Submitted:
09 September 2024
Posted:
10 September 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
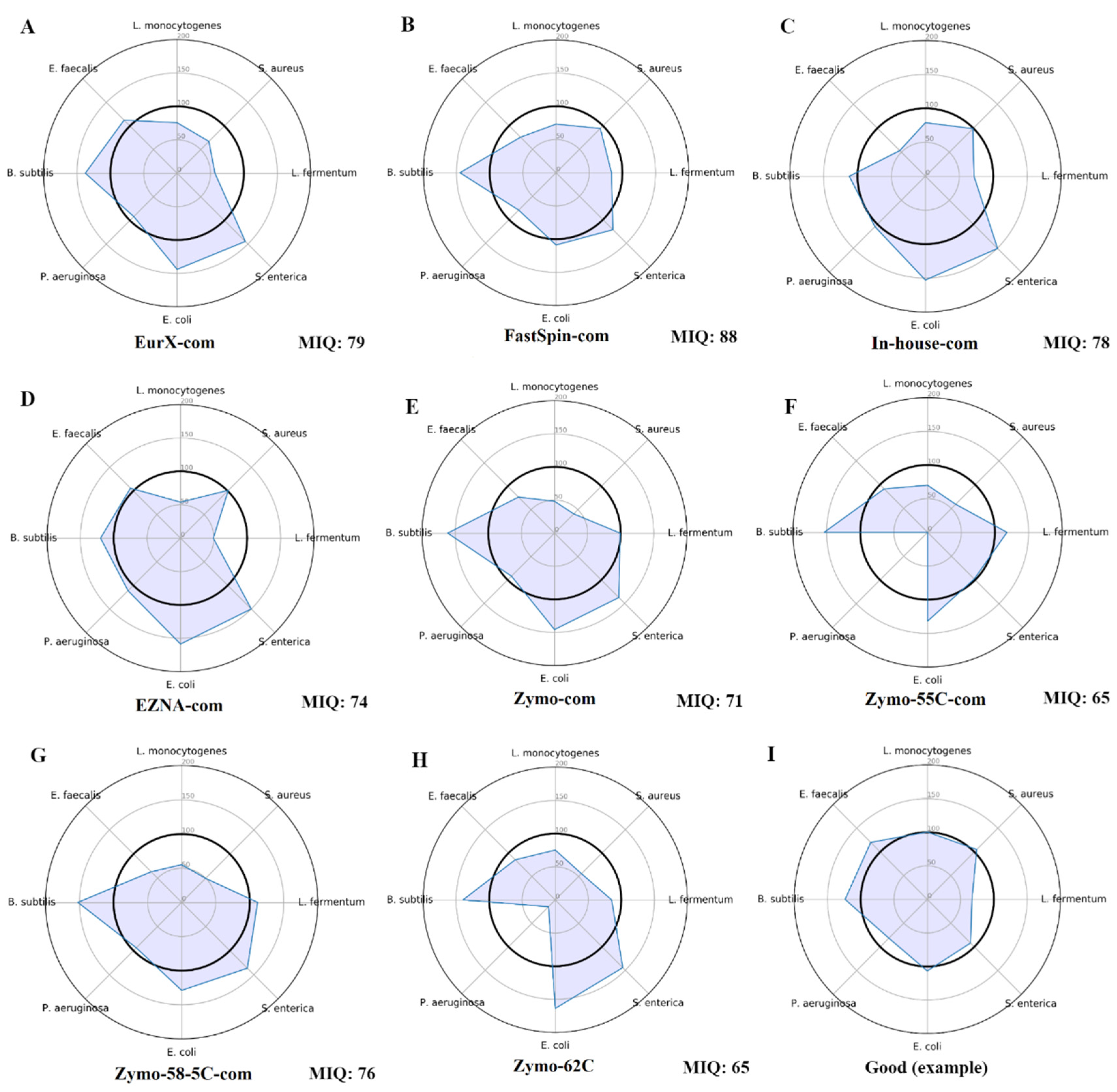
2.1. MIQ Score Variability in DNA Extraction Protocols
2.2. Bias Introduced by 16S rRNA DBs and/or DNA Extraction Kits on Closed-Reference Clustered Mock Samples
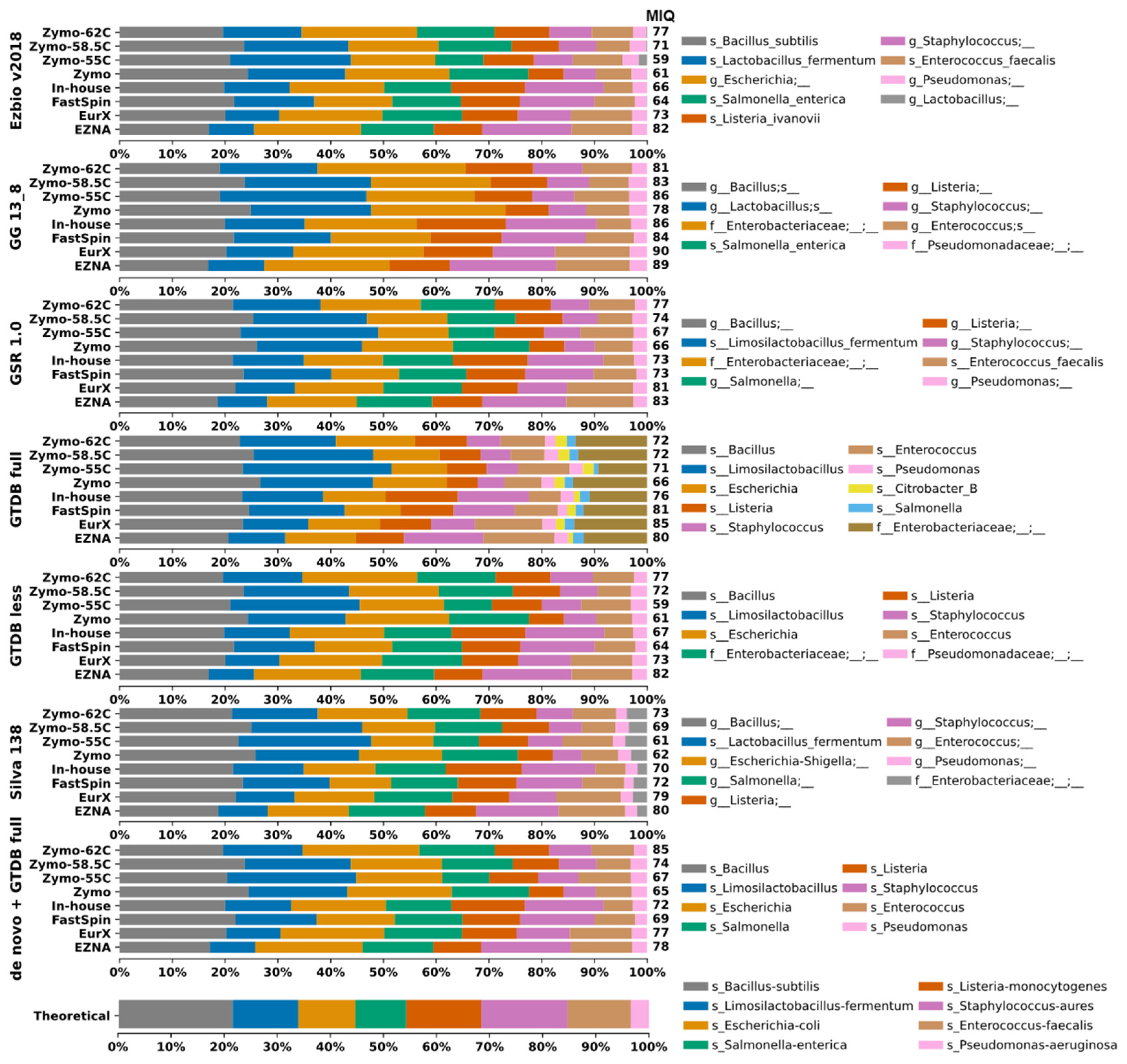
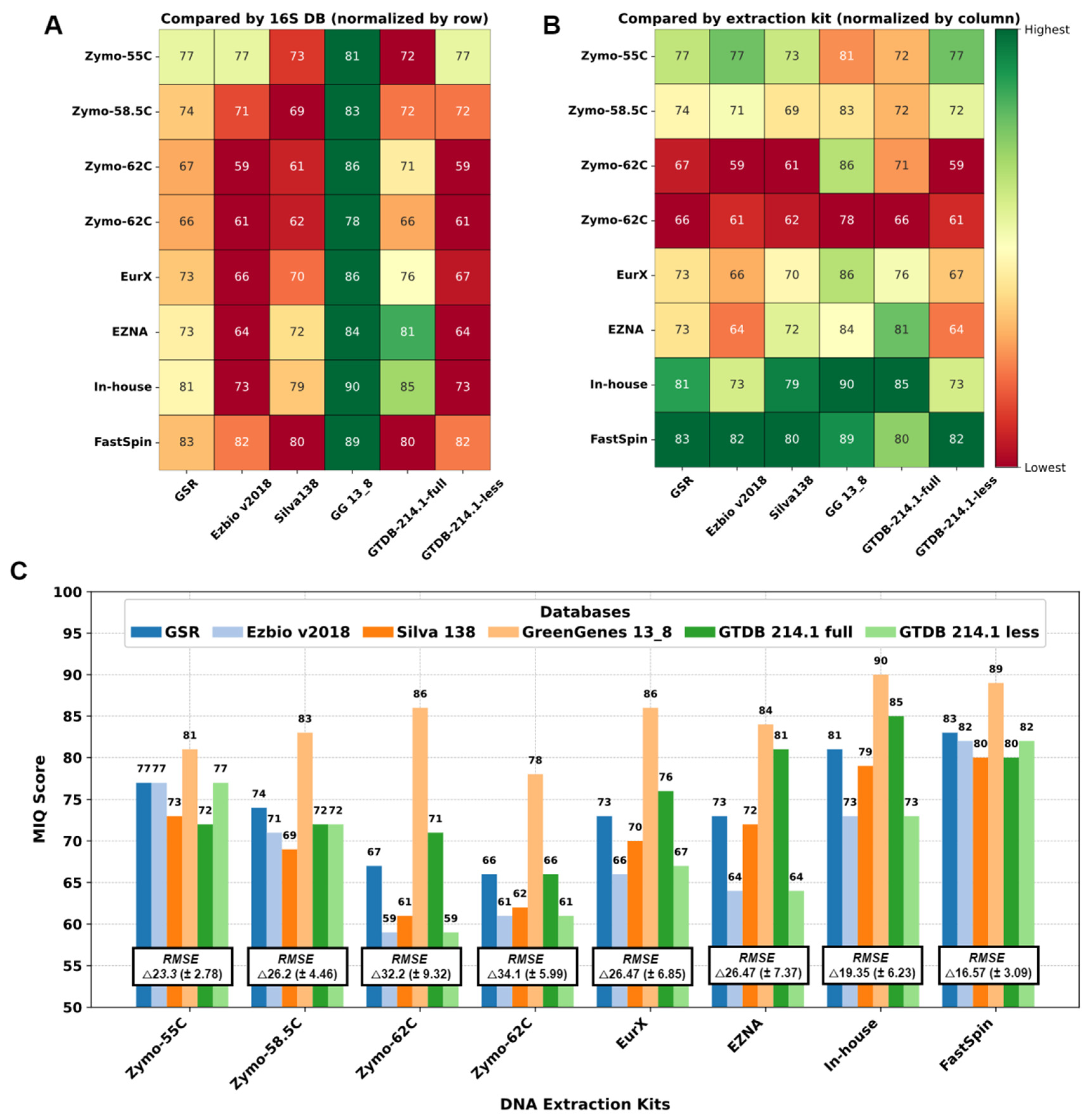
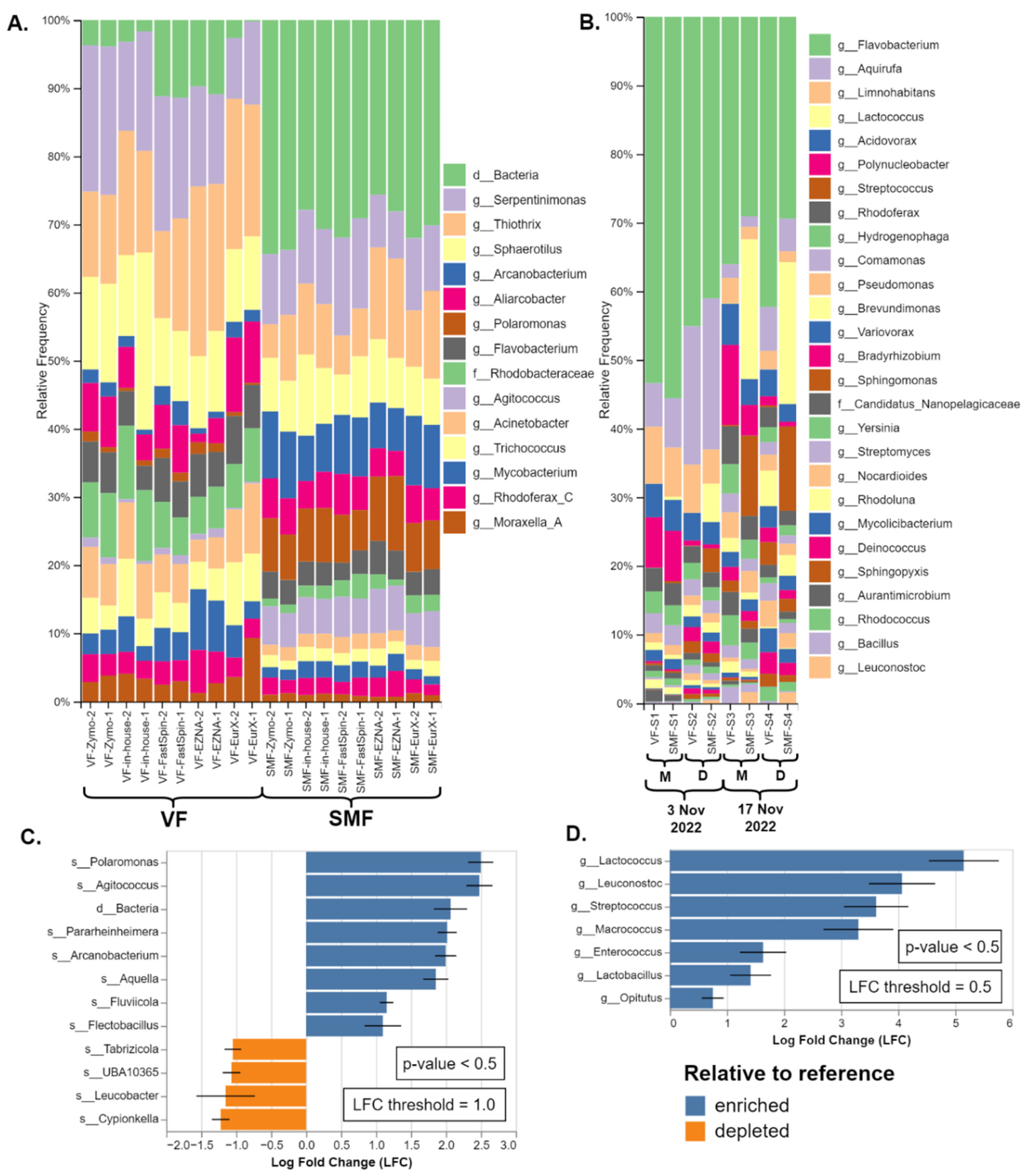
2.3. Taxa Identification Efficiency of 16S DBs on de novo Clustered Mock Samples
2.4. Evaluation of Skimmed Milk Flocculation
3. Discussion
4. Materials and Methods
4.1. Samples
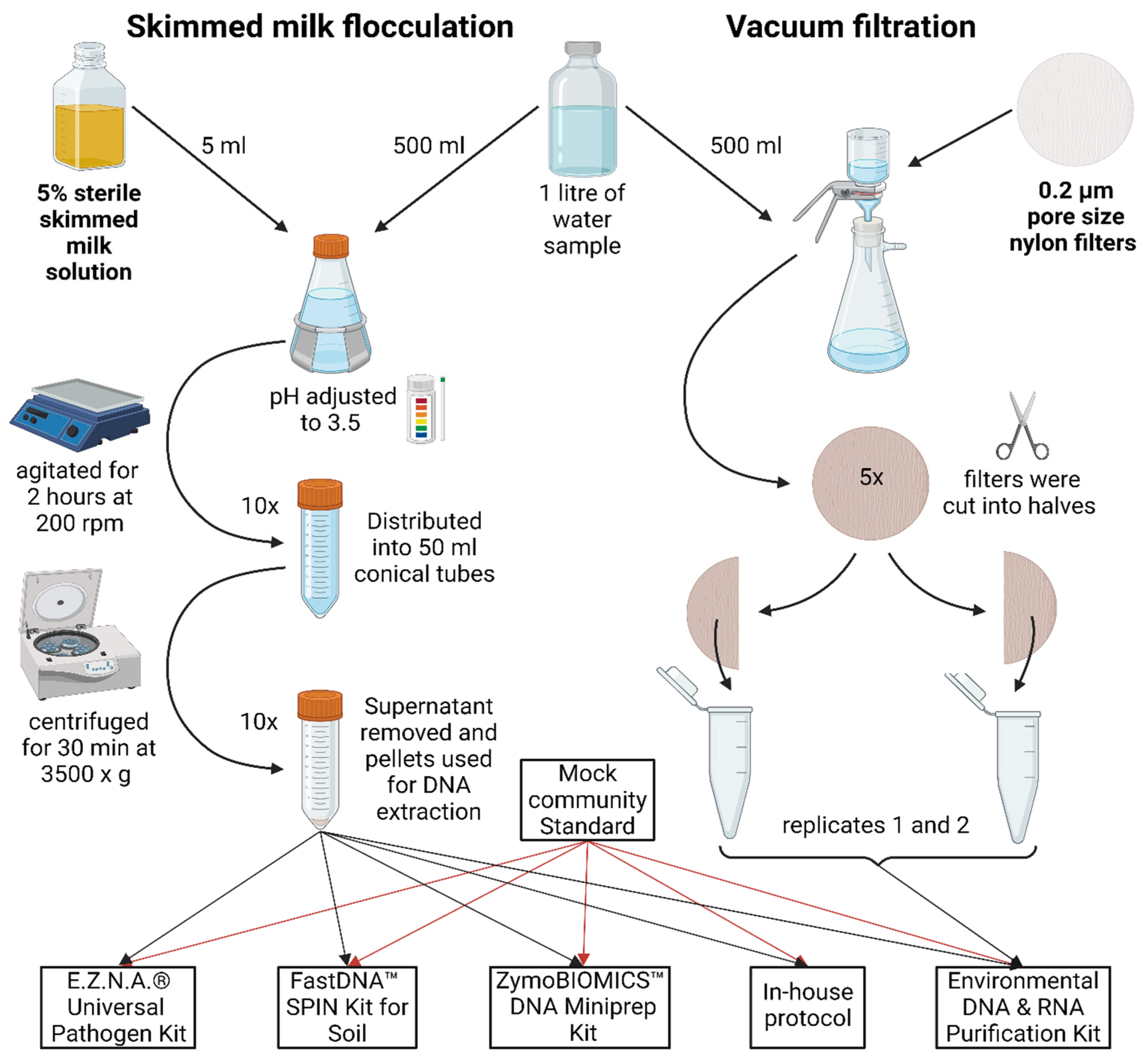
4.2. Skim Milk Flocculation
4.3. Vacuum Filtration
4.4. DNA Extraction
4.5. 16S rRNA and “Shotgun” Metagenomics
4.6. Bioinformatic Analysis
5. Conclusion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgement
Conflicts of Interest
References
- B. Calgua, A. Mengewein, A. Grunert, et al., “Development and application of a one-step low cost procedure to concentrate viruses from seawater samples,” J. Virol. Methods, vol. 153, no. 2, pp. 79–83, Nov. 2008 (PMID:18765255) URL: https://pubmed.ncbi.nlm.nih.gov/18765255/.
- C. F. Nnadozie, J. Lin, and R. Govinden, “Selective isolation of bacteria for metagenomic analysis: Impact of membrane characteristics on bacterial filterability,” Biotechnol. Prog., vol. 31, no. 4, pp. 853–866, Jul. 2015 (PMID:26018114) URL: https://onlinelibrary.wiley.com/doi/full/10.1002/btpr.2109.
- S. E. Philo, A. Q. W. Ong, E. K. Keim, et al., “Development and Validation of the Skimmed Milk Pellet Extraction Protocol for SARS-CoV-2 Wastewater Surveillance,” Food Environ. Virol., vol. 14, no. 4, Dec. 2022 (PMID:35143035) URL: https://pubmed.ncbi.nlm.nih.gov/35143035/.
- K. Yanaç, J. Francis, J. Zambrano-Alvarado, Q. Yuan, and M. Uyaguari-Díaz, “Concentration of Virus Particles from Environmental Water and Wastewater Samples Using Skimmed Milk Flocculation and Ultrafiltration,” JoVE (Journal Vis. Exp., vol. 2023, no. 193, p. e65058, Mar. 2023 (PMID:37010296) URL: https://www.jove.com/v/65058/concentration-virus-particles-from-environmental-water-wastewater.
- A. S. F. Assis, M. H. Otenio, B. P. Drumond, T. M. Fumian, M. P. Miagostovich, and M. L. da Rosa e Silva, “Optimization of the skimmed-milk flocculation method for recovery of adenovirus from sludge,” Sci. Total Environ., vol. 583, pp. 163–168, Apr. 2017 (PMID:28094048) URL: https://pubmed.ncbi.nlm.nih.gov/28094048/.
- E. Borgmästars, S. Persson, M. Hellmér, M. Simonsson, and R. Eriksson, “Comparison of Skimmed Milk and Lanthanum Flocculation for Concentration of Pathogenic Viruses in Water,” Food Environ. Virol., vol. 13, no. 3, pp. 380–389, Sep. 2021 (PMID:33974212) URL: https://link.springer.com/article/10.1007/s12560-021-09477-x.
- D. Abraham, V. R. Mohan, and G. Kang, “Skimmed Milk Flocculation Technique for Waste Water,” May 2021 URL: https://www.protocols.io/view/skimmed-milk-flocculation-technique-for-waste-wate-buzmnx46.
- E. Gonzales-Gustavson, Y. Cárdenas-Youngs, M. Calvo, et al., “Characterization of the efficiency and uncertainty of skimmed milk flocculation for the simultaneous concentration and quantification of water-borne viruses, bacteria and protozoa,” J. Microbiol. Methods, vol. 134, pp. 46–53, Mar. 2017 (PMID:28093213).
- P. I. Costea, G. Zeller, S. Sunagawa, et al., “Towards standards for human fecal sample processing in metagenomic studies,” Nat. Biotechnol. 2017 3511, vol. 35, no. 11, pp. 1069–1076, Oct. 2017 (PMID:28967887) URL: https://www.nature.com/articles/nbt.3960.
- A. Wesolowska-Andersen, M. I. Bahl, V. Carvalho, et al., “Choice of bacterial DNA extraction method from fecal material influences community structure as evaluated by metagenomic analysis,” Microbiome, vol. 2, no. 1, pp. 1–11, Jun. 2014 URL: https://microbiomejournal.biomedcentral.com/articles/10.1186/2049-2618-2-19.
- M. Y. Lim, E. J. Song, S. H. Kim, J. Lee, and Y. Do Nam, “Comparison of DNA extraction methods for human gut microbial community profiling,” Syst. Appl. Microbiol., vol. 41, no. 2, pp. 151–157, Mar. 2018 (PMID:29305057) URL: https://pubmed.ncbi.nlm.nih.gov/29305057/.
- J. P. Shaffer, C. S. Carpenter, C. Martino, et al., “A Comparison of Six DNA Extraction Protocols for 16S, ITS and Shotgun Metagenomic Sequencing of Microbial Communities,” Biotechniques, vol. 73, no. 1, pp. 34–46, Jun. 2022 (PMID:35713407) URL: https://www.tandfonline.com/doi/abs/10.2144/btn-2022-0032.
- C. Elie, M. Perret, H. Hage, et al., “Comparison of DNA extraction methods for 16S rRNA gene sequencing in the analysis of the human gut microbiome,” Sci. Reports 2023 131, vol. 13, no. 1, pp. 1–12, Jun. 2023 (PMID:37355726) URL: https://www.nature.com/articles/s41598-023-33959-6.
- K. Dixit, D. Davray, D. Chaudhari, et al., “Benchmarking of 16S rRNA gene databases using known strain sequences,” Bioinformation, vol. 17, no. 3, p. 377, Mar. 2021 (PMID:34092959) URL: /pmc/articles/PMC8131573/.
- A. Hiergeist, J. Ruelle, S. Emler, and A. Gessner, “Reliability of species detection in 16S microbiome analysis: Comparison of five widely used pipelines and recommendations for a more standardized approach,” PLoS One, vol. 18, no. 2, p. e0280870, Feb. 2023 (PMID:36795699) URL: https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0280870.
- L.-A. G. Molano, S. Vega-Abellaneda, and C. Manichanh, “GSR-DB: a manually curated and optimized taxonomical database for 16S rRNA amplicon analysis,” mSystems, vol. 9, no. 2, Feb. 2024 (PMID:38189256) URL: https://journals.asm.org/doi/10.1128/msystems.00950-23.
- B. Gil-Pulido, E. Tarpey, W. Finnegan, X. Zhan, A. D. W. Dobson, and N. O’Leary, “Dominance of the genus Polaromonas in the microbial ecology of an Intermittently Aerated Sequencing Batch Reactor (IASBR) treating dairy processing wastewater under varying aeration rates,” J. Dairy Res., vol. 85, no. 3, pp. 388–390, Aug. 2018 (PMID:30088464) URL: https://www.cambridge.org/core/journals/journal-of-dairy-research/article/abs/dominance-of-the-genus-polaromonas-in-the-microbial-ecology-of-an-intermittently-aerated-sequencing-batch-reactor-iasbr-treating-dairy-processing-wastewater-under-varying-aeration-rates/408F705A2F5326C109608BE1B424098F.
- L. Yuan, H. Dai, G. He, Z. Yang, and X. Jiao, “Invited review: Current perspectives for analyzing the dairy biofilms by integrated multiomics,” J. Dairy Sci., vol. 106, no. 12, pp. 8181–8192, Dec. 2023 (PMID:37641326).
- P. D. Franzmann and V. B. D. Skerman, “Agitococcus lubricus gen. nov. sp. nov., a lipolytic, twitching Coccus from freshwater,” Int. J. Syst. Bacteriol., vol. 31, no. 2, pp. 177–183, Apr. 1981 URL: https://www.microbiologyresearch.org/content/journal/ijsem/10.1099/00207713-31-2-177.
- S. Mazhar, K. N. Kilcawley, C. Hill, and O. McAuliffe, “A Systems-Wide Analysis of Proteolytic and Lipolytic Pathways Uncovers The Flavor-Forming Potential of The Gram-Positive Bacterium Macrococcus caseolyticus subsp. caseolyticus,” Front. Microbiol., vol. 11, Jul. 2020 URL: /pmc/articles/PMC7358451/.
- J. E. Keller, S. Schwendener, J. Neuenschwander, G. Overesch, and V. Perreten, “Prevalence and characterization of methicillin-resistant Macrococcus spp. in food producing animals and meat in Switzerland in 2019,” Schweiz. Arch. Tierheilkd., vol. 164, no. 2, pp. 153–164, 2022 (PMID:35103598).
- K. B. E., B. Lasse, M. Patrick, et al., “Impact of Sample Type and DNA Isolation Procedure on Genomic Inference of Microbiome Composition,” mSystems, vol. 1, no. 5, p. 10.1128/msystems.00095-16, Oct. 2016. [CrossRef]
- F. Fouhy, A. G. Clooney, C. Stanton, M. J. Claesson, and P. D. Cotter, “16S rRNA gene sequencing of mock microbial populations-impact of DNA extraction method, primer choice and sequencing platform,” BMC Microbiol., vol. 16, no. 1, pp. 1–13, Jun. 2016 (PMID:27342980) URL: https://link.springer.com/articles/10.1186/s12866-016-0738-z.
- N. A. Kennedy, A. W. Walker, S. H. Berry, et al., “The Impact of Different DNA Extraction Kits and Laboratories upon the Assessment of Human Gut Microbiota Composition by 16S rRNA Gene Sequencing,” PLoS One, vol. 9, no. 2, p. e88982, Feb. 2014 (PMID:24586470) URL: https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0088982.
- R. C. Edgar, “Accuracy of taxonomy prediction for 16S rRNA and fungal ITS sequences,” PeerJ, vol. 2018, no. 4, 2018 URL: /pmc/articles/PMC5910792/.
- B. J. Callahan, P. J. McMurdie, and S. P. Holmes, “Exact sequence variants should replace operational taxonomic units in marker-gene data analysis,” ISME J. 2017 1112, vol. 11, no. 12, pp. 2639–2643, Jul. 2017 (PMID:28731476) URL: https://www.nature.com/articles/ismej2017119.
- M. Chiarello, M. McCauley, S. Villéger, and C. R. Jackson, “Ranking the biases: The choice of OTUs vs. ASVs in 16S rRNA amplicon data analysis has stronger effects on diversity measures than rarefaction and OTU identity threshold,” PLoS One, vol. 17, no. 2, p. e0264443, Feb. 2022 (PMID:35202411) URL: https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0264443.
- P. Harkes, A. K. A. Suleiman, S. J. J. van den Elsen, et al., “Conventional and organic soil management as divergent drivers of resident and active fractions of major soil food web constituents,” Sci. Reports 2019 91, vol. 9, no. 1, pp. 1–15, Sep. 2019 (PMID:31534146) URL: https://www.nature.com/articles/s41598-019-49854-y.
- S. Takahashi, J. Tomita, K. Nishioka, T. Hisada, and M. Nishijima, “Development of a Prokaryotic Universal Primer for Simultaneous Analysis of Bacteria and Archaea Using Next-Generation Sequencing,” PLoS One, vol. 9, no. 8, p. e105592, Aug. 2014 (PMID:25144201) URL: https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0105592.
- E. Bolyen, J. R. Rideout, M. R. Dillon, et al., “Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2,” Nat. Biotechnol. 2019 378, vol. 37, no. 8, pp. 852–857, Jul. 2019 (PMID:31341288) URL: https://www.nature.com/articles/s41587-019-0209-9.
- N. A. Bokulich, B. D. Kaehler, J. R. Rideout, et al., “Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin,” Microbiome, vol. 6, no. 1, pp. 1–17, May 2018 (PMID:29773078) URL: https://microbiomejournal.biomedcentral.com/articles/10.1186/s40168-018-0470-z.
- D. E. Wood, J. Lu, and B. Langmead, “Improved metagenomic analysis with Kraken 2,” Genome Biol., vol. 20, no. 1, pp. 1–13, Nov. 2019 (PMID:31779668) URL: https://genomebiology.biomedcentral.com/articles/10.1186/s13059-019-1891-0.
- J. Lu, F. P. Breitwieser, P. Thielen, and S. L. Salzberg, “Bracken: Estimating species abundance in metagenomics data,” PeerJ Comput. Sci., vol. 2017, no. 1, p. e104, Jan. 2017 URL: https://peerj.com/articles/cs-104.
- J. Lu, N. Rincon, D. E. Wood, et al., “Metagenome analysis using the Kraken software suite,” Nat. Protoc. 2022 1712, vol. 17, no. 12, pp. 2815–2839, Sep. 2022 (PMID:36171387) URL: https://www.nature.com/articles/s41596-022-00738-y.
- H. Lin and S. Das Peddada, “Analysis of compositions of microbiomes with bias correction,” Nat. Commun., vol. 11, no. 1, Dec. 2020 (PMID:32665548) URL: /pmc/articles/PMC7360769/.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



