Submitted:
24 September 2024
Posted:
25 September 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
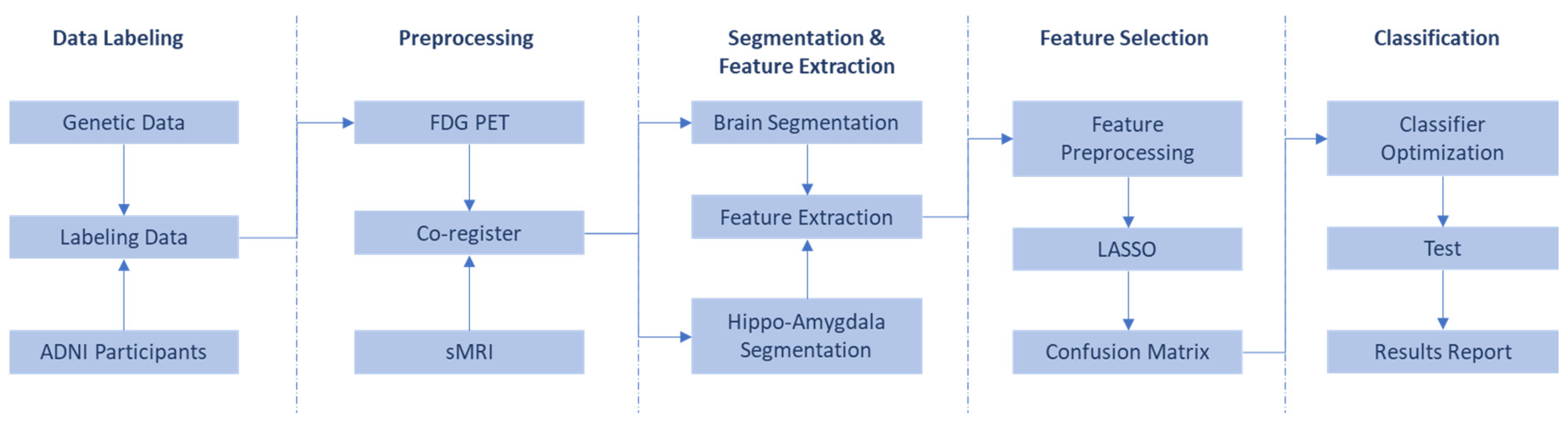
2. Method
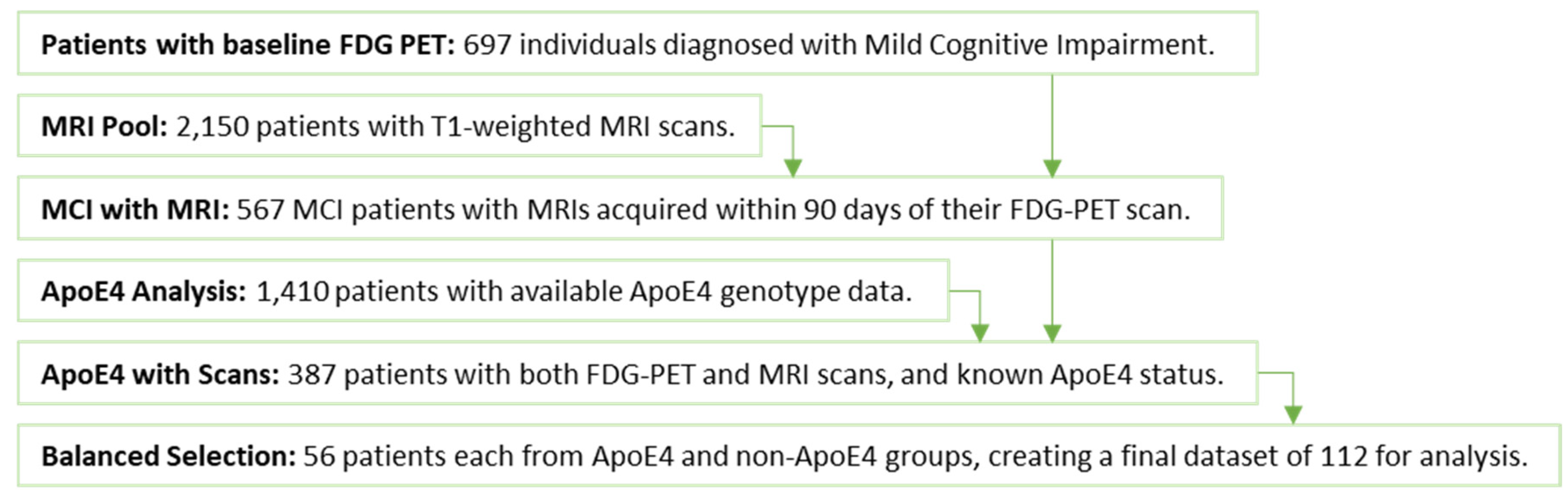
2.1. ADNI and Participants
2.2. ApoE Genotyping
2.3. FDG PET Acquisition
2.4. MRI Acquisition
2.5. Co-Registration
2.6. Segmentation
2.7. Feature Extraction
2.8. Feature Selection
2.9. Classification
3. Results
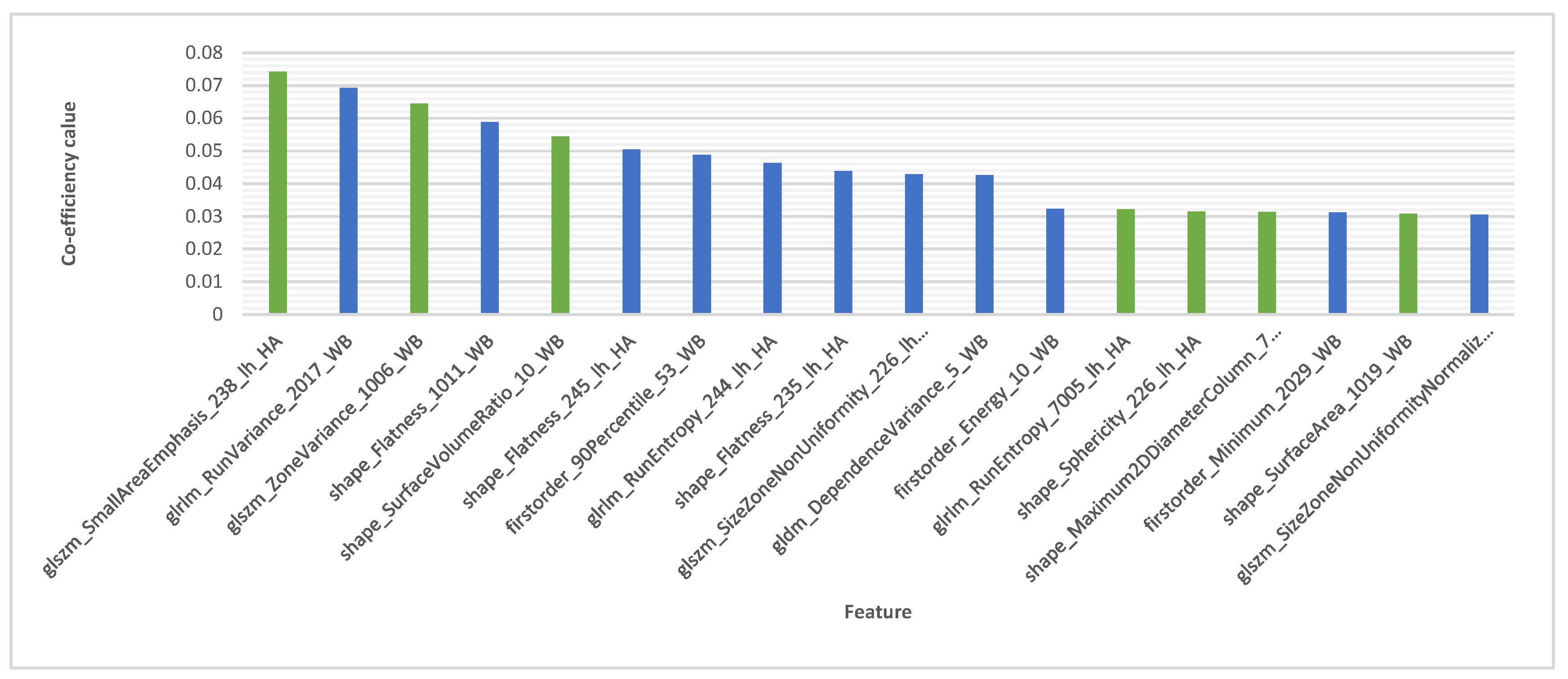
3.1. Feature Extraction
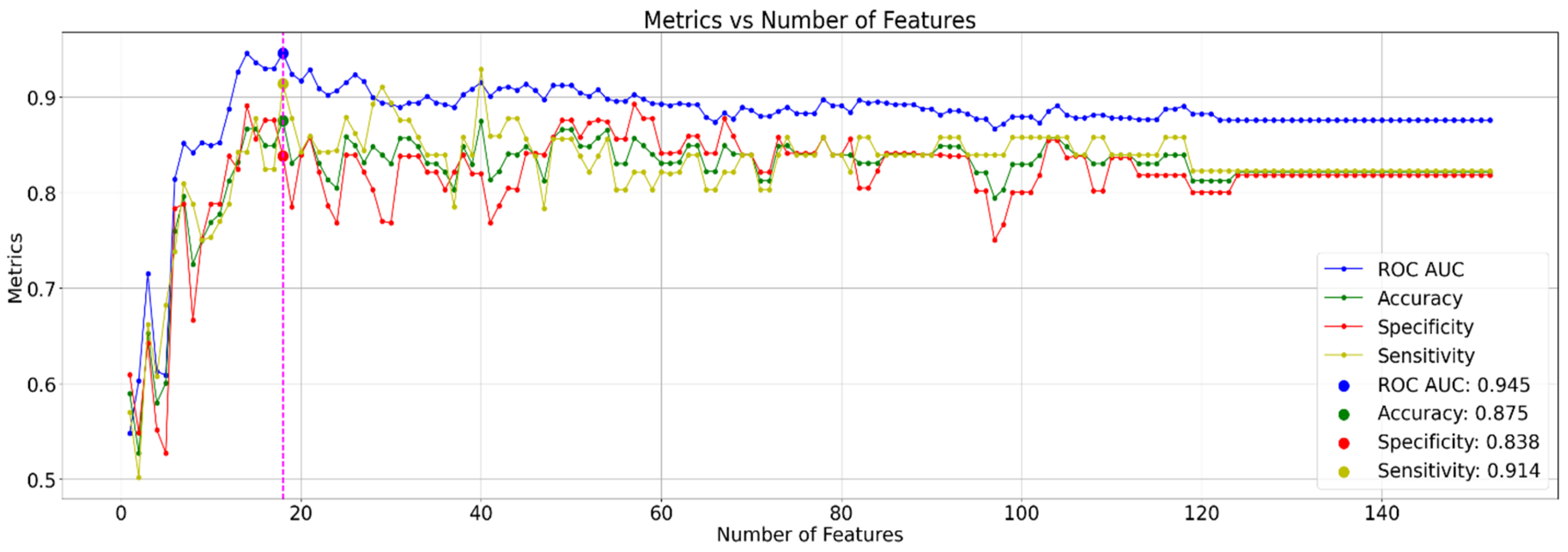
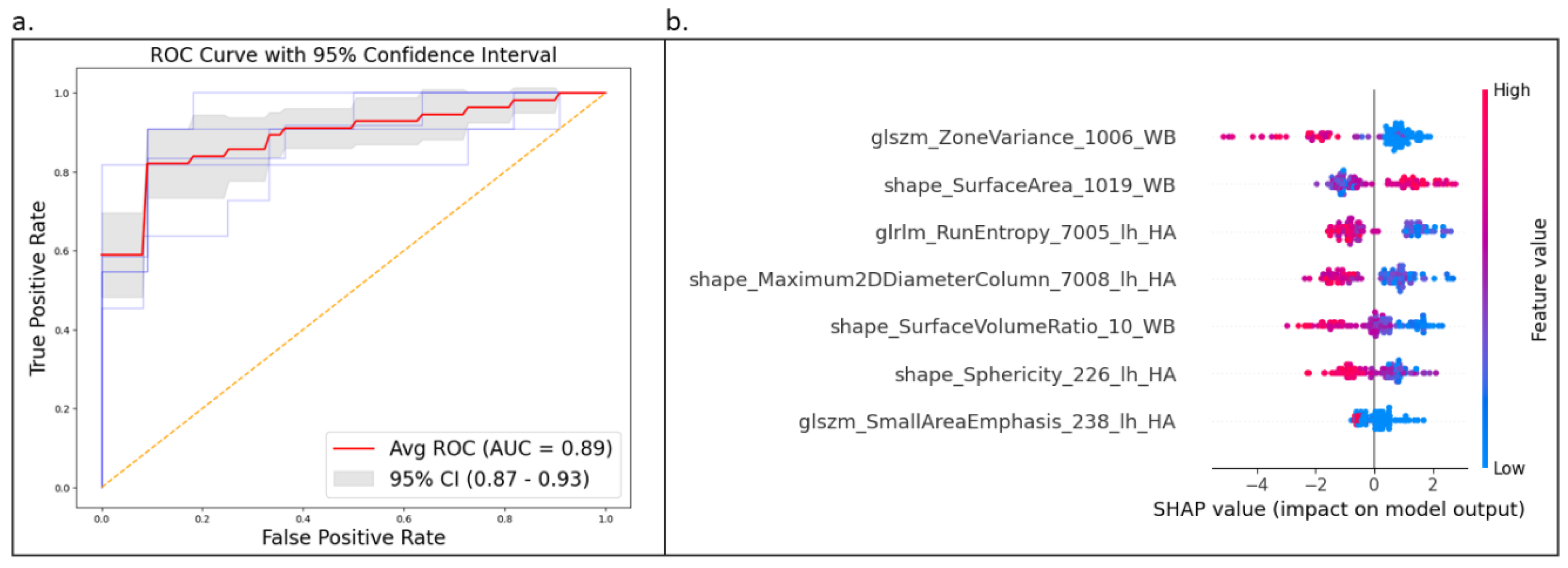
3.2. Classification
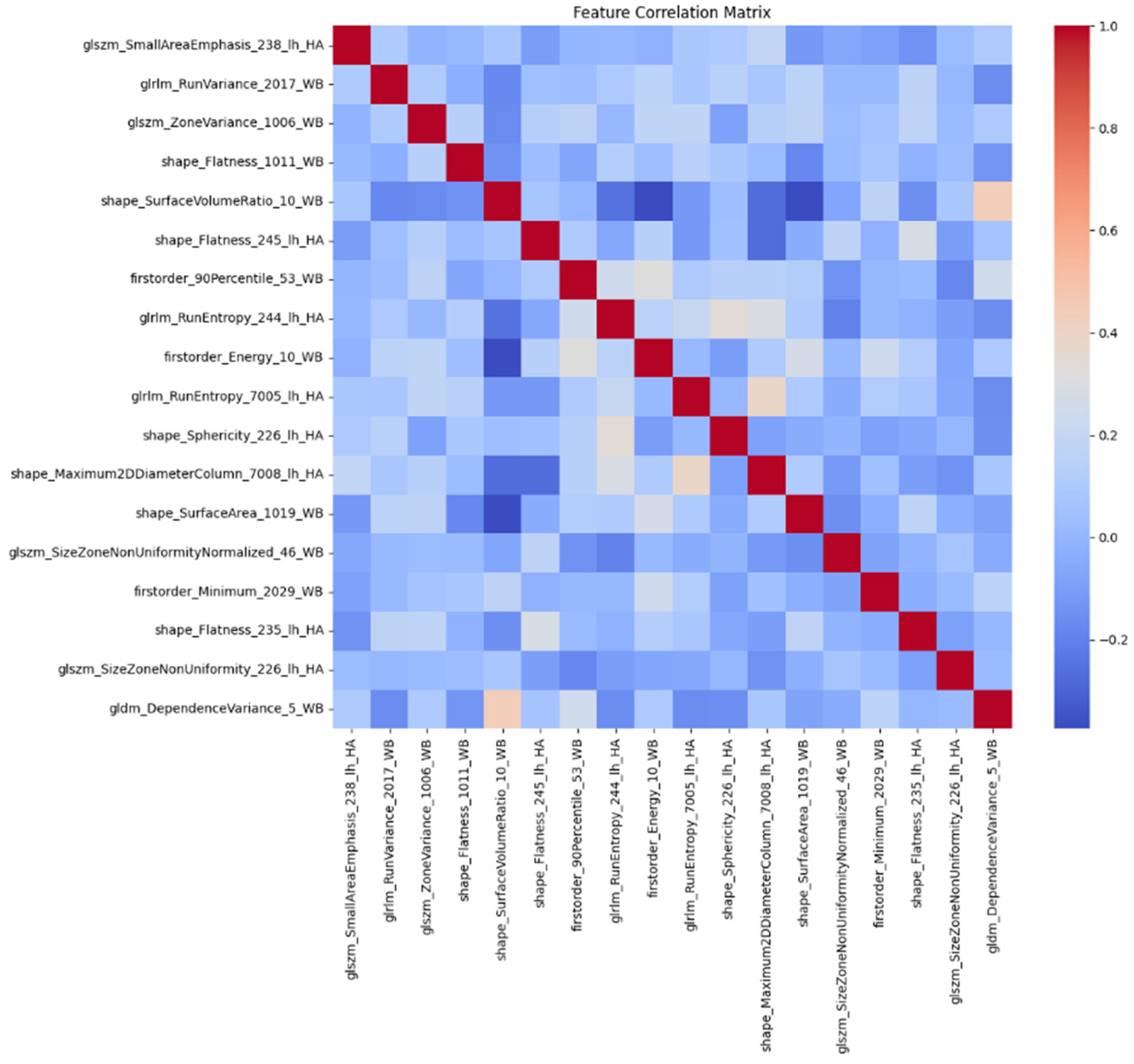
3.3. Additional Feature Refinement
3.4. Classification with Reduced Feature Set
4. Discussion
4.1. Findings
4.2. Explanation of the Relationship between Regions and APOE Genotype
4.3. Features Explanation
4.4. Interpretability
4.5. Clinical Significance
4.6. Limitations
4.7. Future study
5. Conclusions
Acknowledgments
References
- Grover, A.; Joshi, A. An Overview of Chronic Disease Models: A Systematic Literature Review. Glob J Health Sci. 2014, 7. [Google Scholar] [CrossRef] [PubMed]
- Chapman, D.P.; Williams, S.M.; Strine, T.W.; Anda, R.F.; Moore, M.J. Dementia and its implications for public health. Prev Chronic Dis. 2006, 3, A34. [Google Scholar]
- World Health Organization. Dementia: Key facts’. [Online]. Available online: https://www.who.int/news-room/fact-sheets/detail/dementia (accessed on 23 July 2024).
- ‘Dementia, vs. Alzheimer’s Disease: What Is the Difference? [Online]. Available online: https://www.alz.org/ (accessed on 23 July 2024).
- Battista, P.; Salvatore, C.; Berlingeri, M.; Cerasa, A.; Castiglioni, I. Artificial intelligence and neuropsychological measures: The case of Alzheimer’s disease. Neurosci Biobehav Rev. 2020, 114, 211–228. [Google Scholar] [CrossRef]
- Collin-Castonguay, P.; Gourdeau, D.; Potvin, O.; Duchesne, S. Synthetic FDG-PET hypometabolism sensitivity validation in AD. Alzheimer’s & Dementia 2023, 19. [Google Scholar] [CrossRef]
- Pereira, H.R.; Ferreira, H.A. Exploring the Potential of Radiomics Features of the Hippocampus in Alzheimer’s Disease Considering Standard versus Parallel Imaging. Alzheimer’s & Dementia 2023 2023, 19. [Google Scholar] [CrossRef]
- Varriano, G.; Guerriero, P.; Santone, A.; Mercaldo, F.; Brunese, L. Explainability of radiomics through formal methods. Comput Methods Programs Biomed. 2022, 220, 106824. [Google Scholar] [CrossRef] [PubMed]
- Rogers, W.; et al. Radiomics: from qualitative to quantitative imaging. Br J Radiol 2020, 93. [Google Scholar] [CrossRef] [PubMed]
- Rasi, R.; Guvenis, A. Predicting amyloid positivity from FDG-PET images using radiomics: A parsimonious model. Comput Methods Programs Biomed 2024, 247, 108098. [Google Scholar] [CrossRef]
- Yuksel, C.; Rasi, R.; Guvenis, A. A New Method for Diagnosing Alzheimer’s Disease and Monitoring Its Severity Using FDG-PET. In 2022 Medical Technologies Congress (TIPTEKNO); IEEE, 2022; pp. 1–4. [Google Scholar] [CrossRef]
- Rasi, R.; Güvenis, A. A Platform for the Radiomic Analysis of Brain FDG PET Images: Detecting Alzheimer’s Disease. In Proceedings of the Bioinformatics and Biomedical Engineering—10th International Work-Conference, IWBBIO 2023, Meloneras, Proceedings, Part I. Gran Canaria, Spain, 12–14 July 2023; Rojas, I., Valenzuela, O., Ruiz, F.R., Herrera, L.J., Ortuño, F.M., Eds.; Lecture Notes in Computer Science 1319. Springer; pp. 244–255. [Google Scholar] [CrossRef]
- Yi, L.X.; Zeng, L.; Wang, Q.; Tan, E.K.; Zhou, Z.D. Reelin links Apolipoprotein E4, Tau, and Amyloid-β in Alzheimer’s disease. Ageing Res Rev. 2024, 98, 102339. [Google Scholar] [CrossRef]
- Hampel, H.; et al. The Amyloid-β Pathway in Alzheimer’s Disease. Mol Psychiatry 2021, 26, 5481–5503. [Google Scholar] [CrossRef]
- Koutsodendris, N.; Nelson, M.R.; Rao, A.; Huang, Y. Apolipoprotein E and Alzheimer’s Disease: Findings, Hypotheses, and Potential Mechanisms. Annual Review of Pathology: Mechanisms of Disease 2022, 17, 73–99. [Google Scholar] [CrossRef] [PubMed]
- Corder, E.H.; et al. Gene Dose of Apolipoprotein E Type 4 Allele and the Risk of Alzheimer’s Disease in Late Onset Families. Science (1979) 1993, 261, 921–923. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.J.; Hyman, B.T.; Serrano-Pozo, A. Multifaceted roles of APOE in Alzheimer disease. Nat Rev Neurol 2024. [CrossRef] [PubMed]
- Chen, Y.; Strickland, M.R.; Soranno, A.; Holtzman, D.M. Apolipoprotein E: Structural Insights and Links to Alzheimer Disease Pathogenesis. Neuron 2021, 109, 205–221. [Google Scholar] [CrossRef] [PubMed]
- Genin, E.; et al. APOE and Alzheimer disease: a major gene with semi-dominant inheritance. Mol Psychiatry 2011, 16, 903–907. [Google Scholar] [CrossRef]
- Jagust, W.J.; et al. The Alzheimer’s Disease Neuroimaging Initiative 2 PET Core: 2015. Alzheimer’s & Dementia 2015, 11, 757–771. [Google Scholar] [CrossRef]
- Wyman, B.T.; et al. Standardization of analysis sets for reporting results from ADNI MRI data. Alzheimer’s & Dementia 2013, 9, 332–337. [Google Scholar] [CrossRef]
- Avants, B.; Epstein, C.; Grossman, M.; Gee, J. Symmetric diffeomorphic image registration with cross-correlation: Evaluating automated labeling of elderly and neurodegenerative brain. Med Image Anal 2008, 12, 26–41. [Google Scholar] [CrossRef]
- Fischl, B. FreeSurfer. Neuroimage 2012, 62, 774–781. [Google Scholar] [CrossRef]
- Kahhale, I.; Buser, N.J.; Madan, C.R.; Hanson, J.L. Quantifying numerical and spatial reliability of hippocampal and amygdala subdivisions in FreeSurfer. Brain Inform. 2023, 10, 9. [Google Scholar] [CrossRef]
- Sämann, P.G.; et al. <scp>FreeSurfer</scp> -based segmentation of hippocampal subfields: A review of methods and applications, with a novel quality control procedure for <scp>ENIGMA</scp> studies and other collaborative efforts. Hum Brain Mapp 2022, 43, 207–233. [Google Scholar] [CrossRef] [PubMed]
- Hatt, M.; Vallieres, M.; Visvikis, D.; Zwanenburg, A. IBSI: an international community radiomics standardization initiative. Soc Nuclear Med. 2018.
- van Griethuysen, J.J.M.; et al. Computational Radiomics System to Decode the Radiographic Phenotype. Cancer Res. 2017, 77, e104–e107. [Google Scholar] [CrossRef]
- Tibshirani, R. Regression Shrinkage and Selection Via the Lasso. J R Stat Soc Series B Stat Methodol 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Wieczorek, J.; Guerin, C.; McMahon, T. K -fold cross-validation for complex sample surveys. Stat 2022, 11. [Google Scholar] [CrossRef]
- Impagliazzo, R.; Lei, R.; Pitassi, T.; Sorrell, J. Reproducibility in learning. In Proceedings of the 54th Annual ACM SIGACT Symposium on Theory of Computing; ACM: New York, NY, USA, 2022; pp. 818–831. [Google Scholar] [CrossRef]
- O’Dwyer, L.; et al. Reduced Hippocampal Volume in Healthy Young ApoE4 Carriers: An MRI Study. PLoS ONE 2012, 7, e48895. [Google Scholar] [CrossRef]
- Pievani, M.; Galluzzi, S.; Thompson, P.M.; Rasser, P.E.; Bonetti, M.; Frisoni, G.B. APOE4 is associated with greater atrophy of the hippocampal formation in Alzheimer’s disease. Neuroimage 2011, 55, 909–919. [Google Scholar] [CrossRef]
- Miranda, A.M.; et al. Effects of APOE4 allelic dosage on lipidomic signatures in the entorhinal cortex of aged mice. Transl Psychiatry 2022, 12, 129. [Google Scholar] [CrossRef]
- La Joie, R.; et al. Association of APOE4 and Clinical Variability in Alzheimer Disease With the Pattern of Tau- and Amyloid-PET. Neurology 2021, 96. [Google Scholar] [CrossRef]
- Young, C.B.; et al. APOE effects on regional tau in preclinical Alzheimer’s disease. Mol Neurodegener 2023, 18, 1. [Google Scholar] [CrossRef]
- Novellino, F.; López, M.E.; Vaccaro, M.G.; Miguel, Y.; Delgado, M.L.; Maestu, F. Association Between Hippocampus, Thalamus, and Caudate in Mild Cognitive Impairment APOEε4 Carriers: A Structural Covariance MRI Study. Front Neurol 2019, 10. [Google Scholar] [CrossRef] [PubMed]

| Group | No. Case | Age | CDRSB | MMSE | ADAS13 | RAVLT Forgetting |
| (Male/ Female) | (mean ± SD) | (mean ± SD) | (mean ± SD) | (mean ± SD) | (mean ± SD) | |
| non-ε4 carrier | 33 / 23 | 71.34 ± 7.6 | 1.61 ± 1.14 | 27.64 ± 1.87 | 15.49 ± 7.03 | 4.29 ± 4.33 |
| Two ε4 carrier | 31 / 25 | 69.35 ± 6.66 | 1.5 ± 0.87 | 27.46 ± 2.08 | 16.65 ± 6.17 | 5.39 ± 2.5 |
| Total | 64 / 48 | 70.35 ± 7.2 | 1.55 ± 1.01 | 27.55 ± 1.97 | 16.07 ± 6.61 | 4.84 ± 3.57 |
| FS ID | Region (left/right hand) | Feature | homozygous ApoE4 carriers | non-ε4 carriers | P_Value | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| mean | SD | min | max | mean | SD | min | max | ||||
| 238 | CA1-body (Hippocampus) (lh) | glszm_SmallAreaEmphasis* | 0.3000 | 1.2963 | -0.3682 | 3.6735 | -0.3000 | 0.4157 | -0.3681 | 2.6631 | 0.0016 |
| 2017 | paracentral (rh) | glrlm_RunVariance | -0.1289 | 0.9366 | -2.3246 | 2.3972 | 0.1289 | 1.0608 | -2.0647 | 3.1278 | 0.1756 |
| 1006 | entorhinal (lh) | glszm_ZoneVariance* | 0.2701 | 1.1090 | -0.6391 | 2.7400 | -0.2701 | 0.8111 | -0.6391 | 3.4895 | 0.0040 |
| 1011 | lateraloccipital (lh) | shape_Flatness | 0.1819 | 1.0437 | -1.8295 | 3.0743 | -0.1819 | 0.9379 | -2.7474 | 2.1383 | 0.0549 |
| 10 | Thalamus (lh) | shape_SurfaceVolumeRatio* | 0.2874 | 1.0451 | -1.8291 | 2.9666 | -0.2874 | 0.8809 | -2.8339 | 2.2949 | 0.0021 |
| 245 | molecular_layer_HP-head (Hippocampus) (lh) | shape_Flatness | -0.2439 | 0.7830 | -1.8315 | 2.1238 | 0.2439 | 1.1411 | -1.7182 | 5.4304 | 0.0097 |
| 53 | Hippocampus (rh) | firstorder_90Percentile | 0.2082 | 0.9508 | -2.2811 | 2.8583 | -0.2082 | 1.0218 | -2.1932 | 2.3466 | 0.0276 |
| 244 | GC-ML-DG-body (Hippocampus) (lh) | glrlm_RunEntropy | 0.1656 | 1.0613 | -2.0553 | 2.6429 | -0.1656 | 0.9242 | -2.1335 | 1.5815 | 0.0811 |
| 235 | subiculum-head (Hippocampus) (lh) | shape_Flatness | -0.1804 | 0.9948 | -2.0349 | 2.6071 | 0.1804 | 0.9902 | -1.8604 | 2.5275 | 0.0571 |
| 226 | Hippocampal_tail (Hippocampus) (lh) | glszm_SizeZoneNonUniformity | 0.1602 | 1.2777 | -0.3627 | 6.7370 | -0.1602 | 0.5930 | -0.3627 | 1.6094 | 0.0926 |
| 5 | Inferior Lateral Ventricle (lh) | gldm_DependenceVariance | 0.2497 | 0.8670 | -3.1855 | 1.8362 | -0.2497 | 1.0760 | -4.4154 | 1.2877 | 0.0080 |
| 10 | Thalamus (lh) | firstorder_Energy | -0.1668 | 0.9803 | -1.8968 | 2.1634 | 0.1668 | 1.0093 | -3.4538 | 2.1225 | 0.0789 |
| 7005 | Central-nucleus (Amygdala) (lh) | glrlm_RunEntropy* | 0.3171 | 0.9355 | -1.8487 | 2.4870 | -0.3171 | 0.9780 | -2.3047 | 1.4931 | 0.0007 |
| 226 | Hippocampal_tail (Hippocampus) (lh) | shape_Sphericity* | 0.1124 | 1.0234 | -2.0888 | 1.7519 | -0.1124 | 0.9815 | -2.4489 | 1.6912 | 0.2379 |
| 7008 | Accessory-Basal-nucleus (Amygdala) (lh) |
shape_Maximum2DDiameter Column* |
0.2723 | 1.0032 | -1.8863 | 2.2889 | -0.2723 | 0.9375 | -1.8863 | 2.2889 | 0.0037 |
| 2029 | superiorparietal (rh) | firstorder_Minimum | 0.1898 | 0.9722 | -1.7638 | 2.2018 | -0.1898 | 1.0089 | -1.9986 | 2.3255 | 0.0451 |
| 1019 | parsorbitalis (lh) | shape_SurfaceArea* | -0.2675 | 0.9132 | -2.3284 | 1.4640 | 0.2675 | 1.0280 | -2.1684 | 2.6100 | 0.0044 |
| 46 | Cerebellar White Matter (rh) | glszm_SizeZoneNonUniformityNormalized | -0.2310 | 0.8696 | -1.4481 | 1.6651 | 0.2310 | 1.0824 | -1.4481 | 1.6651 | 0.0144 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




