The study created an approach to diagnose different stages of AMD by analyzing macular images. It identified four main stages: no disease, early, intermediate, and late. Additionally, two late-stage development scenarios were identified: GA and SF. A series of OCT images was then generated and pre-processed. A basic classifier model was then trained and tested on these images to identify various challenges that affect the accurate classification of AMD stages and development options.
2.1. Dataset Structure
To develop the algorithm, we utilized a diverse set of 1928 OCT images of the macular region of patients with AMD. These images covered broad spectrum of AMD stages obtained during optical coherence tomography of the retina on Avanti XR (Optovue; USA) and REVO NX (Optopol; Poland) devices at the Optimed Laser Vision Restoration Center (Ufa, Russia).
The list of classes under consideration included the following cases with the corresponding code designations:
The imbalance of classes resulting from the disproportionate sizes of classes SI and VI during direct classification necessitated additional solutions outlined below.
To enhance the algorithm’s accuracy, we have added a new label. This label is specifically used to classify conclusions about the nearest extreme states of stage P, such as S and V. However, these labels are not assigned to the entire class P but only to a carefully chosen group of highly representative examples that illustrate the proximity of P to stages S and V.
It was also decided to enhance the algorithm by enabling it to identify the progression of stage P based on its proximity to stages S and V. By labeling only a portion of the class; users can utilize their expert judgment to personalize the analysis of the intermediate class, reducing potential disagreements among experts. To determine the minimum threshold for the number of examples of class P, we considered several options for the proportion of labeled and unlabeled data: 1/6, 1/4, and 1/3. In the section on algorithm implementation, the choice of the minimum proportion of labeled data was analyzed in terms of its accuracy in identifying the progression of intermediate class AMD on test samples.
The preprocessing of the grayscale OCT images in the dataset involved several essential steps:
Normalizing pixel brightness levels to remove any color distortions;
Generating new image samples by randomly flipping them horizontally, ensuring that all possible C-scan image positions were accounted for;
Resizing images to a standard 64 by 128-pixel format to ensure consistent visualization of images obtained from different tomographs, given the prominent horizontal orientation of the retina.
2.2. Developing a Classifier Structure and Identifying the Problem of Direct Classification of Medical Data
The modern approach to image analysis relies on processing images at the level of individual pixels. To analyze an image, the brightness of each pixel is determined, as well as its location in the entire image and other clusters of pixels [
50]. These features, obtained through analysis, form a vector representation of the image in a feature space. The more distinct the vectors representing one class are from those of others, the greater the classification effectiveness will be [
51].
Convolutional neural networks (CNNs) are effective for extracting features from images. These networks can separate image vectors in feature space when there are enough representative examples for each class in classification problems [
52,
53]. However, additional steps are required when the number of images is limited or there is a class imbalance to ensure the stability of deep computer vision algorithms [
54,
55]. To address these challenges, we selected a base classification model and then modernized it step by step, evaluating the effectiveness of each change.
A four-layer CNN-based encoder served as the base classification model for the analysis. To ensure consistent training conditions for all ML models throughout the study, the following training parameters were established:
Number of training epochs: 80;
Optimizer: Adam algorithm;
Error function: cross entropy.
We trained and tested the base encoder model using the transformed dataset to classify six classes, which include the main stages and two late-stage scenarios. To assess the performance of our classifiers, we utilized the following set of metrics:
Precision: measures the proportion of correct positive predictions among all positive predictions made by the model, including false positives. High precision indicates a significant probability that the answer is correct in the case of positive predictions for a given class [
56];
Sensitivity: is the ratio of correctly identified positive cases to the total number of positive cases, including false negative cases. A high sensitivity value indicates that the model is more efficient at correctly identifying positive cases [
56];
Specificity: measures the ratio of correctly identified negative cases to the total number of negative cases. A high specificity indicates that the model accurately identifies negative cases [
56];
F1-Score: is the harmonic mean of precision and recall. It measures the model’s overall performance, considering both false positives and false negatives, especially when dealing with class imbalance [
56].
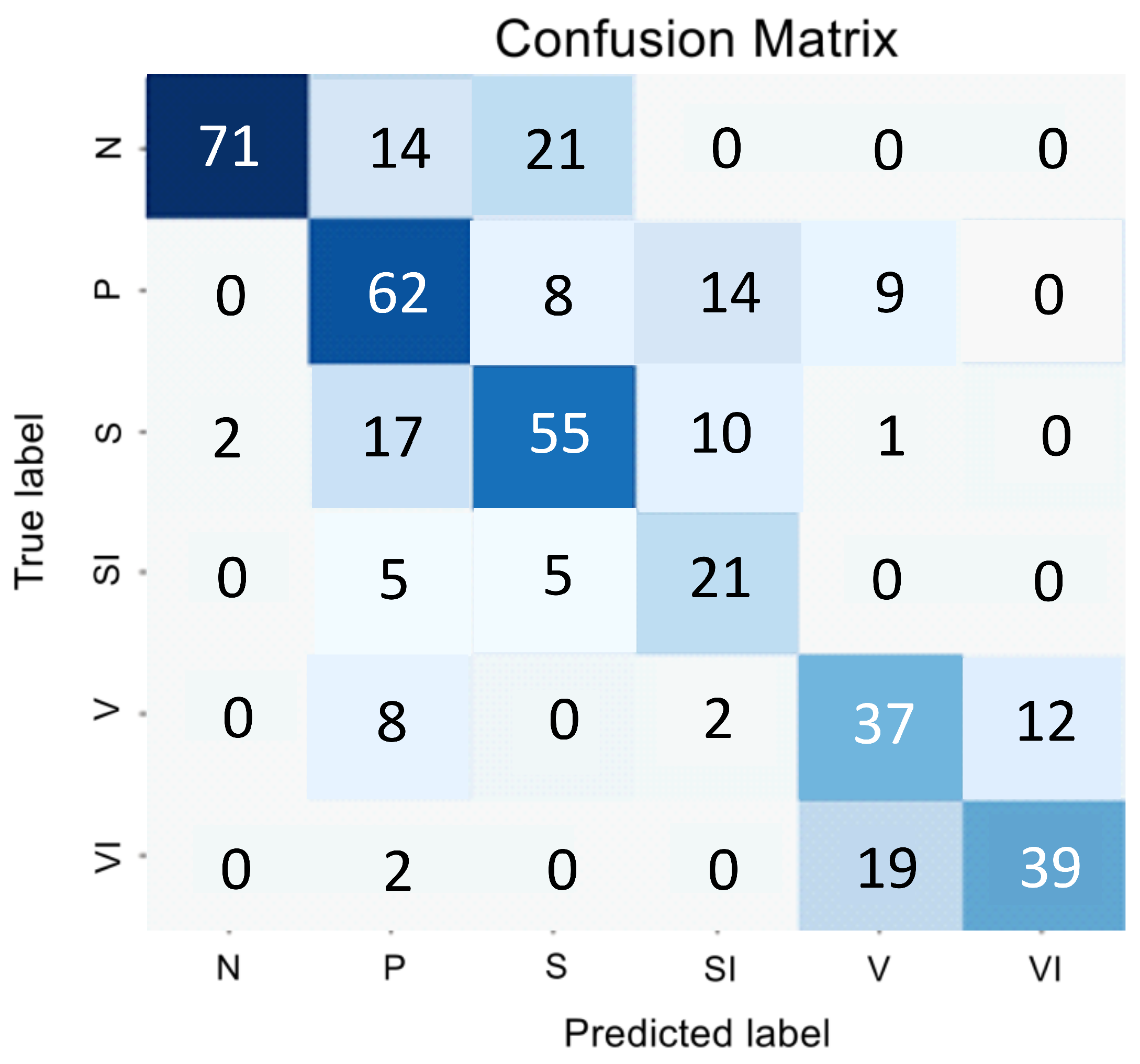
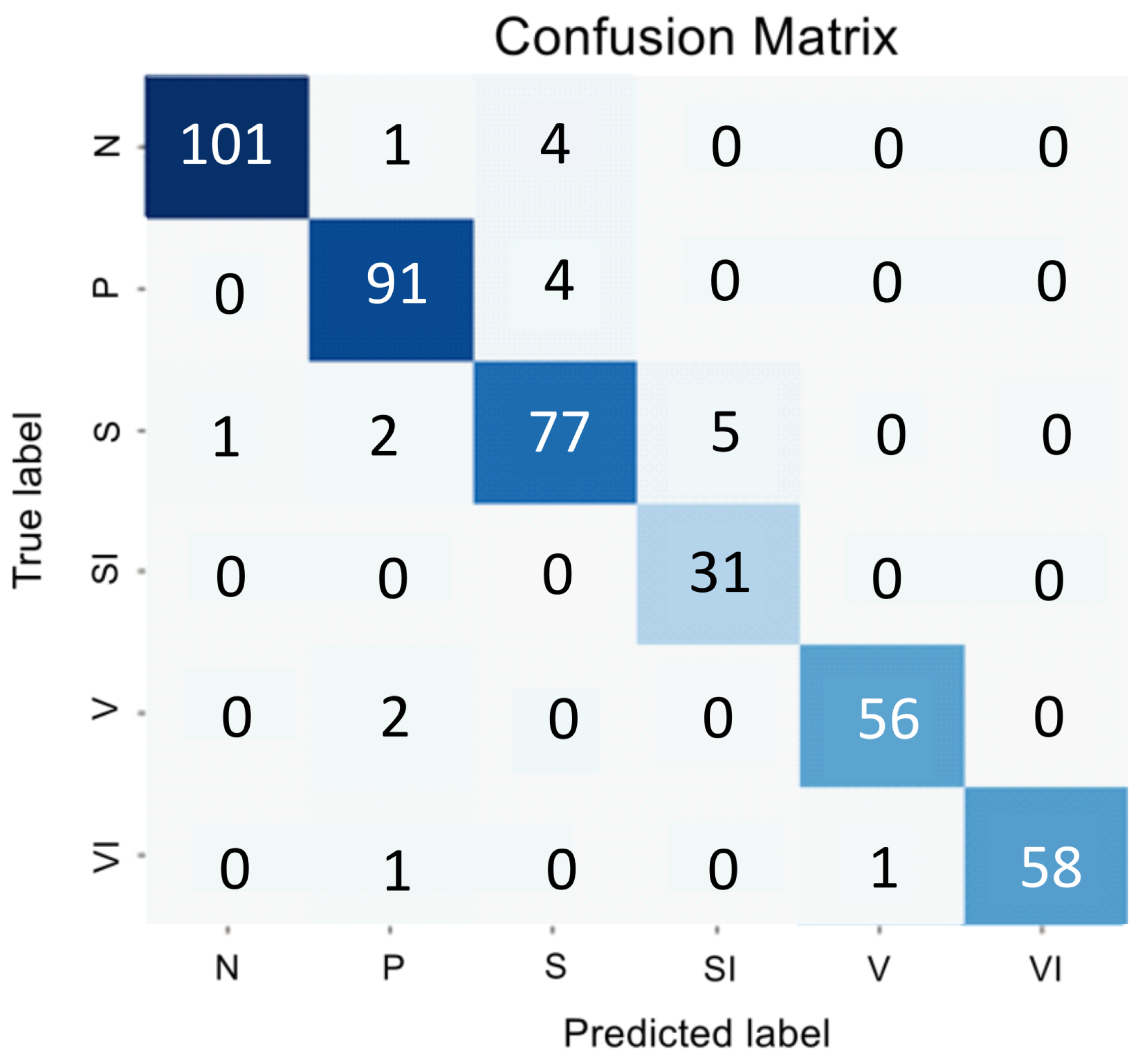
Figure 1 and
Table 1 presents the encoder testing results as an error matrix.
The analysis of the confusion matrix and metrics for the direct classifier showed that the model’s performance was unsatisfactory due to the difficulty in detecting the intermediate stage and the imbalanced nature of the SI and VI classes, which have similar features to the early and intermediate AMD stages. The model showed conservatism, with a high precision value but a low sensitivity value. It meant that while the model was unlikely to make errors in selecting the N and VI classes, it was highly likely to miss many actual instances of these classes.
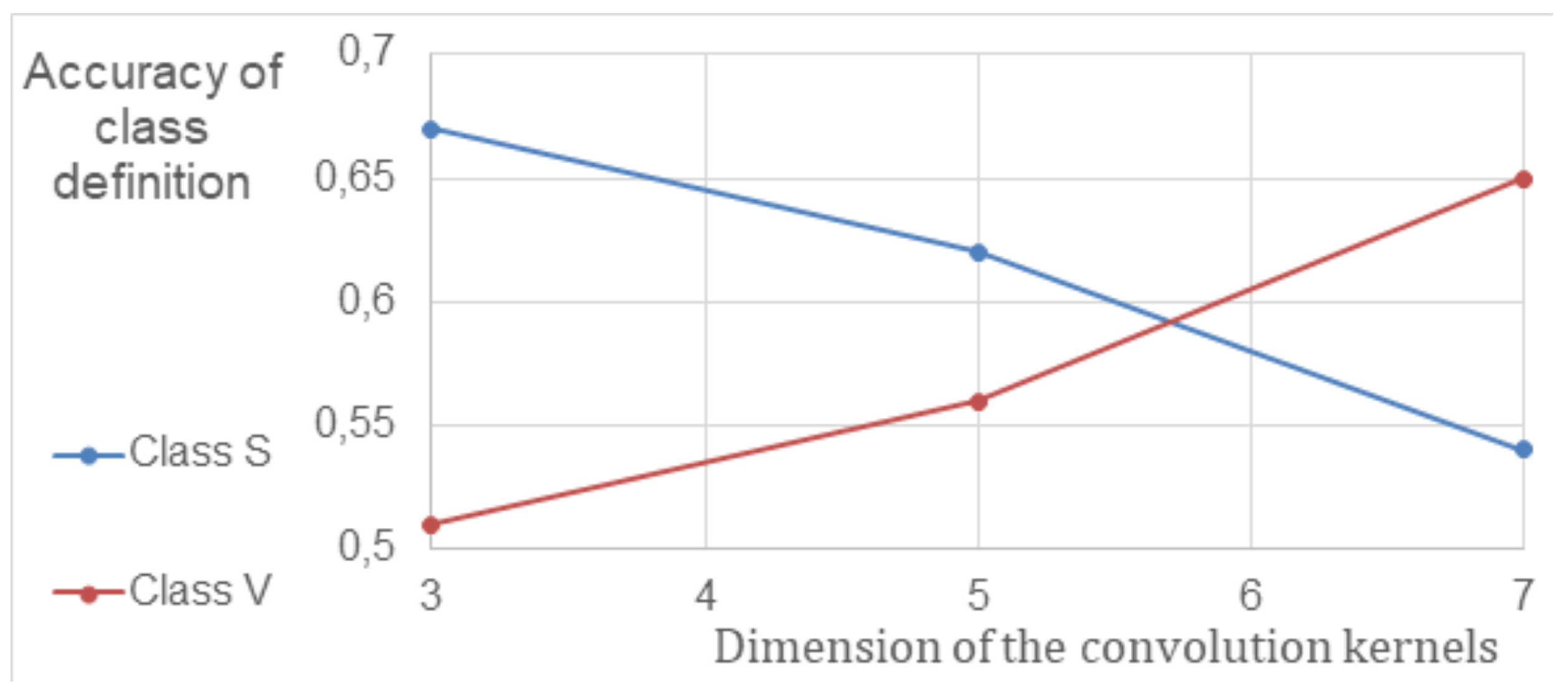
To improve the efficiency of CNN, the parameters of convolutional layers and their impact on classification accuracy were studied. Since biomarkers in different stages of AMD vary in size, the impact of convolution kernel size was investigated in terms of accuracy in separating S and V classes. A neural network with convolutional and fully connected layers was used for the study. The results, showing the relationship between the model’s sensitivity (true positive rate, TPR) and convolution kernel size, are presented in
Figure 2 after cross-validation. The data indicates that the sensitivity of CNNs with an average kernel size increased by at least 8% compared to the previously used 5 × 5 kernel size.
The results of the cross-validation process showed a connection between the levels of biomarkers linked to different stages of AMD and the model’s sensitivity when using different convolution parameters. These findings were used to create a framework that simultaneously processes information through multiple convolution layers, each with its unique convolution kernels.
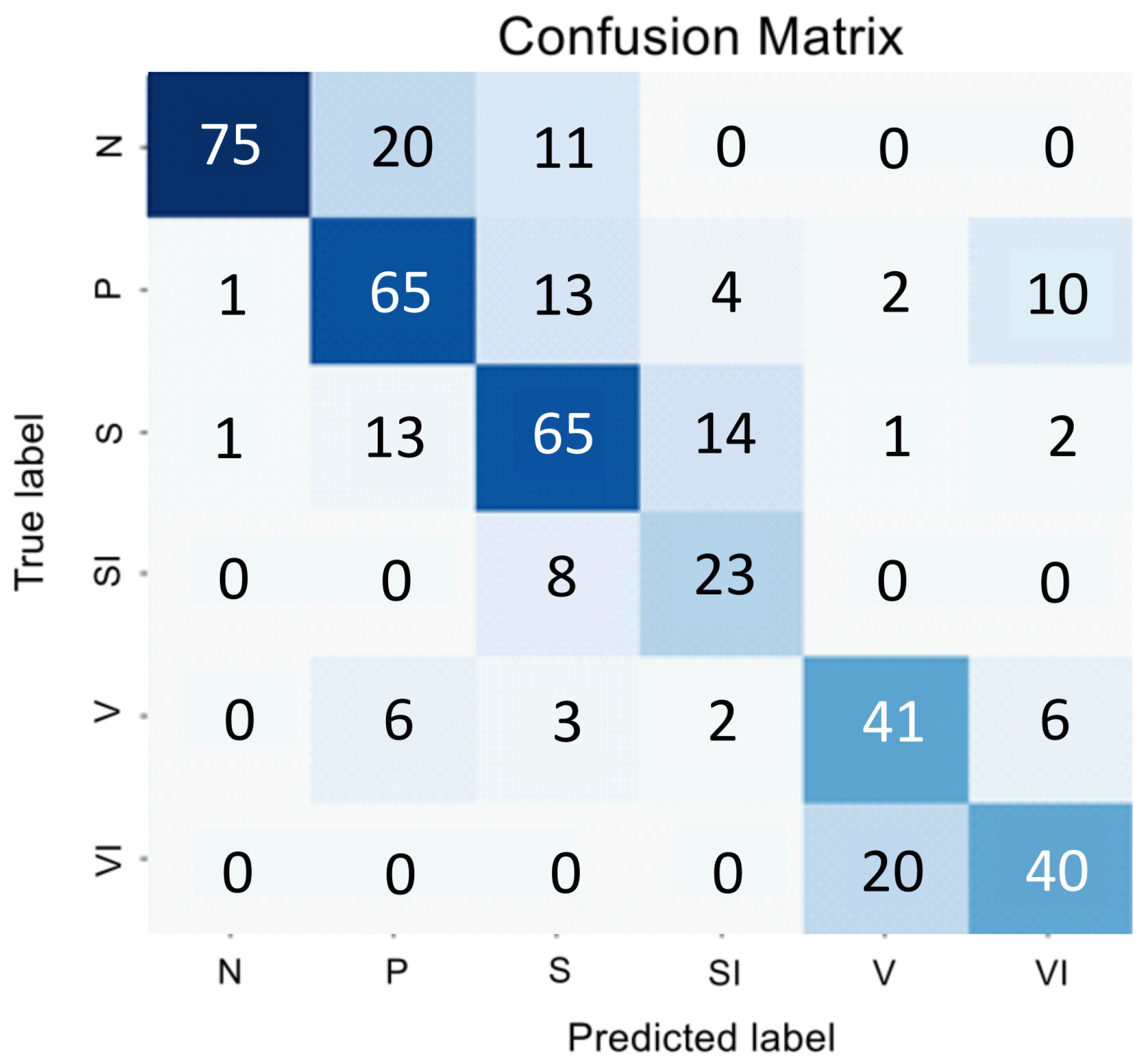
We explored how the encoder’s performance varies when employing parallel convolution layers at each stage. The results of this study are presented in a confusion matrix shown in
Figure 3 and summarized in
Table 2.
The changes made to the encoder architecture have greatly improved the classifier’s performance, resulting in a higher overall F1 Score. Despite these improvements, the challenge of addressing a significant class imbalance and accurately distinguishing class P from the background classes S and V persists.
In order to tackle this problem, we incorporated elements of a hierarchical approach into the structure of the encoder. The goal was to break down the complex task of categorizing six nonequilibrium classes into more manageable sub-tasks. The components of the classifier were then identified as follows:
The main task of the global classifier model is to classify the four main stages;
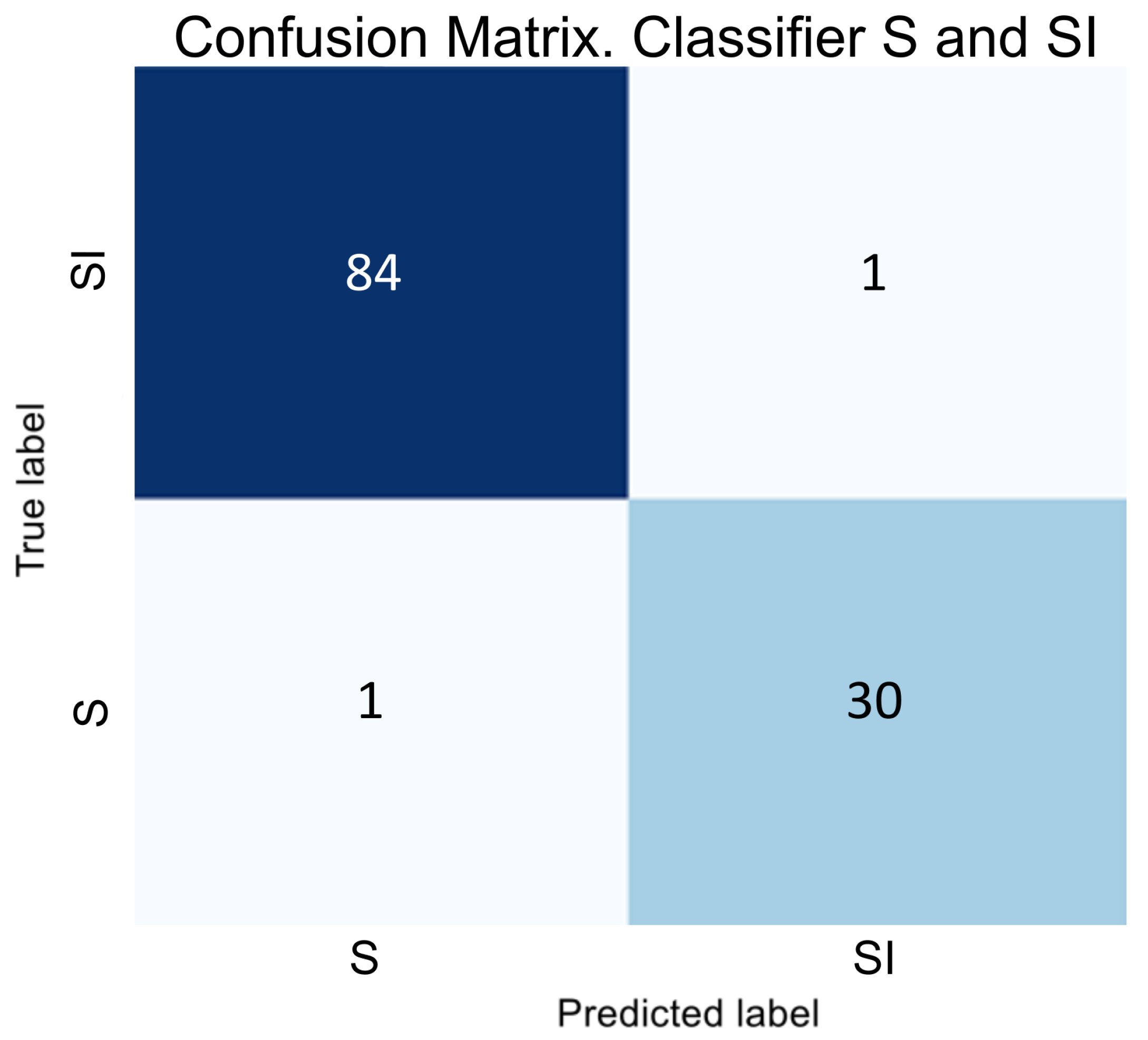
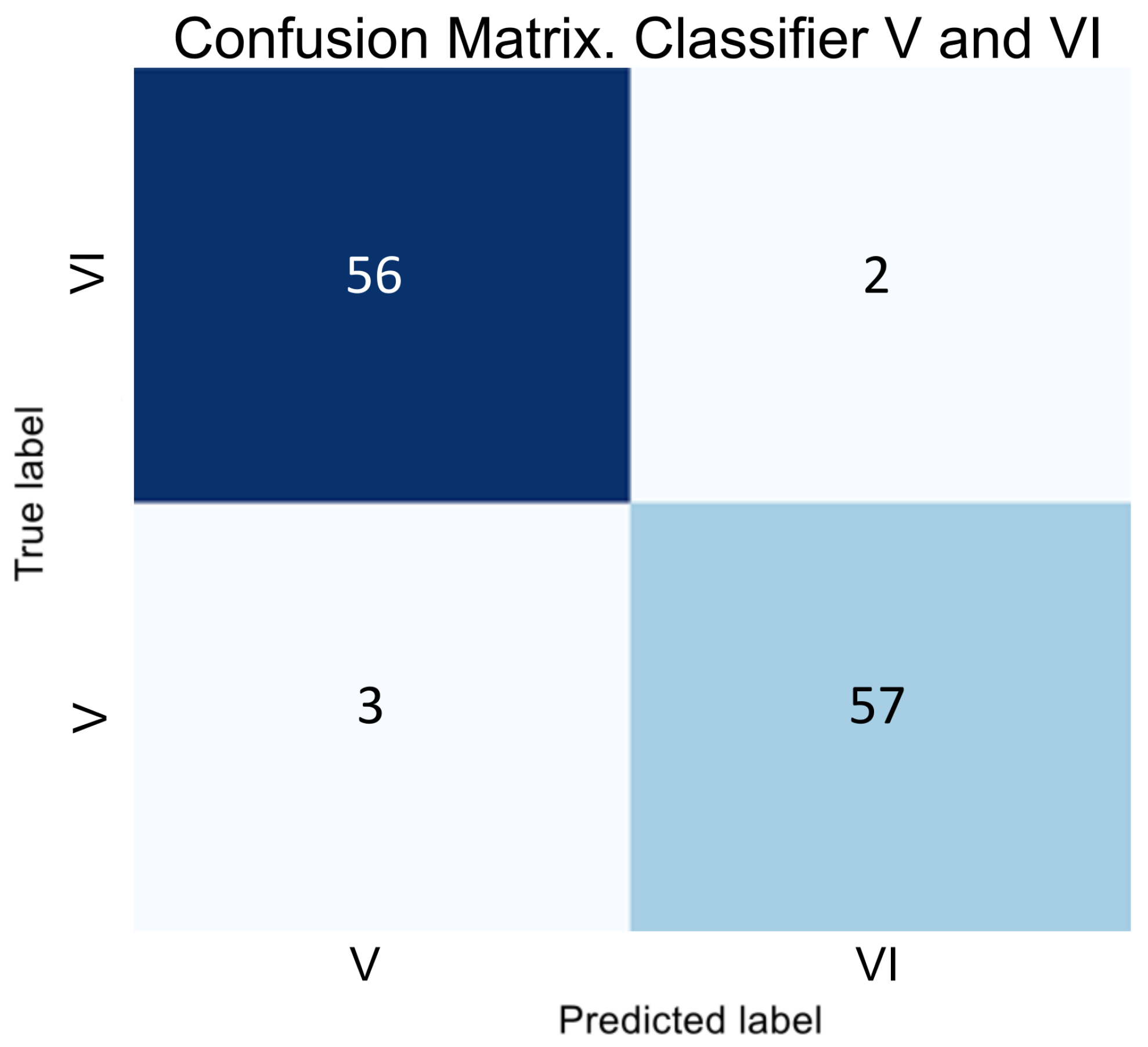
Two binary classifiers address the class imbalance issue by determining classes SI and VI;
We developed a regression model to assess the degree of proximity of class P to classes S and V.
When analyzing the data in
Figure 3, we can conclude that despite SI being a subclass of V, the visual correlation of GA and the early stage of AMD is much stronger than with the late stage. Therefore, we decided to classify SI as a subclass specifically to S, and the subsequent transformation stage is forced to correlate these samples with V. Thus, if a sample belongs to classes S or V, then two binary classifiers determine whether it is part of class S or SI, and V or VI.
It is important to note that the image features extracted by the global model using different convolutional layers can act as input to binary classifiers. It significantly reduces the computational complexity required for high accuracy, sensitivity, and specificity. This is achieved by passing information directly from the global model to these classifiers from intermediate layers. This approach was also utilized to create input data for the regression algorithm, which determines the degree of development of stage P.
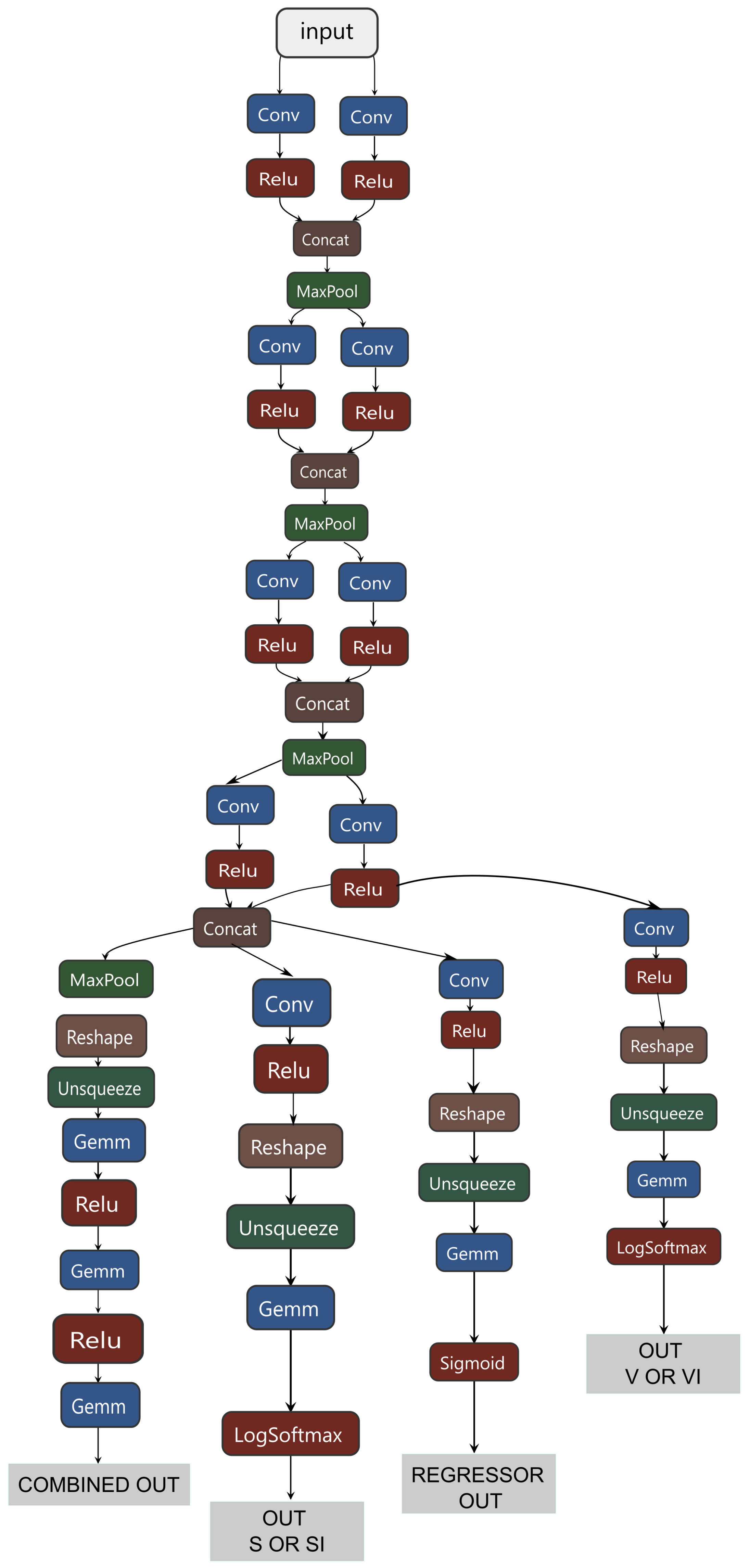
Thus, a hybrid classifier integrates several models, including a global classifier, two binary classifiers for processing SI and VI subclasses, and a regressor for processing the intermediate stage P. The general scheme of its structure is shown in
Appendix A.1. Common information data were allocated for the SI subclass and the regression of class P, effectively identifying small and large predictors. Only large predictors were allocated for the VI subclass, proving to be the most effective approach.
The hybrid classifier’s training process involved several stages. First, only the global classifier was trained. Then, the binary classifiers and the regressor were trained, and the global classifier processed the input data. To train the global classifier, classes S and V included examples from both their samples and from classes SI and VI, respectively. In the next stage, binary classifiers S and V determined classes SI and VI.
The algorithm for determining the severity of intermediate AMD included a regressor operating in tandem with the Label Propagation (LPA) algorithm [
57]. The training was conducted in three iterations using three different versions of the labeled dataset (1/6, 1/4, and 1/3). Accuracy was evaluated using a predetermined test dataset, which accounted for 10% of the total labeled data volume.