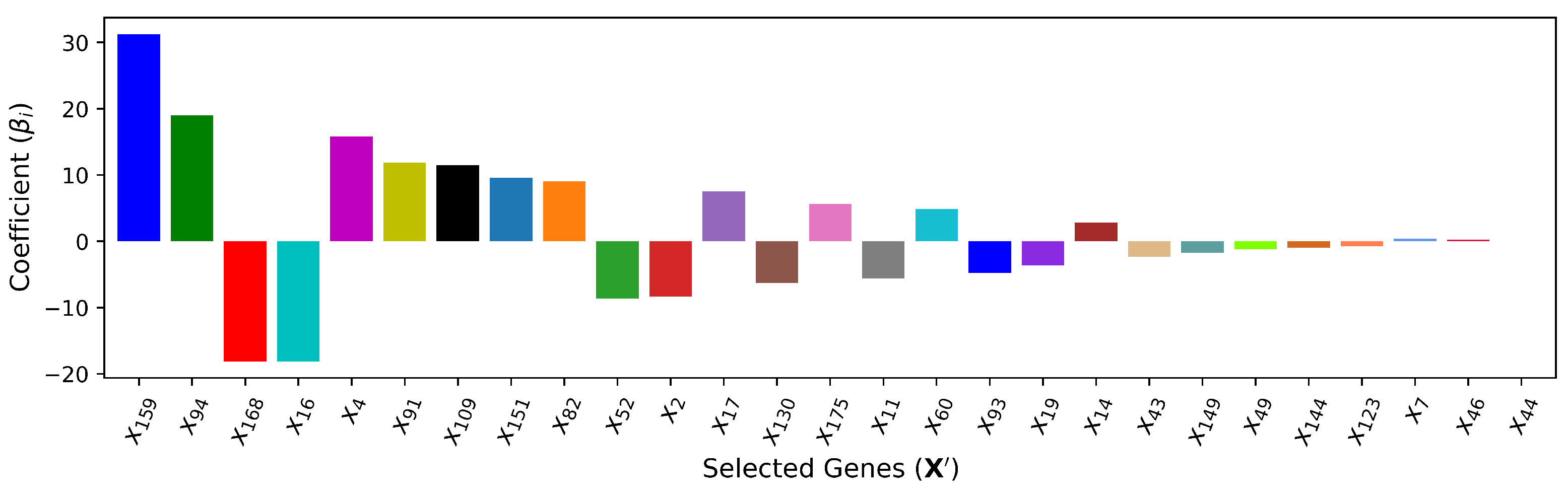
5.2. Cardiac Gene Selection
6. Conclusion
Acknowledgments
Conflicts of Interest
Appendix A
| X | Gene | X | Gene | X | Gene | X | Gene | |||
|---|---|---|---|---|---|---|---|---|---|---|
| ABO | ACO2 | ACOT2 | ACOX2 | |||||||
| ACTN2 | AK1 | ANK1 | ANKRD2 | |||||||
| ATP5G1 | ATP5G3 | BAG3 | BRP44L | |||||||
| BSG | C1QA | C1QB | CA4 | |||||||
| CABC1 | CAND2 | CASQ2 | CCL15 | |||||||
| CD151 | CD320 | CFD | CHST7 | |||||||
| CKM | CKMT2 | CLEC3B | CLTB | |||||||
| COQ9 | COX5A | COX5B | COX6A2 | |||||||
| COX7A1 | COX8A | CPT1B | CRIP2 | |||||||
| CRYAB | CRYM | CSDC2 | CSRP3 | |||||||
| CTTN | CYC1 | DCHS1 | DCI | |||||||
| DES | DEXI | DMPK | DSPP | |||||||
| ECHDC3 | ECSIT | EEF1A2 | EFEMP2 | |||||||
| ENDOG | ERCC1 | FABP3 | FAHD2A | |||||||
| FARS2 | FHL2 | FLJ22222 | FLNC | |||||||
| FXYD1 | GADD45GIP1 | GAMT | GATA4 | |||||||
| GATA6 | GOT1 | GPC1 | GYS1 | |||||||
| HOMER3 | HRC | HSPB1 | HSPB2 | |||||||
| HSPB3 | HSPB6 | HSPB7 | HSPB8 | |||||||
| HSPG2 | ICAM2 | IDH2 | IFI27 | |||||||
| IGFBP2 | IGFBP7 | IL11RA | ILVBL | |||||||
| INMT | ITGA7 | ITGB1BP2 | ITGB1BP3 | |||||||
| KCNH2 | LDB3 | LGALS3BP | MAPKAPK3 | |||||||
| MB | MCOLN1 | MRAS | MRPL12 | |||||||
| MRPL34 | MRPL41 | MRPS12 | MSRB2 | |||||||
| MYBPC3 | MYH6 | MYH7 | MYL2 | |||||||
| MYL3 | MYL4 | MYL7 | MYL9 | |||||||
| MYLC2PL | MYOM1 | MYOM2 | MYOZ2 | |||||||
| NDUFA11 | NDUFA7 | NDUFB10 | NDUFB7 | |||||||
| NDUFS7 | NDUFS8 | NKX2-5 | NOL3 | |||||||
| NPPA | NPPB | NRAP | OGDH | |||||||
| OPLAH | PCTK3 | PDE4DIP | PDK2 | |||||||
| PDLIM5 | PGAM2 | PGM1 | PHPT1 | |||||||
| PLA2G5 | PLEKHF1 | PLN | POLR2I | |||||||
| POLRMT | POMGNT1 | POPDC2 | PPAPDC3 | |||||||
| PPP1R13L | PPP1R1A | PPP2R3B | PTGDS | |||||||
| PTP4A3 | PTPLA | PTRF | PXMP2 | |||||||
| RAMP1 | RAMP3 | RASIP1 | RBPMS | |||||||
| RGS3 | RRAS | S100A1 | SEPW1 | |||||||
| SGCG | SH3RF2 | SIVA | SLC25A11 | |||||||
| SLC25A4 | SLC29A1 | SLC4A3 | SMPX | |||||||
| SMTN | SNTA1 | STAB1 | STOML1 | |||||||
| STOML2 | SYNPO2L | TACC2 | TAX1BP3 | |||||||
| TCAP | TIMM8B | TM7SF2 | TMEM159 | |||||||
| TNNC1 | TNNI3 | TNNT2 | TNXA | |||||||
| TNXB | TPM1 | TSPAN4 | UQCR | |||||||
| UQCRC1 | VAMP5 | VEGFB | VWF | |||||||
| WDR13 | ||||||||||
References
- Virani, S.S.; Alonso, A.; Aparicio, H.J.; Benjamin, E.J.; Bittencourt, M.S.; Callaway, C.W.; Carson, A.P.; Chamberlain, A.M.; Cheng, S.; Delling, F.N.; Elkind, M.S.; Evenson, K.R.; Ferguson, J.F.; Gupta, D.K.; Khan, S.S.; Kissela, B.M.; Knutson, K.L.; Lee, C.D.; Lewis, T.T.; Liu, J.; Loop, M.S.; Lutsey, P.L.; Ma, J.; Mackey, J.; Martin, S.S.; Matchar, D.B.; Mussolino, M.E.; Navaneethan, S.D.; Perak, A.M.; Roth, G.A.; Samad, Z.; Satou, G.M.; Schroeder, E.B.; Shah, S.H.; Shay, C.M.; Stokes, A.; VanWagner, L.B.; Wang, N.Y.; Tsao, C.W. Heart Disease and Stroke Statistics – 2021 Update. Circulation 2021, 143, e254–e743. [CrossRef]
- Fryar, C.D.; Chen, T.C.; Li, X. Prevalence of uncontrolled risk factors for cardiovascular disease: United States, 1999-2010. NCHS Data Brief 2012, pp. 1–8.
- Centers for Disease Control and Prevention. Underlying Cause of Death, 1999-2019 Request, accessed August 2021.
- Sinnecker, D.; Laugwitz, K.L.; Moretti, A. Induced pluripotent stem cell-derived cardiomyocytes for drug development and toxicity testing. Pharmacol Ther 2014, 143, 246–252. [CrossRef]
- Jung, G.; Bernstein, D. hiPSC Modeling of Inherited Cardiomyopathies. Curr Treat Options Cardiovasc Med 2014, 16, 320. [CrossRef]
- Nunes, S.S.; Miklas, J.W.; Liu, J.; Aschar-Sobbi, R.; Xiao, Y.; Zhang, B.; Jiang, J.; Massé, S.; Gagliardi, M.; Hsieh, A.; Thavandiran, N.; Laflamme, M.A.; Nanthakumar, K.; Gross, G.J.; Backx, P.H.; Keller, G.; Radisic, M. Biowire: A platform for maturation of human pluripotent stem cell-derived cardiomyocytes. Nature Methods 2013, 10, 781–787. [CrossRef]
- Shah, N.; Morsi, Y.; Manasseh, R. From mechanical stimulation to biological pathways in the regulation of stem cell fate. Cell Biochemistry and Function 2014, 32, 309–325. [CrossRef]
- Venkatesh, S.; Baljinnyam, E.; Tong, M.; Kashihara, T.; Yan, L.; Liu, T.; Li, H.; Xie, L.H.; Nakamura, M.; ichi Oka, S.; Suzuki, C.K.; Fraidenraich, D.; Sadoshima, J. Proteomic analysis of mitochondrial biogenesis in cardiomyocytes differentiated from human induced pluripotent stem cells. Am J Physiol Regul Integr Comp Physiol 2021, 320, R547 – R562.
- Lundy, S.D.; Zhu, W.Z.; Regnier, M.; Laflamme, M.A. Structural and functional maturation of cardiomyocytes derived from human pluripotent stem cells. Stem Cells Dev 2013, 22, 1991–2002. [CrossRef]
- Ahmed, R.E.; Anzai, T.; Chanthra, N.; Uosaki, H. A Brief Review of Current Maturation Methods for Human Induced Pluripotent Stem Cells-Derived Cardiomyocytes. Frontiers in Cell and Developmental Biology 2020, 8, 178. [CrossRef]
- Solomatine, D.P.; Ostfeld, A. Data-driven modelling: some past experiences and new approaches. Journal of Hydroinformatics 2008, 10, 3–22. [CrossRef]
- Ren, Y.; Cui, Q.; Zhao, X.; Wang, Y.; Huang, X.; Ni, W. Data-Driven Intelligent Management of Energy Constrained Autonomous Vehicles in Smart Cities. Cognitive Radio-Oriented Wireless Networks, 2021, pp. 112–125.
- Kamel, E.; Sheikh, S.; Huang, X. Data-driven predictive models for residential building energy use based on the segregation of heating and cooling days. Energy 2020, 206, 118045. [CrossRef]
- Babiarz, J.E.; Ravon, M.; Sridhar, S.; Ravindran, P.; Swanson, B.; Bitter, H.; Weiser, T.; Chiao, E.; Certa, U.; Kolaja, K.L. miRNA expression profiling of differentiating human-induced pluripotent stem cell (hiPSC)-derived cardiomyocytes, accessed July 2021, [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE35672].
- Alfons Staerk. What do Lord Kelvin and Peter Drucker have in common?, 2021.
- Peters, N.S.; Severs, N.J.; Rothery, S.M.; Lincoln, C.; Yacoub, M.H.; Green, C.R. Spatiotemporal relation between gap junctions and fascia adherens junctions during postnatal development of human ventricular myocardium. Circulation 1994, 90, 713–725. [CrossRef]
- Laflamme, M.A.; Murry, C.E. Heart regeneration. Nature 2011, 473, 326–335. [CrossRef]
- Jiang, Y.; Park, P.; Hong, S.M.; Ban, K. Maturation of Cardiomyocytes Derived from Human Pluripotent Stem Cells: Current Strategies and Limitations. Mol Cells 2018, 41, 613–621. [CrossRef]
- Denning, C.; Borgdorff, V.; Crutchley, J.; Firth, K.S.A.; George, V.; Kalra, S.; Kondrashov, A.; Hoang, M.D.; Mosqueira, D.; Patel, A.; Prodanov, L.; Rajamohan, D.; Skarnes, W.C.; Smith, J.G.W.; Young, L.E. Cardiomyocytes from human pluripotent stem cells: From laboratory curiosity to industrial biomedical platform. Biochim Biophys Acta 2016, 1863, 1728–1748. [CrossRef]
- Liu, J.; Lieu, D.K.; Siu, C.W.; Fu, J.D.; Tse, H.F.; Li, R.A. Facilitated maturation of Ca2+ handling properties of human embryonic stem cell-derived cardiomyocytes by calsequestrin expression. American journal of physiology. Cell physiology 2009.
- van den Berg, C.W.; Okawa, S.; Chuva de Sousa Lopes, S.M.; van Iperen, L.; Passier, R.; Braam, S.R.; Tertoolen, L.G.; del Sol, A.; Davis, R.P.; Mummery, C.L. Transcriptome of human foetal heart compared with cardiomyocytes from pluripotent stem cells. Development 2015, 142, 3231–3238, [https://journals.biologists.com/dev/article-pdf/142/18/3231/1838606/dev123810.pdf]. [CrossRef]
- Babiarz, J.E.; Ravon, M.; Sridhar, S.; Ravindran, P.; Swanson, B.; Bitter, H.; Weiser, T.; Chiao, E.; Certa, U.; Kolaja, K.L. Determination of the human cardiomyocyte mRNA and miRNA differentiation network by fine-scale profiling. Stem Cells Dev 2012, 21, 1956–1965. [CrossRef]
- Su, A.I.; Wiltshire, T.; Batalov, S.; Lapp, H.; Ching, K.A.; Block, D.; Zhang, J.; Soden, R.; Hayakawa, M.; Kreiman, G.; Cooke, M.P.; Walker, J.R.; Hogenesch, J.B. A gene atlas of the mouse and human protein-encoding transcriptomes. Proceedings of the National Academy of Sciences 2004, 101, 6062–6067. [CrossRef]
- Illumina Inc.. Illumina HumanWG-6 v3.0 expression beadchip, 2021.
- Siegel, A.F. Chapter 12 - Multiple Regression: Predicting One Variable From Several Others. In Practical Business Statistics; Academic Press, 2016; pp. 355–418. [CrossRef]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 2016, pp. 785–794. [CrossRef]
- Zhang, X.; Wang, W.; Li, F.; Voiculescu, I. Stretchable impedance sensor for mammalian cell proliferation measurements. Lab Chip 2017, 17, 2054–2066. [CrossRef]
| Gene | Description | Adult 1 | Adult 2 | Day 120 | Day 0 | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| ACTC1 | Actin, alpha, cardiac muscle 1 | 15.64 | 15.60 | 15.48 | 9.74 | |||||
| MYH7 | Myosin light chain 7 | 15.62 | 15.62 | 14.02 | 6.82 | |||||
| CRYAB | Crystallin alpha B | 15.52 | 15.48 | 13.81 | 6.80 | |||||
| TNNC1 | Troponin C1, slow skeletal and cardiac type | 15.50 | 15.30 | 14.57 | 7.46 | |||||
| MYL2 | Myosin light chain 2 | 15.43 | 15.38 | 14.56 | 6.99 | |||||
| MYL3 | Myosin light chain 3 | 15.15 | 15.18 | 14.60 | 6.86 | |||||
| MYH6 | Myosin light chain 6 | 14.71 | 15.57 | 15.01 | 6.99 | |||||
| MB | Myoglobin | 14.59 | 14.50 | 13.67 | 6.90 | |||||
| MYBPC3 | Myosin binding protein C, cardiac | 14.54 | 14.71 | 13.96 | 6.85 | |||||
| TNNT2 | Troponin T2, cardiac type | 14.51 | 14.08 | 13.72 | 7.48 | |||||
| TNNI3 | Troponin I3, cardiac type | 14.37 | 14.06 | 12.32 | 7.95 | |||||
| CKMT2 | Creatine kinase, mitochondrial 2 | 14.22 | 14.29 | 12.45 | 7.16 | |||||
| NPPA | Natriuretic peptide A | 14.17 | 14.95 | 15.22 | 6.84 | |||||
| CASQ2 | Calsequestrin 2 | 14.08 | 13.69 | 12.41 | 6.92 | |||||
| HRC | Histidine rich calcium binding protein | 14.02 | 13.96 | 11.81 | 7.38 | |||||
| MYL7 | Myoslin light chain 7 | 13.65 | 14.24 | 15.07 | 7.11 | |||||
| ACTN2 | Actinin alpha 2 | 12.15 | 11.31 | 10.59 | 7.48 | |||||
| NKX2-5 | NK2 homeobox 5 | 11.10 | 11.03 | 10.71 | 6.76 | |||||
| PLN | Phospholamban | 10.79 | 8.28 | 11.50 | 6.88 | |||||
| LDB3 | LIM domain binding 3 | 9.15 | 8.76 | 8.92 | 6.86 | |||||
| KCNH2 | Potassium voltage-gated channel subfamily H member 2 | 8.16 | 8.52 | 8.07 | 7.10 | |||||
| Time t | Time t | |||
|---|---|---|---|---|
| Day 0 | 0.08 | Day 28 | 0.55 | |
| Day 3 | 0.09 | Day 35 | 0.58 | |
| Day 7 | 0.12 | Day 45 | 0.59 | |
| Day 10 | 0.23 | Day 60 | 0.61 | |
| Day 14 | 0.37 | Day 90 | 0.61 | |
| Day 20 | 0.49 | Day 120 | 0.65 |
| Data Set | Number of Records | Percentage | ||
|---|---|---|---|---|
| Training | 85% | |||
| Testing | 15% |
| Method | Genes in Ranked by Importance | RMSE (Day) | Score | |||||
|---|---|---|---|---|---|---|---|---|
| (Linear) | 2 | , | 6.837 | 0.934 | ||||
| (Linear) | 39 | , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , | 5.216 | 0.962 | ||||
| (Linear) | 40 | , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , | 5.521 | 0.957 | ||||
| (XGBoost) | 27 | , , , , , , , , , , , , , , , , , , , , , , , , , , | 4.461 | 0.972 | ||||
| (Linear) | 27 | , , , , , , , , , , , , , , , , , , , , , , , , , , | 8.724 | 0.892 | ||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




