Submitted:
15 October 2024
Posted:
16 October 2024
You are already at the latest version
Abstract
Keywords:
I. Introduction
II. Related Work
III. Methods
A. System Architecture
B. Model Architecture
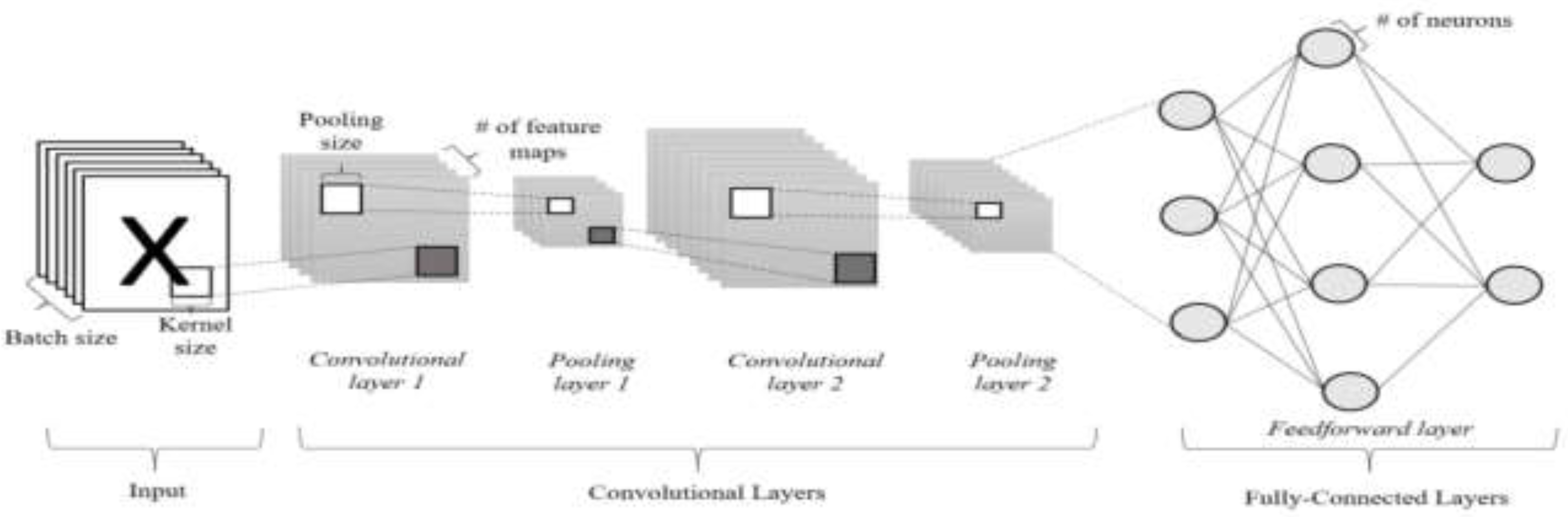
IV. Experiment
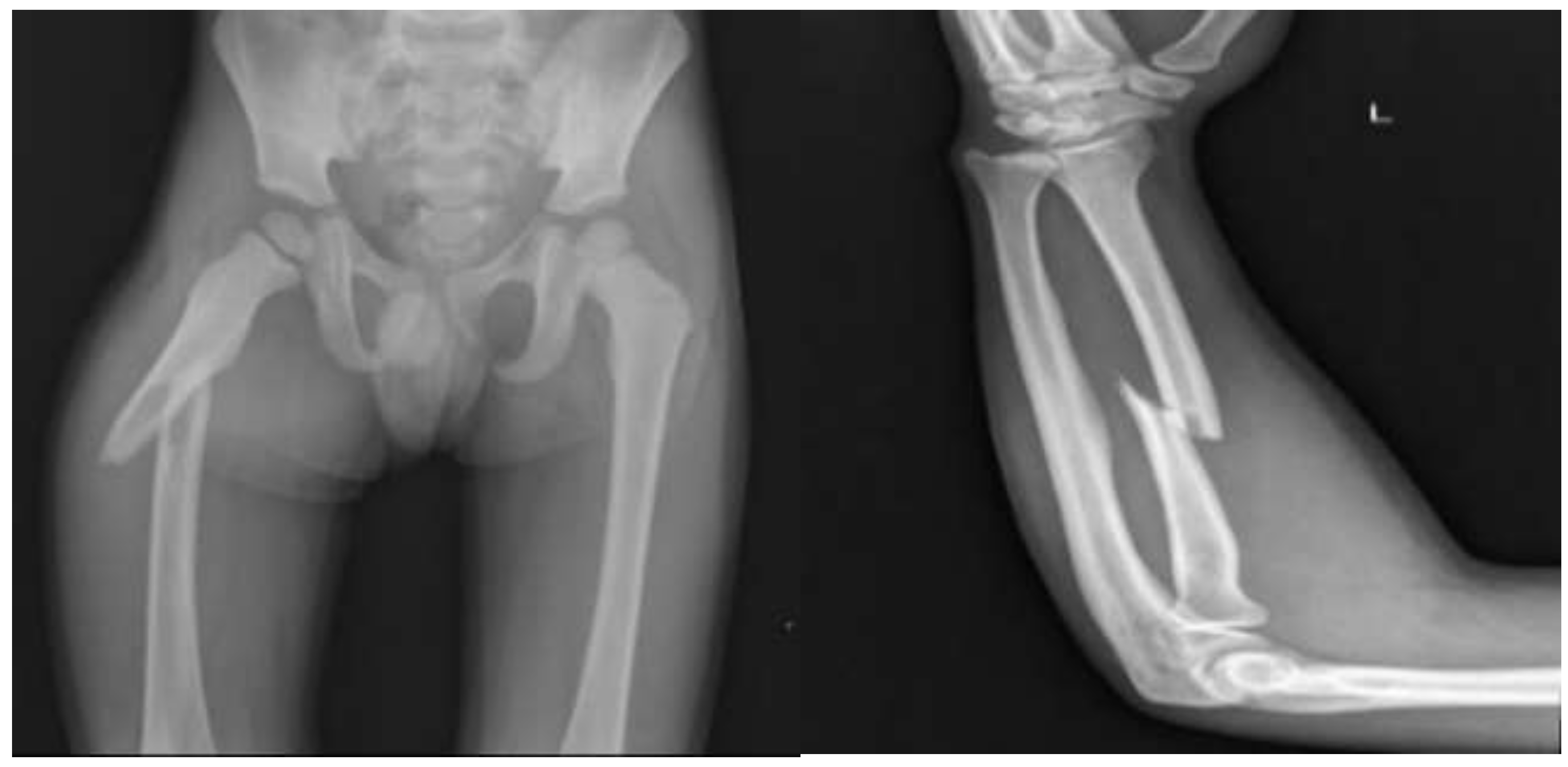
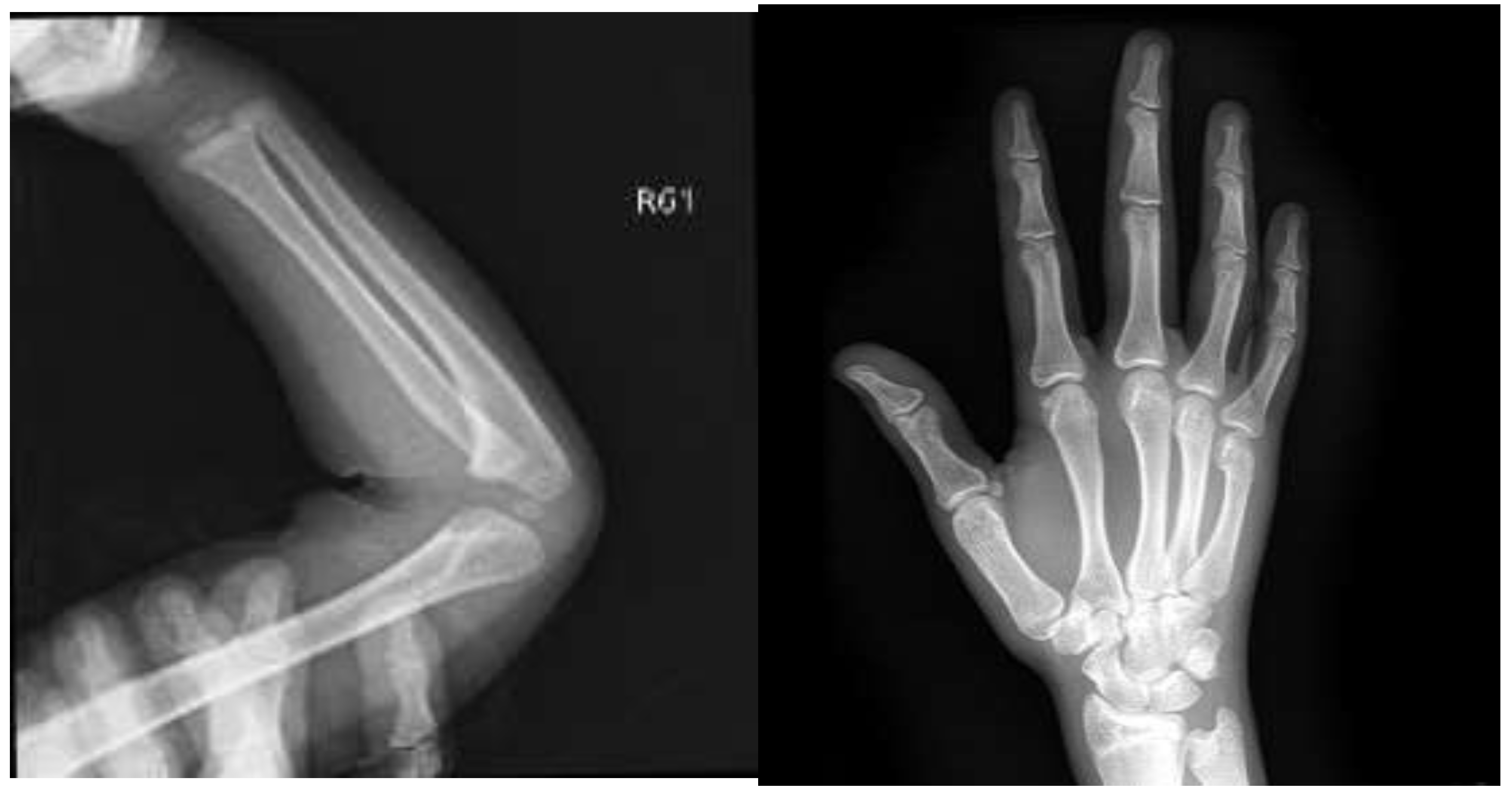
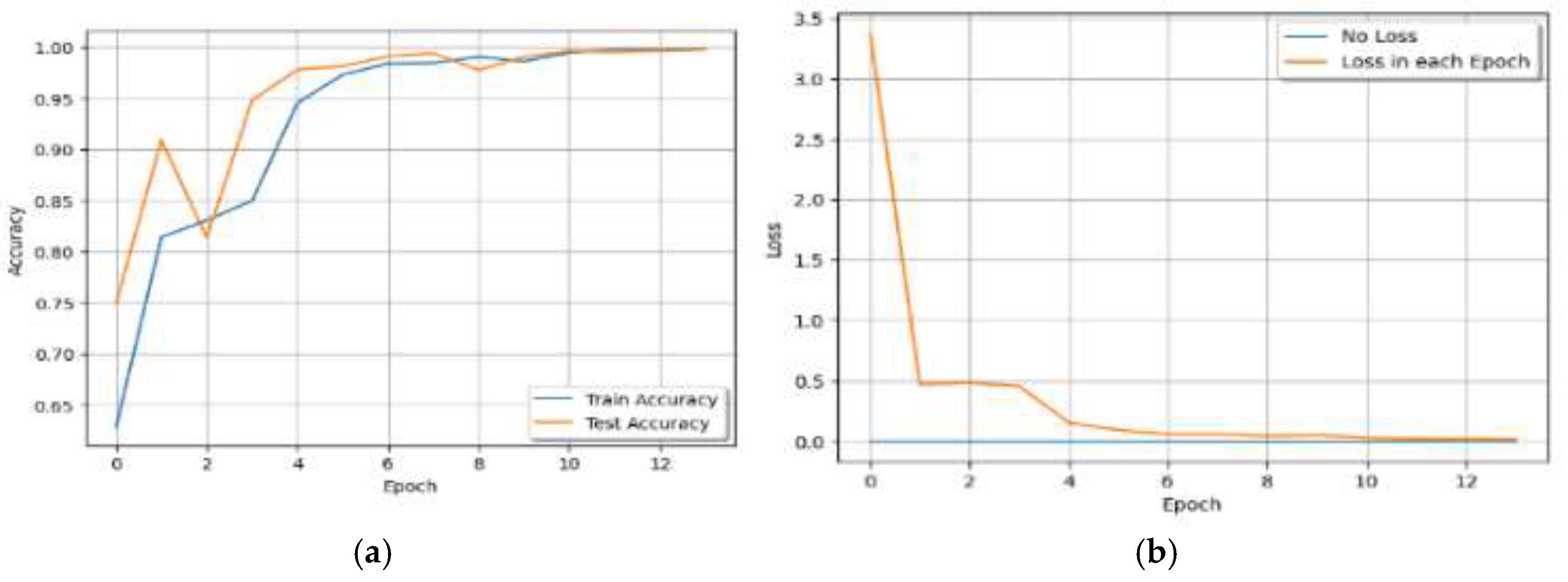
V. Results
| Epoch | Train Loss | Train Accuracy | Test Accuracy |
|---|---|---|---|
| 0 | 3.3721 | 0.62841292 | 0.74952284 |
| 1 | 0.4713 | 0.81388012 | 0.90917256 |
| 2 | 0.4813 | 0.830523224 | 0.813292915 |
| 3 | 0.4554 | 0.849559447 | 0.948018412 |
| 4 | 0.1532 | 0.94528445 | 0.97799483 |
| 5 | 0.0916 | 0.973022952 | 0.981250701 |
| 6 | 0.0580 | 0.984009572 | 0.991018300 |
| 7 | 0.0573 | 0.98455346 | 0.99404962 |
| 8 | 0.0409 | 0.990536277 | 0.977658021 |
| 9 | 0.0484 | 0.98607636 | 0.99056921 |
| 10 | 0.0263 | 0.99456107 | 0.99607050 |
| 11 | 0.0188 | 0.99749809 | 0.99517233 |
| 12 | 0.0176 | 0.99738931 | 0.99640732 |
| 13 | 0.0155 | 0.99793321 | 0.99793321 |
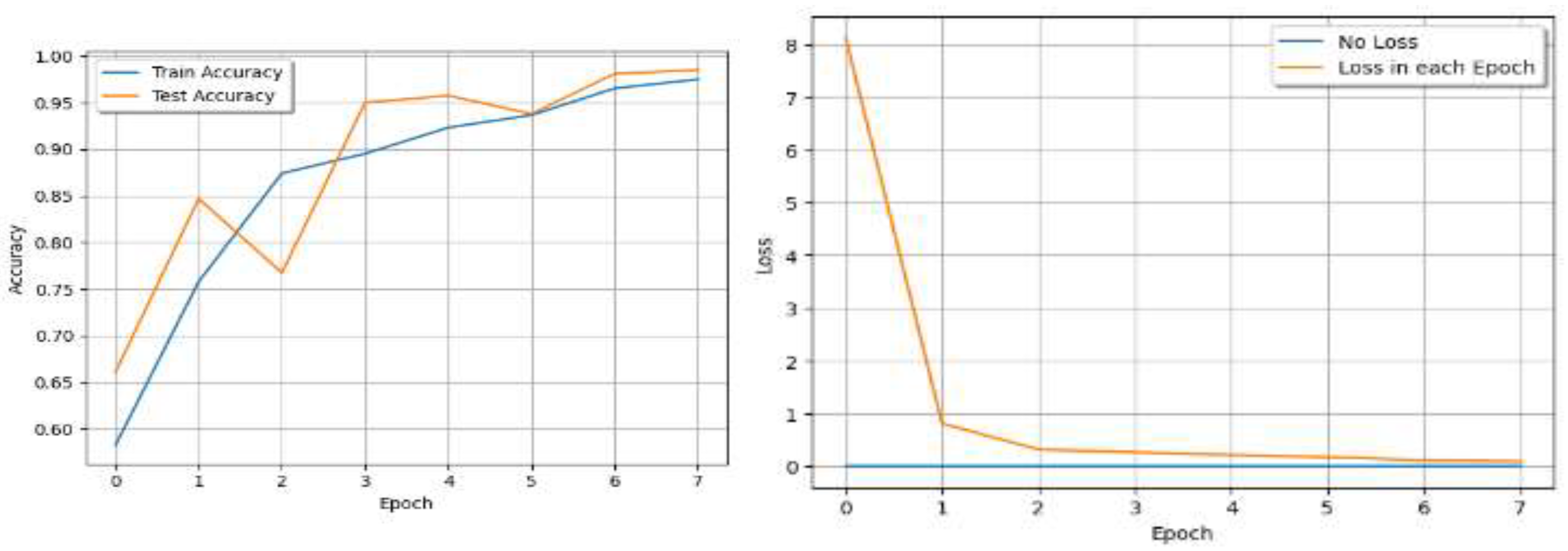
VI. Discussion
VII. Compared with the Other Results
VIII. Description of Shortcomings and Improvements
IX. Conclusion
References
- Burr DB. Introduction - bone turnover and fracture risk. J Musculoskeletal Neuronal Interact 2003, 3, 408–409. [Google Scholar]
- Pranata YD, Wang K, Wang J, Idram I, Lai J, Liu J, Hsieh I. Deep learning and surf for automated classification and detection of calcaneus fractures in ct images. Comput Methods Progr Biomed 2019, 171, 27–37. [Google Scholar]
- Urakawa T, Tanaka Y, Goto S, Matsuzawa H, Watanabe K, Endo N. Detecting intertrochanteric hip fractures with orthopedist-level accuracy using a deep convolutional neural network. Skeletal Radiol 2019, 48, 239–244. [Google Scholar]
- Dhahir BM, Hameed IH, Jaber AR. Prospective and retrospective study of fractures according to trauma mechanism and type of bone fracture. Res J Pharm Technol 2017, 10, 1994–2002. [Google Scholar]
- Bandyopadhyay O, Biswas A, Bhattacharya BB. Long-bone fracture detection in digital X-ray images based on digital-geometric techniques. Comput Methods Progr Biomed 2016, 123, 2–14. [Google Scholar]
- Cao Y, Wang H, Moradi M, Prasanna P, Syeda-Mahmood TF. Fracture detection in X-ray images through stacked random forests feature fusion. In: Biomedical imaging (ISBI), 2015 IEEE 12th international symposium on. IEEE; 2015. p. 801–5.
- Umadevi N, Geethalakshmi S. Multiple classification system for fracture detection in human bone X-ray images. In: Computing communication & networking technologies (ICCCNT), 2012 third international conference on. IEEE; 2012. p. 1–8.
- Lum VLF, Leow WK, Chen Y, Howe TS, Png MA. Combining classifiers for bone fracture detection in X-ray images. In: Image processing, 2005. ICIP 2005. IEEE international conference on, vol. 1. IEEE; 2005. I–1149.
- Liang J, Pan B-C, Huang Y-H, Fan X-Y. Fracture identification of X-ray image. In: Wavelet analysis and pattern recognition (ICWAPR), 2010 international conference on. IEEE; 2010. p. 67–73.
- Al-Ayyoub M, Hmeidi I, Rababah H. Detecting hand bone fractures in X-ray images. JMPT (J Manip Physiol Ther) 2013, 4, 155–168. [Google Scholar]
- Chai HY, Wee LK, Swee TT, Salleh S-H, Ariff A, et al. Gray-level co-occurrence matrix bone fracture detection. Am J Appl Sci 2011, 8, 26. [Google Scholar] [CrossRef]
- T. T. Peng, et al., report Detection of femur fractures in X-ray images, Master of Science Thesis, National University of Singapore. & T. Anu, M. M. R. Raman, Detection of bone fracture using image processing methods. Int J Comput Appl.
- Mahendran S, Baboo SS. An enhanced tibia fracture detection tool using image processing and classification fusion techniques in X-ray images. Global J Comput Sci Technol 2011, 11, 23–28. [Google Scholar]
- Building and Training a Custom Convolutional Neural Network with PyTorch using Cow Teat Image Dataset, Veerabhadra Rao Marellapudi Yeshiva University, NYC, NY vmarella@mail.yu.edu November 9, 2023.
- Bone Fracture Detection in X-ray Images using Convolutional Neural Network, Book on ResearchGate April 2022, All content following this page was uploaded by Rinisha Bagaria on 21 June 2022. Received March 29, 2020, accepted April 15, 2020, date of publication April 20, 2020, date of current version May 4, 2020.
- Muratsu (3, Syoji Kobashi (1 1) Graduate School of Engineering, University An automated fracture detection from pelvic CT images with 3-D convolutional neural networks Naoto Yamamoto (1, Rashedur Rahman (1, Naomi Yagi (1, 2), Keigo Hayashi (3, Akihiro Maruo (3, Hirotsugu of Hyogo, Himeji, Japan 2) Himeji Dokkyo University, Himeji, Japan 3) Steel Memorial Hirohata Hospital, Himeji, Japan. 3-2019.
- Gulshan, V. , et al. (2016). Development and Validation of a Deep Learning Algorithm for Detection of Diabetic Retinopathy in Retinal Fundus Photographs. & „Deep Learning” by Ian Goodfellow, Yoshua Bengio, and Aaron Courville. & „Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow” by Aurélien Géron.
- Saito, T. , & Rehmsmeier, M. (2015). The Precision-Recall Plot is More Informative than the ROC Plot when Evaluating Binary Classifiers on Imbalanced Datasets. & Bradley, A. P. (1997). The use of the area under the ROC curve in the evaluation of machine learning algorithms.
- Srivastava, N. , et al. (2014). Dropout: A Simple Way to Prevent Neural Networks from Overfitting. & Loshchilov, I., & Hutter, F. (2017). SGDR: Stochastic gradient descent with warm restarts.
- Caruana, R. , et al. (2008). Intelligible models for healthcare: Predicting pneumonia risk and hospital 30-day readmission. & Berrar, D. (2019). Cross-Validation. & Litjens, G., et al. (2017). A survey on deep learning in medical image analysis. & Razavian, A. S., et al. (2014). CNN Features Off-the-Shelf: An Astounding Baseline for Recognition.
- D. C. Ciresan, et al., „Deep neural networks segment neuronal membranes in electron microscopy images.” Advances in Neural Information Processing Systems (NIPS), 2012.
- Y. LeCun, L. Bottou, Y. Bengio, and P. Haffner, „Gradient-based learning applied to document recognition,” Proceedings of the IEEE, 1998. & S. Ren, K. He, R. Girshick, and J. Sun, „Faster R-CNN: Toward Real-Time Object Detection with Region Proposal Networks,”. arXiv:1506.01497, 2015.
- A. Esteva, et al., „A guide to deep learning in healthcare.” Nature Medicine, 2019. & T. G. Dietterich, „Ensemble methods in machine learning,” Multiple Classifier Systems, 2000.
- N. Srivastava, et al., „Dropout: A Simple Way to Prevent Neural Networks from Overfitting,” Journal of Machine Learning Research, 2014. & S. Raschka, et alal., „Model Evaluation, Model Selection, and Algorithm Selection in Machine Learning,” arXiv:1811.12808, 2018.
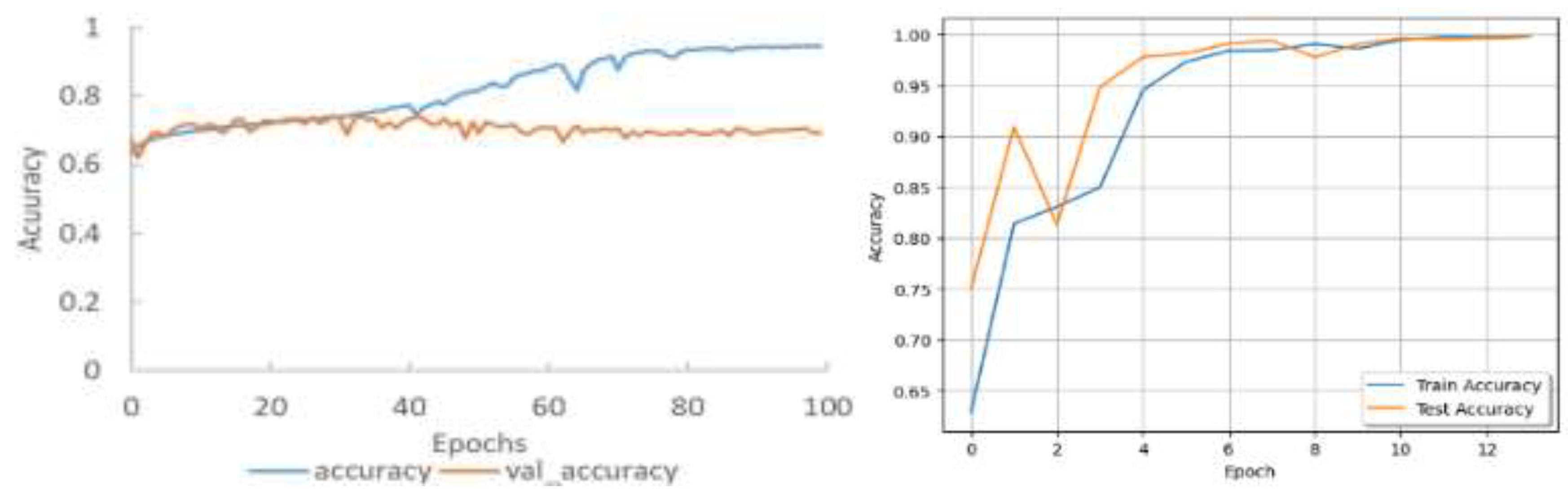
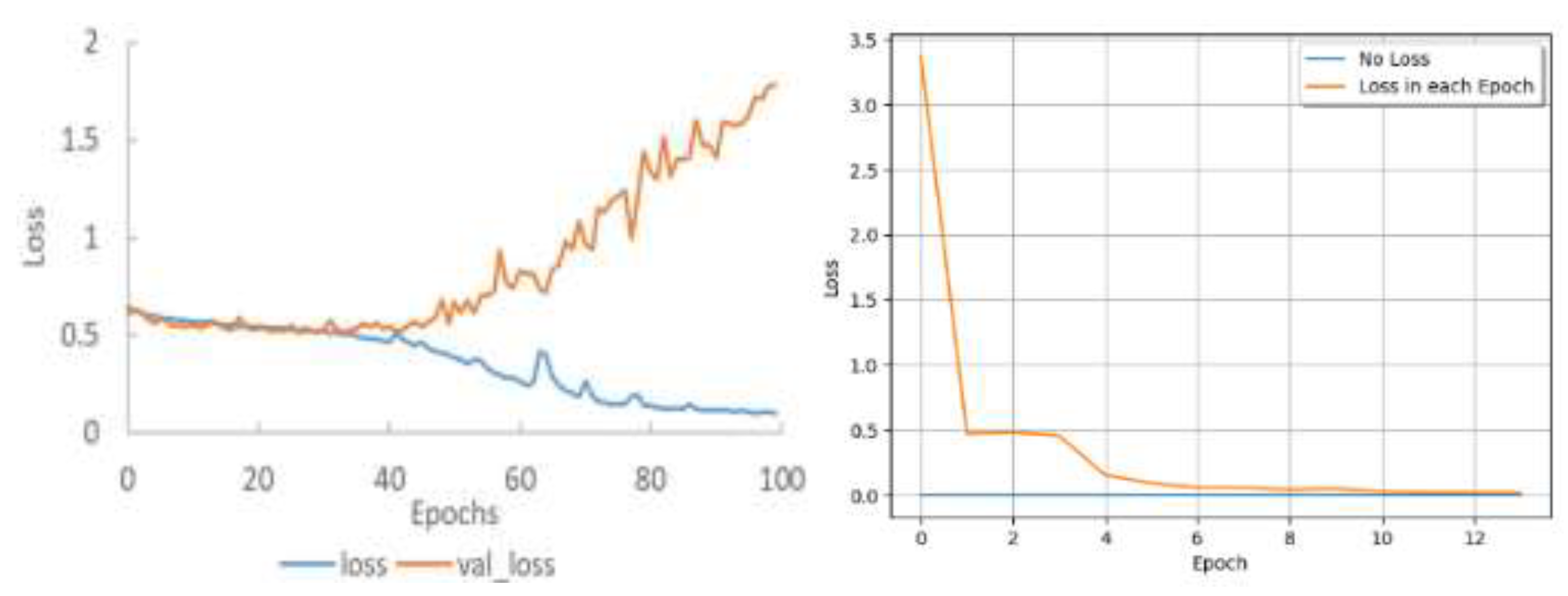
| Epoch | Train Loss | Train Accuracy | ||
| Rao’s result | Our result | Rao’s result | Our result | |
| 1 | 1.0686 | 0.4713 | 55.53% | 81.38% |
| 2 | 1.0481 | 0.4813 | 52.83% | 83.05% |
| 3 | 0.9955 | 0.4554 | 55.18% | 84.95% |
| 4 | 0.9264 | 0.1532 | 59.97% | 94.52% |
| 5 | 0.9029 | 0.0916 | 61.36% | 97.30% |
| 6 | 0.9105 | 0.0580 | 60.84% | 98.40% |
| 7 | 0.8952 | 0.0573 | 61.01% | 98.45% |
| 8 | 0.8948 | 0.0409 | 61.18% | 99.05% |
| 9 | 0.8859 | 0.0484 | 61.88% | 98.60% |
| 10 | 0.8875 | 0.0263 | 62.05% | 99.45% |
| 11 | 0.8824 | 0.0188 | 62.75% | 99.74% |
| Epoch | Batch size | Accuracy (%) | AUC | Specificity |
| 10 20 20 20 20 20 |
10 32 32 32 32 32 |
87.20% 76.20% 88.00% 89.90% 89.00% 86.82% |
0.8244 0.6589 0.8286 0.8088 0.8417 0.6819 |
87.20% 76.20% 88.00% 89.90% 89.00% 86.82% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




