1. Introduction
Camellia oleifera Abel, a species within the Dungarunga group or shrub category, holds a place in the esteemed Camellia family, scientifically known as Theaceae. It has high values in economic, edible, environmental and medicinal field [
1]. In 2019, there were 4.4 million hectares of land used to grow C.oleifera in China, in which production exceeded 2.4 million tons [
2]. C.oleifera cultivation is still facing great hazards from diseases and insect pests, some of which usually lead to 20%-40% reduction in fruit percentage and 10%--30% loss of seeds. It may cause the branches to wilt, even the entire plant to die in severe cases. Meanwhile, serious economic losses will occur, as disease spread rapidly and are difficult in control. At present, the control measures for diseases and insect pests of C. oleifera predominantly adopted are chemical interventions, and complemented by tea plantation sanitation. However, diseases are characterized by a lengthy incubation period and rapid dissemination, which is a challenge to determine the optimal timing for conventional control measures. Conrad et al. [
3] emphasize that manual detection of diseases is costly and inefficient. There is an pressing need for a fast, and accurate detection and segmentation method. However, conventional image processing techniques struggle to swiftly and accurately identify multiple diseases in intricate environments [
4].
With the modernization, and intelligence of agriculture, accurately identifying disease categories and taking corresponding measures are significant. In agriculture, the rapid advancement and adoption of machine learning have proven to be a powerful and efficient solution [
5]. Conventional mechanical learning methods, K-mean clustering, Markov Random Fields, Random Forests, and Support Vector Machines had been applied in the field of image segmentation for many years [
6]. However, they exhibited inefficiencies in computation, limited scalability, and poor generalization performance, especially for handling complex data and tasks. Thanks to the groundbreaking advancements in deep learning, remarkable strides have been achieved in the field of plant disease segmentation [
7,
8]. Deep learning models focused on the efficient processing and full utilizing for the contextual information of images, through automatic feature extraction. Convolutional Neural Networks (CNNs), as a important component of deep learning, have emerged as the preferred network architecture for numerous model. Lu et al. [
9] reviewed the application of CNN on diseases and insect pests of plant leaf. The convolutional layers were used to effectively capture local information in CNN. By stacking multiple layers of convolutional and pooling layers, CNN can learned hierarchical features, then efficiently comprehended both global and local contexts within the image. Consequently, the realm of image segmentation had witnessed the emergence of numerous innovative CNN-based architectures. Ramcharan et al. [
10] applied transfer learning to train neural networks for the recognition about diseases of cassava, a significant economic crop. Lu et al. [
11] introduced an automatic diagnosis system utilizing a supervised deep learning framework for identifying wheat disease. Moreover, it was packed into a real-time mobile application. Liu et al. [
12] introduced an innovative approach for grape leaf disease classification, which relied on an improved CNN. In the network, the Inception structure and dense connectivity strategy were introduced to boost the extraction of multidimensional features. For detecting five common leaf diseases of apple trees, the Inception and Rainbow were combined in a new deep learning network by Jiang et al. [
13]. Wang et al. [
14] developed a new model that combined U-Net and DeeplabV3+, specifically tailored for cucumber foliar disease segmentation in challenging background. Deng et al. [
15] proposed a cross-layer attentional mechanism combined with a multiscale convolutional module, which could greatly solved the problem of fuzzy edges and tiny disease in tomato foliar diseases.
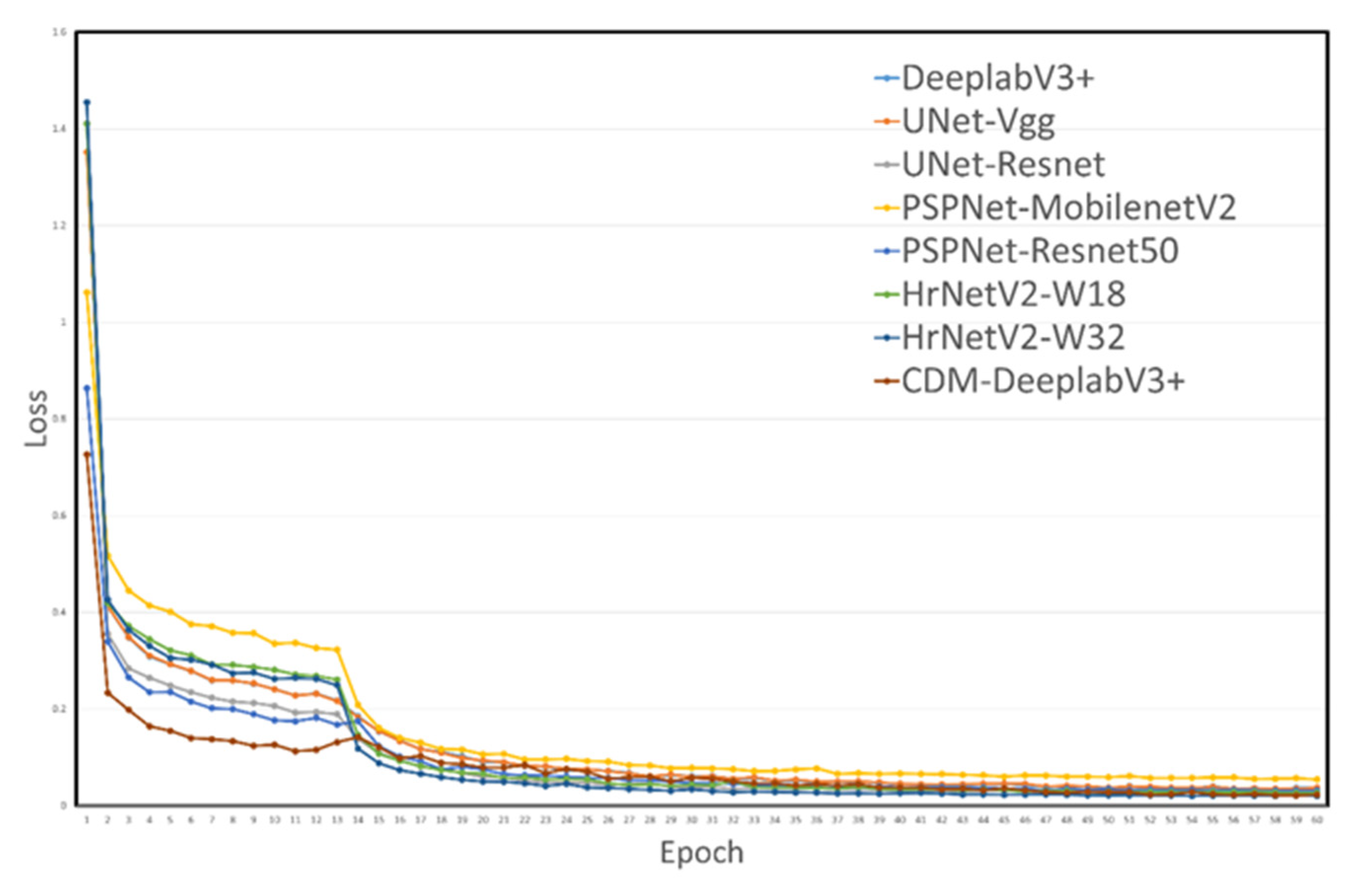
Based on the above studies, we can infer that the segmentation of diseases and insect pests on important economic crops contributed to the development of agriculture. C. oleifera is also considered an economically significant crop, but the current relevant research mainly focused on pathological aspect. Meanwhile, DeeplabV3+ was a leading model of semantic segmentation model, which had the following advantages by comparing with other models such as UNet, UNet++, PSPNet, HrNetV2, et[
16]:
• The Encoder-Decoder architecture enhanced feature extraction by utilizing deep networks within the encoder, while the decoder improved the precision of segmentation outcomes through a process of upsampling.
• The use of enhanced dilated convolutions was strategically employed to effectively gather information from various spatial scales, thereby significantly expanding the receptive field of convolutional layers. It allowed the model to capture a broader context.
• The ASPP module enhanced segmentation accuracy through the multiple dilated convolutions of different rates.
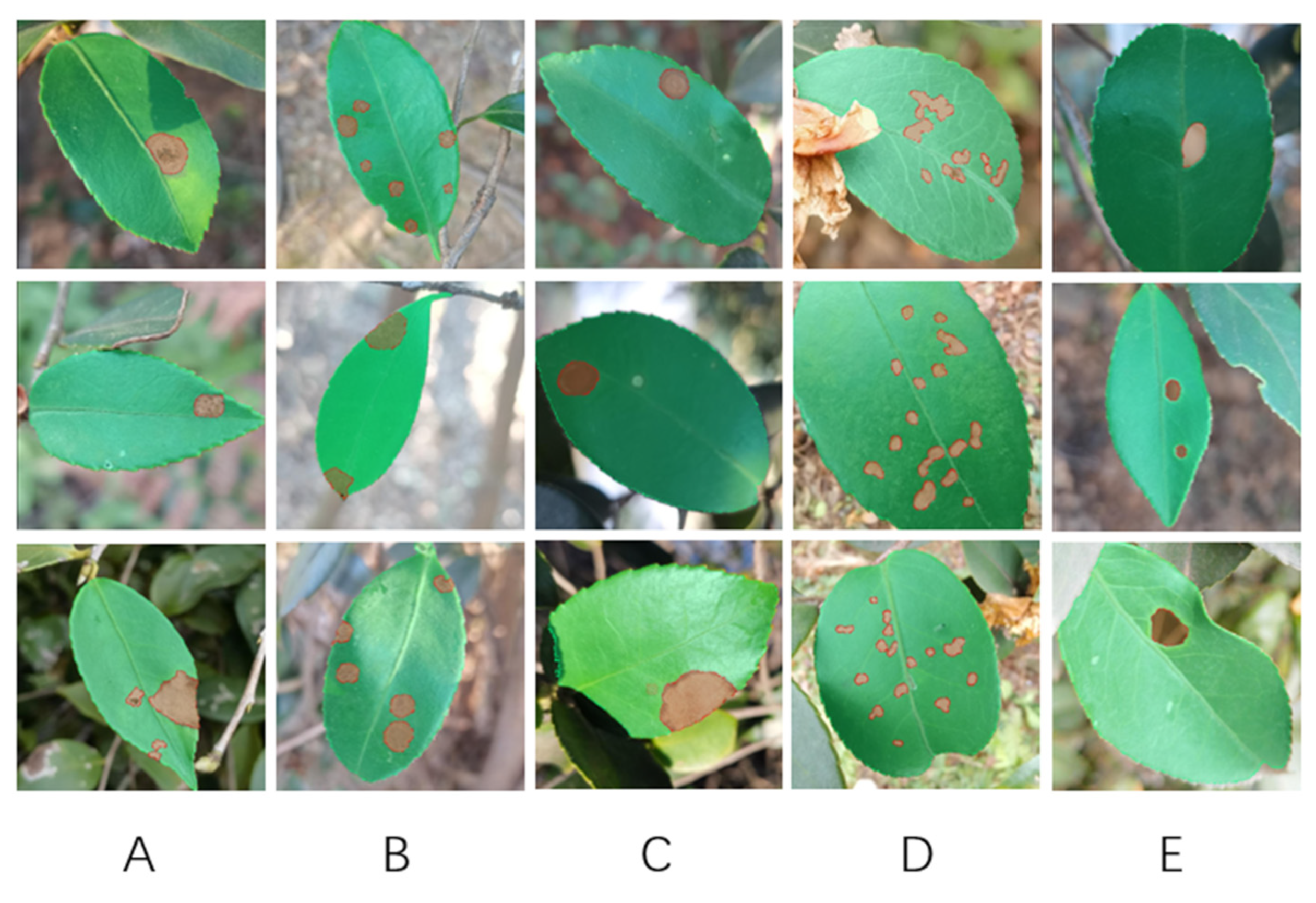
Due to the lack of public dataset, we built own dataset of C. oleifera diseases and insect pests. The dataset was comprised of 1264 images and 5 different diseases, which all were shot in their natural environments. Then, we utilized the DeeplabV3+ model directly on self-build dataset, but encountered the following challenges:
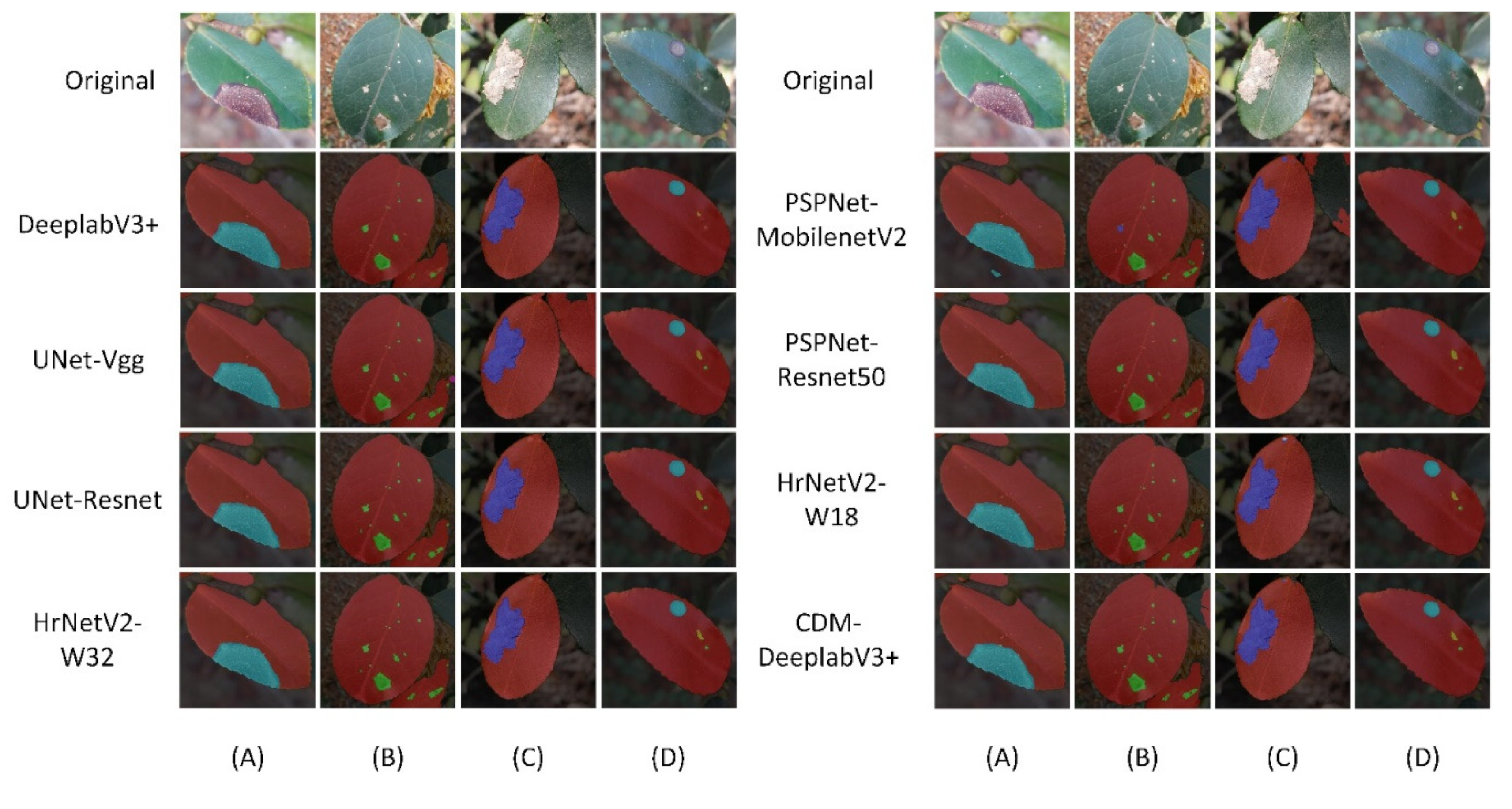
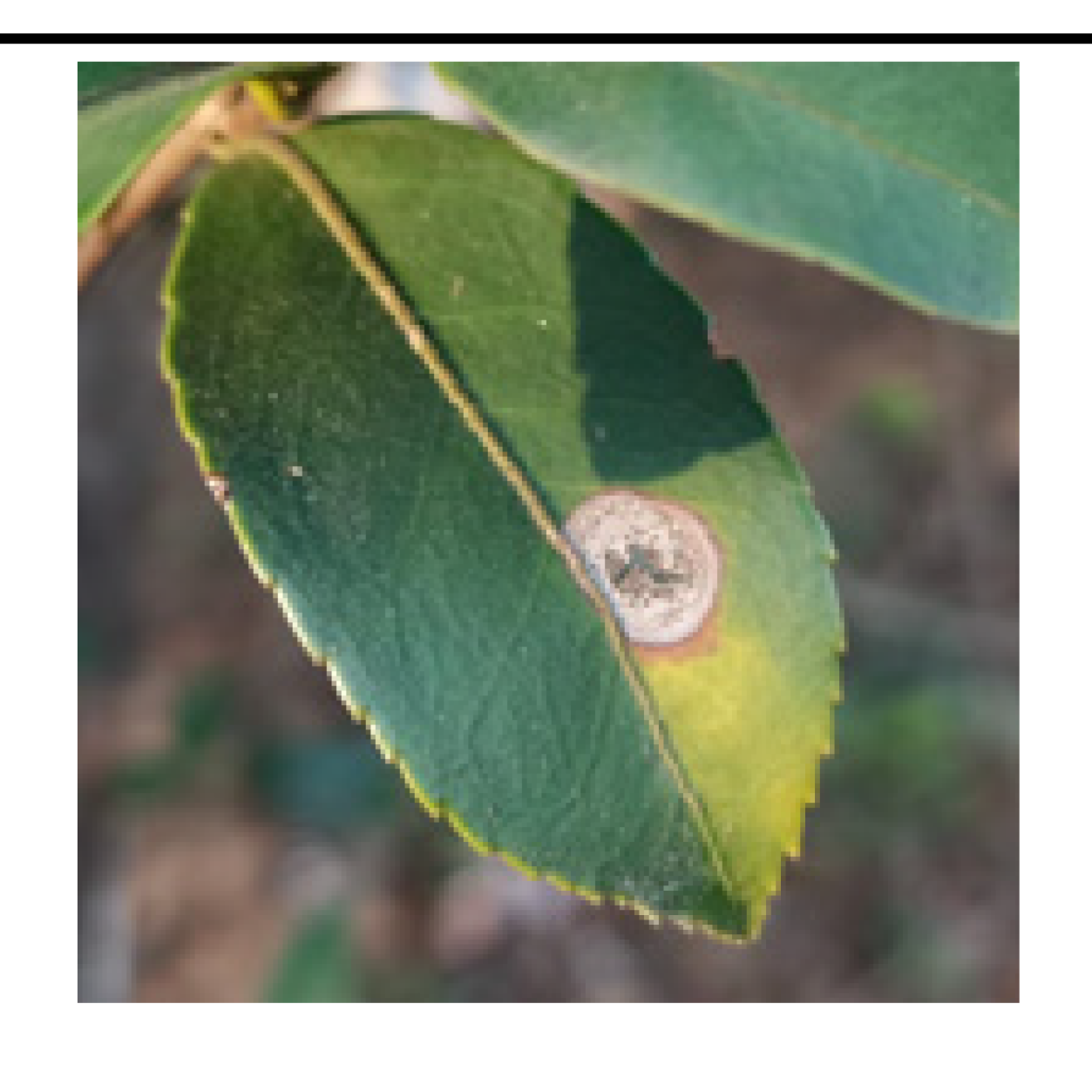
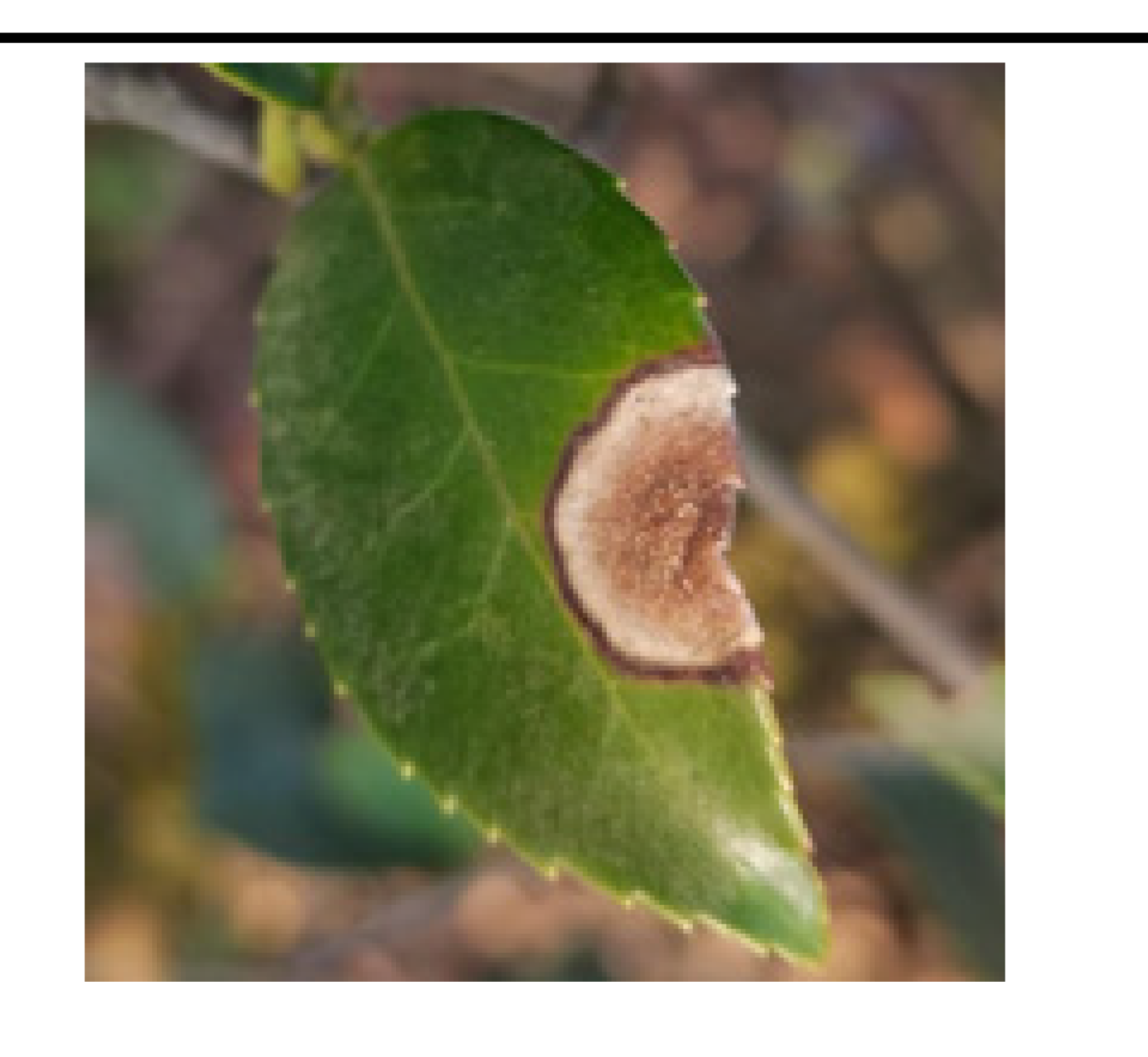
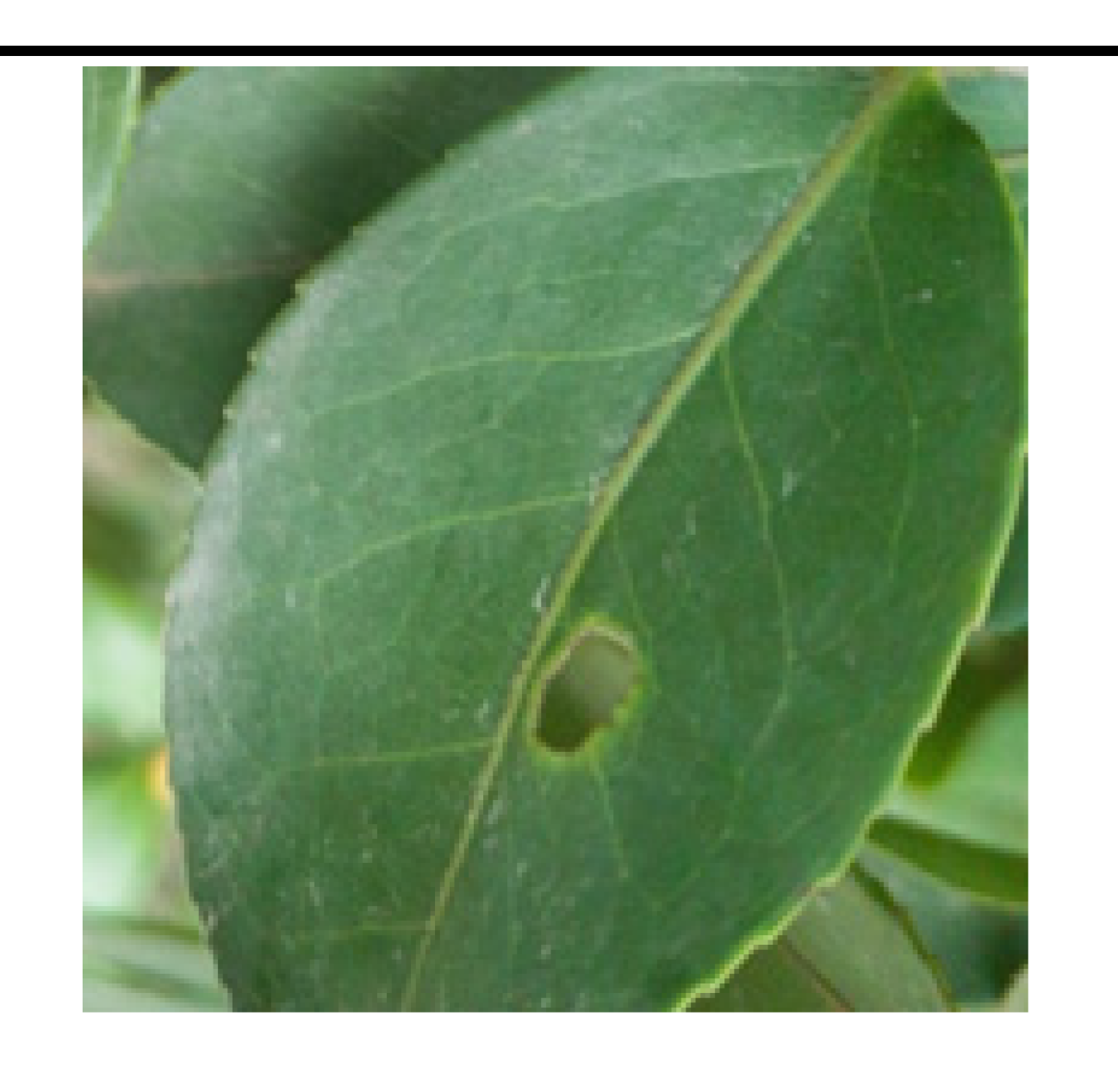
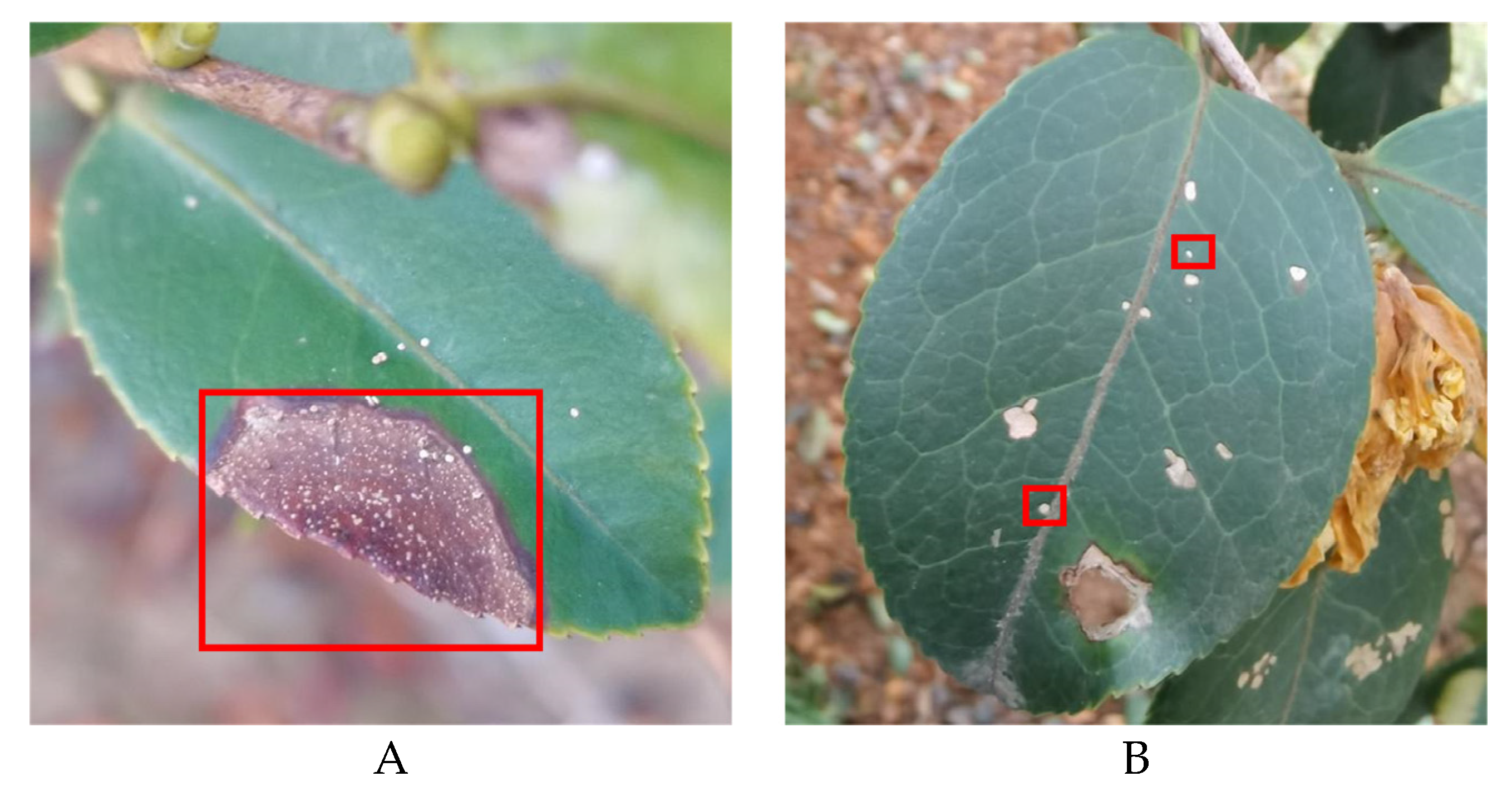
• The boundary of diseased leaves between the background was fuzzy, as shown in
Figure 1. (A). This presented a challenge for the conventional DeeplabV3+ network. It struggled to effectively differentiate between background and disease features, resulting in a notable reduction in feature extraction accuracy.
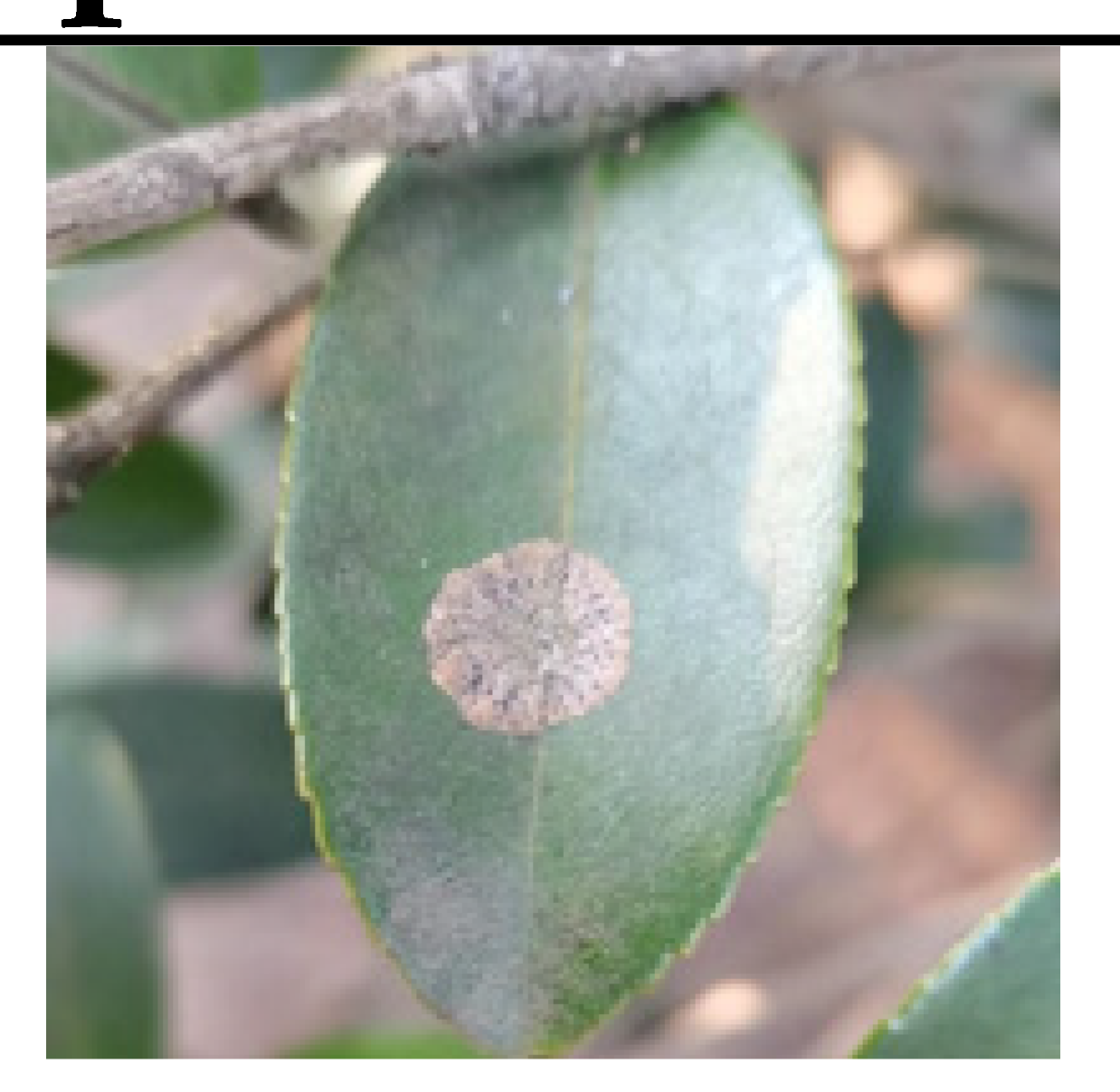
• The gridding of dilated convolutions increased the loss of tiny disease information. The average pooling layer also lost edge and texture information. This presented a difficulty for the model in accurately segmenting these small diseased areas, as shown in
Figure 1B.
• DeeplabV3+ adopted Xception as its backbone, which possessed a large number of parameters. The backbone produced complex computations and long segmentation duration. Consequently, it was unfit to be applied in practical crop production.
To solving the problem of fuzzy boundary between features and background, Khadidos et al. [
17] introduced an approach that relied on level set evolution. The method assigned weights to energy terms based on their significance in boundary delineation, which served the dual purpose of improving both the precision of feature extraction and the accuracy of boundary segmentation. Xia et al.[
18] integrated the Reverse Edge Attention Module (REAM) into their encoder architecture of model, strategically placing it between consecutive layers. We proposed a CBAM-ASPP module to solve not only the similarity of color between diseases and background, but also the presence of blurry boundaries. To improve the ability of capture features, we expanded convolution layers by assigning various dilation rate which captured features from different scales. Additionally, we removed the average pooling layer to better retain edge and texture information. By connecting Convolutional Block Attention Module (CBAM) with dilated convolutions, the model was directed to concentrate on salient features and distinguish the edges of diseased leaves more effectively.
The feature extraction of tiny disease also became a challenge. Gao et al.[
19] introduced the H-SimAM Attention Mechanism which aimed to concentrate on potential symptoms within the complex background. Xiao et al. [
20] developed the Context Enhancement Module and Feature Purification Module to enhance the detection accuracy, particularly for smaller targets. Due to the distinctive encoder-decoder architecture of DeeplabV3+, we have introduced the DANet attention mechanism to create an enhanced encoder. By focusing attention on feature maps via spatial and channel dimensions, the DANet mitigated impact of complex background and challenges of tiny object shapes.
The large model parameters and complex computations was also a serious problem. Liang et al.[
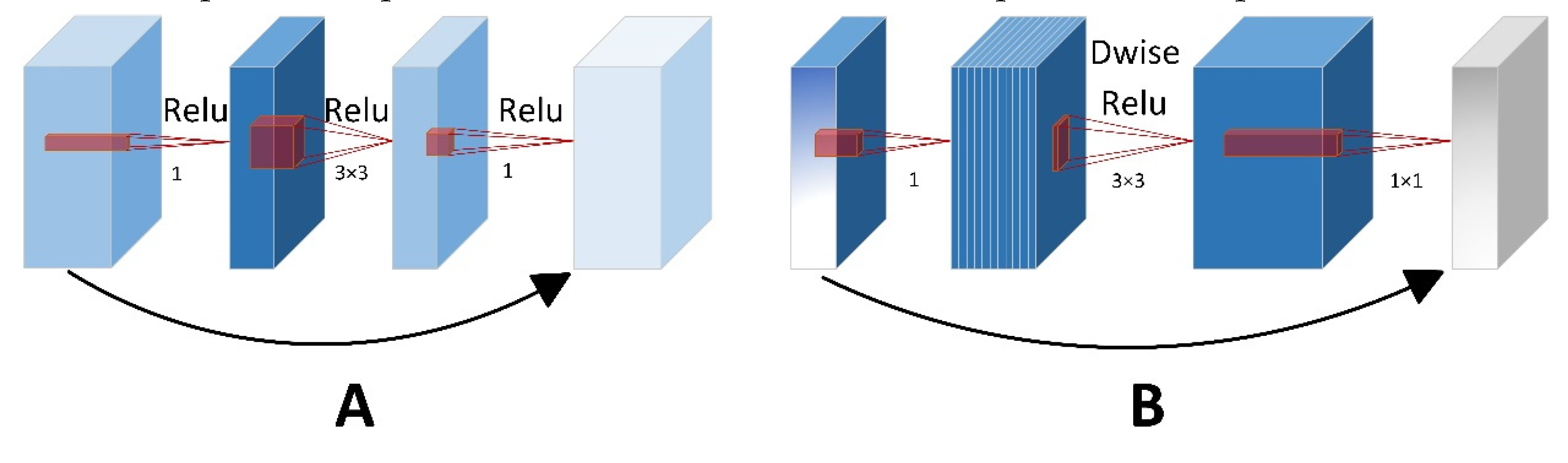
21] discussed that lightweight design based on repetitive feature maps mainly concentrated on lightweighting the cheap operations. However, these methods often neglected generalization capabilities of network and huge computational demands in convolutional layers. To address this, the authors implemented a group convolution technique aimed at decreasing the computational intensityof floating-point operations. MobileNetV2 was adopted as the backbone due to its suitability for mobile and embedded device. The key role of MobileNetV2 contributed to the design of inverted residual block, which was integrated into depthwise separable convolutions and linear bottleneck layers. These innovations enabled the model to effectively decrease parameters without decreasing accuracy of segmentation detection. As a result, MobileNetV2 provided more practical values for mobile devices in agriculture.
The key contributions were encapsulated as follows:
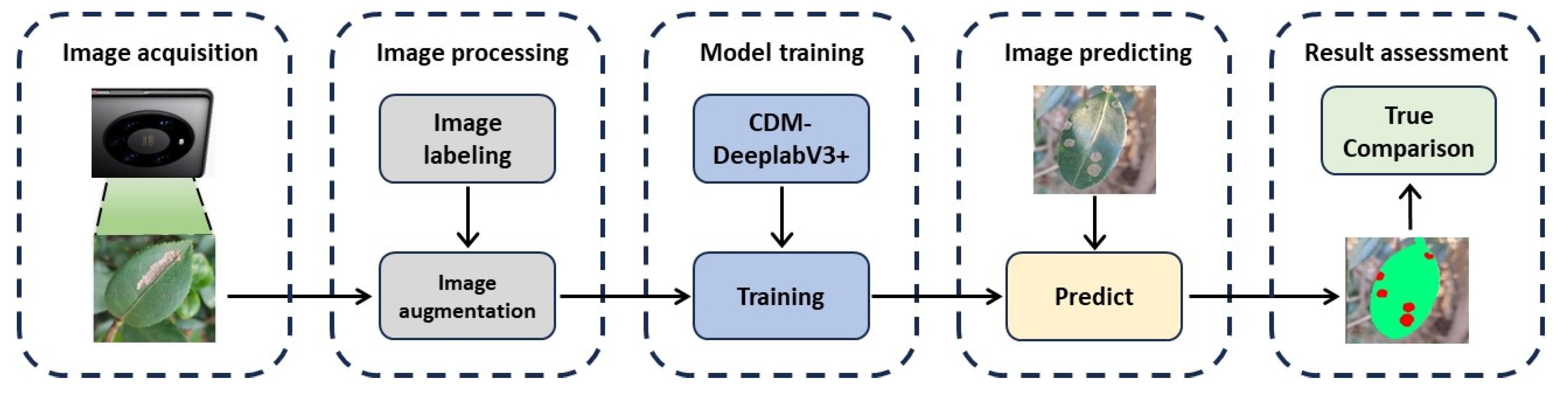
1. We designed an experimental simulation method for insect pest and disease segmentation based on CDM-DeeplabV3+ network and employed a self-build dataset containing 1265 images that had five of diseases and insect pests of C. oleifera. All the data analysis in this research were concluded from this proprietary dataset.
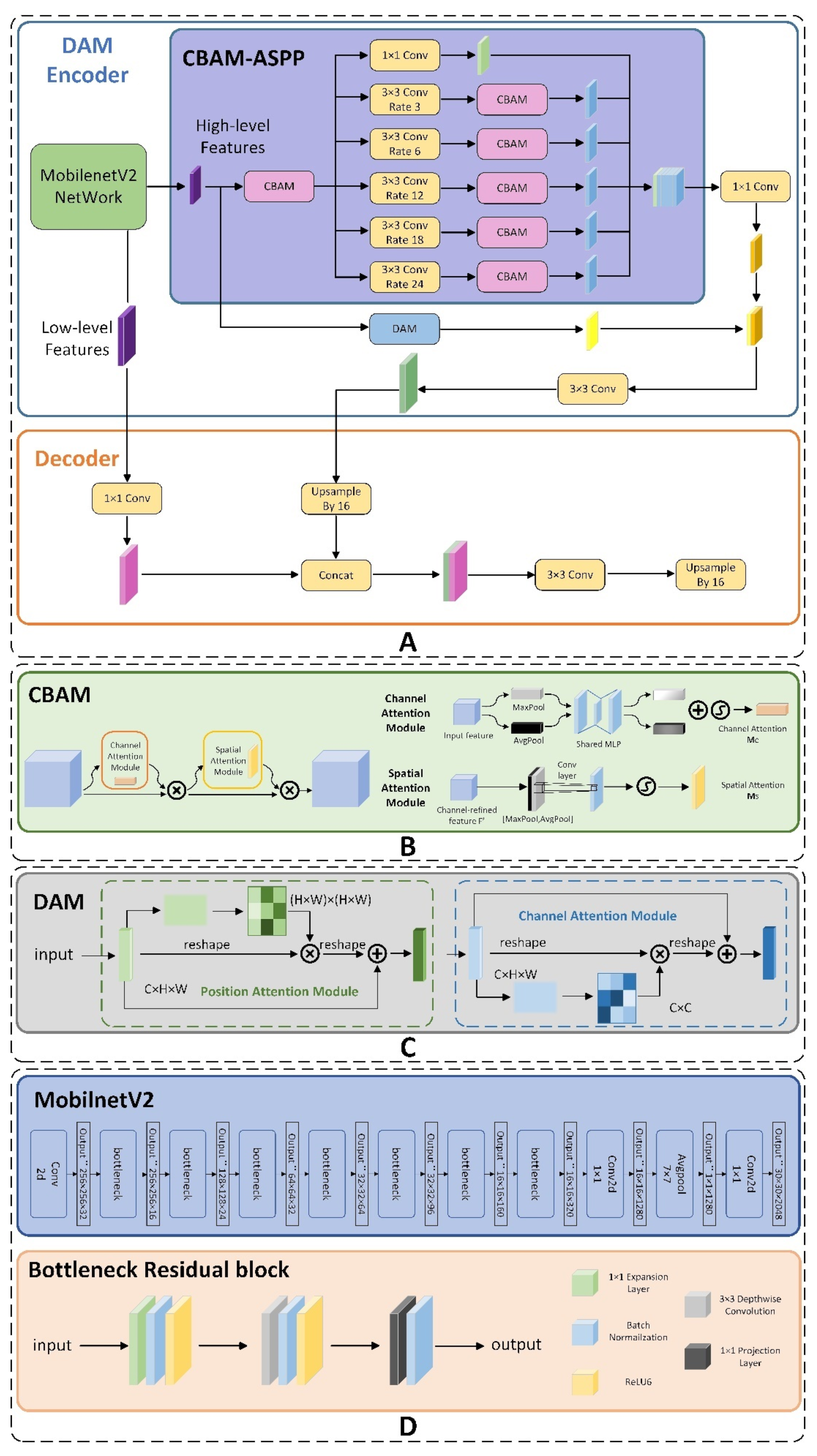
2. A Residual Attention ASPP (CBAM-ASPP) was proposed, which extended dilated convolution amount to five and integrates CBAM modules to form a new Attention Convolution Unit. This unit significantly enhanced the extraction of key semantic information across varying scales. Additionally, the module removed the use of the average pooling layer and redirected attention towards edge and texture features, thereby reinforcing the performance of network to accurately segment edge features of leaf diseases.
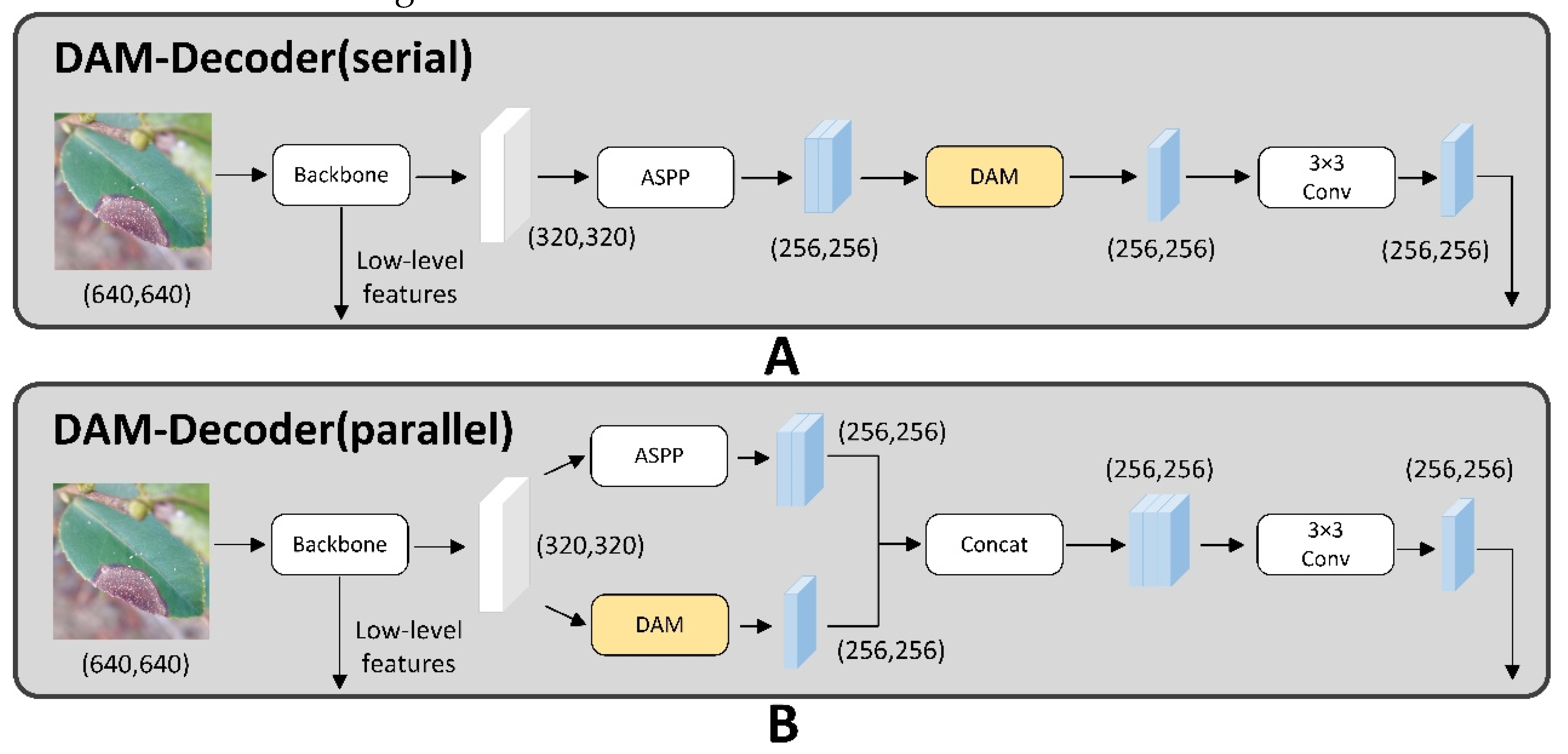
3. A Dual-Attention Encoder Module (DAM- Encoder) was applied to improve feature extraction capabilities along both spatial and channel dimensions. Within the encoder, we integrated the CBAM-ASPP and DANet modules in a parallel configuration. It captured minutely detailed features from diverse perspectives, effectively addressing the problem of losing tiny features.
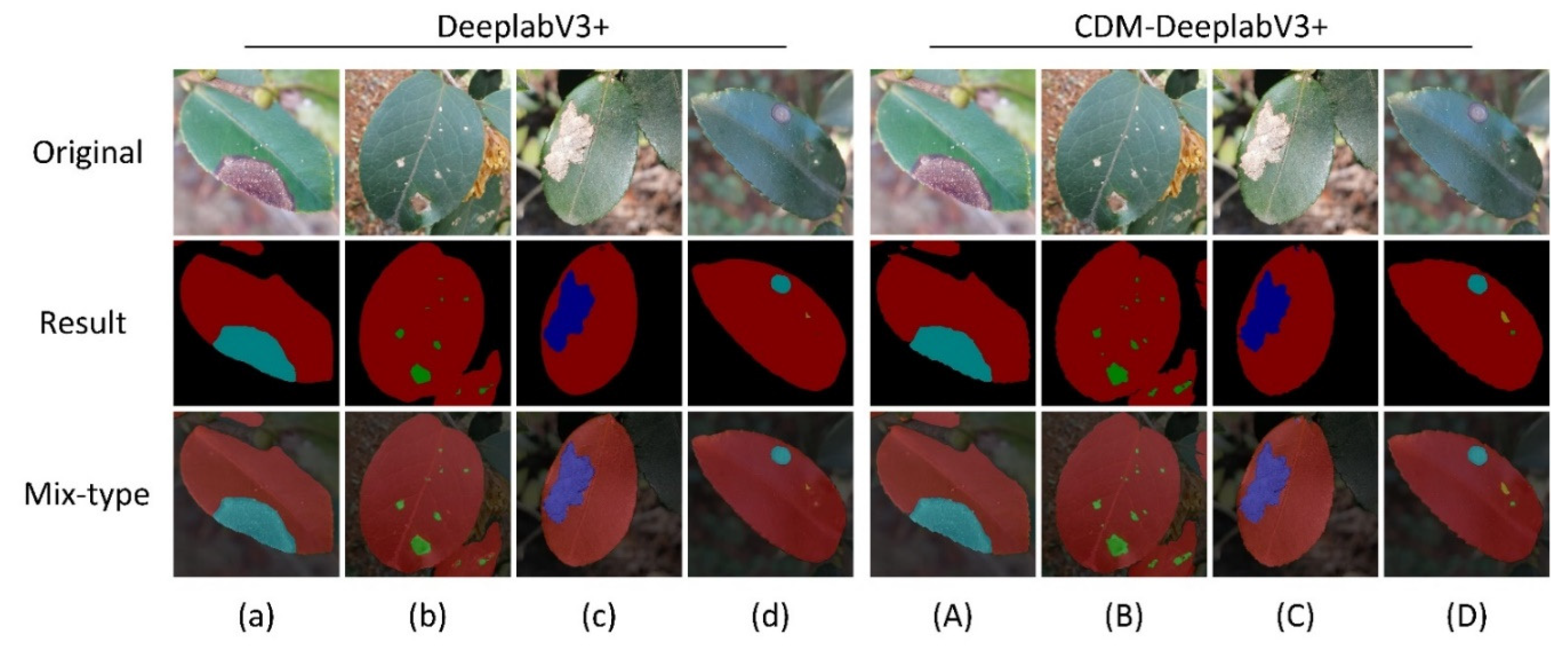
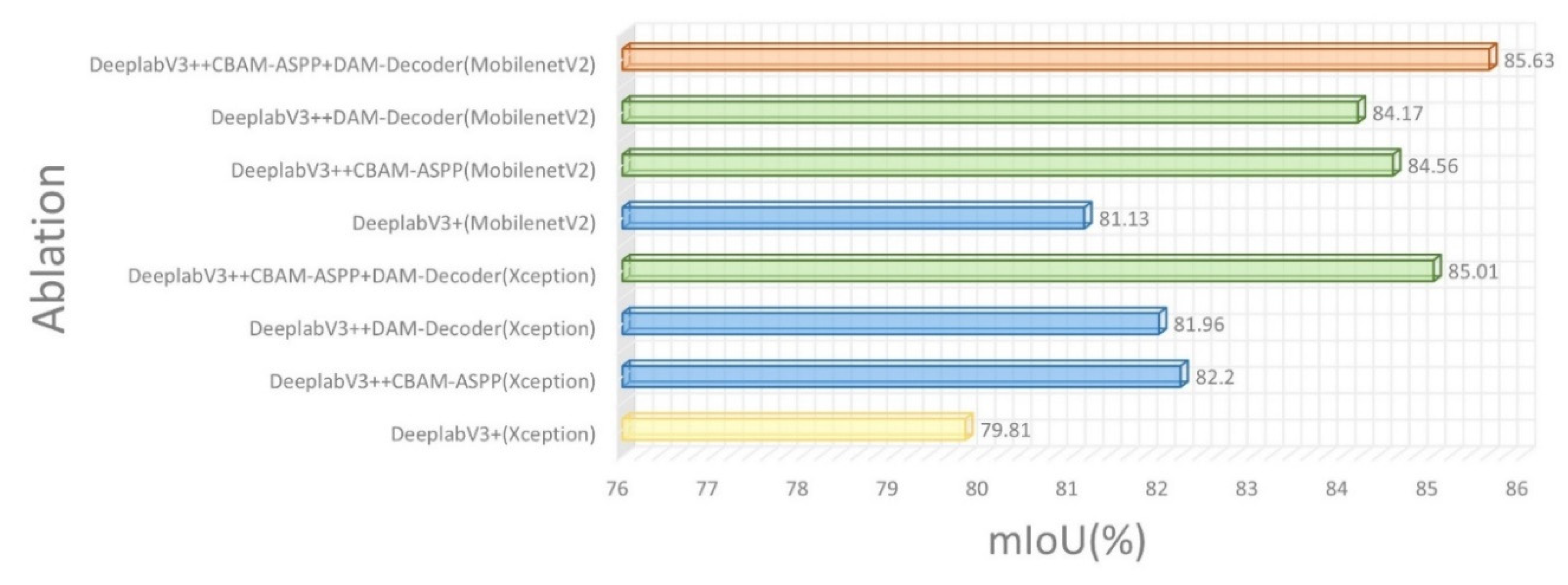
4. We changed the backbone to MobilenetV2. Unlike Xeception, MobilenetV2 introduced the inverted residual block, which substantially decreased the number of parameters maintaining the precision of segmentation detection. The experimental results show that CDM-DeeplabV3+ achieved an mIoU of 85.63%, which had an increase of 5.82% to original model, and model parameters reduction by almost 5 times compared to original model.