Submitted:
10 September 2024
Posted:
12 September 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Probiotic Preparation
| Bacteria | Identification | Source | Provided By |
|---|---|---|---|
| Enterotoxigenic Escherichia coli | F4 (K88) | Swine rectal isolate | Paul Ebner/Animal Sciences Purdue University |
| Enterotoxigenic Escherichia coli | O78:H11 | Human fecal isolate | ATCC 35401 |
| Listeria monocytogenes | F4244 | Clinical isolate, Human Central Nervous System (CNS) isolate | CDC, Our lab collection |
| Campylobacter jejuni | ATCC 29428 | Human fecal isolate | ATCC |
| Salmonella enterica serovar Typhimurium | ST-1 | Our collection | |
| Lactobacillus acidophilus | LabMAX-3 | CH2 Animal Solutions | CH2 Animal Solutions |
| Lacticaseibacillus casei | LabMAX-3 | CH2 Animal Solutions | CH2 Animal Solutions |
| Enterococcus faecium | LabMAX-3 | CH2 Animal Solutions | CH2 Animal Solutions |
| Lactobacillus acidophilus | NRRL 31910 | ATCC | Our lab collection |
| Lacticaseibacillus casei | ATCC 334 | Cheese | Our collection |
| Enterococcus faecium | ATCC 8459 | Cheese | Dharmendra Mishra, Purdue University |
| Pediococcus acidilactici | H | Fermented sausage | Our Lab collection [48] |
| Staphylococcus aureus | ATCC 25923 (Rosenbach) | Clinical Isolate | Our Lab collection |
2.2. PCR Confirmation of Probiotic Strains
2.3. Pathogen Propagation
2.4. Antimicrobial Activity Testing on Agar Plates
2.5. Biofilm Formation by Probiotics
2.6. MDCK Cell Line Preparation
2.7. Treatment of MDCK with Lipopolysaccharide
2.8. Probiotic Adhesion to MDCK Cell Line by Plate Counting
2.9. Adhesion and Biofilm Analysis by Giemsa Staining
2.10. Lactate Dehydrogenase Assay
2.11. Cytokine ELISA
2.12. Data Analysis
3. Results and Discussion
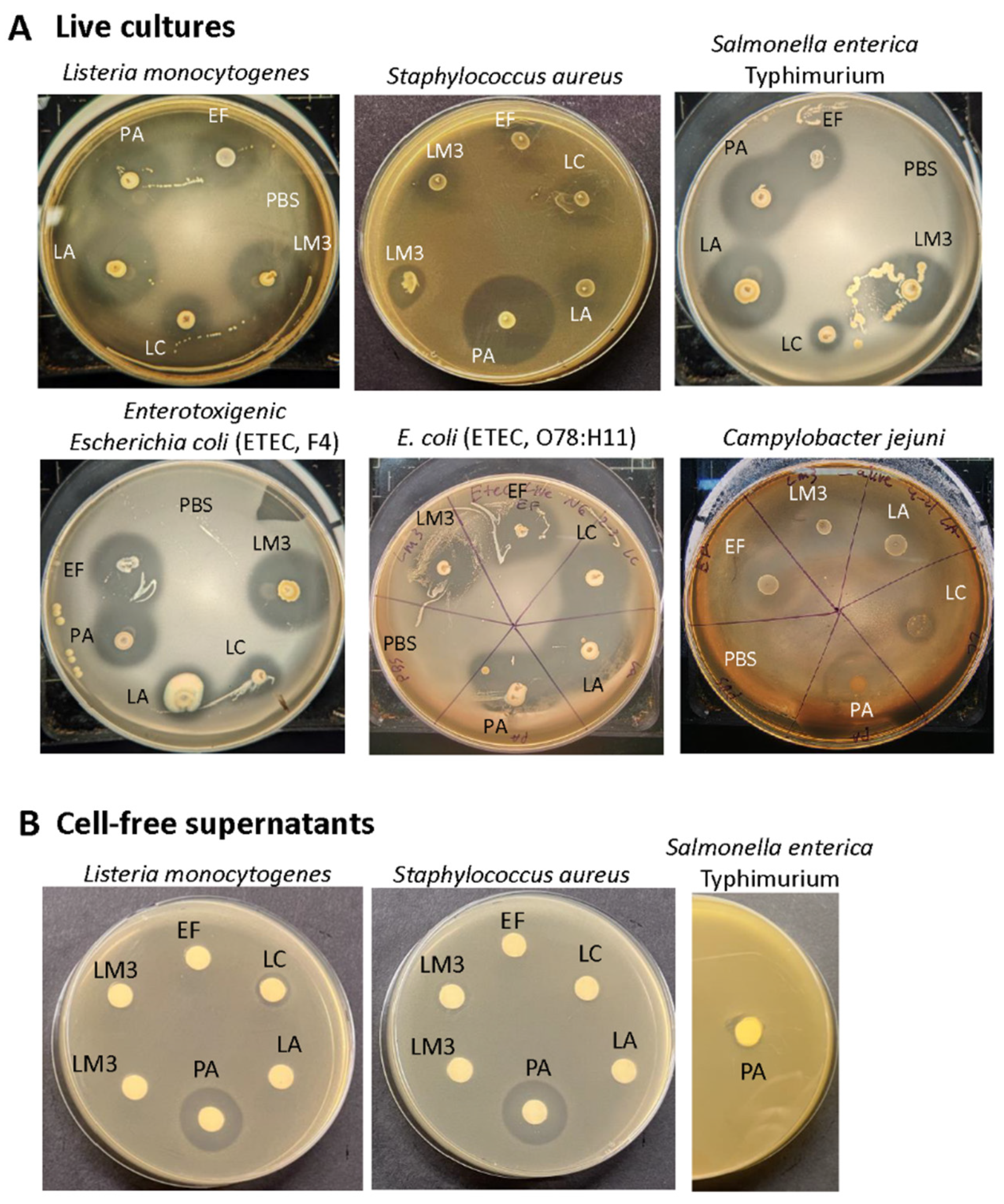
3.1. Antimicrobial Activity of Probiotics Against Pathogens
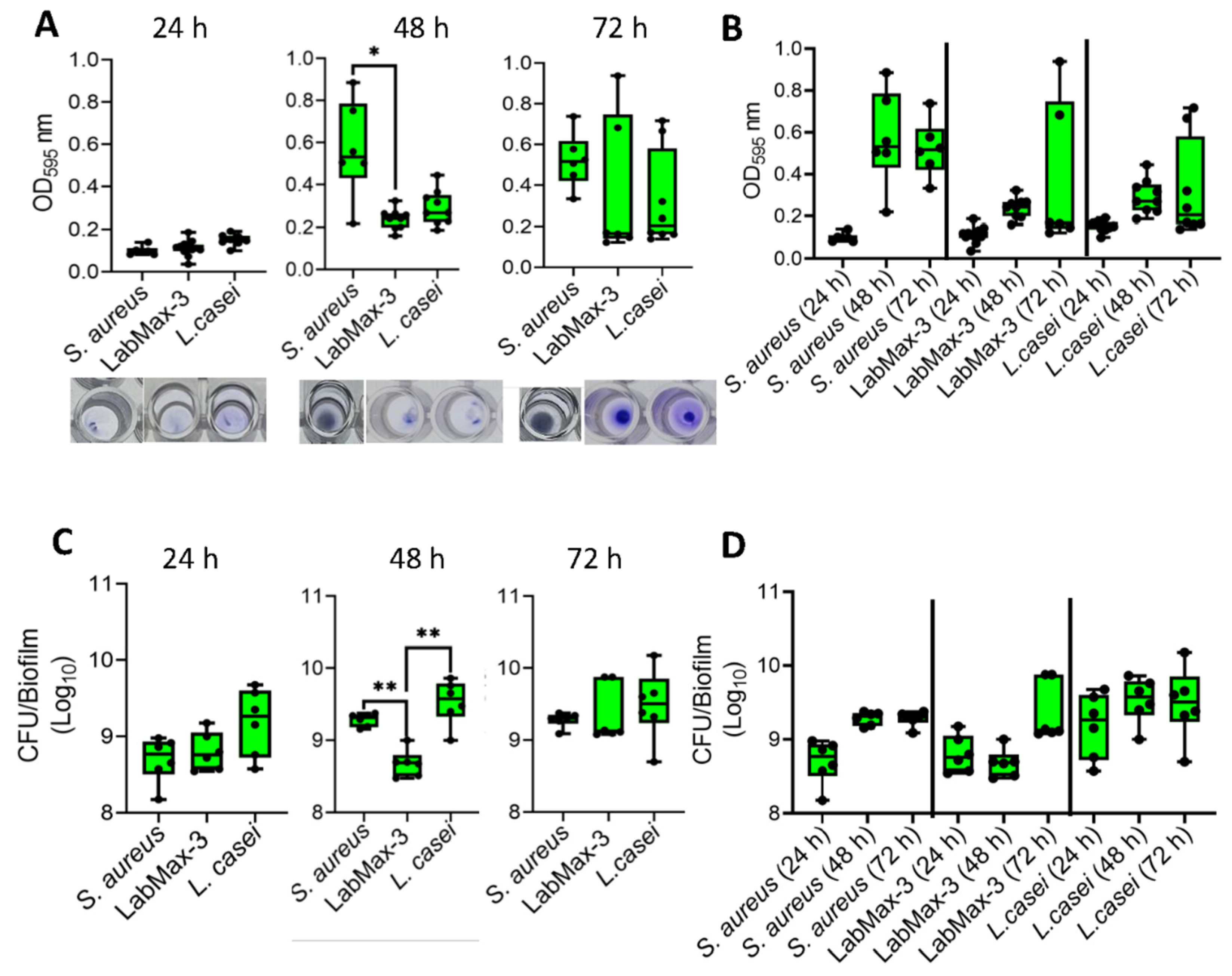
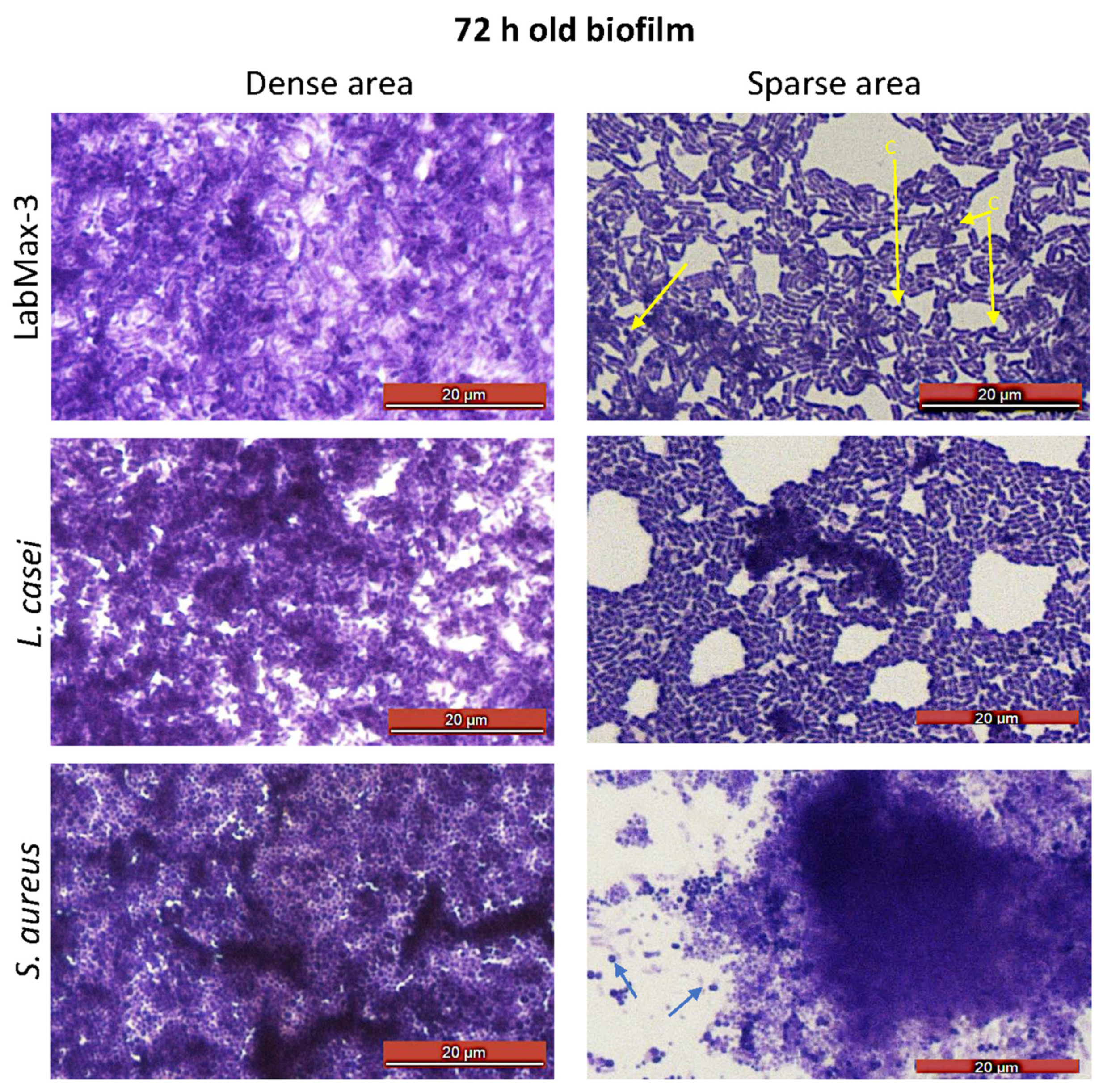
3.2. LabMAX-3 Forms Biofilm on Abiotic Surface
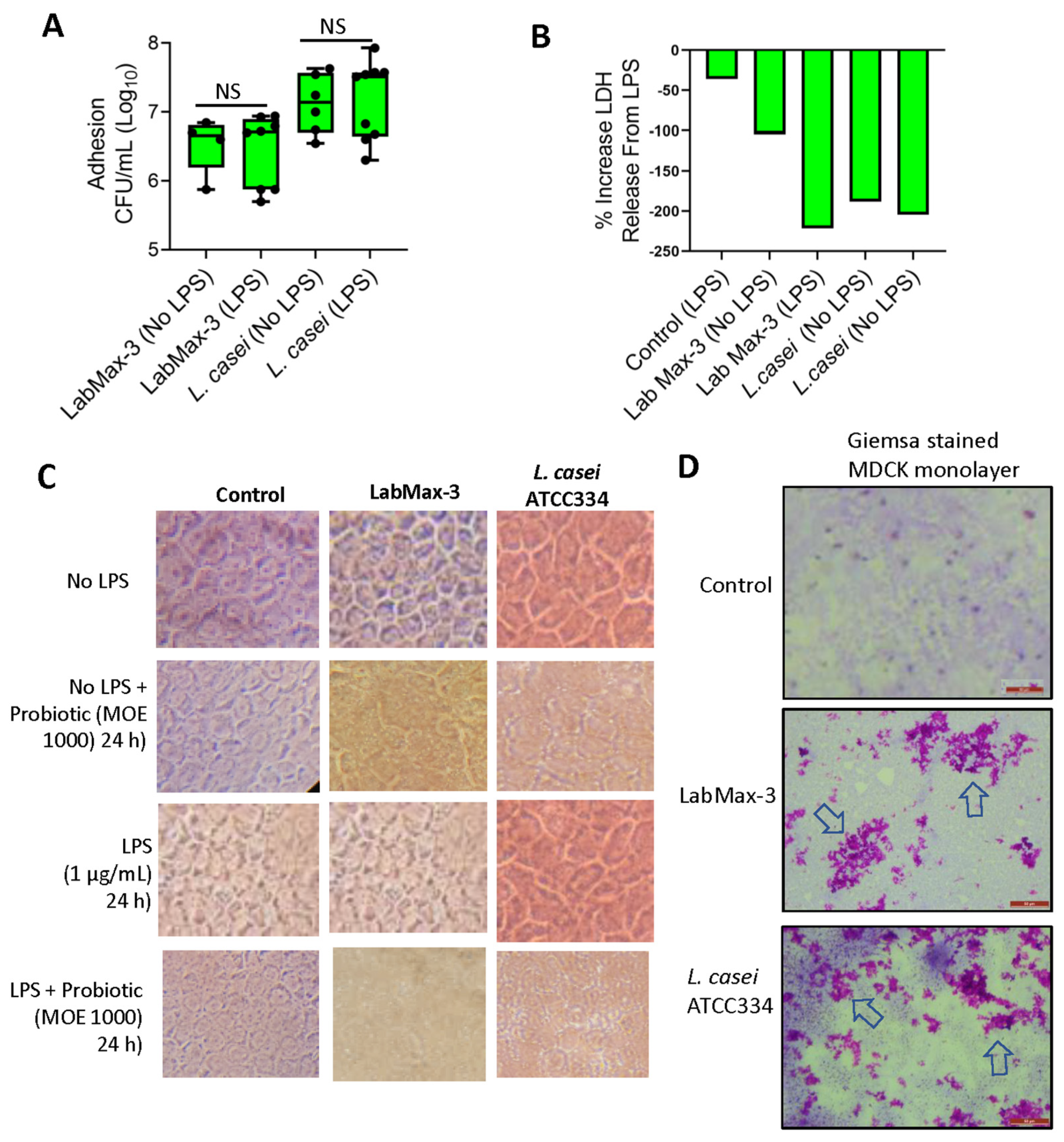
3.3. LabMAX-3 Probiotic Blend Adhesion and Biofilm Formation on MDCK Cells
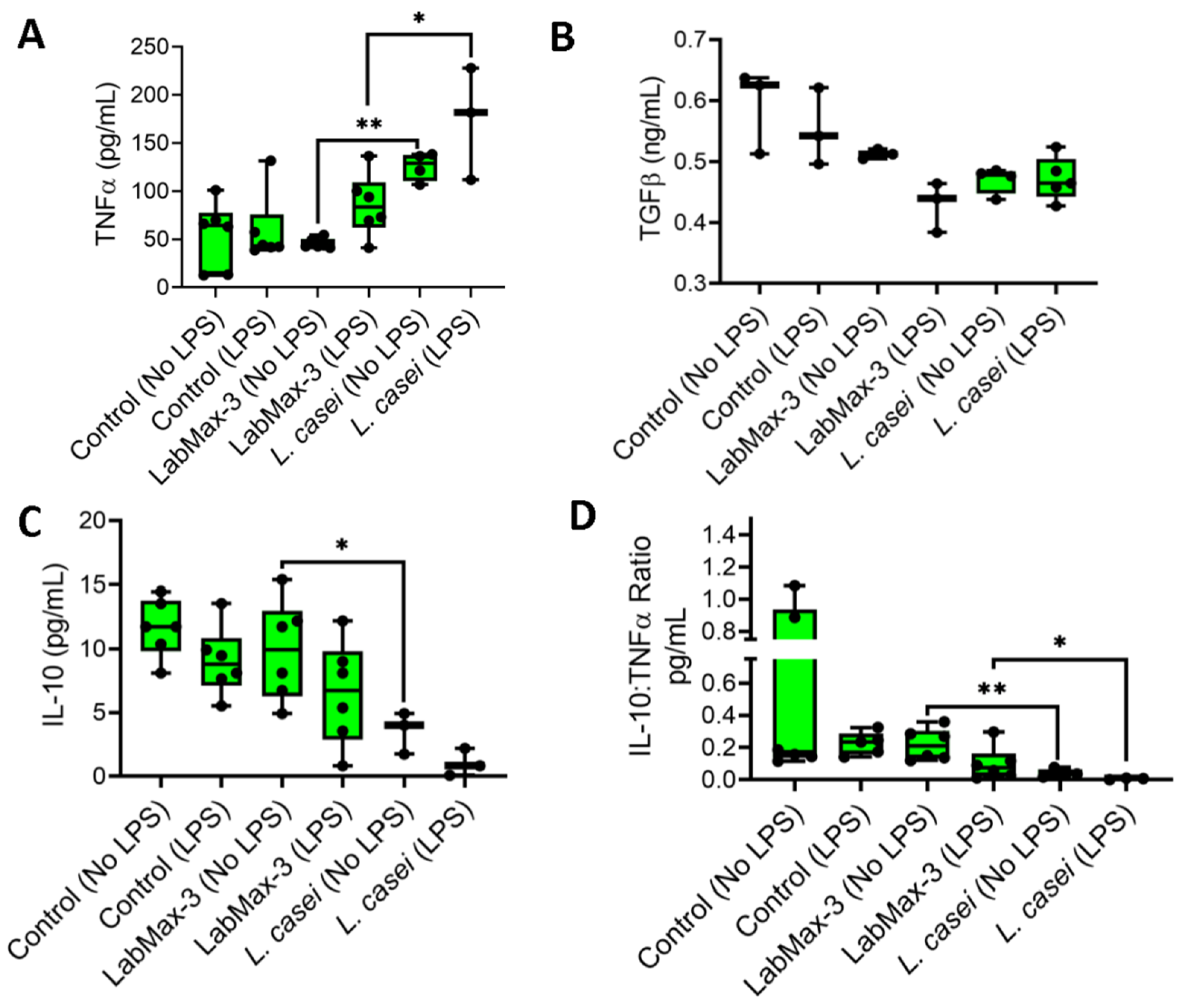
3.4. Anti-inflammatory Response of LabMAX-3 to MDCK Cells
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Amalaradjou, M. A. R.; Bhunia, A. K., Modern approaches in probiotics research to control foodborne pathogens. Adv. Food Nutr. Res. 2012, 67, 185-239. [CrossRef]
- Schmitz, S. S., Value of Probiotics in Canine and Feline Gastroenterology. Vet Clin North Am Small Anim Pract 2021, 51, (1), 171-217. [CrossRef]
- Cristofori, F.; Dargenio, V. N.; Dargenio, C.; Miniello, V. L.; Barone, M.; Francavilla, R., Anti-inflammatory and immunomodulatory effects of probiotics in gut inflammation: a door to the body. Front. Immunol. 2021, 12, 578386. [CrossRef] [PubMed]
- Hill, C.; Guarner, F.; Reid, G.; Gibson, G. R.; Merenstein, D. J.; Pot, B.; Morelli, L.; Canani, R. B.; Flint, H. J.; Salminen, S.; Calder, P. C.; Sanders, M. E., Expert consensus document: The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, (8), 506-514. [CrossRef]
- Lee, J. Y.; Tsolis, R. M.; Bäumler, A. J., The microbiome and gut homeostasis. Science 2022, 377, (6601), eabp9960. [CrossRef]
- Wieërs, G.; Belkhir, L.; Enaud, R.; Leclercq, S.; Philippart de Foy, J. M.; Dequenne, I.; de Timary, P.; Cani, P. D., How Probiotics Affect the Microbiota. Front Cell Infect Microbiol 2019, 9, 454. [CrossRef]
- Zou, J.; Reddivari, L.; Shi, Z.; Li, S.; Wang, Y.; Bretin, A.; Ngo, V. L.; Flythe, M.; Pellizzon, M.; Chassaing, B.; Gewirtz, A. T., Inulin Fermentable Fiber Ameliorates Type I Diabetes via IL22 and Short-Chain Fatty Acids in Experimental Models. Cell Mol Gastroenterol Hepatol 2021, 12, (3), 983-1000. [CrossRef]
- Pilla, R.; Suchodolski, J. S., The Gut Microbiome of Dogs and Cats, and the Influence of Diet. Vet Clin North Am Small Anim Pract 2021, 51, (3), 605-621. [CrossRef]
- White, R.; Atherly, T.; Guard, B.; Rossi, G.; Wang, C.; Mosher, C.; Webb, C.; Hill, S.; Ackermann, M.; Sciabarra, P.; Allenspach, K.; Suchodolski, J.; Jergens, A. E., Randomized, controlled trial evaluating the effect of multi-strain probiotic on the mucosal microbiota in canine idiopathic inflammatory bowel disease. Gut Microbes 2017, 8, (5), 451-466. [CrossRef]
- Drolia, R.; Amalaradjou, M. A. R.; Ryan, V.; Tenguria, S.; Liu, D.; Bai, X.; Xu, L.; Singh, A. K.; Cox, A. D.; Bernal-Crespo, V.; Schaber, J. A.; Applegate, B. M.; Vemulapalli, R.; Bhunia, A. K., Receptor-targeted engineered probiotics mitigate lethal Listeria infection. Nat. Commun. 2020, 11, (1), 6344. [CrossRef] [PubMed]
- Ryan, V. E.; Bailey, T. W.; Liu, D.; Vemulapalli, T.; Cooper, B.; Cox, A. D.; Bhunia, A. K., Listeria adhesion protein-expressing bioengineered probiotics prevent fetoplacental transmission of Listeria monocytogenes in a pregnant Guinea pig model. Microb Pathog 2021, 151, 104752. [CrossRef] [PubMed]
- Suez, J.; Zmora, N.; Segal, E.; Elinav, E., The pros, cons, and many unknowns of probiotics. Nat. Med. 2019, 25, (5), 716-729. [CrossRef]
- Bhunia, A. K., Microbes as a tool to defend against antibiotic resistance in food animal production. Indian J. Anim. Hlth 2019, 58, (2), 01-18. [CrossRef]
- Levy, M.; Kolodziejczyk, A. A.; Thaiss, C. A.; Elinav, E., Dysbiosis and the immune system. Nat Rev Immunol 2017, 17, (4), 219-232. [CrossRef]
- Kong, C.; Gao, R.; Yan, X.; Huang, L.; Qin, H., Probiotics improve gut microbiota dysbiosis in obese mice fed a high-fat or high-sucrose diet. Nutrition 2019, 60, 175-184. [CrossRef]
- Suchodolski, J. S.; Camacho, J.; Steiner, J. M., Analysis of bacterial diversity in the canine duodenum, jejunum, ileum, and colon by comparative 16S rRNA gene analysis. FEMS Microbiol. Ecol. 2008, 66, (3), 567-578. [CrossRef]
- Pilla, R.; Suchodolski, J. S., The Role of the Canine Gut Microbiome and Metabolome in Health and Gastrointestinal Disease. Front Vet Sci 2019, 6, 498. [CrossRef]
- Lambertini, E.; Buchanan, R. L.; Narrod, C.; Ford, R. M.; Baker, R. C.; Pradhan, A. K., Quantitative assessment of human and pet exposure to Salmonella associated with dry pet foods. Int. J. Food Microbiol. 2016, 216, 79-90. [CrossRef] [PubMed]
- Antunes, P.; Novais, C.; Peixe, L.; Freitas, A. R., Pet Food Safety: Emerging bacterial hazards and implications for Public Health. Curr. Opin. Food Sci. 2024, 101165. [CrossRef]
- Nemser, S. M.; Doran, T.; Grabenstein, M.; McConnell, T.; McGrath, T.; Pamboukian, R.; Smith, A. C.; Achen, M.; Danzeisen, G.; Kim, S., Investigation of Listeria, Salmonella, and toxigenic Escherichia coli in various pet foods. Foodborne Pathog. Dis. 2014, 11, (9), 706-709. [CrossRef]
- Sola-Oladokun, B.; Culligan, E. P.; Sleator, R. D., Engineered probiotics: Applications and biological containment. Annu. Rev. Food Sci. Technol. 2017, 8, (1), 353-370. [CrossRef]
- Tan, L.; Fu, J.; Feng, F.; Liu, X.; Cui, Z.; Li, B.; Han, Y.; Zheng, Y.; Yeung, K. W. K.; Li, Z.; Zhu, S.; Liang, Y.; Feng, X.; Wang, X.; Wu, S., Engineered probiotics biofilm enhances osseointegration via immunoregulation and anti-infection. Sci. Adv. 2020, 6, (46). [CrossRef]
- Zhou, J.; Li, M.; Chen, Q.; Li, X.; Chen, L.; Dong, Z.; Zhu, W.; Yang, Y.; Liu, Z.; Chen, Q., Programmable probiotics modulate inflammation and gut microbiota for inflammatory bowel disease treatment after effective oral delivery. Nat Commun 2022, 13, (1), 3432. [CrossRef]
- Piewngam, P.; Zheng, Y.; Nguyen, T. H.; Dickey, S. W.; Joo, H. S.; Villaruz, A. E.; Glose, K. A.; Fisher, E. L.; Hunt, R. L.; Li, B.; Chiou, J.; Pharkjaksu, S.; Khongthong, S.; Cheung, G. Y. C.; Kiratisin, P.; Otto, M., Pathogen elimination by probiotic Bacillus via signalling interference. Nature 2018, 562, (7728), 532-537. [CrossRef]
- Piewngam, P.; Khongthong, S.; Roekngam, N.; Theapparat, Y.; Sunpaweravong, S.; Faroongsarng, D.; Otto, M., Probiotic for pathogen-specific Staphylococcus aureus decolonisation in Thailand: a phase 2, double-blind, randomised, placebo-controlled trial. Lancet Microbe 2023, 4, (2), e75-e83. [CrossRef]
- Al-Shamiri, M. M.; Wang, J.; Zhang, S.; Li, P.; Odhiambo, W. O.; Chen, Y.; Han, B.; Yang, E.; Xun, M.; Han, L.; Han, S., Probiotic Lactobacillus Species and Their Biosurfactants Eliminate Acinetobacter baumannii Biofilm in Various Manners. Microbiol Spectr 2023, 11, (2), e0461422. [CrossRef]
- Burkholder, K. M.; Bhunia, A. K., Salmonella enterica serovar Typhimurium adhesion and cytotoxicity during epithelial cell stress is reduced by Lactobacillus rhamnosus GG. Gut Pathog 2009, 1, (1), 14. [CrossRef]
- Nathan, V. B.; Lu, H.; Horn, N. L.; Drolia, R.; Bhunia, A. K., Sequestration of zearalenone using microorganisms blend in vitro. Lett Appl Microbiol 2023, 76, (2), ovad020. [CrossRef]
- Ragoubi, C.; Quintieri, L.; Greco, D.; Mehrez, A.; Maatouk, I.; D’Ascanio, V.; Landoulsi, A.; Avantaggiato, G., Mycotoxin removal by Lactobacillus spp. and their application in animal liquid feed. Toxins 2021, 13, (3), 185. [CrossRef]
- Liu, L.; Xie, M.; Wei, D., Biological detoxification of mycotoxins: Current status and future advances. Int. J. Mol. Sci. 2022, 23, (3), 1064. [CrossRef]
- Grześkowiak, Ł.; Endo, A.; Beasley, S.; Salminen, S., Microbiota and probiotics in canine and feline welfare. Anaerobe 2015, 34, 14-23. [CrossRef]
- Mathipa, M. G.; Bhunia, A. K.; Thantsha, M. S., Internalin AB-expressing recombinant Lactobacillus casei protects Caco-2 cells from Listeria monocytogenes-induced damages under simulated intestinal conditions. PLoS One 2019, 14, (7), e0220321. [CrossRef] [PubMed]
- Mathipa, M. G.; Thantsha, M. S.; Bhunia, A. K., Lactobacillus casei expressing Internalins A and B reduces Listeria monocytogenes interaction with Caco-2 cells in vitro. Microb. Biotechnol. 2019, 12, (4), 715-729. [CrossRef]
- Amalaradjou, M. A.; Bhunia, A. K., Bioengineered probiotics, a strategic approach to control enteric infections. Bioengineered 2013, 4, (6), 379-87. [CrossRef]
- Cruz, K. C. P.; Enekegho, L. O.; Stuart, D. T., Bioengineered Probiotics: Synthetic Biology Can Provide Live Cell Therapeutics for the Treatment of Foodborne Diseases. Front. Bioeng. Biotechnol. 2022, 10, 890479. [CrossRef]
- Plaza-Diaz, J.; Ruiz-Ojeda, F. J.; Gil-Campos, M.; Gil, A., Mechanisms of Action of Probiotics. Adv. Nutr. 2019, 10, (suppl_1), S49-S66. [CrossRef]
- Bai, X.; Liu, D.; Xu, L.; Tenguria, S.; Drolia, R.; Gallina, N. L. F.; Cox, A. D.; Koo, O.-K.; Bhunia, A. K., Biofilm-isolated Listeria monocytogenes exhibits reduced systemic dissemination at the early (12–24 h) stage of infection in a mouse model. npj Biofilms and Microbiomes 2021, 7, (1), 1-16. [CrossRef] [PubMed]
- Zhu, X.; Liu, D.; Singh, A. K.; Drolia, R.; Bai, X.; Tenguria, S.; Bhunia, A. K., Tunicamycin mediated inhibition of wall teichoic acid affect Staphylococcus aureus and Listeria monocytogenes cell morphology, biofilm formation and virulence. Front. Microbiol. 2018, 9, (July), 1352. [CrossRef] [PubMed]
- Motta, J. P.; Wallace, J. L.; Buret, A. G.; Deraison, C.; Vergnolle, N., Gastrointestinal biofilms in health and disease. Nat Rev Gastroenterol Hepatol 2021, 18, (5), 314-334. [CrossRef]
- Azimi, S.; Klementiev, A. D.; Whiteley, M.; Diggle, S. P., Bacterial Quorum Sensing During Infection. Annu Rev Microbiol 2020, 74, 201-219. [CrossRef]
- Mukherjee, S.; Bassler, B. L., Bacterial quorum sensing in complex and dynamically changing environments. Nat Rev Microbiol 2019, 17, (6), 371-382. [CrossRef]
- Zhang, H.; Yeh, C.; Jin, Z.; Ding, L.; Liu, B. Y.; Zhang, L.; Dannelly, H. K., Prospective study of probiotic supplementation results in immune stimulation and improvement of upper respiratory infection rate. Synth Syst Biotechnol 2018, 3, (2), 113-120. [CrossRef]
- Bai, X.; Gallina, N. L. F.; Bhunia, A. K., Microbial Biofilms in Food Safety and Public Health Domains. In Reference Module in Food Science, Elsevier: 2023. [CrossRef]
- Sambanthamoorthy, K.; Feng, X.; Patel, R.; Patel, S.; Paranavitana, C., Antimicrobial and antibiofilm potential of biosurfactants isolated from lactobacilli against multi-drug-resistant pathogens. BMC Microbiol 2014, 14, 197. [CrossRef]
- Nataraj, B. H.; Ramesh, C.; Mallappa, R. H., Functional group characterization of lactic bacterial biosurfactants and evaluation of antagonistic actions against clinical isolates of methicillin-resistant Staphylococcus aureus. Lett Appl Microbiol 2021, 73, (3), 372-382. [CrossRef]
- Irvine, J. D.; Takahashi, L.; Lockhart, K.; Cheong, J.; Tolan, J. W.; Selick, H. E.; Grove, J. R., MDCK (Madin–Darby canine kidney) cells: a tool for membrane permeability screening. J. Pharmaceut. Sci. 1999, 88, (1), 28-33. [CrossRef] [PubMed]
- Capellini, F. M.; Vencia, W.; Amadori, M.; Mignone, G.; Parisi, E.; Masiello, L.; Vivaldi, B.; Ferrari, A.; Razzuoli, E., Characterization of MDCK cells and evaluation of their ability to respond to infectious and non-infectious stressors. Cytotechnology 2020, 72, 97-109. [CrossRef]
- Bhunia, A. K.; Johnson, M. C.; Ray, B., Purification, characterization and antimicrobial spectrum of a bacteriocin produced by Pediococcus acidilactici. J. Appl. Bacteriol. 1988, 65, (4), 261-268. [CrossRef]
- Kim, E.; Yang, S. M.; Lim, B.; Park, S. H.; Rackerby, B.; Kim, H. Y., Design of PCR assays to specifically detect and identify 37 Lactobacillus species in a single 96 well plate. BMC Microbiol 2020, 20, (1), 96. [CrossRef]
- Belloso Daza, M. V.; Almeida-Santos, A. C.; Novais, C.; Read, A.; Alves, V.; Cocconcelli, P. S.; Freitas, A. R.; Peixe, L., Distinction between Enterococcus faecium and Enterococcus lactis by a gluP PCR-Based Assay for Accurate Identification and Diagnostics. Microbiol Spectr 2022, 10, (6), e0326822. [CrossRef]
- Bai, X.; Xu, L.; Singh, A. K.; Qiu, X.; Liu, M.; Abuzeid, A.; El-Khateib, T.; Bhunia, A. K., Inactivation of Polymicrobial Biofilms of Foodborne Pathogens Using Epsilon Poly-L-Lysin Conjugated Chitosan Nanoparticles. Foods 2022, 11, (4), 569. [CrossRef]
- Roberts, P. H.; Davis, K. C.; Garstka, W. R.; Bhunia, A. K., Lactate dehydrogenase release assay from Vero cells to distinguish verotoxin producing Escherichia coli from non-verotoxin producing strains. J. Microbiol. Methods 2001, 43, (3), 171-181. [CrossRef]
- Bermudez-Brito, M.; Plaza-Díaz, J.; Muñoz-Quezada, S.; Gómez-Llorente, C.; Gil, A., Probiotic mechanisms of action. Annals Nutr. Metabol. 2012, 61, (2), 160-174. [CrossRef]
- Aleman, R. S.; Yadav, A., Systematic review of probiotics and their potential for developing functional nondairy foods. Appl. Microbiol. 2023, 4, (1), 47-69. [CrossRef]
- Rossi, G.; Pengo, G.; Caldin, M.; Palumbo Piccionello, A.; Steiner, J. M.; Cohen, N. D.; Jergens, A. E.; Suchodolski, J. S., Comparison of microbiological, histological, and immunomodulatory parameters in response to treatment with either combination therapy with prednisone and metronidazole or probiotic VSL# 3 strains in dogs with idiopathic inflammatory bowel disease. Plos One 2014, 9, (4), e94699.
- Ziese, A.-L.; Suchodolski, J. S.; Hartmann, K.; Busch, K.; Anderson, A.; Sarwar, F.; Sindern, N.; Unterer, S., Effect of probiotic treatment on the clinical course, intestinal microbiome, and toxigenic Clostridium perfringens in dogs with acute hemorrhagic diarrhea. 2018, 13, (9), e0204691.
- Panja, K.; Areerat, S.; Chundang, P.; Palaseweenun, P.; Akrimajirachoote, N.; Sitdhipol, J.; Thaveethaptaikul, P.; Chonpathompikunlert, P.; Niwasabutra, K.; Phapugrangkul, P.; Kovitvadhi, A., Influence of dietary supplementation with new Lactobacillus strains on hematology, serum biochemistry, nutritional status, digestibility, enzyme activities, and immunity in dogs. Vet World 2023, 16, (4), 834-843. [CrossRef]
- Xu, H.; Huang, W.; Hou, Q.; Kwok, L.-Y.; Laga, W.; Wang, Y.; Ma, H.; Sun, Z.; Zhang, H., Oral administration of compound probiotics improved canine feed intake, weight gain, immunity and intestinal microbiota. Front. Immunol. 2019, 10, 394673. [CrossRef] [PubMed]
- Johnson-Henry, K. C.; Donato, K. A.; Shen-Tu, G.; Gordanpour, A.; Sherman, P. A., Lactobacillus rhamnosus strain GG prevents enterohemorrhagic Escherichia coli O157 : H7-Induced changes in epithelial barrier function. Infect. Immun. 2008, 76, (4), 1340-1348. [CrossRef]
- Gao, J.; Sadiq, F. A.; Zheng, Y.; Zhao, J.; He, G.; Sang, Y., Biofilm-based delivery approaches and specific enrichment strategies of probiotics in the human gut. Gut Microbes 2022, 14, (1), 2126274. [CrossRef]
- Freeman, D. J.; Falkiner, F. R.; Keane, C. T., New method for detecting slime production by coagulase negative staphylococci. J Clin Pathol 1989, 42, (8), 872-4. [CrossRef]
- Salas-Jara, M. J.; Ilabaca, A.; Vega, M.; García, A., Biofilm Forming Lactobacillus: New Challenges for the Development of Probiotics. Microorganisms 2016, 4, (3). [CrossRef]
- Popović, N.; Dinić, M.; Tolinački, M.; Mihajlović, S.; Terzić-Vidojević, A.; Bojić, S.; Djokić, J.; Golić, N.; Veljović, K., New Insight into Biofilm Formation Ability, the Presence of Virulence Genes and Probiotic Potential of Enterococcus sp. Dairy Isolates. front. Microbiol. 2018, 9. [CrossRef] [PubMed]
- Bustamante, M.; Oomah, B. D.; Oliveira, W. P.; Burgos-Díaz, C.; Rubilar, M.; Shene, C., Probiotics and prebiotics potential for the care of skin, female urogenital tract, and respiratory tract. Folia Microbiol 2020, 65, 245-264. [CrossRef]
- Gorreja, F.; Walker, W. A., The potential role of adherence factors in probiotic function in the gastrointestinal tract of adults and pediatrics: a narrative review of experimental and human studies. Gut Microbes 2022, 14, (1), 2149214. [CrossRef]
- Sebastián Domingo, J. J., Review of the role of probiotics in gastrointestinal diseases in adults. Gastroenterol. Hepatol. 2017, 40, (6), 417-429. [CrossRef]
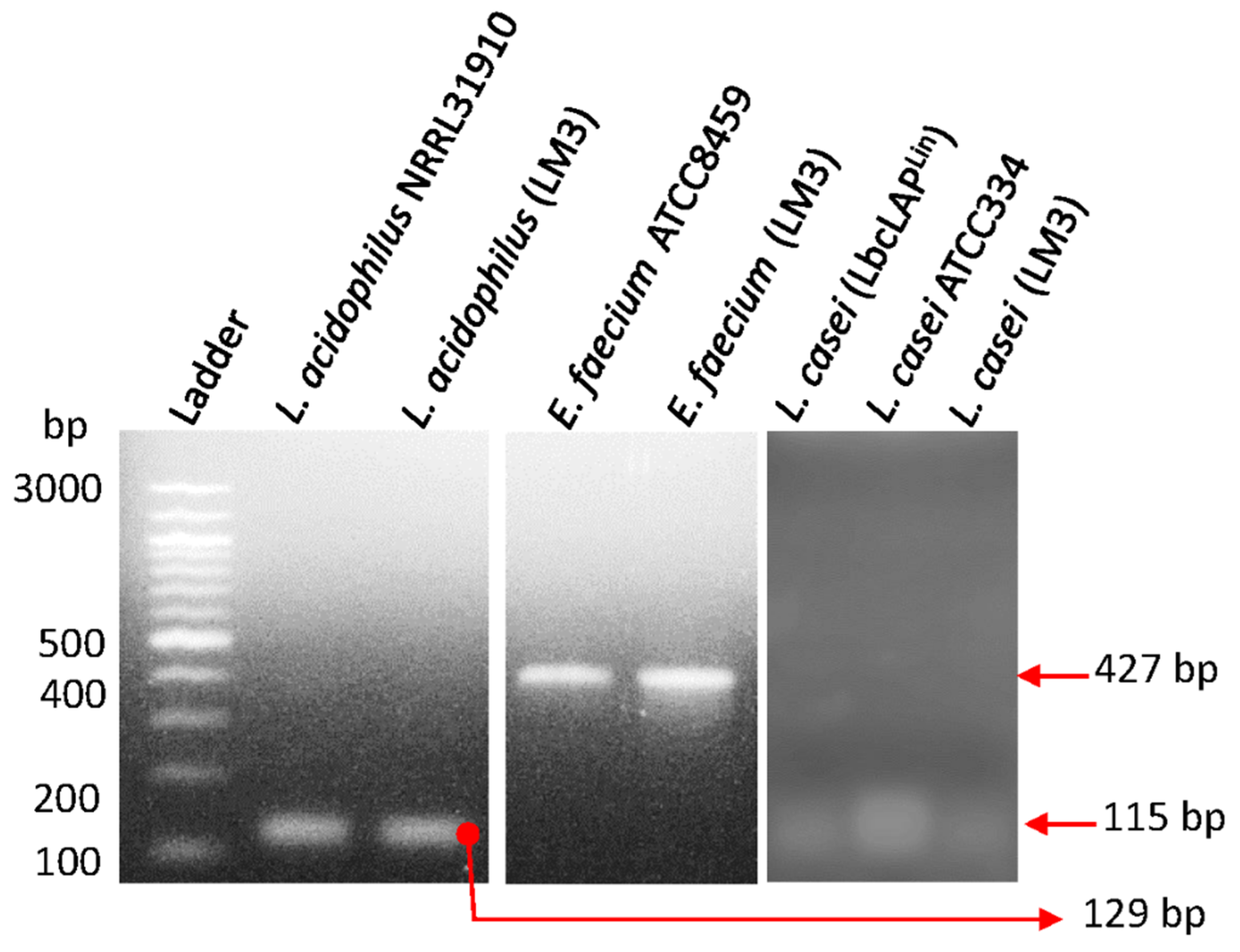
| Organism | Primer | Target gene | Product size (bp) | Ref |
|---|---|---|---|---|
| L. acidophilus | F-CCT TTC TAA GGA AGC GAA GGA T R-ACG CTT GGT ATT CCA AAT CGC |
16S-23S | 129 | [49] |
| L. casei | F-CCA CAA TCC TTG GCT GTT CT R-GCT TGA GGC GAT TGT AAT CC |
Putative protein | 115 | [49] |
| E. faecium | F-GCGTGCATGGTTAAGACGAC R-CTGCTGGATCGCTGGGTTAT |
Rhomboid protease GluP (serine protease) | 427 | [50] |
| Bacteria | Strain | Avg zone of inhibition ± SEM (mm) | |||||
|---|---|---|---|---|---|---|---|
| LabMAX-3* | LC* | LA* | EF* | PA | LC (ATCC 334) | ||
| Listeria monocytogenes | F4244 | 15.83±1.64 | 8.50±0.00 | 19.00±1.44 | 17.67±0.72 | 20.67±0.44 | 11.00±0.57 |
| Staphylococcus aureus | ATCC 25923 | 7.33±0.67 | 10.67±0.67 | 9.33±0.67 | 7.33±0.67 | 20.67±0.67 | 18.33±0.88 |
| Salmonella enterica serovar Typhimurium | ST-1 | 22.67±1.59 | 10.33±0.33 | 15.50±0.50 | 18.00±1.26 | 19.83±0.33 | 9.33±2.19 |
| Enterotoxigenic Escherichia coli | F4 (K88) | 19.83±1.69 | 8.17±0.88 | 21.17±0.17 | 19.50±1.041 | 22.67±0.17 | 13.67±2.73 |
| Enterotoxigenic Escherichia coli | O78:H11 | 6.00±0.57 | 9.33±0.67 | 10.00±1.00 | 9.00±0.00 | 11.67±0.33 | 11.67±3.28 |
| Campylobacter jejuni⁋ | ATCC 29428 | 15.00 | 10.00 | 0.0 | 13.50 | 22.00 | NA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




