Submitted:
28 October 2024
Posted:
28 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Search for Genes in T. rubrum Homologous to Genes Related to Oxidative Stress in Other Trichophyton Species
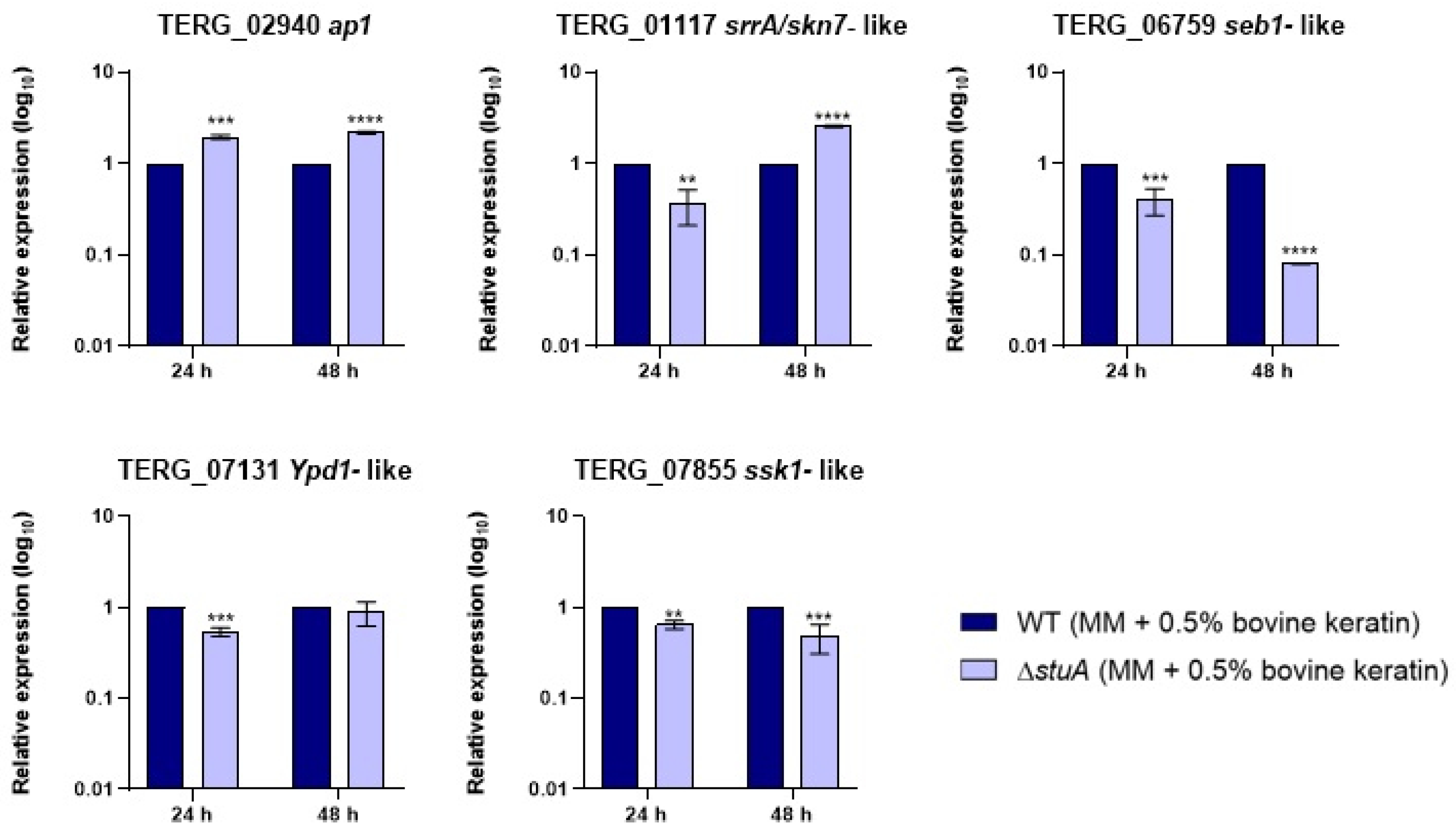
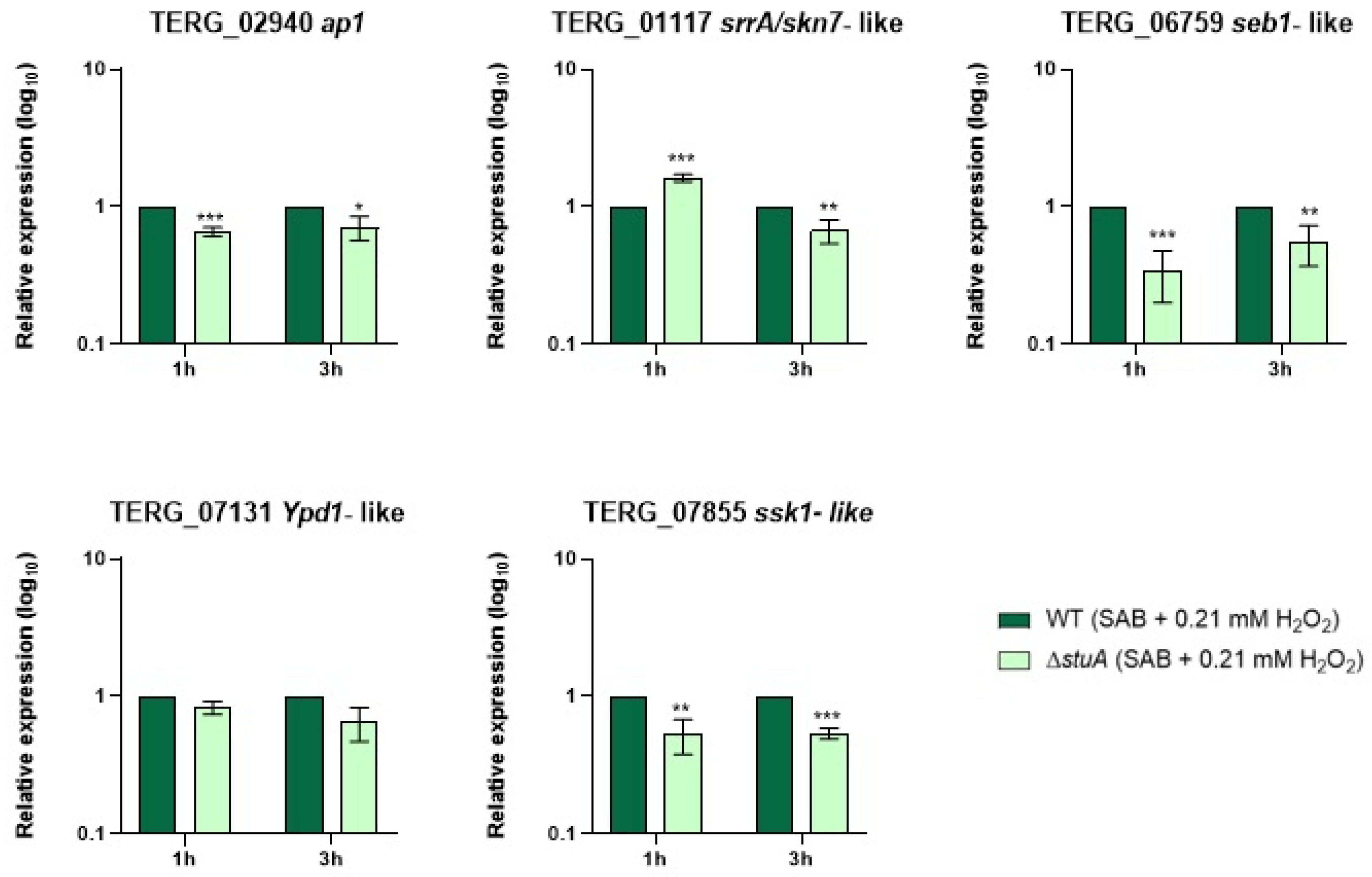
2.2. The Transcription Factor StuA Regulates Genes Related to Oxidative Stress Pathways in T. rubrum
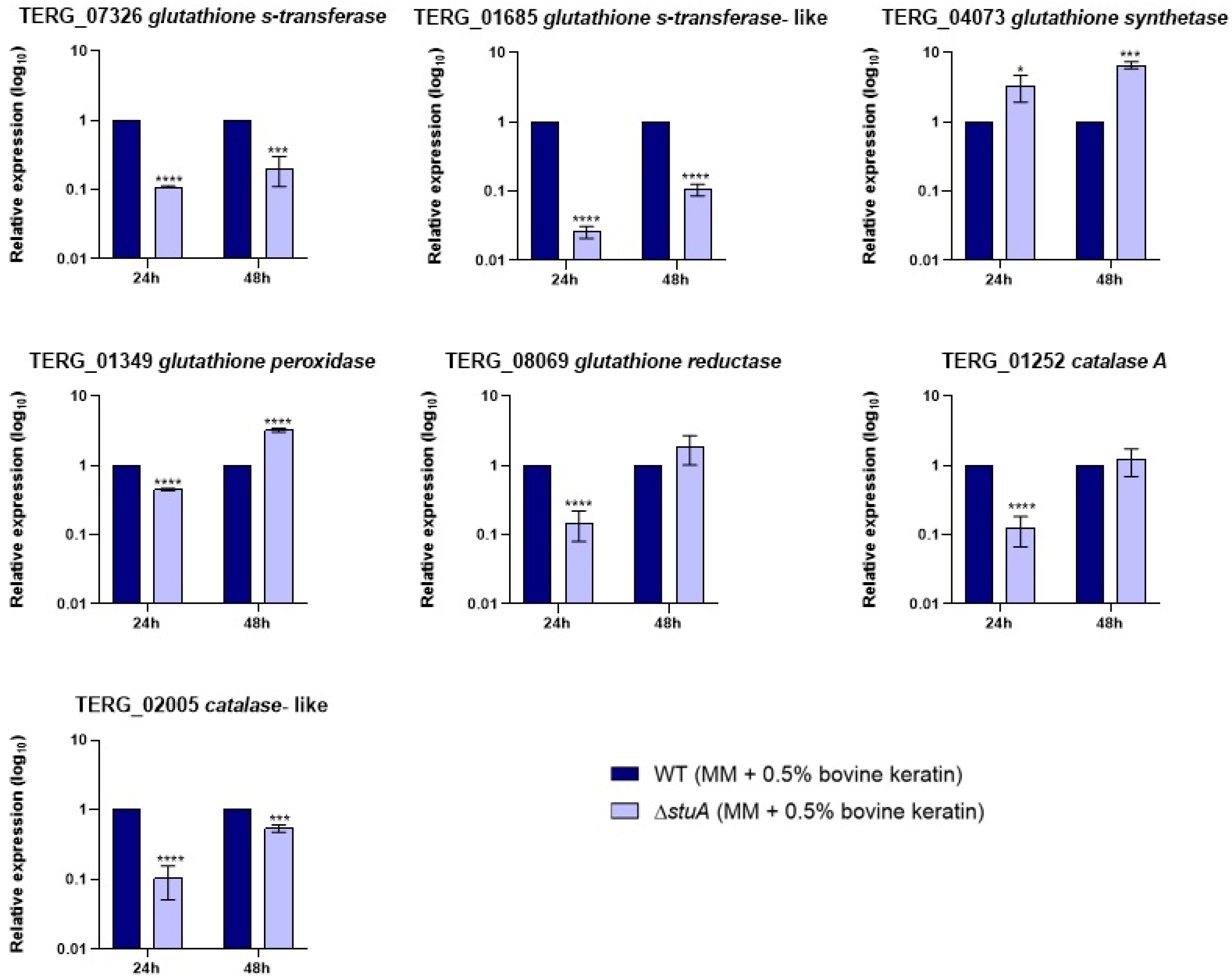
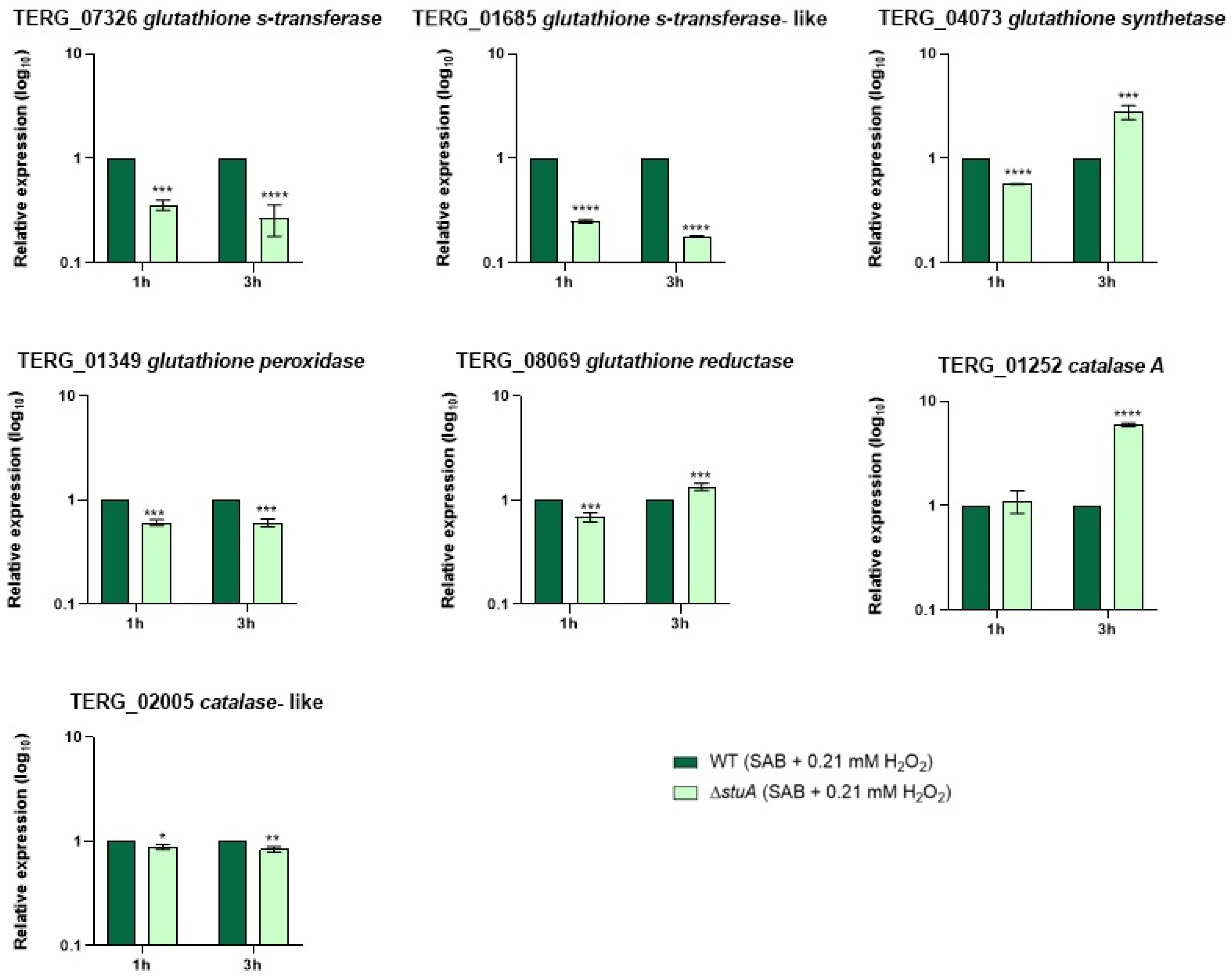
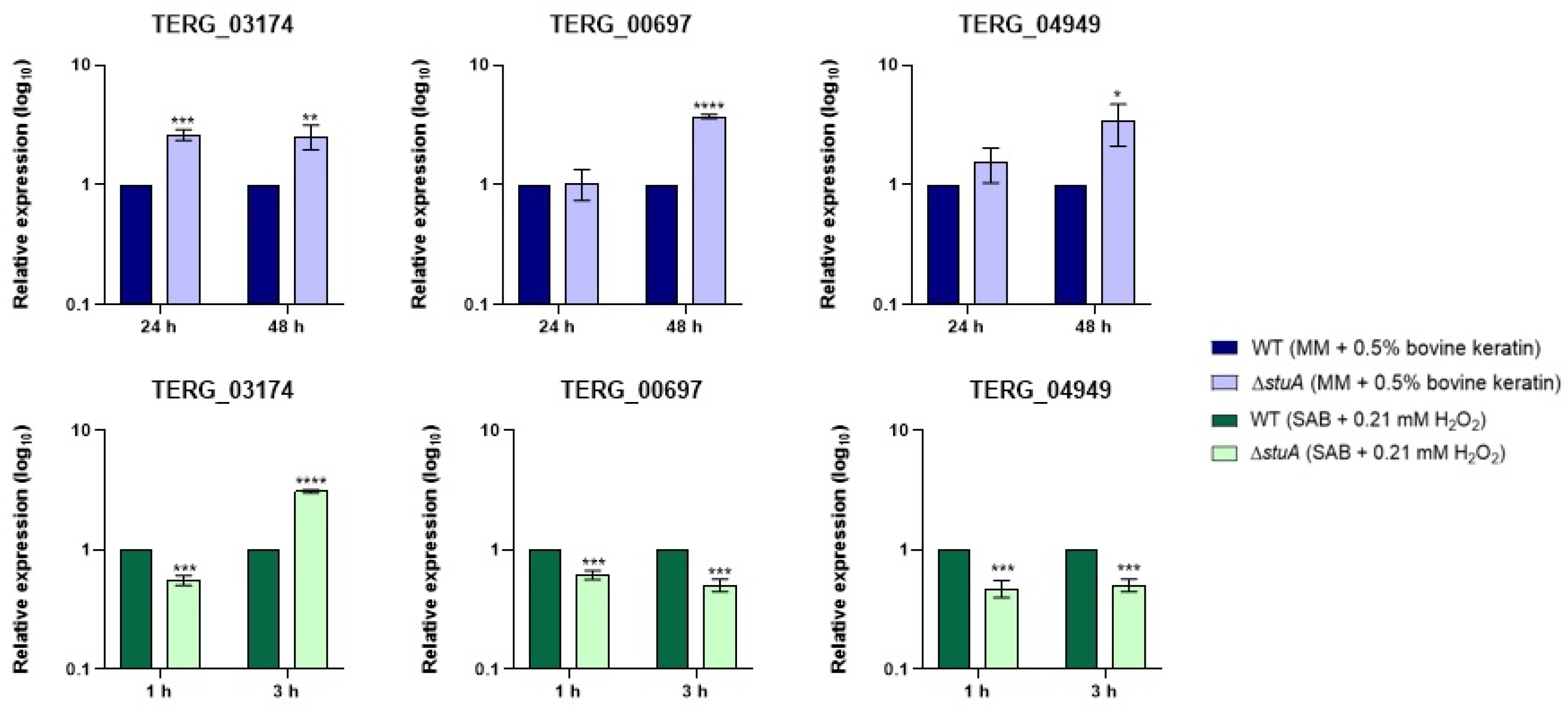
2.3. Transcript Levels of Glutathione S-Transferase and a Catalase-like Gene Are Reduced in the Mutant Strain
2.4. The Transcription Factor StuA as a Consensus-Binding Site for a Gene Encoding a Glutathione S-Transferase in T. rubrum
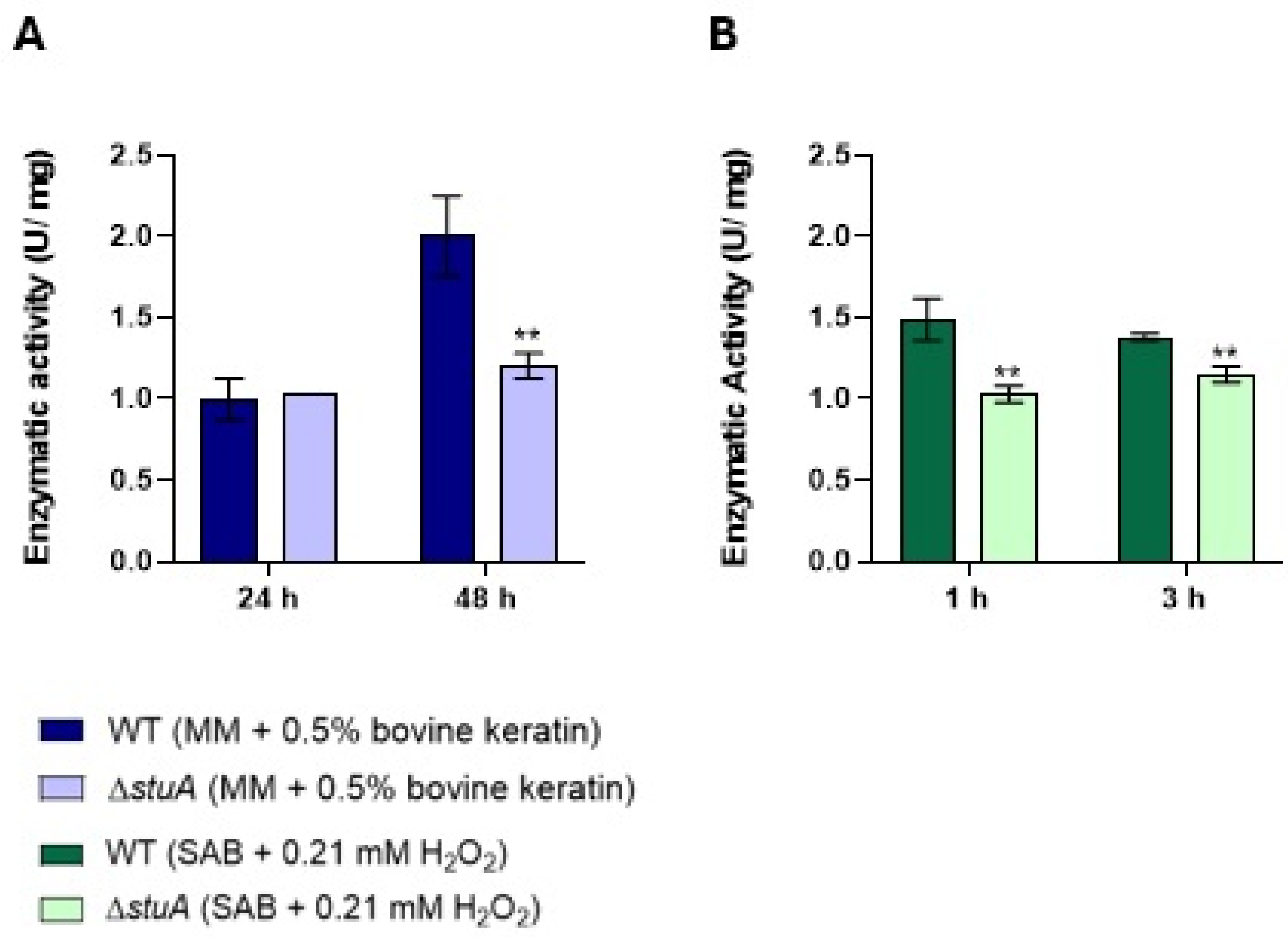
2.5. The Mutant ΔstuA Exhibited Lower Catalase Activity
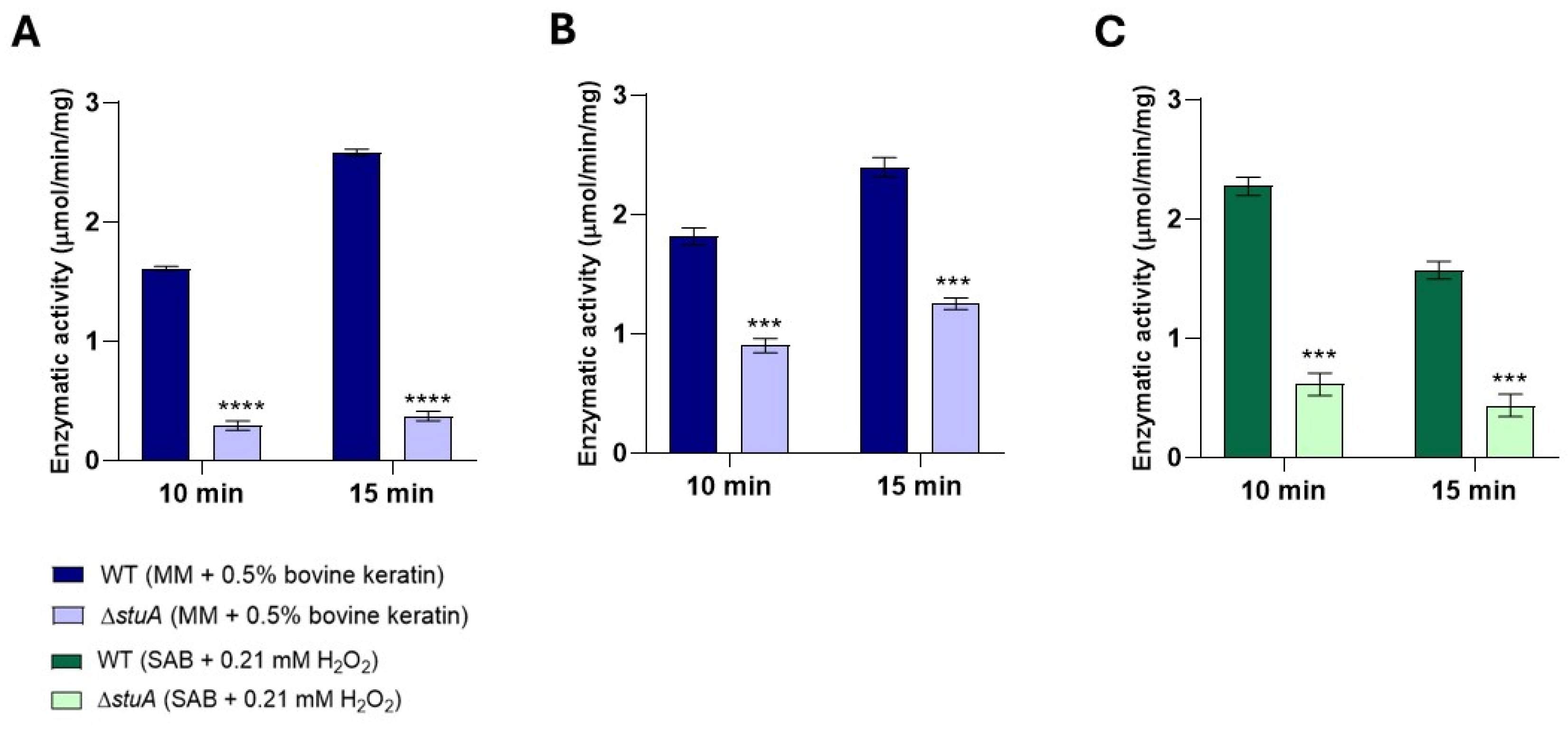
2.6. The ΔstuA Mutant Exhibited Reduced Glutathione S-Transferase Activity During Cultivation in Keratin and Under Hydrogen Peroxide Exposure
2.7. Genes Involved in Iron Metabolism Are Downregulated When the ΔstuA Strain is Exposed to Hydrogen Peroxide
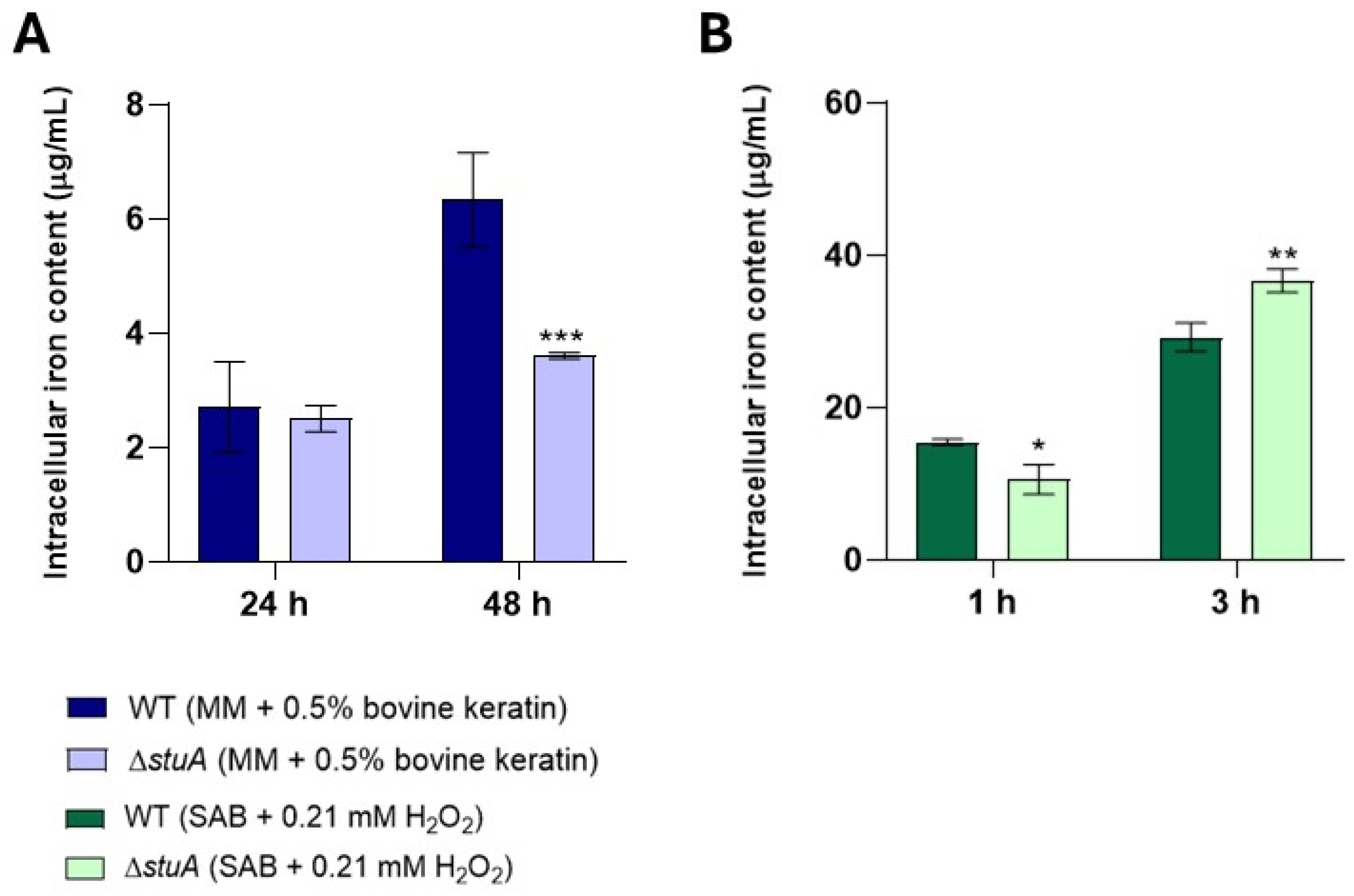
2.8. The Intracellular Iron Content Is Higher in the ΔstuA Strain After Three Hours of Exposure to Hydrogen Peroxide
3. Discussion
4. Materials and Methods
4.1. Search for Oxidative Stress-Related Gene Homologs in T. rubrum CBS118892
4.2. Fungal Strains and Culture Conditions
4.3. Oxidative Stress Induction with Hydrogen Peroxide
4.4. RNA Extraction and cDNA Synthesis
4.5. Reverse Transcription-Quantitative Polymerase Chain Reaction Analyses
4.6. Metabolic Assays
4.6.1. Catalase Activity Assay
4.6.2. Intracellular Iron Assay
4.6.3. GST Enzymatic Activity Assay
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Petrucelli, M.F.; Abreu, M.H.; Cantelli, B.A.M.; Segura, G.G.; Nishimura, F.G.; Bitencourt, T.A.; Fachin, A. Epidemiology and Diagnostic Perspectives of Dermatophytoses. Journal of Fungi 2020, 6, 310. [Google Scholar] [CrossRef] [PubMed]
- Ciesielska, A.; Kawa, A.; Kanarek, K.; Sobon, A.; Szewczyk, R. Metabolomic Analysis of Trichophyton rubrum and Microsporum canis During Keratin Degradation. Scientific Reports 2021, 11, 3959. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Rossi, N.M.; Peres, N.T.A.; Bitencourt, T.A.; Martins, M.P.; Rossi, A. State-of-the-Art Dermatophyte Infections: Epidemiology Aspects, Pathophysiology, and Resistance Mechanisms. Journal of Fungi (Basel) 2021, 7, 629. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Rossi, N.M.; Bitencourt, T.A.; Peres, N.T.A.; Lang, E.A.S.; Gomes, E.V.; Quaresemin, N.R.; Martins, M.P.; Lopes, L.; Rossi, A. Dermatophyte Resistance to Antifungal Drugs: Mechanisms and Prospectus. Frontiers in Microbiology 2018, 9, 1108. [Google Scholar] [CrossRef]
- Yaakoub, H.; Mina, S.; Calenda, A.; Bouchara, J.-P.; Papon, N. Oxidative Stress Response Pathways in Fungi. Cellular and Molecular Life Sciences 2022, 79, 333. [Google Scholar] [CrossRef]
- Zadrąg-Tęcza, R.; Maślanka, R.; Bednarska, S.; Kwolek-Mirek, M. Response Mechanisms to Oxidative Stress in Yeast and Filamentous Fungi, Stress Response Mechanisms in Fungi: Theoretical and Practical Aspects; Springer International Publishing: Cham, 2018; pp. 1–34. [Google Scholar]
- Berndt, C.; Lillig, C.H. Glutathione, Glutaredoxins, and Iron. Antioxidants & Redox Signaling 2017, 27, 1235–1251. [Google Scholar]
- Wangsanut, T.; Pongpom, M. The Role of the Glutathione System in Stress Adaptation, Morphogenesis and Virulence of Pathogenic Fungi. International Journal of Molecular Science 2022, 23, 10645. [Google Scholar] [CrossRef]
- Yaakoub, H.; Sanchez, N.S.; Ongay-Larios, L.; Courdavault, V.; Calenda, A.; Bouchara, J.; Coria, R.; Papon, N. The High Osmolarity Glycerol (HOG) Pathway in Fungi. Critical Reviews in Microbiology 2022, 48, 657–695. [Google Scholar] [CrossRef]
- Weidemller, P.; Kholmatov, M.; Petsalaki, E.; Zaugg, J.B. Transcription Factors: Bridge Between Cell Signaling and Gene Regulation. Proteomics 2021, 21, 2000034. [Google Scholar] [CrossRef]
- Shelest, E. Transcription Factors in Fungi. FEMS microbiology letters 2008, 286, 145–151. [Google Scholar] [CrossRef]
- Mulford, K.E.; Fassler, J.S. Association of the Skn7 and Yap1 Transcription Factors in the Saccharomyces cerevisiae Oxidative Stress Response. Eukaryotic Cell 2011, 10, 761–769. [Google Scholar] [CrossRef] [PubMed]
- Bitencourt, T.A.; Neves-Da-Rocha, J.; Martins, M.P.; Sanches, P.R.; Lang, E.A.S.; Bortolossi, J.C.; Rossi, A.; Martinez-Rossi, N.M. StuA-Regulated Processes in the Dermatophyte Trichophyton rubrum: Transcription Profile, Cell-Cell Adhesion, and Immunomodulation. Frontiers in Cellular and Infection Microbiology 2021, 11. [Google Scholar] [CrossRef] [PubMed]
- Sheppard, D. C.; Doedt, T.; Chiang, L.Y.; Kim, H.S.; Chen, D.; Nierman, W.C.; Filler, S.G. The Aspergillus fumigatus StuA Protein Governs the Up-Regulation of a Discrete Transcriptional Program During the Acquisition of Developmental Competence. Molecular biology of the cell 2005, 16, 5866–5879. [Google Scholar] [CrossRef] [PubMed]
- Elza, A.S.; Bitencourt, T.A.; Peres, N.T.A.; Lopes, L.; Silva, L.G.; Cazzaniga, R.A.; Rossi, A.; Martinez-Rossi, N.M. The stuA Gene Controls Development, Adaptation, Stress Tolerance, and Virulence of the dermatophyte Trichophyton rubrum. Microbiological Research 2020, 241, 126592. [Google Scholar]
- Martins-Santana, L.; Petrucelli, M.F.; Sanches, P.R.; Almeida, F.; Martinez-Rossi, N.M.; Rossi, A. The StuA Transcription Factor and Alternative Splicing Mechanisms Drive the Levels of MAPK Hog1 Transcripts in the Dermatophyte Trichophyton rubrum. Mycopathologia 2024, 189, 37. [Google Scholar] [CrossRef]
- Petrucelli, M.F.; Martins-Santana, L.; Sanches, P.R.; Oliveira, V.M.; Rossi, A.; Martinez-Rossi, N.M. The Transcription Factor StuA Regulates the Glyoxylate Cycle in the Dermatophyte Trichophyton rubrum under Carbon Starvation. International Journal of Molecular Sciences 2024, 25, 405. [Google Scholar] [CrossRef]
- Liao, B.; Ye, X.; Chen, X.; Zhou, Y.; Cheng, L.; Zhou, X.; Ren, B. The Two-Component Signal Transduction System and its Regulation in Candida albicans. Virulence 2021, 12, 1884–1899. [Google Scholar]
- Mendoza-Martínez, A.E.; Cano-Domínguez, N.; Aguirre, J. Yap1 Homologs Mediate More than the Redox Regulation of the Antioxidant Response in Filamentous Fungi. Fungal Biology 2020, 124, 253–262. [Google Scholar] [CrossRef]
- Peres, N.T.A.; Lang, E.A.; Bitencourt, T.A.; Oliveira, V.M.; Fachin, A.L.; Rossi, A.; Martinez-Rossi, N.M. The bZIP Ap1 Transcription Factor is a Negative Regulator of Virulence Attributes of the Anthropophilic Dermatophyte Trichophyton rubrum. Current Research in Microbial Sciences 2022, 3, 100132. [Google Scholar] [CrossRef]
- Freitas, F.Z.; Virgilio, S.; Cupertino, F.B.; Kowbel, D.J.; Fioramonte, M.; Gozzo, F.C.; Glass, L.; Bertoloni, M.C. The SEB-1 Transcription Factor Binds to the STRE Motif in Neurospora crassa and Regulates a Variety of Cellular Processes Including the Stress Response and Reserve Carbohydrate Metabolism. G3 (Genes, Genomes, Genetics 2016, 6, 1327–1343. [Google Scholar] [CrossRef]
- Wu, Y.; Xu, L.; Yin, Z.; Feng, H.; Huang, L. Transcription Factor VmSeb1 is Rrequired for the Growth, Development, and Virulence in Valsa mali. Microbial Pathogenesis 2018, 123, 132–138. [Google Scholar] [CrossRef] [PubMed]
- Scherer, M.; Wei, H.; Liese, R.; Fischer, R. Aspergillus nidulans Catalase-Peroxidase Gene (cpeA) is Transcriptionally Induced During Sexual Development Through the Transcription Factor StuA. Eukaryotic Cell 2002, 1, 725–735. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.R.; Reindl, K.M. Glutathione S-Transferases in Cancer. Antioxidants 2021, 10, 701. [Google Scholar] [CrossRef] [PubMed]
- Fraser, J.A.; Davis, M.A.; Hynes, M.J. A Gene from Aspergillus nidulans With Similarity to URE2 of Saccharomyces cerevisiae Encodes a Glutathione S-transferase Which Contributes to Heavy Metal and Xenobiotic Resistance. Applied and Environmental Microbiology 2002, 68, 2802–2808. [Google Scholar] [CrossRef]
- Lok, H.C.; Richardson, V.; Kalinowski, D. S.; Kovacevic, Z.; Lane, D.J.R.; Richardson, D.R. Glutathione S-transferase and MRP1 Form an Integrated System Involved in the Storage and Transport of Dinitrosyl–dithiolato Iron Complexes in Cells. Free Radical Biology and Medicine 2014, 75, 14–29. [Google Scholar] [CrossRef]
- Gao, X.; Hu, W.; Qian, D.; Bai, X.; He, H.; Li, L.; Sun, S. The Mechanisms of Ferroptosis Under Hypoxia. Cellular and Molecular Neurobiology 2023, 43, 3329–3341. [Google Scholar] [CrossRef]
- Stelitano, G.; Cocorullo, M.; Mori, M.; Villa, S.; Meneghetti, F.; Chiarelli, L.R. Iron Acquisition and Metabolism as a Promising Target for Antimicrobials (Bottlenecks and Opportunities): Where Do We Stand? International Journal of Molecular Sciences 2023, 24, 6181. [Google Scholar] [CrossRef]
- Kröber, A.; Scherlach, K.; Hortschansky, P.; Shelest, E.; Staib, P.; Kniemeyer, O.; Brakhage, A.A. HapX Mediates Iron Homeostasis in the Pathogenic Dermatophyte Arthroderma benhamiae but Is Dispensable for Virulence. Plos One 2016, 11, e0150701. [Google Scholar] [CrossRef]
- Pradhan, A.; Herrero-de-Dios, C.; Belmonte, R.; Budge, S.; Garcia, A.L.; Kolmogorova, A.; Lee, K.K.; Martin, B.D.; Ribeiro, A.; Bebes, A.; Yuecel, R.; Gow, N.A.R.; Munro, C.A.; MacCallum, D.M.; Quinn, J.; Brown, A.J. Elevated Catalase Expression in a Fungal Pathogen is a Double-Edged Sword of Iron. 2017, 13, e1006405. [Google Scholar] [CrossRef]
- Cove, D.J. The Induction and Repression of Nitrate Reductase in the Fungus Aspergillus nidulans. Biochimica et Biophysica Acta (BBA) - Enzymology and Biological Oxidation 1966, 113, 51-56.
- Jacobson, L.S.; Mclntyre, L.; Mykusz, J. Assessment of Real-Time PCR Cycle Threshold Values in Microsporum canis Culture-Positive and Culture-Negative Cats in an Animal Shelter: a Field Study. Journal of feline medicine and surgery 2018, 20, 108–113. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2- ΔΔCT method. methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Jacob, T.R.; Peres, N.T.A.; Persinoti, G.F.; Silva, L.G.; Mazucato, M.; Rossi, A.; Martinez-Rossi, N.M. rpb2 is a Reliable Reference Gene for Quantitative Gene Expression Analysis in the Dermatophyte Trichophyton rubrum. Medical Mycology 2012, 50, 368–377. [Google Scholar] [CrossRef] [PubMed]
- Swift, M.L. GraphPad Prism, Data Analysis, and Scientific Graphing. Journal of chemical information and computer sciences 1997, 37, 411–412. [Google Scholar] [CrossRef]

| Gene ID | Gene Product Name | Homologous |
|---|---|---|
| TERG_01117 | Stress response transcription factor SrrA/Skn7 | ARB_07479 (Trichophyton benhamiae) |
| TERG_06759 | C2H2 transcription factor (Seb1) | ARB_00366 (Trichophyton benhamiae) |
| TERG_07131 | Phosphotransmitter protein Ypd1 | TEQG_03843 (Trichophyton equinum) |
| TERG_07855 | Response regulator ssk1 | TRV_02413 (Trichophyton verrucosum) |
| TERG_01685 | Glutathione S-transferase | ARB_02229 (Trichophyton benhamiae) |
| TERG_02005 | Catalase | TESG_07337 (Trichophyton tonsurans) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




