Submitted:
06 October 2024
Posted:
07 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction: Hepatocellular Carcinoma
1.1. Available Therapeutic Approaches for HCC
1.2. HCC Diagnoses
1.2.1. Instrumental Diagnoses
1.2.2. Biochemical Diagnoses
1.2.3. Considerations about HCC Diagnoses/Monitoring
2. Non-Coding RNAs
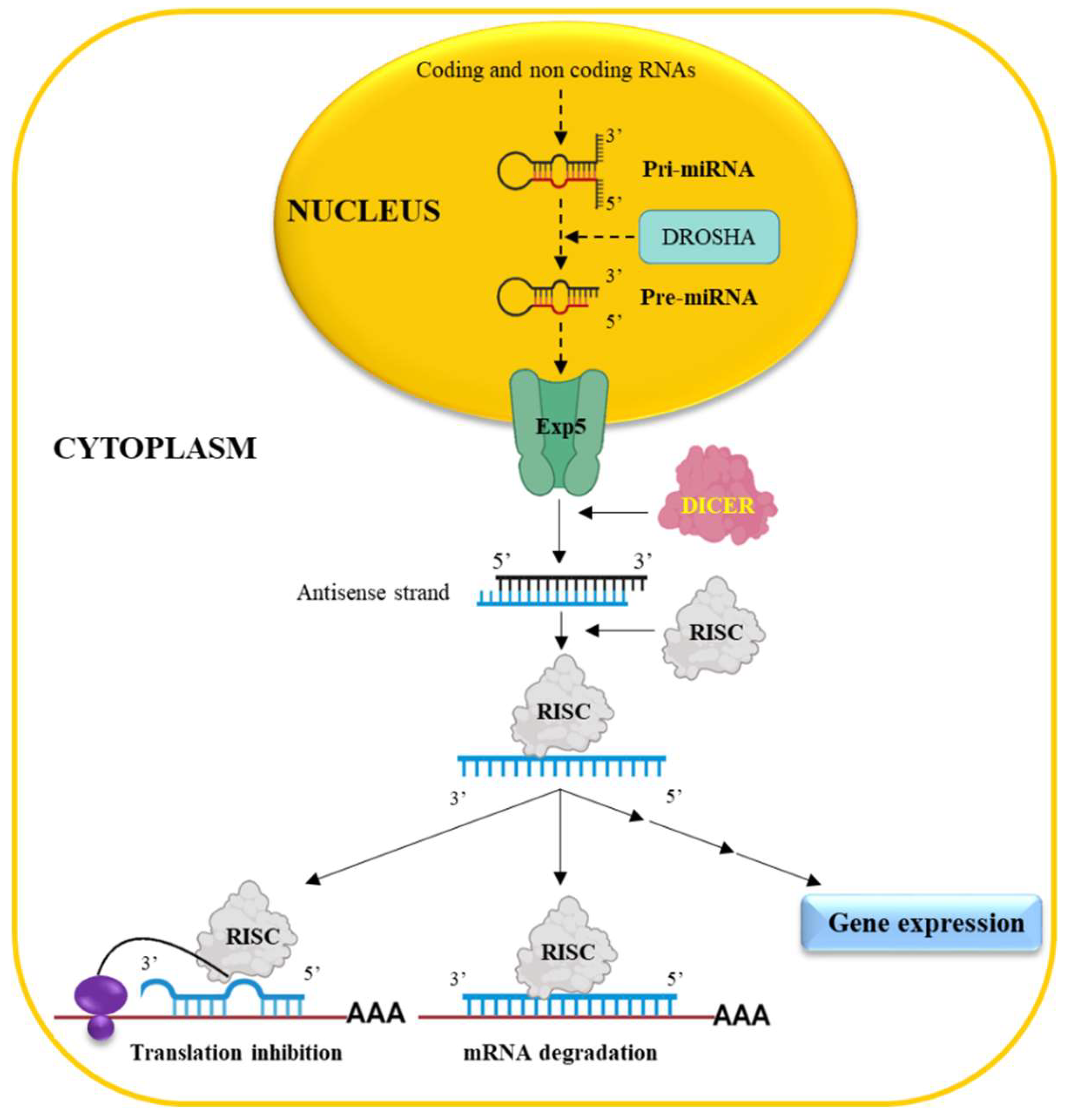
2.1. miRNAs
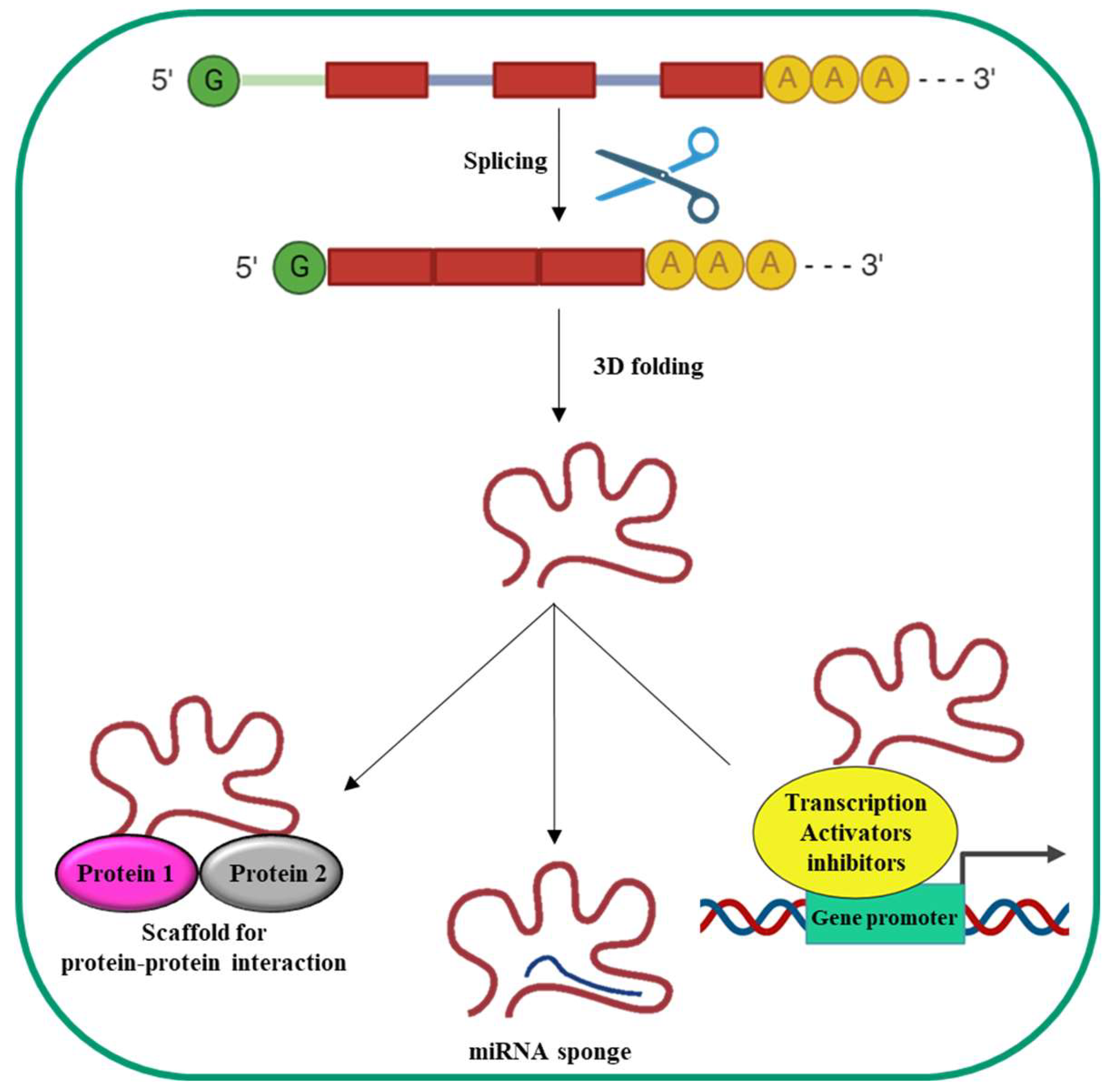
2.2. Long Non-Coding RNA
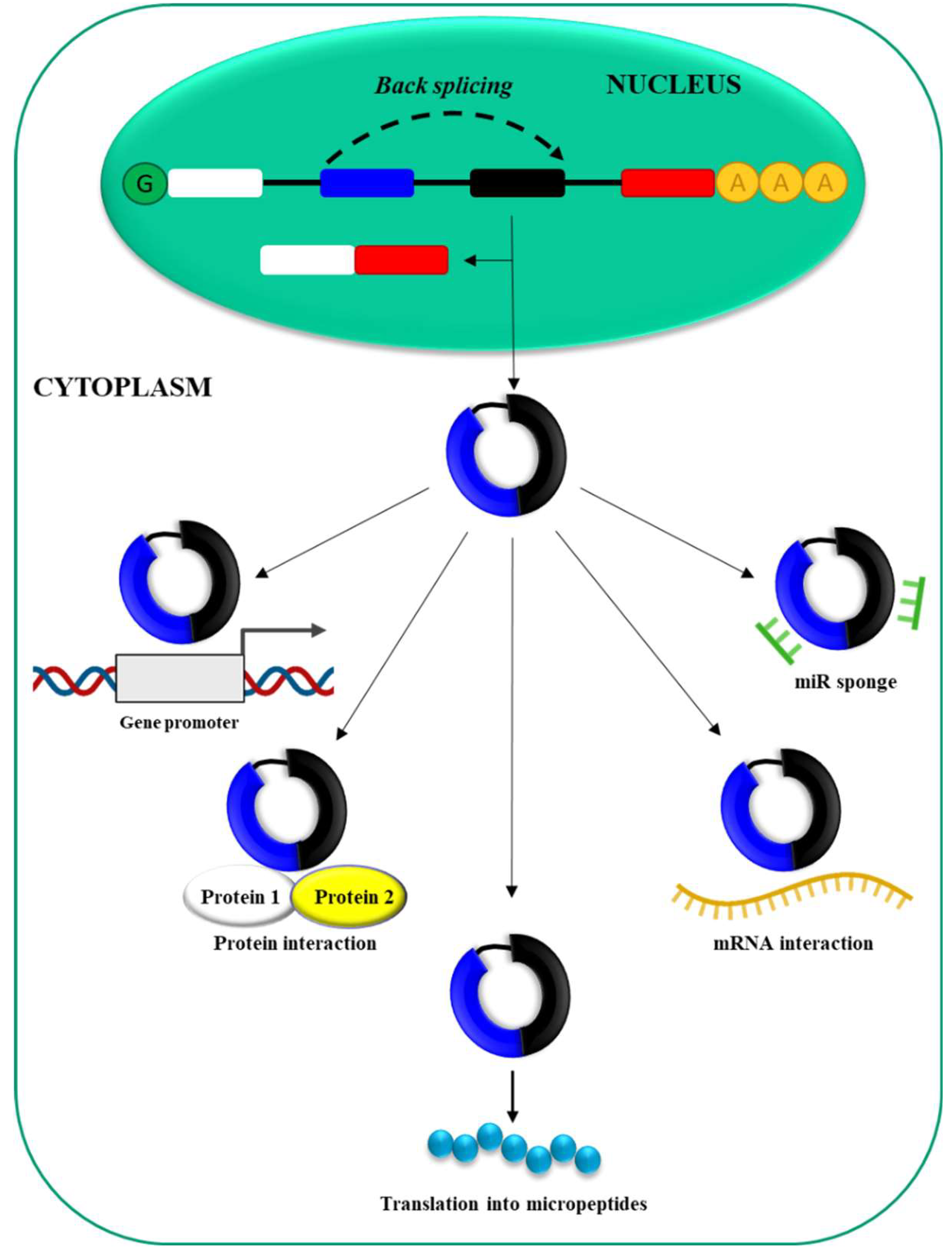
2.3. circRNAs
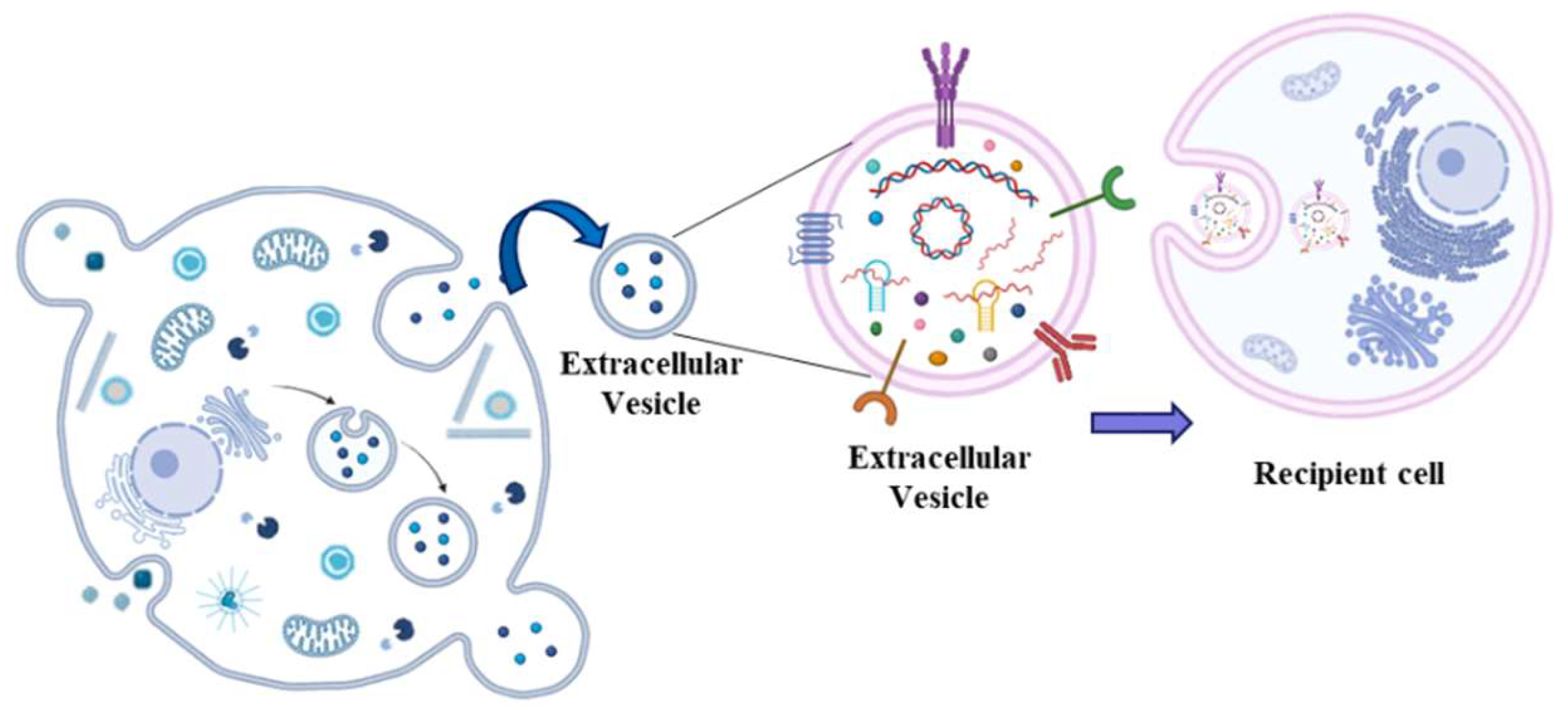
3. Extracellular Vesicles
4. ncRNAs Employment as Biomarkers for HCC
4.1. miRNA
4.1.1. miR-221 and miR-18a
4.1.2. miR-487a
4.1.3. miR-33a
4.1.4. miR-105-1
4.1.5. miR-138-5p
4.1.6. miR-21
4.1.7. miR-122
4.1.8. miR 139-5p
4.2. LncRNAs
4.2.1. LncRNAs HULC and MALATI
4.2.2. LncRNAs Linc00152
4.2.3. lncRNA HOTAIR
4.3. CircRNAs
4.3.1. Circ_0001445/ cSMARCA5
4.3.2. circ_0001649
4.3.4. circ_0003570
4.3.5. Other circRNAs
5. Conclusion
Author Contributions
Funding
Conflicts of Interest
References
- Dasgupta, P.; Henshaw,C.; Youlden,D.R.; Clark,P.J.; Aitken,J.F.; Baade,P.D. Global Trends in Incidence Rates of Primary Adult Liver Cancers: A Systematic Review and Meta-Analysis. Front Oncol. 2020, 10, 171. [CrossRef]
- Rumgay, H.; Arnold,M.; Ferlay,J.; Lesi,O.; Cabasag,C.J.; Vignat,J.; Laversanne,M.; McGlynn,K.A.; Soerjomataram,I. Global burden of primary liver cancer in 2020 and predictions to 2040. J Hepatol. 2022, 77, 1598-1606. [CrossRef]
- Siegel, R. L.; Miller,K.D.; Fuchs,H.E.; Jemal,A. Cancer statistics, 2022. CA Cancer J Clin. 2022, 72, 7-33.
- Rumgay, H.; Ferlay,J.; de,M.C.; Georges,D.; Ibrahim,A.S.; Zheng,R.; Wei,W.; Lemmens,V.E.P.P.; Soerjomataram,I. Global, regional and national burden of primary liver cancer by subtype. Eur. J Cancer. 2022, 161, 108-118. [CrossRef]
- Llovet, J. M.; Kelley,R.K.; Villanueva,A.; Singal,A.G.; Pikarsky,E.; Roayaie,S.; Lencioni,R.; Koike,K.; Zucman-Rossi,J.; Finn,R.S. Hepatocellular carcinoma. Nat. Rev Dis. Primers. 2021, 7, 6.
- Knox, J. J.; Cleary,S.P.; Dawson,L.A. Localized and systemic approaches to treating hepatocellular carcinoma. J. Clin. Oncol. 2015, 33, 1835-1844. [CrossRef]
- Affo, S.; Yu,L.X.; Schwabe,R.F. The Role of Cancer-Associated Fibroblasts and Fibrosis in Liver Cancer. Annu. Rev Pathol. 2017, 12, 153-186. [CrossRef]
- Schwabe, R. F. and Luedde,T. Apoptosis and necroptosis in the liver: a matter of life and death. Nat. Rev Gastroenterol. Hepatol. 2018, 15, 738-752. [CrossRef]
- Petrick, J. L.; Florio,A.A.; Znaor,A.; Ruggieri,D.; Laversanne,M.; Alvarez,C.S.; Ferlay,J.; Valery,P.C.; Bray,F.; McGlynn,K.A. International trends in hepatocellular carcinoma incidence, 1978-2012. Int. J Cancer. 2020, 147, 317-330. [CrossRef]
- Chidambaranathan-Reghupaty, S.; Fisher,P.B.; Sarkar,D. Hepatocellular carcinoma (HCC): Epidemiology, etiology and molecular classification. Adv. Cancer Res. 2021, 149, 1-61.
- EASL Clinical Practice Guidelines: Management of hepatocellular carcinoma. J Hepatol. 2018, 69, 182-236.
- Mazzaferro, V.; Regalia,E.; Doci,R.; Andreola,S.; Pulvirenti,A.; Bozzetti,F.; Montalto,F.; Ammatuna,M.; Morabito,A.; Gennari,L. Liver transplantation for the treatment of small hepatocellular carcinomas in patients with cirrhosis. N. Engl. J Med. 1996, 334, 693-699. [CrossRef]
- Ducreux, M.; Abou-Alfa,G.K.; Bekaii-Saab,T.; Berlin,J.; Cervantes,A.; de,B.T.; Eng,C.; Galle,P.; Gill,S.; Gruenberger,T.; Haustermans,K.; Lamarca,A.; Laurent-Puig,P.; Llovet,J.M.; Lordick,F.; Macarulla,T.; Mukherji,D.; Muro,K.; Obermannova,R.; O’Connor,J.M.; O’Reilly,E.M.; Osterlund,P.; Philip,P.; Prager,G.; Ruiz-Garcia,E.; Sangro,B.; Seufferlein,T.; Tabernero,J.; Verslype,C.; Wasan,H.; Van,C.E. The management of hepatocellular carcinoma. Current expert opinion and recommendations derived from the 24th ESMO/World Congress on Gastrointestinal Cancer, Barcelona, 2022. ESMO. Open. 2023, 8, 101567.
- Llovet, J. M.; de,B.T.; Kulik,L.; Haber,P.K.; Greten,T.F.; Meyer,T.; Lencioni,R. Locoregional therapies in the era of molecular and immune treatments for hepatocellular carcinoma. Nat. Rev Gastroenterol. Hepatol. 2021, 18, 293-313. [CrossRef]
- Hanna, R. F.; Miloushev,V.Z.; Tang,A.; Finklestone,L.A.; Brejt,S.Z.; Sandhu,R.S.; Santillan,C.S.; Wolfson,T.; Gamst,A.; Sirlin,C.B. Comparative 13-year meta-analysis of the sensitivity and positive predictive value of ultrasound, CT, and MRI for detecting hepatocellular carcinoma. Abdom. Radiol. (NY). 2016, 41, 71-90. [CrossRef]
- Chen, D. S.; Sung,J.L.; Sheu,J.C.; Lai,M.Y.; How,S.W.; Hsu,H.C.; Lee,C.S.; Wei,T.C. Serum alpha-fetoprotein in the early stage of human hepatocellular carcinoma. Gastroenterology. 1984, 86, 1404-1409. [CrossRef]
- Tzartzeva, K.; Obi,J.; Rich,N.E.; Parikh,N.D.; Marrero,J.A.; Yopp,A.; Waljee,A.K.; Singal,A.G. Surveillance Imaging and Alpha Fetoprotein for Early Detection of Hepatocellular Carcinoma in Patients With Cirrhosis: A Meta-analysis. Gastroenterology. 2018, 154, 1706-1718. [CrossRef]
- Yu, N. C.; Chaudhari,V.; Raman,S.S.; Lassman,C.; Tong,M.J.; Busuttil,R.W.; Lu,D.S. CT and MRI improve detection of hepatocellular carcinoma, compared with ultrasound alone, in patients with cirrhosis. Clin. Gastroenterol. Hepatol. 2011, 9, 161-167. [CrossRef]
- Gupta, P.; Soundararajan,R.; Patel,A.; Kumar,M.; Sharma,V.; Kalra,N. Abbreviated MRI for hepatocellular carcinoma screening: A systematic review and meta-analysis. J Hepatol. 2021, 75, 108-119. [CrossRef]
- Lehrich, B. M.; Zhang,J.; Monga,S.P.; Dhanasekaran,R. Battle of the biopsies: Role of tissue and liquid biopsy in hepatocellular carcinoma. J Hepatol. 2024, 80, 515-530. [CrossRef]
- Piccinino, F.; Sagnelli,E.; Pasquale,G.; Giusti,G. Complications following percutaneous liver biopsy. A multicentre retrospective study on 68,276 biopsies. J Hepatol. 1986, 2, 165-173. [CrossRef]
- Scott, E. C.; Baines,A.C.; Gong,Y.; Moore,R., Jr.; Pamuk,G.E.; Saber,H.; Subedee,A.; Thompson,M.D.; Xiao,W.; Pazdur,R.; Rao,V.A.; Schneider,J.; Beaver,J.A. Trends in the approval of cancer therapies by the FDA in the twenty-first century. Nat. Rev Drug Discov. 2023, 22, 625-640. [CrossRef]
- Tayob, N.; Kanwal,F.; Alsarraj,A.; Hernaez,R.; El-Serag,H.B. The Performance of AFP, AFP-3, DCP as Biomarkers for Detection of Hepatocellular Carcinoma (HCC): A Phase 3 Biomarker Study in the United States. Clin. Gastroenterol. Hepatol. 2023, 21, 415-423. [CrossRef]
- Cui, K.; Ou,Y.; Shen,Y.; Li,S.; Sun,Z. Clinical value of circulating tumor cells for the diagnosis and prognosis of hepatocellular carcinoma (HCC): A systematic review and meta-analysis. Medicine (Baltimore). 2020, 99, e22242.
- Tao, K.; Bian,Z.; Zhang,Q.; Guo,X.; Yin,C.; Wang,Y.; Zhou,K.; Wan,S.; Shi,M.; Bao,D.; Yang,C.; Xing,J. Machine learning-based genome-wide interrogation of somatic copy number aberrations in circulating tumor DNA for early detection of hepatocellular carcinoma. EBioMedicine. 2020, 56, 102811. [CrossRef]
- Nemeth, K.; Bayraktar,R.; Ferracin,M.; Calin,G.A. Non-coding RNAs in disease: from mechanisms to therapeutics. Nat. Rev Genet. 2024, 25, 211-232. [CrossRef]
- Uszczynska-Ratajczak, B.; Lagarde,J.; Frankish,A.; Guigo,R.; Johnson,R. Towards a complete map of the human long non-coding RNA transcriptome. Nat. Rev Genet. 2018, 19, 535-548. [CrossRef]
- Elbashir, S. M.; Harborth,J.; Lendeckel,W.; Yalcin,A.; Weber,K.; Tuschl,T. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature. 2001, 411, 494-498. [CrossRef]
- Kozomara, A.; Birgaoanu,M.; Griffiths-Jones,S. miRBase: from microRNA sequences to function. Nucleic Acids Res. 2019, 47, D155-D162. [CrossRef]
- Lee, Y.; Jeon,K.; Lee,J.T.; Kim,S.; Kim,V.N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 2002, 21, 4663-4670. [CrossRef]
- Aukerman, M. J. and Sakai,H. Regulation of flowering time and floral organ identity by a MicroRNA and its APETALA2-like target genes. Plant Cell. 2003, 15, 2730-2741. [CrossRef]
- Carthew, R. W. and Sontheimer,E.J. Origins and Mechanisms of miRNAs and siRNAs. Cell. 2009, 136, 642-655. [CrossRef]
- Bracht, J.; Hunter,S.; Eachus,R.; Weeks,P.; Pasquinelli,A.E. Trans-splicing and polyadenylation of let-7 microRNA primary transcripts. RNA. 2004, 10, 1586-1594. [CrossRef]
- Lee, Y.; Ahn,C.; Han,J.; Choi,H.; Kim,J.; Yim,J.; Lee,J.; Provost,P.; Radmark,O.; Kim,S.; Kim,V.N. The nuclear RNase III Drosha initiates microRNA processing. Nature. 2003, 425, 415-419. [CrossRef]
- Ray, A.; Sarkar,A.; Banerjee,S.; Biswas,K. Non-Canonical Targets of MicroRNAs: Role in Transcriptional Regulation, Disease Pathogenesis and Potential for Therapeutic Targets. Microrna. 2024. [CrossRef]
- Vasudevan, S. Posttranscriptional upregulation by microRNAs. Wiley. Interdiscip. Rev RNA. 2012, 3, 311-330. [CrossRef]
- Bartel, D. P. Metazoan MicroRNAs. Cell. 2018, 173, 20-51.
- Kumar, M. A.; Baba,S.K.; Sadida,H.Q.; Marzooqi,S.A.; Jerobin,J.; Altemani,F.H.; Algehainy,N.; Alanazi,M.A.; Abou-Samra,A.B.; Kumar,R.; Al-Shabeeb Akil,A.S.; Macha,M.A.; Mir,R.; Bhat,A.A. Extracellular vesicles as tools and targets in therapy for diseases. Signal Transduct. Target Ther. 2024, 9, 27. [CrossRef]
- Ladd, A. D.; Duarte,S.; Sahin,I.; Zarrinpar,A. Mechanisms of drug resistance in HCC. Hepatology. 2024, 79, 926-940. [CrossRef]
- Tian, Y.; Zhang,M.; Liu,L.X.; Wang,Z.C.; Liu,B.; Huang,Y.; Wang,X.; Ling,Y.Z.; Wang,F.; Feng,X.; Tu,Y. Exploring non-coding RNA mechanisms in hepatocellular carcinoma: implications for therapy and prognosis. Front Immunol. 2024, 15, 1400744. [CrossRef]
- Mallela, V. R.; Rajtmajerova,M.; Trailin,A.; Liska,V.; Hemminki,K.; Ambrozkiewicz,F. miRNA and lncRNA as potential tissue biomarkers in hepatocellular carcinoma. Noncoding. RNA. Res. 2024, 9, 24-32. [CrossRef]
- Jesenko, T.; Brezar,S.K.; Cemazar,M.; Biasin,A.; Tierno,D.; Scaggiante,B.; Grassi,M.; Grassi,C.; Dapas,B.; Truong,N.H.; Abrami,M.; Zanconati,F.; Bonazza,D.; Rizzolio,F.; Parisi,S.; Pastorin,G.; Grassi,G. Targeting Non-Coding RNAs for the Development of Novel Hepatocellular Carcinoma Therapeutic Approaches. Pharmaceutics. 2023, 15.
- Quinn, J. J. and Chang,H.Y. Unique features of long non-coding RNA biogenesis and function. Nat. Rev Genet. 2016, 17, 47-62. [CrossRef]
- Brannan, C. I.; Dees,E.C.; Ingram,R.S.; Tilghman,S.M. The product of the H19 gene may function as an RNA. Mol. Cell Biol. 1990, 10, 28-36.
- Zhao, L.; Wang,J.; Li,Y.; Song,T.; Wu,Y.; Fang,S.; Bu,D.; Li,H.; Sun,L.; Pei,D.; Zheng,Y.; Huang,J.; Xu,M.; Chen,R.; Zhao,Y.; He,S. NONCODEV6: an updated database dedicated to long non-coding RNA annotation in both animals and plants. Nucleic Acids Res. 2021, 49, D165-D171. [CrossRef]
- Derrien, T.; Johnson,R.; Bussotti,G.; Tanzer,A.; Djebali,S.; Tilgner,H.; Guernec,G.; Martin,D.; Merkel,A.; Knowles,D.G.; Lagarde,J.; Veeravalli,L.; Ruan,X.; Ruan,Y.; Lassmann,T.; Carninci,P.; Brown,J.B.; Lipovich,L.; Gonzalez,J.M.; Thomas,M.; Davis,C.A.; Shiekhattar,R.; Gingeras,T.R.; Hubbard,T.J.; Notredame,C.; Harrow,J.; Guigo,R. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012, 22, 1775-1789.
- Wu, H.; Yang,L.; Chen,L.L. The Diversity of Long Noncoding RNAs and Their Generation. Trends Genet. 2017, 33, 540-552. [CrossRef]
- He, J.; Huang,B.; Zhang,K.; Liu,M.; Xu,T. Long non-coding RNA in cervical cancer: From biology to therapeutic opportunity. Biomed. Pharmacother. 2020, 127, 110209. [CrossRef]
- Esteller, M. Non-coding RNAs in human disease. Nat. Rev Genet. 2011, 12, 861-874. [CrossRef]
- Chen, L. L. The biogenesis and emerging roles of circular RNAs. Nat. Rev Mol. Cell Biol. 2016, 17, 205-211. [CrossRef]
- Li, X.; Yang,L.; Chen,L.L. The Biogenesis, Functions, and Challenges of Circular RNAs. Mol. Cell. 2018, 71, 428-442. [CrossRef]
- Liang, D. and Wilusz,J.E. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 2014, 28, 2233-2247. [CrossRef]
- Pisignano, G.; Michael,D.C.; Visal,T.H.; Pirlog,R.; Ladomery,M.; Calin,G.A. Going circular: history, present, and future of circRNAs in cancer. Oncogene. 2023, 42, 2783-2800. [CrossRef]
- Li, Z.; Huang,C.; Bao,C.; Chen,L.; Lin,M.; Wang,X.; Zhong,G.; Yu,B.; Hu,W.; Dai,L.; Zhu,P.; Chang,Z.; Wu,Q.; Zhao,Y.; Jia,Y.; Xu,P.; Liu,H.; Shan,G. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256-264. [CrossRef]
- Czubak, K.; Sedehizadeh,S.; Kozlowski,P.; Wojciechowska,M. An Overview of Circular RNAs and Their Implications in Myotonic Dystrophy. Int. J Mol. Sci. 2019, 20. [CrossRef]
- Rybak-Wolf, A.; Stottmeister,C.; Glazar,P.; Jens,M.; Pino,N.; Giusti,S.; Hanan,M.; Behm,M.; Bartok,O.; Ashwal-Fluss,R.; Herzog,M.; Schreyer,L.; Papavasileiou,P.; Ivanov,A.; Ohman,M.; Refojo,D.; Kadener,S.; Rajewsky,N. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Mol. Cell. 2015, 58, 870-885.
- Zhang, X. O.; Wang,H.B.; Zhang,Y.; Lu,X.; Chen,L.L.; Yang,L. Complementary sequence-mediated exon circularization. Cell. 2014, 159, 134-147. [CrossRef]
- Memczak, S.; Jens,M.; Elefsinioti,A.; Torti,F.; Krueger,J.; Rybak,A.; Maier,L.; Mackowiak,S.D.; Gregersen,L.H.; Munschauer,M.; Loewer,A.; Ziebold,U.; Landthaler,M.; Kocks,C.; le,N.F.; Rajewsky,N. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013, 495, 333-338. [CrossRef]
- Salzman, J.; Chen,R.E.; Olsen,M.N.; Wang,P.L.; Brown,P.O. Cell-type specific features of circular RNA expression. PLoS. Genet. 2013, 9, e1003777. [CrossRef]
- Guo, T.; Tang,X.H.; Gao,X.Y.; Zhou,Y.; Jin,B.; Deng,Z.Q.; Hu,Y.; Xing,X.F.; Li,Z.Y.; Ji,J.F. A liquid biopsy signature of circulating exosome-derived mRNAs, miRNAs and lncRNAs predict therapeutic efficacy to neoadjuvant chemotherapy in patients with advanced gastric cancer. Mol. Cancer. 2022, 21, 216. [CrossRef]
- Shi, M.; Jia,J.S.; Gao,G.S.; Hua,X. Advances and challenges of exosome-derived noncoding RNAs for hepatocellular carcinoma diagnosis and treatment. Biochem. Biophys. Rep. 2024, 38, 101695. [CrossRef]
- Johnstone, R. M.; Adam,M.; Hammond,J.R.; Orr,L.; Turbide,C. Vesicle formation during reticulocyte maturation. Association of plasma membrane activities with released vesicles (exosomes). J Biol. Chem. 1987, 262, 9412-9420.
- Raposo, G.; Nijman,H.W.; Stoorvogel,W.; Liejendekker,R.; Harding,C.V.; Melief,C.J.; Geuze,H.J. B lymphocytes secrete antigen-presenting vesicles. J Exp. Med. 1996, 183, 1161-1172. [CrossRef]
- Valadi, H.; Ekstrom,K.; Bossios,A.; Sjostrand,M.; Lee,J.J.; Lotvall,J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654-659. [CrossRef]
- Hood, J. L.; San,R.S.; Wickline,S.A. Exosomes released by melanoma cells prepare sentinel lymph nodes for tumor metastasis. Cancer Res. 2011, 71, 3792-3801. [CrossRef]
- Fang, T.; Lv,H.; Lv,G.; Li,T.; Wang,C.; Han,Q.; Yu,L.; Su,B.; Guo,L.; Huang,S.; Cao,D.; Tang,L.; Tang,S.; Wu,M.; Yang,W.; Wang,H. Tumor-derived exosomal miR-1247-3p induces cancer-associated fibroblast activation to foster lung metastasis of liver cancer. Nat. Commun. 2018, 9, 191. [CrossRef]
- Zhou, Y.; Ren,H.; Dai,B.; Li,J.; Shang,L.; Huang,J.; Shi,X. Hepatocellular carcinoma-derived exosomal miRNA-21 contributes to tumor progression by converting hepatocyte stellate cells to cancer-associated fibroblasts. J Exp. Clin. Cancer Res. 2018, 37, 324. [CrossRef]
- Ma, D.; Gao,X.; Liu,Z.; Lu,X.; Ju,H.; Zhang,N. Exosome-transferred long non-coding RNA ASMTL-AS1 contributes to malignant phenotypes in residual hepatocellular carcinoma after insufficient radiofrequency ablation. Cell Prolif. 2020, 53, e12795. [CrossRef]
- Li, X.; Lei,Y.; Wu,M.; Li,N. Regulation of Macrophage Activation and Polarization by HCC-Derived Exosomal lncRNA TUC339. Int. J Mol. Sci. 2018, 19. [CrossRef]
- Chen, W.; Quan,Y.; Fan,S.; Wang,H.; Liang,J.; Huang,L.; Chen,L.; Liu,Q.; He,P.; Ye,Y. Exosome-transmitted circular RNA hsa_circ_0051443 suppresses hepatocellular carcinoma progression. Cancer Lett. 2020, 475, 119-128. [CrossRef]
- Fang, Y.; Zhang,Q.; Chen,Z.; Guo,C.; Wu,J. Clinical significance and immune characteristics analysis of miR-221-3p and its key target genes related to epithelial-mesenchymal transition in breast cancer. Aging (Albany. NY). 2024, 16, 322-347. [CrossRef]
- Eslaminejad, T.; Nematollahi-Mahani,S.N.; Sargazi,M.L.; Ansari,M.; Mirzaie,V. Evaluating the effects of curcumin nano-chitosan on miR-221 and miR-222 expression and Wnt/beta-catenin pathways in MCF-7, MDA-MB-231 and SKBR3 cell lines. Diagn. Pathol. 2024, 19, 35.
- Hu, G.; Zhai,S.; Yu,S.; Huang,Z.; Gao,R. Circular RNA circRHOBTB3 is downregulated in hepatocellular carcinoma and suppresses cell proliferation by inhibiting miR-18a maturation. Infect. Agent. Cancer. 2021, 16, 48. [CrossRef]
- Yun, J. H.; Baek,M.J.; Jung,H.I. Expression of miR-221 and miR-18a in patients with hepatocellular carcinoma and its clinical significance. Korean J Clin. Oncol. 2022, 18, 17-26. [CrossRef]
- Chen, F.; Li,X.F.; Fu,D.S.; Huang,J.G.; Yang,S.E. Clinical potential of miRNA-221 as a novel prognostic biomarker for hepatocellular carcinoma. Cancer Biomark. 2017, 18, 209-214. [CrossRef]
- Yousuf, T.; Dar,S.B.; Bangri,S.A.; Choh,N.A.; Rasool,Z.; Shah,A.; Rather,R.A.; Rah,B.; Bhat,G.R.; Ali,S.; Afroze,D. Diagnostic implication of a circulating serum-based three-microRNA signature in hepatocellular carcinoma. Front Genet. 2022, 13, 929787.
- Liu, W.; Hu,K.; Zhang,F.; Lu,S.; Chen,R.; Ren,Z.; Yin,X. The prognostic significance of microRNA-221 in hepatocellular carcinoma: An updated meta-analysis. Int. J Biol. Markers. 2021, 36, 17246008211032689. [CrossRef]
- Yang, X.; Wang,M.; Lin,B.; Yao,D.; Li,J.; Tang,X.; Li,S.; Liu,Y.; Xie,R.; Yu,S. miR-487a promotes progression of gastric cancer by targeting TIA1. Biochimie. 2018, 154, 119-126.
- Luo, A.; Liu,H.; Huang,C. KLF5-induced miR-487a augments the progression of osteosarcoma cells by targeting NKX3-1 in vitro. Oncol. Lett. 2022, 24, 258.
- Chang, R. M.; Xiao,S.; Lei,X.; Yang,H.; Fang,F.; Yang,L.Y. miRNA-487a Promotes Proliferation and Metastasis in Hepatocellular Carcinoma. Clin. Cancer Res. 2017, 23, 2593-2604. [CrossRef]
- Cirera-Salinas, D.; Pauta,M.; Allen,R.M.; Salerno,A.G.; Ramirez,C.M.; Chamorro-Jorganes,A.; Wanschel,A.C.; Lasuncion,M.A.; Morales-Ruiz,M.; Suarez,Y.; Baldan,A.; Esplugues,E.; Fernandez-Hernando,C. Mir-33 regulates cell proliferation and cell cycle progression. Cell Cycle. 2012, 11, 922-933. [CrossRef]
- Najafi-Shoushtari, S. H.; Kristo,F.; Li,Y.; Shioda,T.; Cohen,D.E.; Gerszten,R.E.; Naar,A.M. MicroRNA-33 and the SREBP host genes cooperate to control cholesterol homeostasis. Science. 2010, 328, 1566-1569. [CrossRef]
- Ramirez, C. M.; Goedeke,L.; Rotllan,N.; Yoon,J.H.; Cirera-Salinas,D.; Mattison,J.A.; Suarez,Y.; de,C.R.; Gorospe,M.; Fernandez-Hernando,C. MicroRNA 33 regulates glucose metabolism. Mol. Cell Biol. 2013, 33, 2891-2902. [CrossRef]
- Zhang, T.; Han,G.; Wang,Y.; Chen,K.; Sun,Y. MicroRNA expression profiles in supraglottic carcinoma. Oncol. Rep. 2014, 31, 2029-2034. [CrossRef]
- Zhou, Y.; Huang,Z.; Wu,S.; Zang,X.; Liu,M.; Shi,J. miR-33a is up-regulated in chemoresistant osteosarcoma and promotes osteosarcoma cell resistance to cisplatin by down-regulating TWIST. J Exp. Clin. Cancer Res. 2014, 33, 12. [CrossRef]
- Huang, C. F.; Sun,C.C.; Zhao,F.; Zhang,Y.D.; Li,D.J. miR-33a levels in hepatic and serum after chronic HBV-induced fibrosis. J Gastroenterol. 2015, 50, 480-490. [CrossRef]
- Xie, R. T.; Cong,X.L.; Zhong,X.M.; Luo,P.; Yang,H.Q.; Lu,G.X.; Luo,P.; Chang,Z.Y.; Sun,R.; Wu,T.M.; Lv,Z.W.; Fu,D.; Ma,Y.S. MicroRNA-33a downregulation is associated with tumorigenesis and poor prognosis in patients with hepatocellular carcinoma. Oncol. Lett. 2018, 15, 4571-4577. [CrossRef]
- Mangiapane, G.; Pascut,D.; Dalla,E.; Antoniali,G.; Degrassi,M.; Croce,L.S.; De,S., V; Piazza,S.; Canarutto,G.; Tiribelli,C.; Tell,G. Clinical Significance of Apurinic/Apyrimidinic Endodeoxyribonuclease 1 and MicroRNA Axis in Hepatocellular Carcinoma. J Clin. Transl. Hepatol. 2023, 11, 1291-1307.
- Mourelatos, Z.; Dostie,J.; Paushkin,S.; Sharma,A.; Charroux,B.; Abel,L.; Rappsilber,J.; Mann,M.; Dreyfuss,G. miRNPs: a novel class of ribonucleoproteins containing numerous microRNAs. Genes Dev. 2002, 16, 720-728.
- Kou, X.; Ding,H.; Li,L.; Chao,H. Hsa-miR-105-1 Regulates Cisplatin-Resistance in Ovarian Carcinoma Cells by Targeting ANXA9. Anal. Cell Pathol. (Amst). 2021, 2021, 6662486.
- Zhou, W.; Fong,M.Y.; Min,Y.; Somlo,G.; Liu,L.; Palomares,M.R.; Yu,Y.; Chow,A.; O’Connor,S.T.; Chin,A.R.; Yen,Y.; Wang,Y.; Marcusson,E.G.; Chu,P.; Wu,J.; Wu,X.; Li,A.X.; Li,Z.; Gao,H.; Ren,X.; Boldin,M.P.; Lin,P.C.; Wang,S.E. Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell. 2014, 25, 501-515.
- Guan, Y.; Chen,L.; Bao,Y.; Li,Z.; Cui,R.; Li,G.; Wang,Y. Identification of low miR-105 expression as a novel poor prognostic predictor for human glioma. Int. J Clin. Exp. Med. 2015, 8, 10855-10864.
- Shen, G.; Rong,X.; Zhao,J.; Yang,X.; Li,H.; Jiang,H.; Zhou,Q.; Ji,T.; Huang,S.; Zhang,J.; Jia,H. MicroRNA-105 suppresses cell proliferation and inhibits PI3K/AKT signaling in human hepatocellular carcinoma. Carcinogenesis. 2014, 35, 2748-2755. [CrossRef]
- Ma, Y. S.; Wu,T.M.; Lv,Z.W.; Lu,G.X.; Cong,X.L.; Xie,R.T.; Yang,H.Q.; Chang,Z.Y.; Sun,R.; Chai,L.; Cai,M.X.; Zhong,X.J.; Zhu,J.; Fu,D. High expression of miR-105-1 positively correlates with clinical prognosis of hepatocellular carcinoma by targeting oncogene NCOA1. Oncotarget. 2017, 8, 11896-11905. [CrossRef]
- Zhou, W.; Zhou,W.; Zeng,Q.; Xiong,J. MicroRNA-138 inhibits hypoxia-induced proliferation of endothelial progenitor cells via inhibition of HIF-1alpha-mediated MAPK and AKT signaling. Exp. Ther. Med. 2017, 13, 1017-1024.
- Liu, H.; Tang,T.; Hu,X.; Tan,W.; Zhou,P.; Zhang,H.; Liu,Y.; Chen,C.; Yang,M.; Zhou,M.; Xuan,S.; Cheng,B.; Yin,W.; Lin,J. miR-138-5p Inhibits Vascular Mimicry by Targeting the HIF-1alpha/VEGFA Pathway in Hepatocellular Carcinoma. J Immunol. Res. 2022, 2022, 7318950.
- Lu, Y.; Liu,Y.; Zhang,K.; Jiang,L. Circular RNA TLK1 Exerts Oncogenic Functions in Hepatocellular Carcinoma by Acting as a ceRNA of miR-138-5p. J Oncol. 2022, 2022, 2415836. [CrossRef]
- Zheng, J. H.; Jian,Z.X.; Jin,H.S.; Chen,S.C.; Wang,G.Y. [Expression of SOX4 gene and early recurrence of hepatocellular carcinoma: their relationship and the clinical significance]. Nan. Fang Yi. Ke. Da. Xue. Xue. Bao. 2010, 30, 818-819.
- Xiao, J. X.; Xu,W.; Fei,X.; Hao,F.; Wang,N.; Chen,Y.; Wang,J. Anillin facilitates cell proliferation and induces tumor growth of hepatocellular carcinoma via miR-138/SOX4 axis regulation. Transl. Oncol. 2020, 13, 100815.
- Jenike, A. E. and Halushka,M.K. miR-21: a non-specific biomarker of all maladies. Biomark. Res. 2021, 9, 18. [CrossRef]
- Mjelle, R.; Dima,S.O.; Bacalbasa,N.; Chawla,K.; Sorop,A.; Cucu,D.; Herlea,V.; Saetrom,P.; Popescu,I. Comprehensive transcriptomic analyses of tissue, serum, and serum exosomes from hepatocellular carcinoma patients. BMC. Cancer. 2019, 19, 1007.
- Stechele, M.; Link,H.; Hirner-Eppeneder,H.; Alunni-Fabbroni,M.; Wildgruber,M.; Salvermoser,L.; Corradini,S.; Schinner,R.; Ben,K.N.; Rossler,D.; Galun,E.; Goldberg,S.N.; Ricke,J.; Kazmierczak,P.M. Circulating miR-21 as a prognostic biomarker in HCC treated by CT-guided high-dose rate brachytherapy. Radiat. Oncol. 2023, 18, 125. [CrossRef]
- Huang, X.; Xiong,Y.; Yang,J.; Yang,G.; Li,J. The Prognostic Significance of miR-21 Expression among Surgically Resected Hepatocellular Carcinoma Patients: Evidence from a Meta-Analysis and Retrospective Cohort Study. Biomed. Res. Int. 2020, 2020, 8848158. [CrossRef]
- Zhang, N.; Hu,Z.; Qiang,Y.; Zhu,X. Circulating miR-130b- and miR-21-based diagnostic markers and therapeutic targets for hepatocellular carcinoma. Mol. Genet. Genomic. Med. 2019, 7, e1012.
- Li, Z. Y.; Xi,Y.; Zhu,W.N.; Zeng,C.; Zhang,Z.Q.; Guo,Z.C.; Hao,D.L.; Liu,G.; Feng,L.; Chen,H.Z.; Chen,F.; Lv,X.; Liu,D.P.; Liang,C.C. Positive regulation of hepatic miR-122 expression by HNF4alpha. J Hepatol. 2011, 55, 602-611. [CrossRef]
- Bandiera, S.; Pfeffer,S.; Baumert,T.F.; Zeisel,M.B. miR-122--a key factor and therapeutic target in liver disease. J Hepatol. 2015, 62, 448-457. [CrossRef]
- Hao, P.; Li,Q.; Zhao,H. Mucin 1 expression is regulated by hsa_circ_0055054/microRNA‑122‑5p and promotes hepatocellular carcinoma development. Oncol. Lett. 2024, 28, 404.
- Zhao, X. F.; Li,N.; Lin,D.D.; Sun,L.B. Circulating MicroRNA-122 for the Diagnosis of Hepatocellular Carcinoma: A Meta-Analysis. Biomed. Res. Int. 2020, 2020, 5353695. [CrossRef]
- Fang, Y.; Yan,D.; Wang,L.; Zhang,J.; He,Q. Circulating microRNAs (miR-16, miR-22, miR-122) expression and early diagnosis of hepatocellular carcinoma. J Clin. Lab Anal. 2022, 36, e24541.
- Pelizzaro, F.; Cardin,R.; Sartori,A.; Imondi,A.; Penzo,B.; Aliberti,C.; Ponzoni,A.; Vitale,A.; Cillo,U.; Farinati,F. Circulating MicroRNA-21 and MicroRNA-122 as Prognostic Biomarkers in Hepatocellular Carcinoma Patients Treated with Transarterial Chemoembolization. Biomedicines. 2021, 9. [CrossRef]
- Ambade, A.; Satishchandran,A.; Szabo,G. Alcoholic hepatitis accelerates early hepatobiliary cancer by increasing stemness and miR-122-mediated HIF-1alpha activation. Sci Rep. 2016, 6, 21340. [CrossRef]
- Krishnan, K.; Steptoe,A.L.; Martin,H.C.; Pattabiraman,D.R.; Nones,K.; Waddell,N.; Mariasegaram,M.; Simpson,P.T.; Lakhani,S.R.; Vlassov,A.; Grimmond,S.M.; Cloonan,N. miR-139-5p is a regulator of metastatic pathways in breast cancer. RNA. 2013, 19, 1767-1780. [CrossRef]
- Wong, C. C.; Wong,C.M.; Tung,E.K.; Au,S.L.; Lee,J.M.; Poon,R.T.; Man,K.; Ng,I.O. The microRNA miR-139 suppresses metastasis and progression of hepatocellular carcinoma by down-regulating Rho-kinase 2. Gastroenterology. 2011, 140, 322-331. [CrossRef]
- Croft, D. R. and Olson,M.F. The Rho GTPase effector ROCK regulates cyclin A, cyclin D1, and p27Kip1 levels by distinct mechanisms. Mol. Cell Biol. 2006, 26, 4612-4627. [CrossRef]
- Huang, D.; Du,X.; Yuan,R.; Chen,L.; Liu,T.; Wen,C.; Huang,M.; Li,M.; Hao,L.; Shao,J. Rock2 promotes the invasion and metastasis of hepatocellular carcinoma by modifying MMP2 ubiquitination and degradation. Biochem. Biophys. Res. Commun. 2014, 453, 49-56. [CrossRef]
- Wang, Z.; Ding,Q.; Li,Y.; Liu,Q.; Wu,W.; Wu,L.; Yu,H. Reanalysis of microRNA expression profiles identifies novel biomarkers for hepatocellular carcinoma prognosis. Tumour. Biol. 2016, 37, 14779-14787. [CrossRef]
- Wang, X.; Gao,J.; Zhou,B.; Xie,J.; Zhou,G.; Chen,Y. Identification of prognostic markers for hepatocellular carcinoma based on miRNA expression profiles. Life Sci. 2019, 232, 116596. [CrossRef]
- Tonon, F.; Cemazar,M.; Kamensek,U.; Zennaro,C.; Pozzato,G.; Caserta,S.; Ascione,F.; Grassi,M.; Guido,S.; Ferrari,C.; Cansolino,L.; Trotta,F.; Kuzmanov,B.G.; Forte,G.; Martino,F.; Perrone,F.; Bomben,R.; Gattei,V.; Elvassore,N.; Murano,E.; Truong,N.H.; Olson,M.; Farra,R.; Grassi,G.; Dapas,B. 5-Azacytidine Downregulates the Proliferation and Migration of Hepatocellular Carcinoma Cells In Vitro and In Vivo by Targeting miR-139-5p/ROCK2 Pathway. Cancers. (Basel). 2022, 14.
- Panzitt, K.; Tschernatsch,M.M.; Guelly,C.; Moustafa,T.; Stradner,M.; Strohmaier,H.M.; Buck,C.R.; Denk,H.; Schroeder,R.; Trauner,M.; Zatloukal,K. Characterization of HULC, a novel gene with striking up-regulation in hepatocellular carcinoma, as noncoding RNA. Gastroenterology. 2007, 132, 330-342. [CrossRef]
- Shah, M. and Sarkar,D. HCC-Related lncRNAs: Roles and Mechanisms. Int. J Mol. Sci. 2024, 25. [CrossRef]
- Lumkul, L.; Jantaree,P.; Jaisamak,K.; Wongkummool,W.; Lapisatepun,W.; Orrapin,S.; Udomruk,S.; Lo,P.L.; Chaiyawat,P. Combinatorial Gene Expression Profiling of Serum HULC, HOTAIR, and UCA1 lncRNAs to Differentiate Hepatocellular Carcinoma from Liver Diseases: A Systematic Review and Meta-Analysis. Int. J Mol. Sci. 2024, 25.
- Wang, B. G.; Lv,Z.; Ding,H.X.; Fang,X.X.; Wen,J.; Xu,Q.; Yuan,Y. The association of lncRNA-HULC polymorphisms with hepatocellular cancer risk and prognosis. Gene. 2018, 670, 148-154.
- Sonohara, F.; Inokawa,Y.; Hayashi,M.; Yamada,S.; Sugimoto,H.; Fujii,T.; Kodera,Y.; Nomoto,S. Prognostic Value of Long Non-Coding RNA HULC and MALAT1 Following the Curative Resection of Hepatocellular Carcinoma. Sci Rep. 2017, 7, 16142. [CrossRef]
- Yang, Z.; Lu,Y.; Xu,Q.; Tang,B.; Park,C.K.; Chen,X. HULC and H19 Played Different Roles in Overall and Disease-Free Survival from Hepatocellular Carcinoma after Curative Hepatectomy: A Preliminary Analysis from Gene Expression Omnibus. Dis. Markers. 2015, 2015, 191029. [CrossRef]
- Ji, P.; Diederichs,S.; Wang,W.; Boing,S.; Metzger,R.; Schneider,P.M.; Tidow,N.; Brandt,B.; Buerger,H.; Bulk,E.; Thomas,M.; Berdel,W.E.; Serve,H.; Muller-Tidow,C. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003, 22, 8031-8041.
- Golam, R. M.; Khalil,M.A.F.; Shaker,O.G.; Ahmed,T.I.; Elguaad,M.K.A.; Hassan,E.A.; El-Ansary,M.R.M.; Ismail,A.; Kandil,Y.I.; Mohammed,O.A.; Doghish,A.S. The clinical significance of long non-coding RNAs MALAT1 and CASC2 in the diagnosis of HCV-related hepatocellular carcinoma. PLoS. One. 2024, 19, e0303314. [CrossRef]
- Liao, X.; Chen,J.; Luo,D.; Luo,B.; Huang,W.; Xie,W. Prognostic value of long non-coding RNA MALAT1 in hepatocellular carcinoma: A study based on multi-omics analysis and RT-PCR validation. Pathol. Oncol. Res. 2022, 28, 1610808. [CrossRef]
- Li, J.; Wang,X.; Tang,J.; Jiang,R.; Zhang,W.; Ji,J.; Sun,B. HULC and Linc00152 Act as Novel Biomarkers in Predicting Diagnosis of Hepatocellular Carcinoma. Cell Physiol Biochem. 2015, 37, 687-696. [CrossRef]
- Liu, G.; Yang,Z.F.; Sun,J.; Sun,B.Y.; Zhou,P.Y.; Zhou,C.; Guan,R.Y.; Wang,Z.T.; Yi,Y.; Qiu,S.J. The LINC00152/miR-205-5p/CXCL11 axis in hepatocellular carcinoma cancer-associated fibroblasts affects cancer cell phenotypes and tumor growth. Cell Oncol. (Dordr. ). 2022, 45, 1435-1449.
- Wang, B.; Yang,S.; Zhao,W. Long Non-Coding RNA NRAD1 and LINC00152 are Highly Expressed and Associated with Prognosis in Patients with Hepatocellular Carcinoma. Onco. Targets. Ther. 2020, 13, 10409-10416. [CrossRef]
- Shehab-Eldeen, S.; Essa,A.; Arafat,E.S.; Sleem,A.S.; Alhosary,A.A.; Darwish,E.; Essa,A.; Al-Omair,O.A.; Al-Khoufi,E.A.; Al Abdulqader,A.K.; Nada,A. Serum LINC00152 and UCA1 in HCV-Induced Hepatocellular Carcinoma: Clinical Significance and Prognostic Value. Biologics. 2023, 17, 137-149. [CrossRef]
- Hajjari, M. and Salavaty,A. HOTAIR: an oncogenic long non-coding RNA in different cancers. Cancer Biol. Med. 2015, 12, 1-9.
- Ishibashi, M.; Kogo,R.; Shibata,K.; Sawada,G.; Takahashi,Y.; Kurashige,J.; Akiyoshi,S.; Sasaki,S.; Iwaya,T.; Sudo,T.; Sugimachi,K.; Mimori,K.; Wakabayashi,G.; Mori,M. Clinical significance of the expression of long non-coding RNA HOTAIR in primary hepatocellular carcinoma. Oncol. Rep. 2013, 29, 946-950. [CrossRef]
- Han, C.; Yang,Y.; Guo,L.; Guan,Q.; Ruan,S. The expression of long non-coding RNA HOTAIR in advanced hepatocellular carcinoma and its prognostic correlation with sunitinib therapy. Arch. Med. Sci. 2022, 18, 71-78. [CrossRef]
- Lou, Z. H.; Xu,K.Y.; Qiao,L.; Su,X.Q.; Ou-Yang,Y.; Miao,L.B.; Liu,F.; Wang,Y.; Fu,A.; Ren,X.H.; Huang,J.S.; Li,Z.Y.; Bao,J.F. Diagnostic Potential of the Serum lncRNAs HOTAIR, BRM and ICR for Hepatocellular Carcinoma. Front Biosci. (Landmark. Ed). 2022, 27, 264. [CrossRef]
- Wen, P.; Qi,X.; Zheng,R. Value of the HOTAIR expression assay in predicting therapy target in hepatocellular carcinoma: A meta-analysis and bioinformatics analysis. Int. J Biol. Markers. 2024, 39, 239-247. [CrossRef]
- Yu, T.; Wang,Y.; Fan,Y.; Fang,N.; Wang,T.; Xu,T.; Shu,Y. CircRNAs in cancer metabolism: a review. J Hematol. Oncol. 2019, 12, 90. [CrossRef]
- Zhang, X.; Zhou,H.; Jing,W.; Luo,P.; Qiu,S.; Liu,X.; Zhu,M.; Liang,C.; Yu,M.; Tu,J. The Circular RNA hsa_circ_0001445 Regulates the Proliferation and Migration of Hepatocellular Carcinoma and May Serve as a Diagnostic Biomarker. Dis. Markers. 2018, 2018, 3073467. [CrossRef]
- Xu, Q.; Zhou,L.; Yang,G.; Meng,F.; Wan,Y.; Wang,L.; Zhang,L. Overexpression of circ_0001445 decelerates hepatocellular carcinoma progression by regulating miR-942-5p/ALX4 axis. Biotechnol. Lett. 2020, 42, 2735-2747. [CrossRef]
- Shi, Y.; Sun,X.; He,X. Overexpression of Aristaless-Like Homeobox-4 Inhibits Proliferation, Invasion, and EMT in Hepatocellular Carcinoma Cells. Oncol. Res. 2017, 25, 11-18. [CrossRef]
- Wang, P.; Xu,L.L.; Zheng,X.B.; Hu,Y.T.; Zhang,J.F.; Ren,S.S.; Hao,X.Y.; Li,L.; Zhang,M.; Xu,M.Q. Correlation between the expressions of circular RNAs in peripheral venous blood and clinicopathological features in hepatocellular carcinoma. Ann. Transl. Med. 2020, 8, 338. [CrossRef]
- Chang, Z.; Song,Y.; Luo,F.; Yang,X.; Cai,Y.; Guo,H. Circular RNA SMARCA5 Promotes a Poor Prognosis and Radiotherapy Resistance for Patients with Hepatocellular Carcinoma. Ann. Clin. Lab Sci. 2023, 53, 573-577.
- Liu, T.; Song,Z.; Gai,Y. Circular RNA circ_0001649 acts as a prognostic biomarker and inhibits NSCLC progression via sponging miR-331-3p and miR-338-5p. Biochem. Biophys. Res. Commun. 2018, 503, 1503-1509. [CrossRef]
- Qin, M.; Liu,G.; Huo,X.; Tao,X.; Sun,X.; Ge,Z.; Yang,J.; Fan,J.; Liu,L.; Qin,W. Hsa_circ_0001649: A circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer Biomark. 2016, 16, 161-169. [CrossRef]
- Zhang, X.; Qiu,S.; Luo,P.; Zhou,H.; Jing,W.; Liang,C.; Tu,J. Down-regulation of hsa_circ_0001649 in hepatocellular carcinoma predicts a poor prognosis. Cancer Biomark. 2018, 22, 135-142. [CrossRef]
- Fu, L.; Wu,S.; Yao,T.; Chen,Q.; Xie,Y.; Ying,S.; Chen,Z.; Xiao,B.; Hu,Y. Decreased expression of hsa_circ_0003570 in hepatocellular carcinoma and its clinical significance. J Clin. Lab Anal. 2018, 32. [CrossRef]
- Zhang, X.; Chen,W.; Guo,D.; Li,Y.; Zhao,Y.; Ren,M.; Lu,G.; Lu,X.; He,S. Circ_0003570 Suppresses the progression of hepatocellular carcinoma through miR-182-5p/STARD13 regulatory axis. Biol. Proced. Online. 2022, 24, 14. [CrossRef]
- Jang, S. Y.; Kim,G.; Tak,W.Y.; Kweon,Y.O.; Lee,Y.R.; Han,Y.S.; Han,J.R.; Park,J.G.; Kang,M.K.; Lee,H.W.; Lee,W.K.; Park,S.Y.; Hur,K. Circular Noncoding RNA hsa_circ_0003570 as a Prognostic Biomarker for Hepatocellular Carcinoma. Genes (Basel). 2022, 13. [CrossRef]
- Kang, M. K.; Kim,G.; Park,J.G.; Jang,S.Y.; Lee,H.W.; Tak,W.Y.; Kweon,Y.O.; Park,S.Y.; Lee,Y.R.; Hur,K. Tissue Circular RNA_0004018 and 0003570 as Novel Prognostic Biomarkers for Hepatitis B-Related Hepatocellular Carcinoma. Genes (Basel). 2023, 14. [CrossRef]
- Jiang, Z.; Shen,L.; Wang,S.; Wu,S.; Hu,Y.; Guo,J.; Fu,L. Hsa_circ_0028502 and hsa_circ_0076251 are potential novel biomarkers for hepatocellular carcinoma. Cancer Med. 2019, 8, 7278-7287. [CrossRef]
- Liu, H.; Lan,T.; Li,H.; Xu,L.; Chen,X.; Liao,H.; Chen,X.; Du,J.; Cai,Y.; Wang,J.; Li,X.; Huang,J.; Yuan,K.; Zeng,Y. Circular RNA circDLC1 inhibits MMP1-mediated liver cancer progression via interaction with HuR. Theranostics. 2021, 11, 1396-1411. [CrossRef]
- Zhang, T.; Jing,B.; Bai,Y.; Zhang,Y.; Yu,H. Circular RNA circTMEM45A Acts as the Sponge of MicroRNA-665 to Promote Hepatocellular Carcinoma Progression. Mol. Ther. Nucleic Acids. 2020, 22, 285-297. [CrossRef]
- Martinez-Quetglas, I.; Pinyol,R.; Dauch,D.; Torrecilla,S.; Tovar,V.; Moeini,A.; Alsinet,C.; Portela,A.; Rodriguez-Carunchio,L.; Sole,M.; Lujambio,A.; Villanueva,A.; Thung,S.; Esteller,M.; Zender,L.; Llovet,J.M. IGF2 Is Up-regulated by Epigenetic Mechanisms in Hepatocellular Carcinomas and Is an Actionable Oncogene Product in Experimental Models. Gastroenterology. 2016, 151, 1192-1205. [CrossRef]
- Chen, D.; Zhang,C.; Lin,J.; Song,X.; Wang,H. Screening differential circular RNA expression profiles reveal that hsa_circ_0128298 is a biomarker in the diagnosis and prognosis of hepatocellular carcinoma. Cancer Manag. Res. 2018, 10, 1275-1283. [CrossRef]
| miRNA | Prognostic significance | Levels in HCC | Ref |
|---|---|---|---|
| miR-221 | Poor | Increased (tissue) Decreased (serum) |
[74,75,76,77] |
| miR-18-a | Poor | Increased (tissue) | [74] |
| miR-487a | Poor | Increased (tissue) | [80] |
| miR-33a | Poor/Good | Reduced (tissue) | [87,88] |
| miR-105-1 | Poor | Reduced (tissue) | [93,94] |
| miR-138-5p | Poor | Reduced (tissue) | [96,97,98] |
| miR-21 | Poor | Increased (Tissue/serum) | [101, 102,103, 104] |
| miR-122 | Poor | Decreased (tissue, serum) | [108,109] |
| miR-139-5p | Poor | Decreased (tissue) | [116,117,118] |
| lncRNA | Prognostic significance | Levels in HCC | References |
|---|---|---|---|
| HULC | Poor | Increased (serum, tissue) |
[119, 121, 122, 123, 128] |
| MALAT1 | Poor | Increased (serum/tissue) | [126,127] |
| Linc00152 | Poor | Increased (plasma, tissue) |
[128,130,131] |
| HOTAIR | Poor | Increased (tissue, serum) |
[133, 134, 135, 136] |
| circRNA | Prognostic significance | Levels in HCC | References |
|---|---|---|---|
| 0001445 | Poor | Decreased (tissue, plasma) Increased (tissue) |
[138,139,141] [142] |
| 0001649 | Poor | Decreased (tissue) | [144,145] |
| 0003570 | Poor | Decreased (tissue) | [146,147,149] |
| 0028502/0076251 | Poor | Decreased (tissue) |
[150] |
| CircDLC1 | Poor | Decreased (tissue) |
[151] |
| 0066659 | Poor | Increased (tissue) |
[152] |
| 0128298 | Poor | Increased (tissue) |
[154] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




