Submitted:
28 August 2024
Posted:
28 August 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
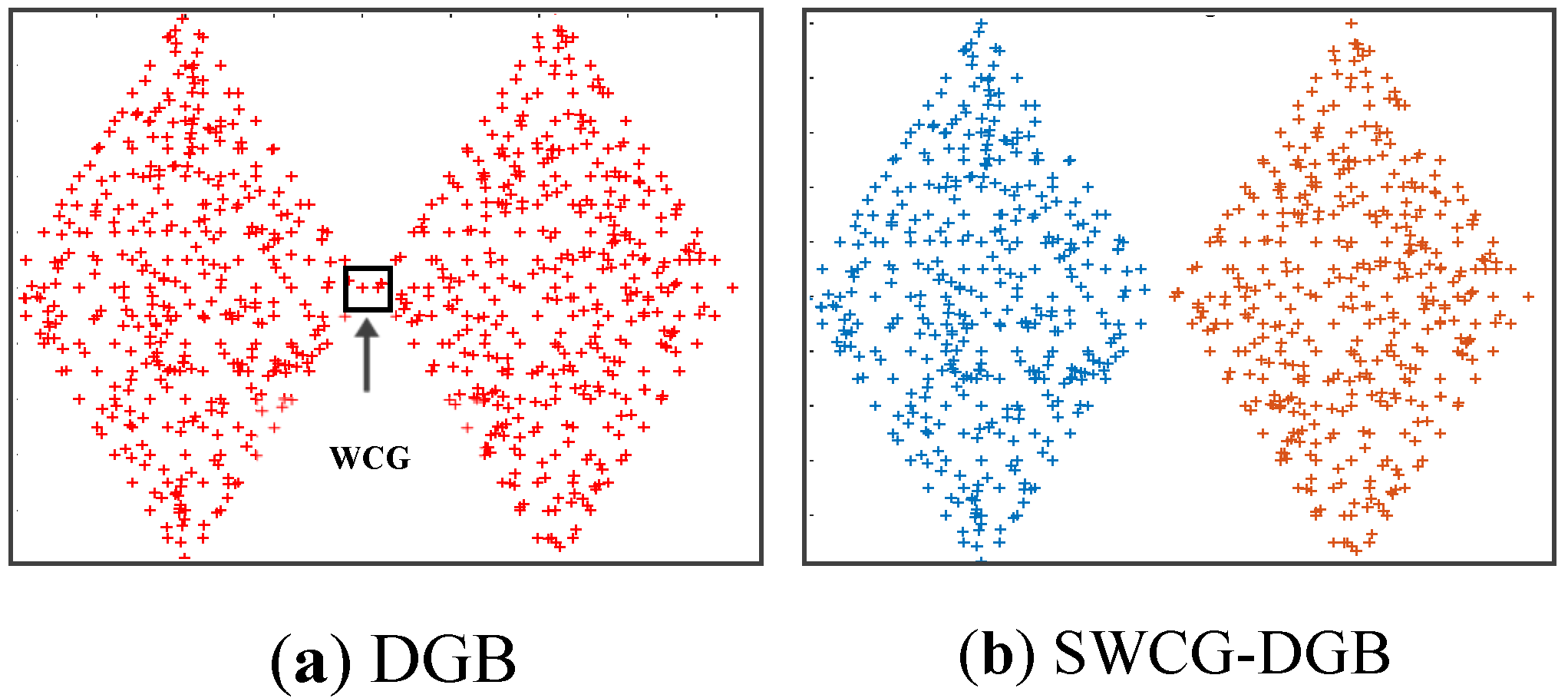
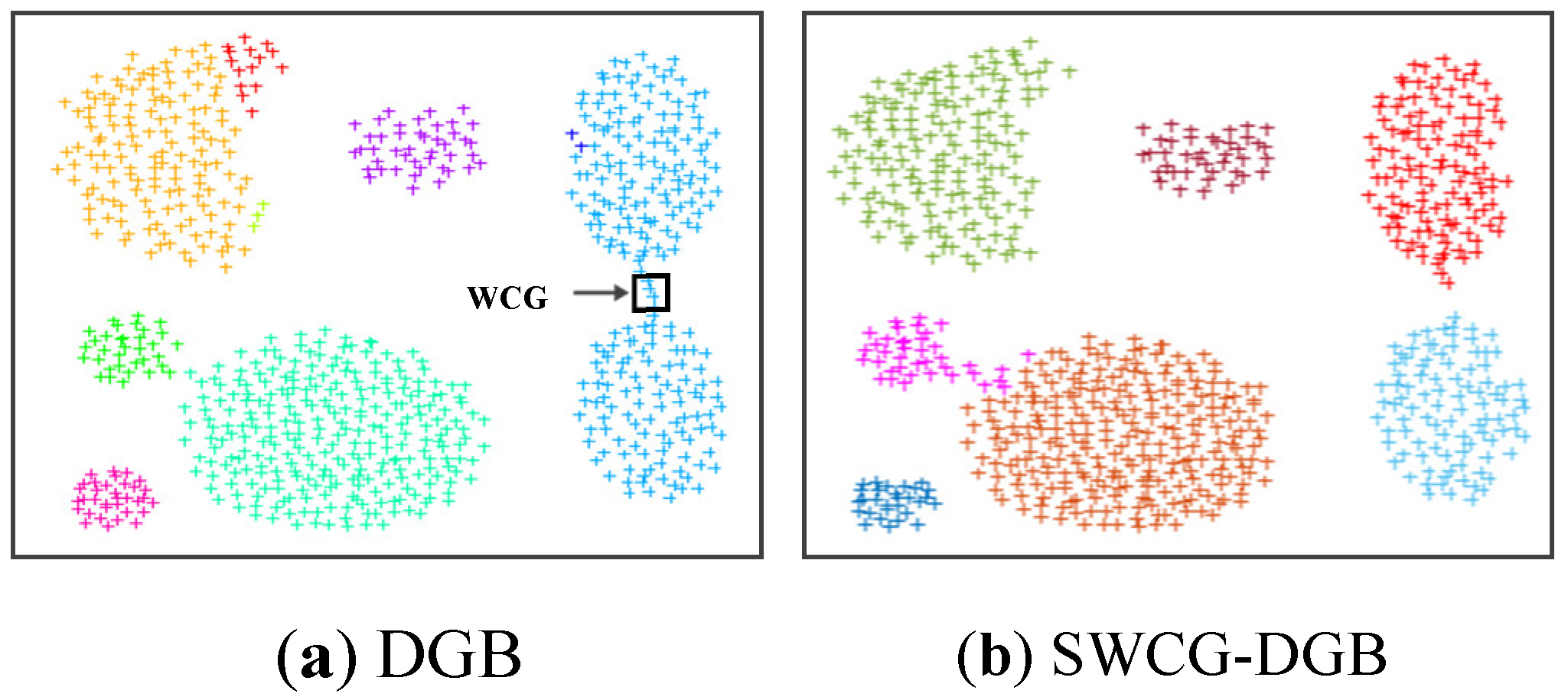
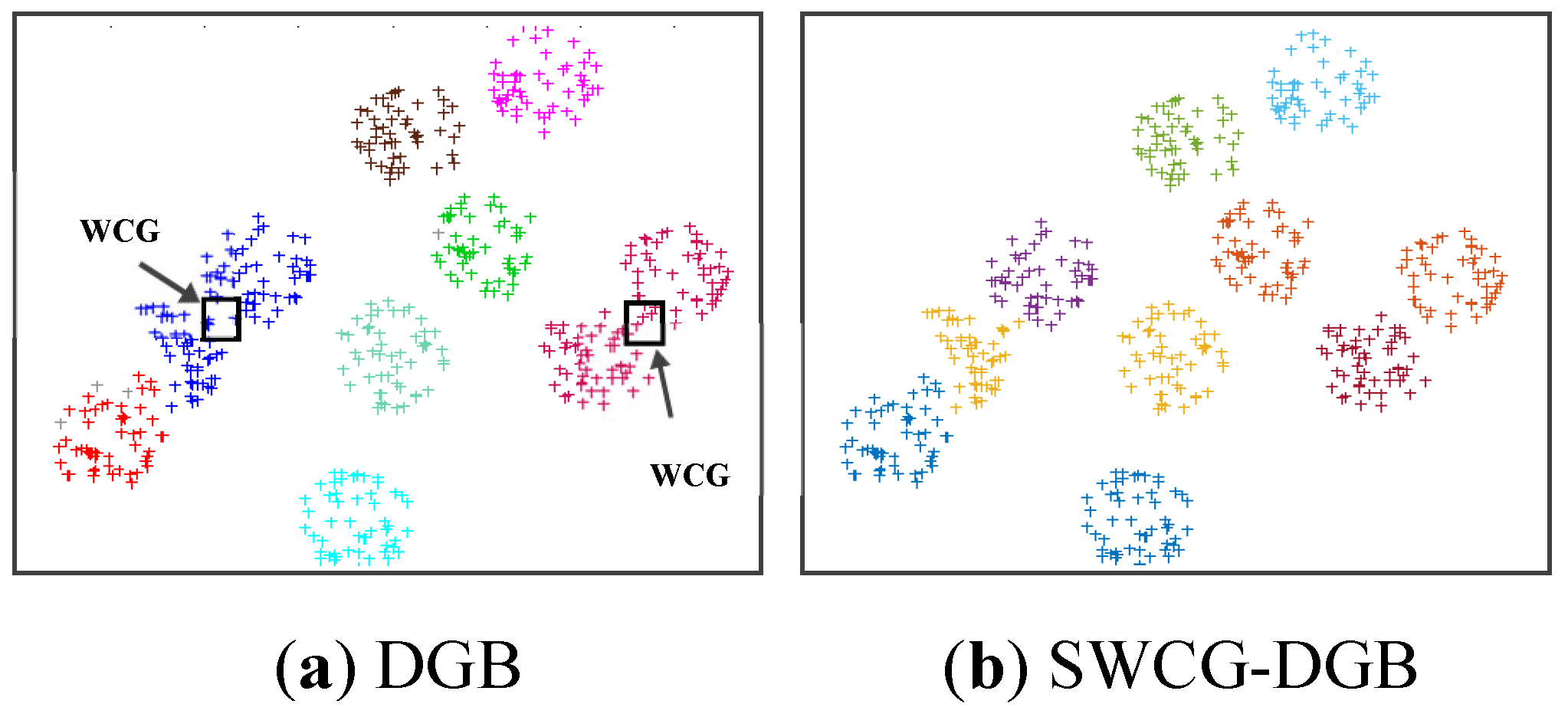
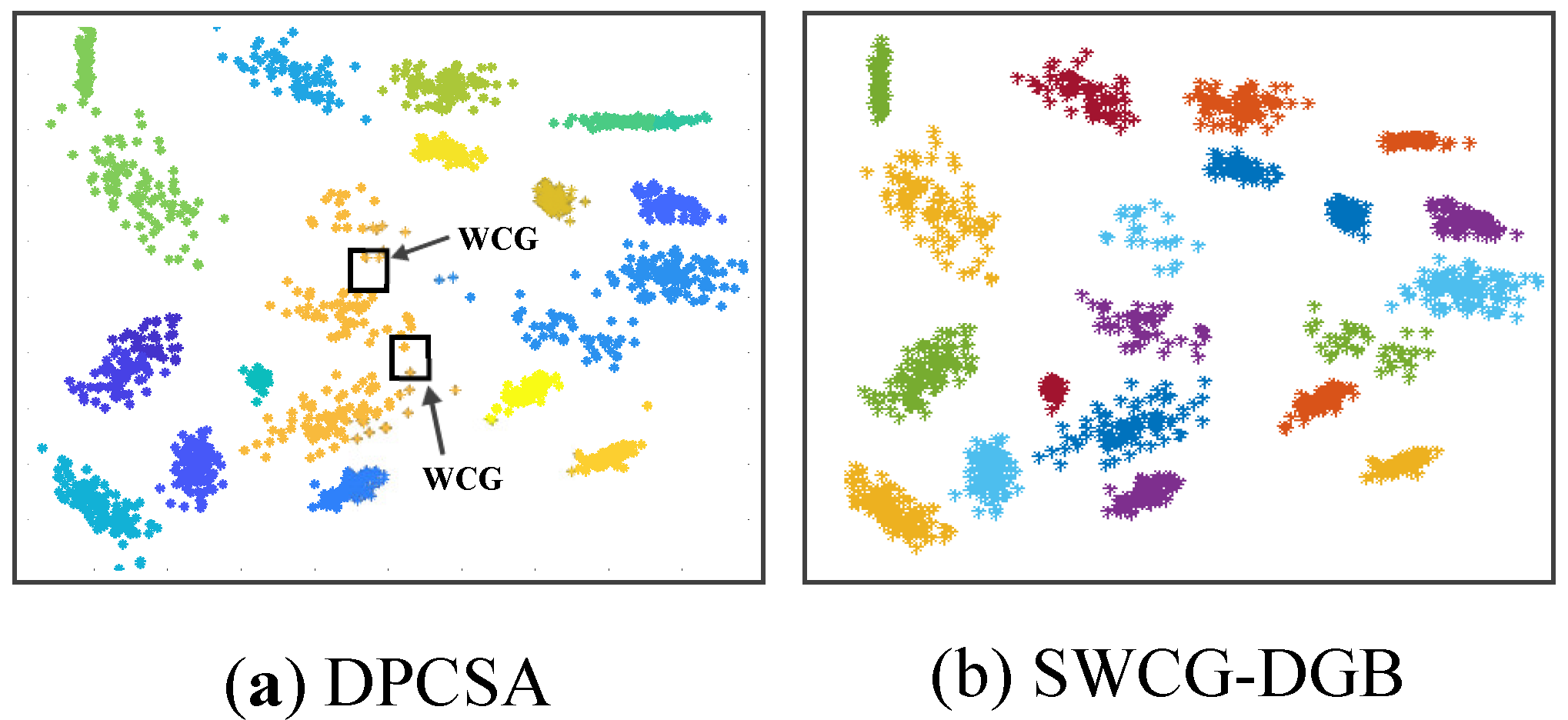
- We point out a common drawback often appearing in the density grid-based clustering approach related to the WCG. Accordingly, we propose a solution for improving the clustering validity, including filtering noise and fine-tuning the generated CDS.
- We propose a buffer for storing features of data points and the grid-cell’s coordinates. It allows detecting WCGs more effectively with a lower computational cost. The reason is that only the data points marked in the buffer are indicated as border or noise points instead of the entire dataset. This enhancement is very efficient for large datasets.
2. Related Concepts
3. Proposed Method
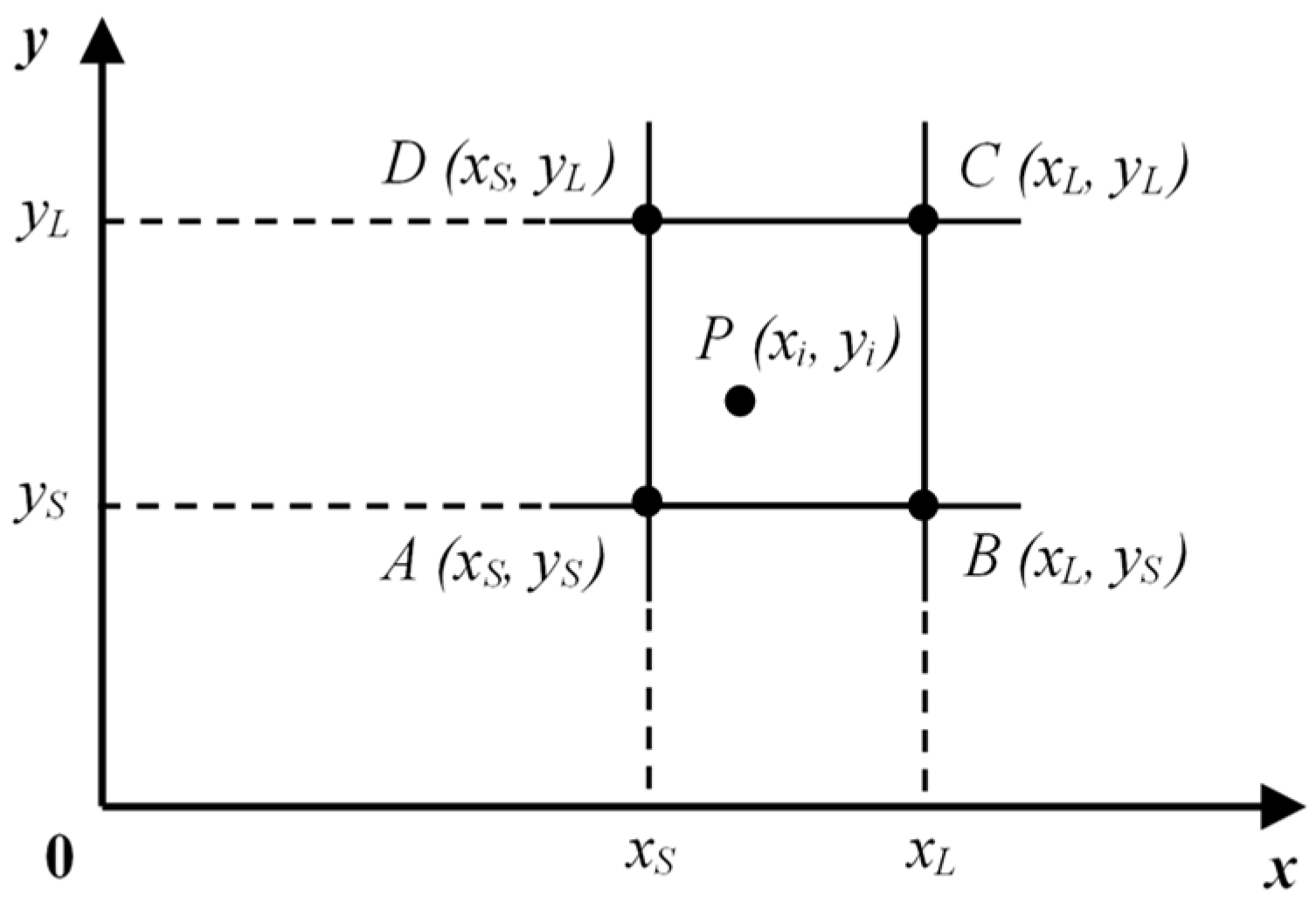
3.1. Initialization
3.2. Finding the Number of Clusters
| Algorithm 1. Searching mountain ridges |
|
Inputs: All data points. Outputs: The updating position of the node in the buffer and CL set. 1: Calculating density of nodes and add them into density_matrix 2: Reshaping density_matrix, finding the nodei has max_density 3: while (max_density > edge_factor ) 4: for i =1 to length(k) of high_density_nodes set 5: (xi, yi)=get_position(rowi, coli) 6: cluster_matrix(rowi, coli)=n (no_cluster) 7: updating node(xj, yj) into the buffer and CL_set 8: for m =k+1 to length of (no_grid*no_grid) 9: if (density of nodem > edge_factor) 10: (xm, ym)=get_position(rowm, colm) 11: if (node(xm, ym) is neighbor of node(xi, yi) ) 12: cluster_matrix(rowm, colm) = cluster_matrix(rowi, coli) 13: density_matrix(rowm, colm)=max_density 14: high_density_nodes=[high_density_nodes m] 15: updating node(xm, ym) into the buffer and CL_set 16: end (if 11) 17: end (if 9) 18: end (for 8) 19: end (for 4) 20: high_density_nodes=[1]; n=n+1 21: end (while 3) |
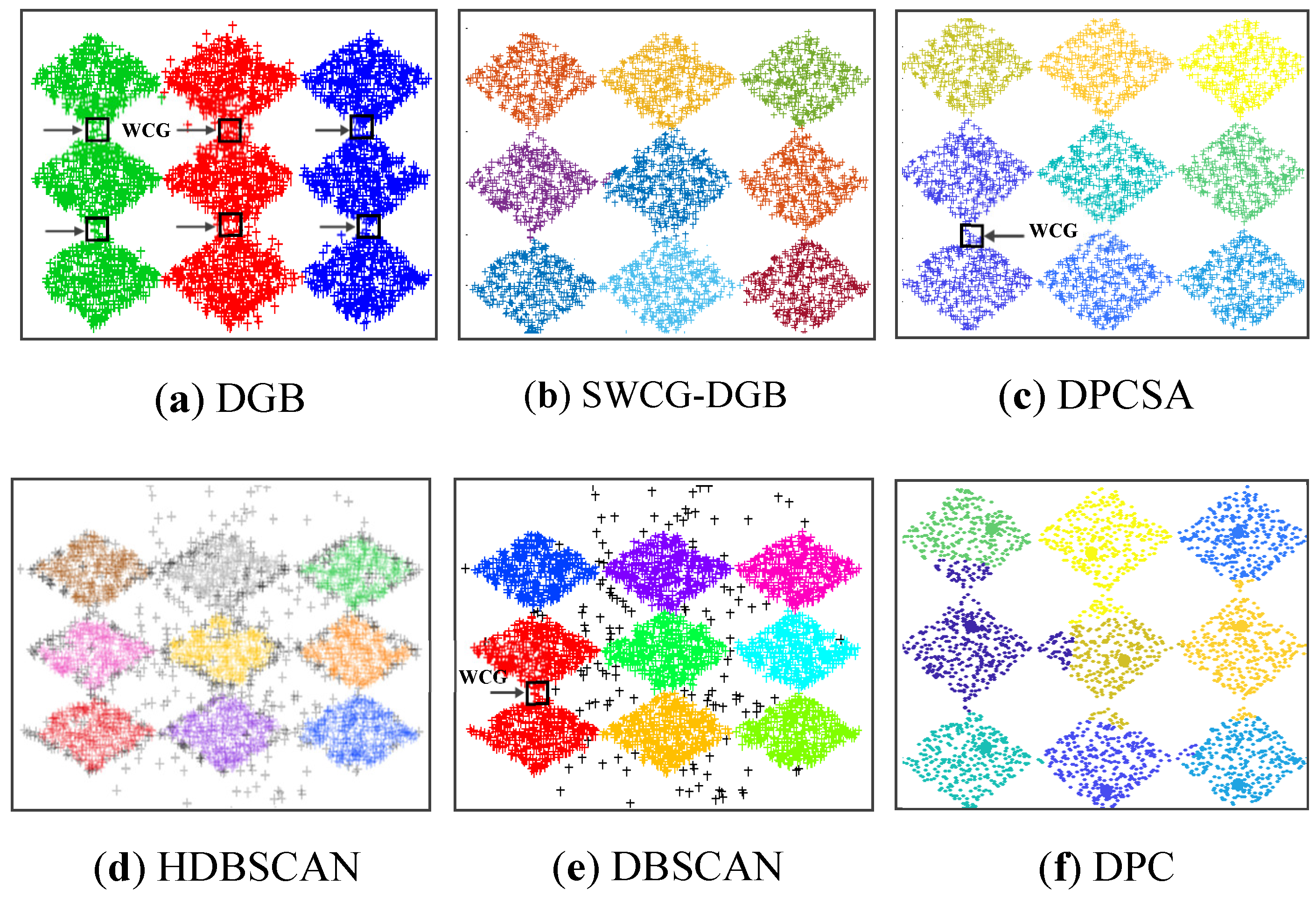
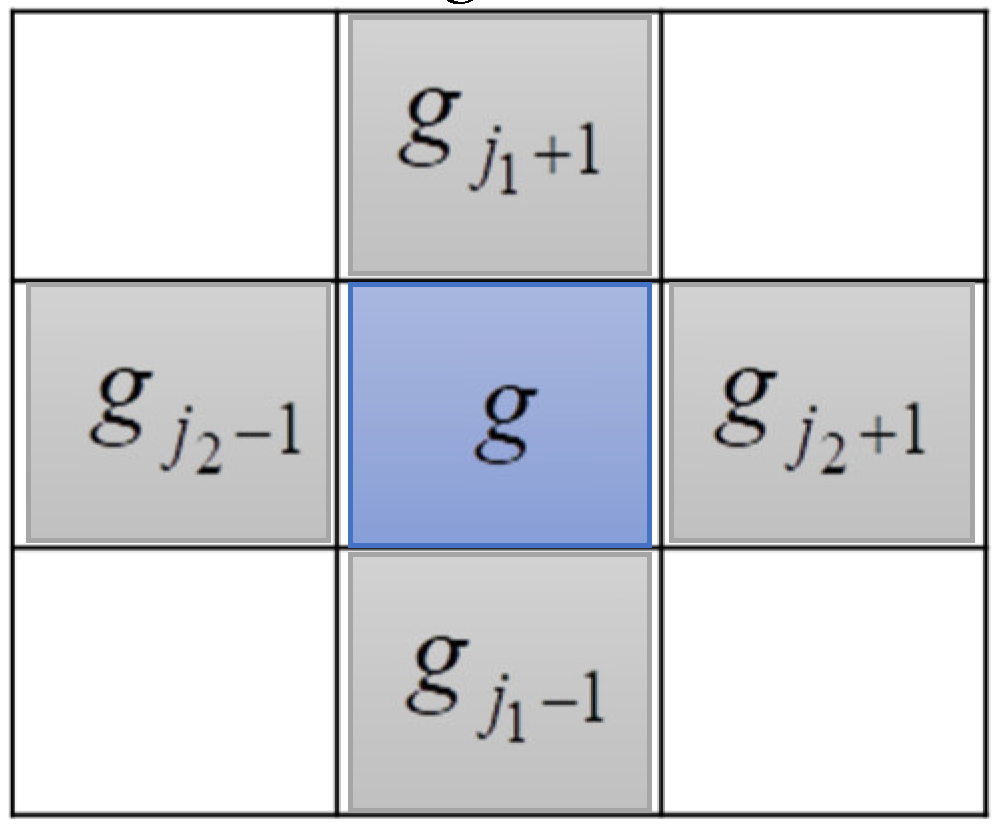
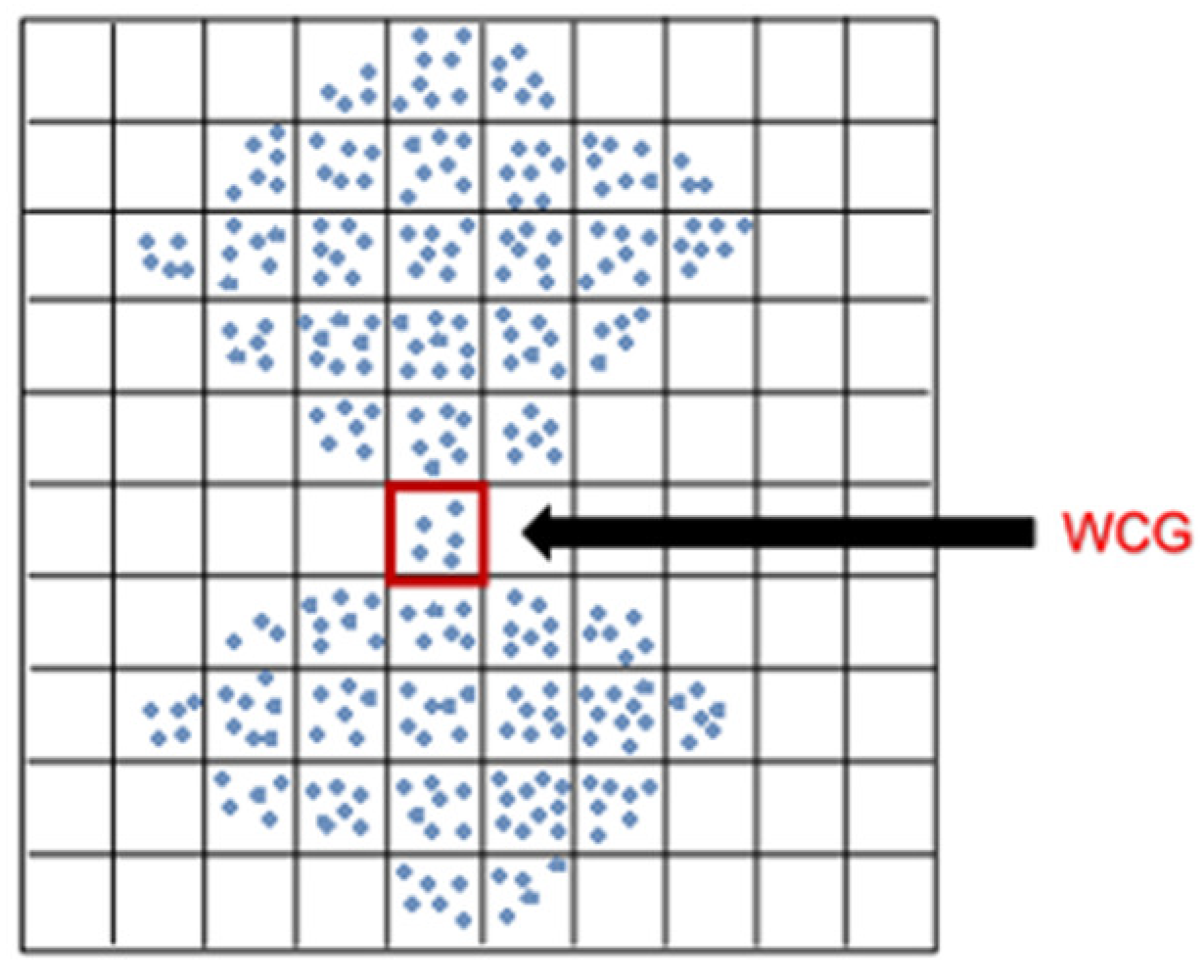
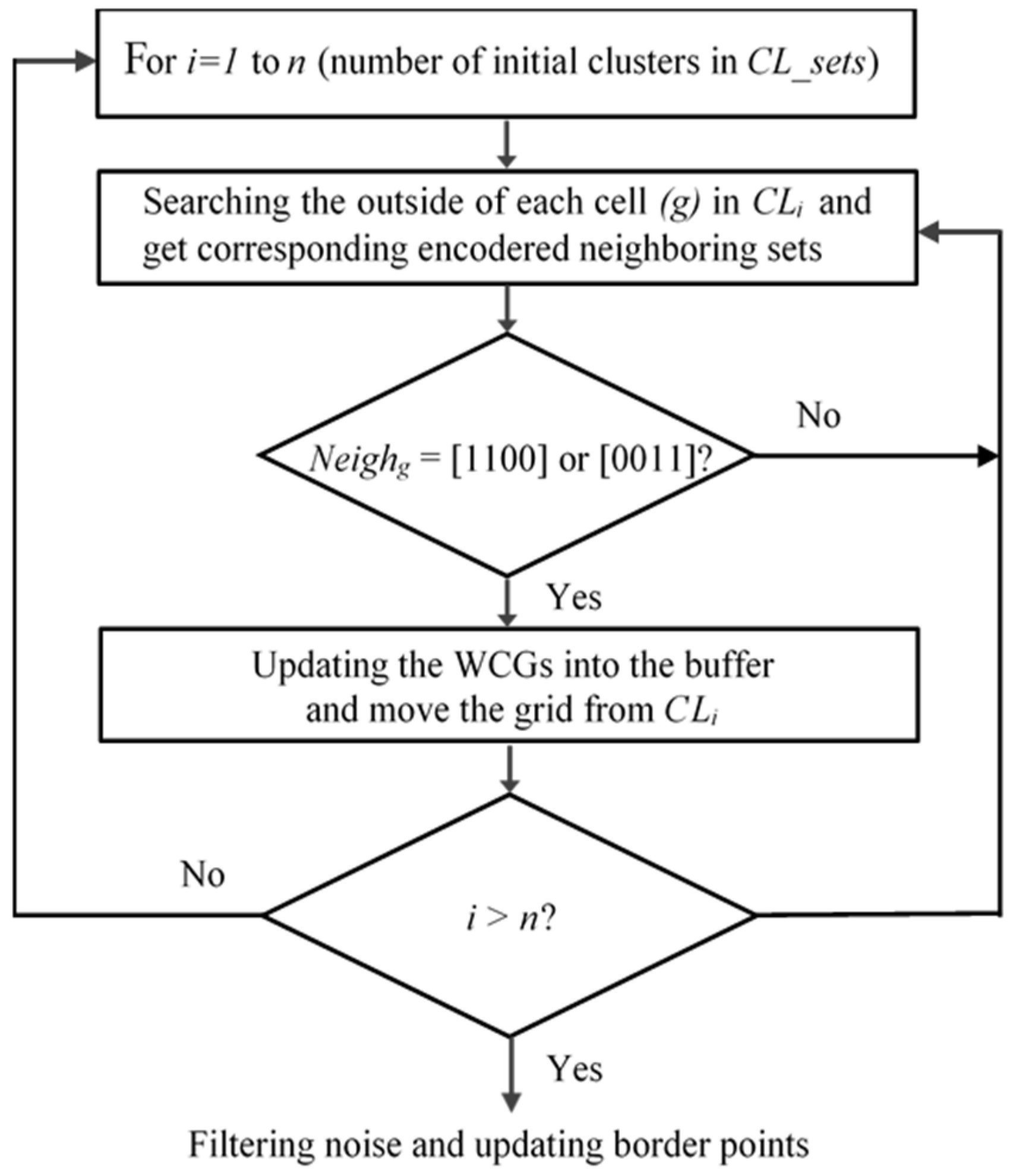
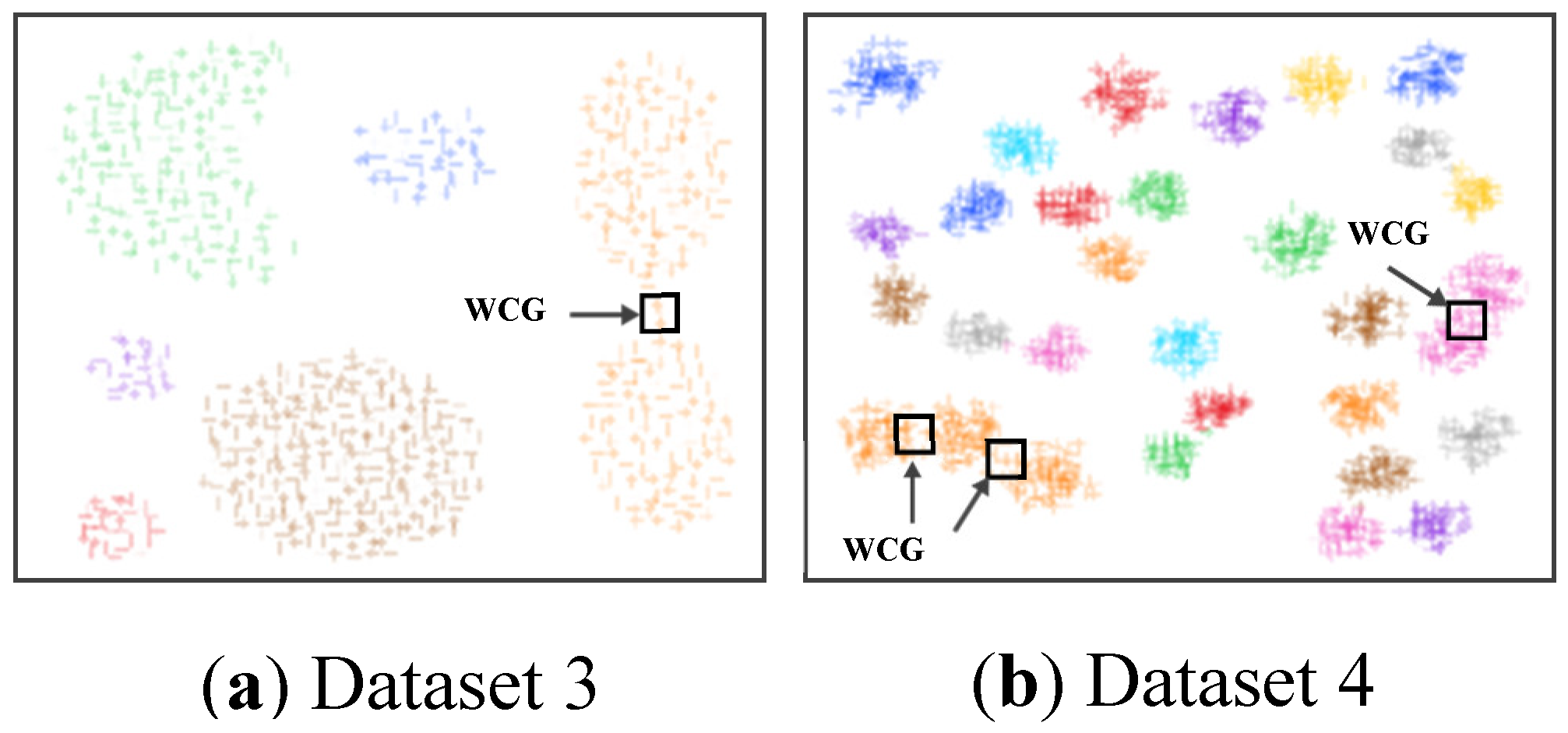
3.3. Searching the Weak-Connected Grids
| Algorithm 2. Searching WCGs |
|
Input: All grids on the CL set. Output: Detecting and moving the WCGs out of each CL set. 1: for i =1 to length_of CL_set 2: for m =1 to length_of CLi 3: if gridm is outside of CLi 4: [Neighm]= get_neighbors of gridm 5: if Neighm =[0; 0; 1; 1] or [1; 1; 0; 0] 6: marking data points of (gridm) as noise in buffer 7: moving data points of (gridm) out of CLi 8: (xm, ym)=get position of (gridm) from buffer 9: density_matrix (rowm, colm) = 0 10: end (if 5) 11: end (if 3) 12: end (for 2) 13: end (for 1) |
3.4. Filtering Noise and Updating Border Points
| Algorithm 3. Filtering noise and updating border points |
|
Input: All points of zero-number_grid set in buffer. Output: Removing noise and updated border points in obtained clusters. 1: for i =1 to length of zero-number_grid set in buffer 2: get_position(rowi, coli); 3: if ( cluster_matrix(rowi, coli)=0 and density_matrix (rowi, coli) > noise_threshold ) 4: for k=1 to 4 5: if cluster_matrix (nodek) > 0 6: satisfactory_node = satisfactory_node+1; 7: sum = sum(satisfactory_node); 8: end (if 5) 9: end (for 4) 10: end (if 3) 11: if sum = 1, get density of satisfactory node 12: if node_ density <noise_threshold 13: cluster_matrix (rowi, coli)=0; 14: gridi = noise; break; 15: else 16: move gridi to corresponding cluster; 17: end (if 12) 18: end (if 11) 19: if sum > 1, get total density of satisfactory nodes 20: if total node_ density <noise_threshold 21: cluster_matrix (rowi, coli)=0; 22: gridi = noise; break; 23: else 24: move gridi to corresponding cluster; 25: end (if 20) 26: end (if 19) 27: end (for 1) |
| Algorithm 4. The SWCG-DGB |
| 1: Normalizing each data point P(xi,yi) following equations (13) and (14). 2: Calculating corner coordinates of each data point P(xi,yi) and update C(xL, yL) into the buffer. 3: Calculating the local density of nodes following equations (4)-(7) and add them into the density_matrix. 4: Finding the mountain ridges (or initial clusters) and map data points corresponding to these clusters into CL_sets (Algorithm 1). 5: Searching the Weak-Connected Grids (Algorithm 2). 6: Filtering Noise and Updating Border Points (Algorithm 3). |
4. Experimental Simulation
4.1. Approach
- The RNC is visually observed to estimate the accuracy.
4.2. Evaluation Measures
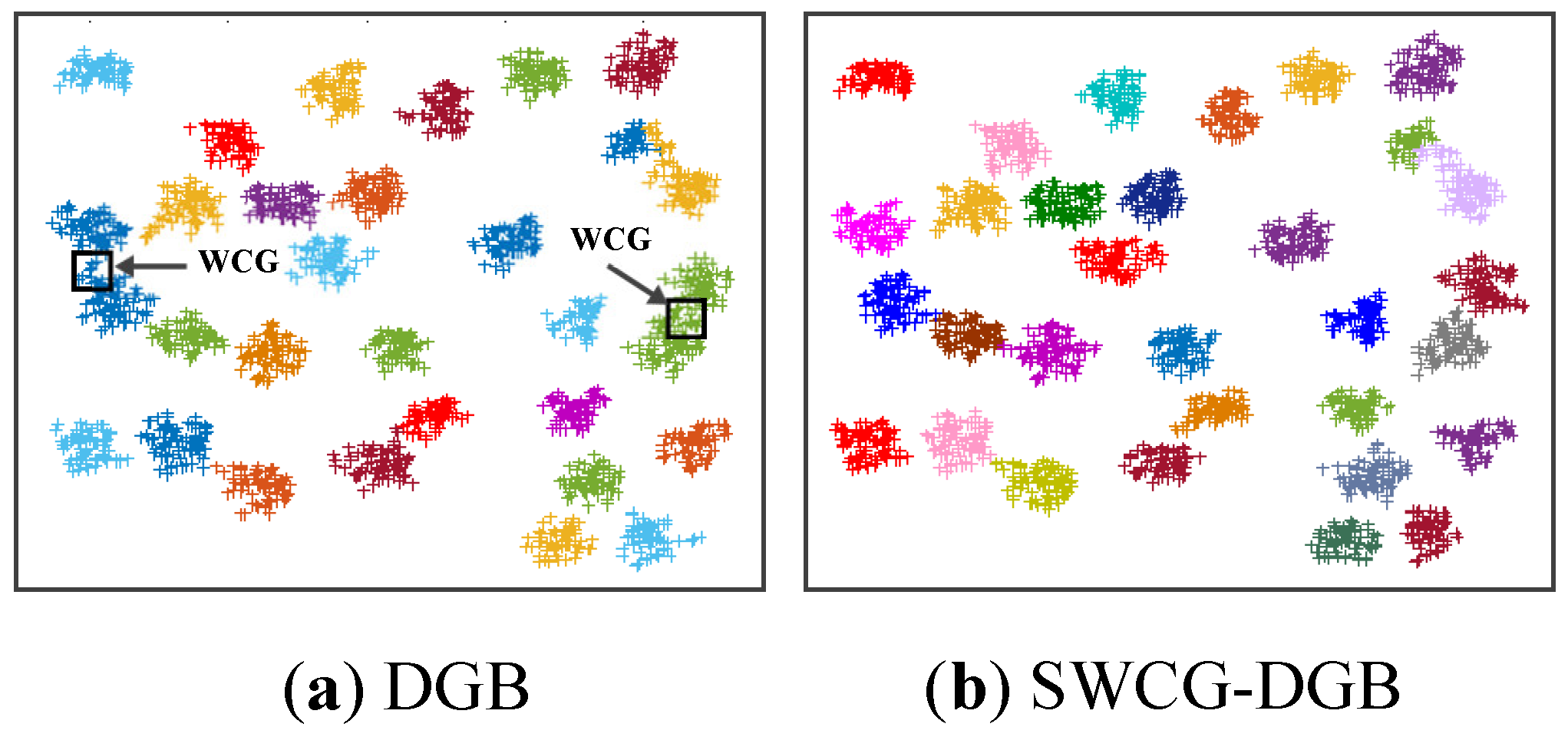
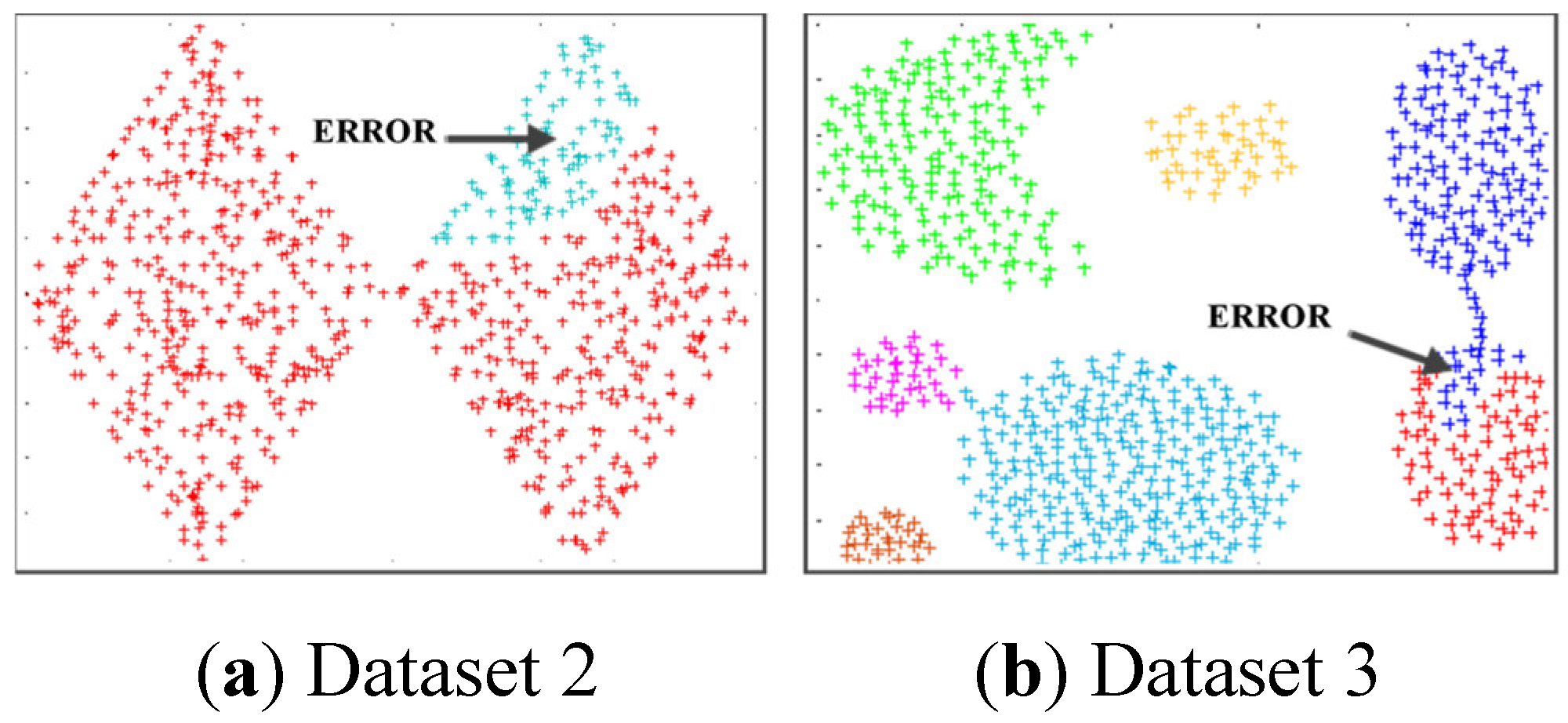
4.3. Simulating Results on Synthetic Datasets
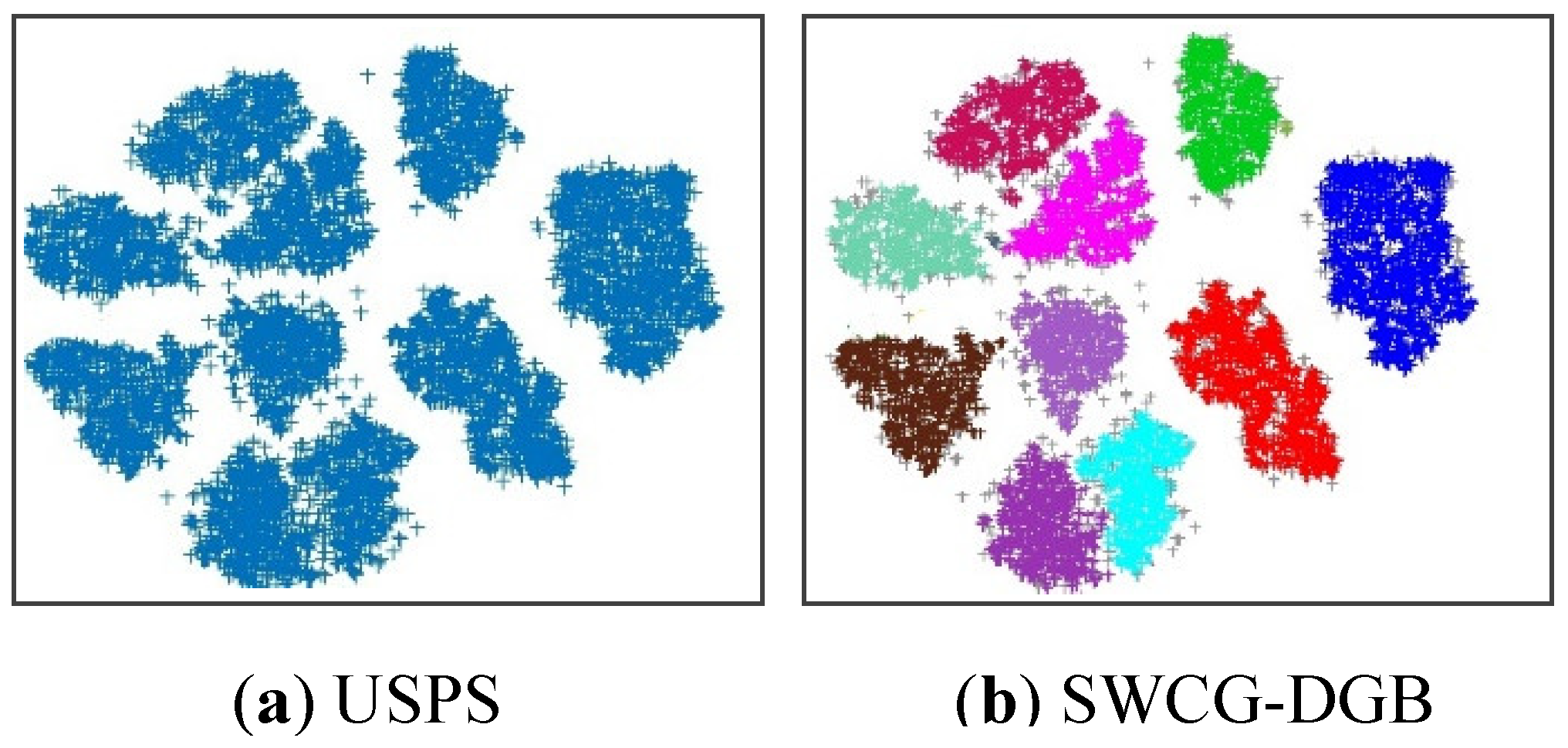
4.3. Simulating Results on Real Datasets
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Salem; Semeh Ben; Sami Naouali; Zied Chtourou. A fast and effective partitional clustering algorithm for large categorical datasets using a k-means based approach. Computers & Elect. Eng. 2018, 68, 463–483. [Google Scholar] [CrossRef]
- Chaira; Tamalika. A novel intuitionistic fuzzy C means clustering algorithm and its application to medical images. Applied soft computing. 2011, 1711–1717. [Google Scholar] [CrossRef]
- Saxena; Amit; et al. A review of clustering techniques and developments. Neurocomputing. 2017, 267, 664–681. [Google Scholar] [CrossRef]
- Li, S.; Li, L.; Yan, J.; He, H. SDE: A novel clustering framework based on sparsity-density entropy. IEEE Transactions on Knowledge and Data Engineering. 2018, 1575–1587. [Google Scholar] [CrossRef]
- Saelens, W.; Cannoodt, R.; Saeys, Y. A comprehensive evaluation of module detection methods for gene expression data. Nature comm. 2018, 1090. [Google Scholar] [CrossRef] [PubMed]
- Ester, M.; Kriegel, H. P.; Sander, J.; Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. kdd. 1996, 226–231. [Google Scholar]
- Cheng; Yizong. Mean shift, mode seeking, and clustering. IEEE transactions on pattern analysis and machine intelligence, 1995; 790–799.
- Georgoulas, G.; Konstantaras, A.; Katsifarakis, E.; Stylios, C. D.; Maravelakis, E.; Vachtsevanos, G. J. “Seismic-mass” density-based algorithm for spatio-temporal clustering. Expert Systems with Applications, 2013; 4183–4189. [Google Scholar]
- Marques; João C. ; Michael B. Orger. “Clusterdv: a simple density-based clustering method that is robust, general and automatic.” Bioinformatics, 2019, 2125-2132.
- Chang; Hong; Dit-Yan Yeung. Robust path-based spectral clustering. Pattern Recognition, 2008; 191–203. [CrossRef]
- Chen; Xinquan. A new clustering algorithm based on near neighbor influence. Expert Systems with applications, 2015; 7746–7758. [CrossRef]
- Huang; Anna. Similarity measures for text document clustering. Proceedings of the sixth new zealand computer science research student conference (NZCSRSC2008), Christchurch, New Zealand, 2008, 9–56.
- Uncu, O.; Gruver, W. A.; Kotak, D. B.; Sabaz, D.; Alibhai, Z.; Ng, C. GRIDBSCAN: GRId density-based spatial clustering of applications with noise. 2006 IEEE Inter. Conf. on Systems, Man and Cybe. vol. 4. IEEE, 2006, 2976-2981.
- Chen, Y.; Tang, S.; Bouguila, N.; Wang, C. , Du, J.; Li, H. A fast clustering algorithm based on pruning unnecessary distance computations in DBSCAN for high-dimensional data. Pattern Recog, 2018, 83, 375–387. [Google Scholar] [CrossRef]
- Marques; J. C.; Orger, M. B. Clusterdv: a simple density-based clustering method that is robust, general and automatic. Bioinformatics, 2019, 2125-2132. [Google Scholar] [CrossRef]
- Campello; R. J.; Moulavi, D.; Sander, J. Density-based clustering based on hierarchical density estimates. Pacific-Asia conf. on knowledge discovery and data mining. Springer, Berlin, Heidelberg, 2013, 160-172.
- McInnes, L.; Healy, J.; Astels, S. hdbscan: Hierarchical density based clustering. J. Open Source Softw, 2017; 205. [Google Scholar] [CrossRef]
- Rodriguez, A.; Laio, A. Clustering by fast search and find of density peaks. Science, 2014; 1492–1496. [Google Scholar] [CrossRef]
- Yu, D.; Liu, G.; Guo, M.; Liu, X.; Yao, S. Density peaks clustering based on weighted local density sequence and nearest neighbor assignment. IEEE Access, 2019; 7, 34301–34317. [Google Scholar] [CrossRef]
- Wu; Bo; Bogdan M. Wilamowski. A fast density and grid based clustering method for data with arbitrary shapes and noise. IEEE Transactions on Industrial Informatics, 2016; 1620–1628. [CrossRef]
- Cheng, C. H.; Fu, A. W.; Zhang, Y. Entropy-based subspace clustering for mining numerical data. Proceedings of the fifth ACM SIGKDD international conference on Knowledge discovery and data mining, 1999; 84–93. [Google Scholar]
- Duan, D.; Li, Y.; Li, R.; Lu, Z. “Incremental K-clique clustering in dynamic social networks. Artificial Intelligence Review, 2012; 129–147. [Google Scholar] [CrossRef]
- Chen, Y. , Tu, L. Density-based clustering for real-time stream data. Proceedings of the 13th ACM SIGKDD Inter. Conf. on Knowledge discovery and data mining, 2007, 133-142.
- Li, H.; Wu, C.; Jing, X.; Wu, L. Fuzzy tracking control for nonlinear networked systems. IEEE Transactions on Cybernetics, 2016; 2020–2031. [Google Scholar] [CrossRef]
- Nguyen; Sy Dzung; Vu Song Thuy Nguyen; Nhat Truong Pham. Determination of The Optimal Number of Clusters: A Fuzzy-set based Method. IEEE Transactions on Fuzzy Systems, 2021. [CrossRef]
- Van Der Maaten; Laurens. Accelerating t-SNE using tree-based algorithms. The journal of machine learning research, 2014; 3221–3245.
- Rand; William M. Objective criteria for the evaluation of clustering methods. Journal of the American Statistical association, 1971; 846–850.
- Hubert, L.; Arabie, P. Comparing partitions journal of classification 2 193–218. Google Scholar, 1985; 128–193. [Google Scholar]
- Strehl, A.; Ghosh, J. Cluster ensembles---a knowledge reuse framework for combining multiple partitions. Journal of machine learning research 3. Dec 2002, 583–617. [Google Scholar]
- https://www.csie.ntu.edu.tw/cjlin/libsvmtools/datasets/multiclass.html.
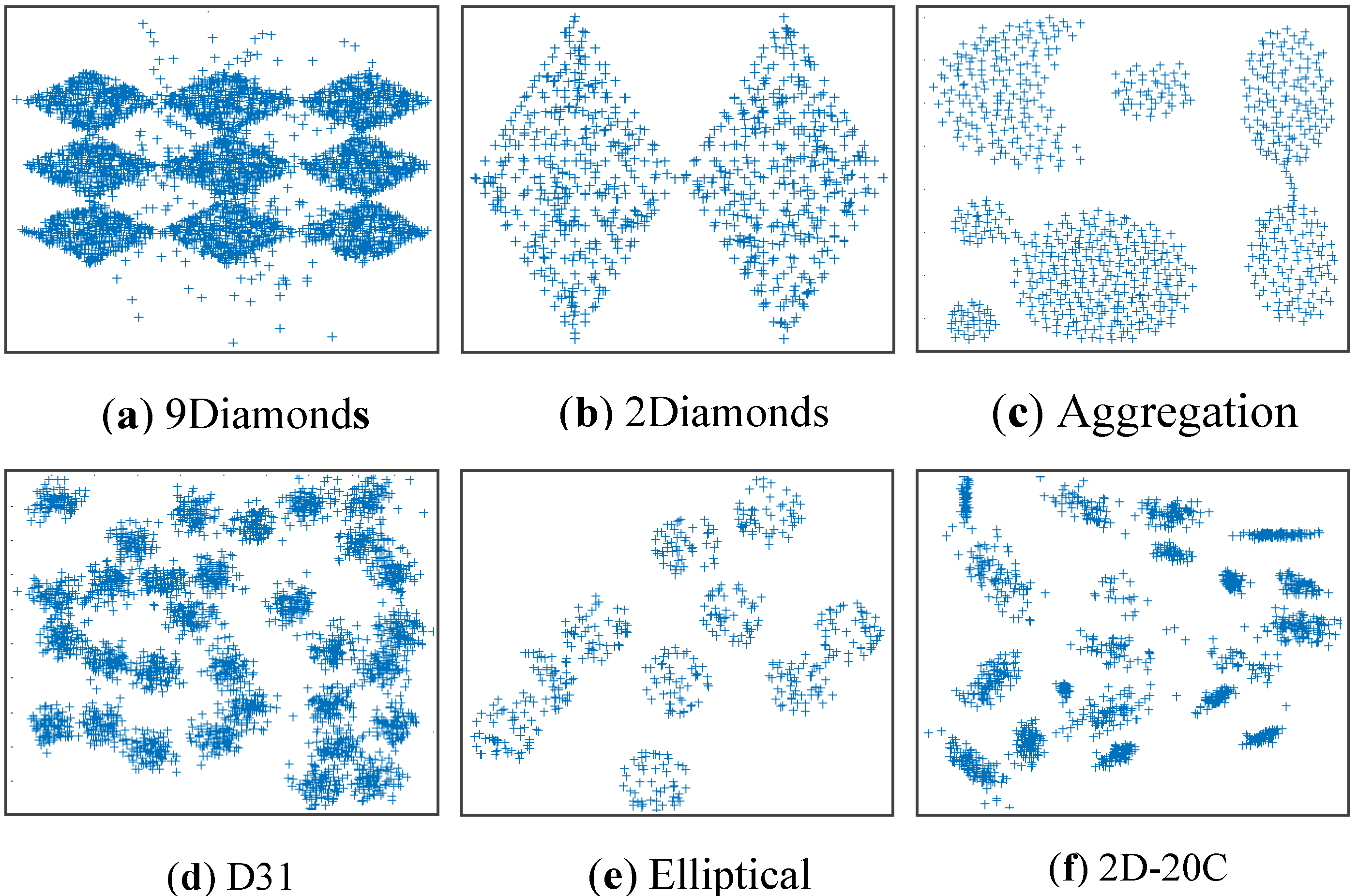





| Datasets | Samples | Dimension | RNC | Noise |
|---|---|---|---|---|
| Dataset 1 (9Diamonds) Dataset 2 (2Diamonds) Dataset 3 (Aggregation) Dataset 4 (D31) Dataset 5 (Elliptical) Dataset 6 (2D-20C) |
3300 800 788 3100 500 1517 |
2 2 2 2 2 2 |
9 2 7 31 10 20 |
Yes (5%) No No No No No |
| Dataset 7 (Iris) Dataset 8 (Seeds) Dataset 9 (USPS) |
150 210 9298 |
4 7 256 |
3 3 10 |
No No No |
| Datasets | HDBSCAN | DBSCAN | DGB | SWCG-DGB |
|---|---|---|---|---|
| minPts | minPts /ε | no_grid | no_grid | |
| Dataset 1 Dataset 2 Dataset 3 Dataset 4 Dataset 5 Dataset 6 |
15 15 15 15 15 15 |
4 / 0.12 3 / 0.6 4 / 1.0 10 / 0.6 10 / 0.6 10 / 0.6 |
20 22 16 55 44 44 |
20 22 16 55 44 44 |
| Dataset 7 Dataset 8 Dataset 9 |
15 15 15 |
6 / 0.16 10 / 0.2 4 / 0.12 |
60 34 66 |
60 34 66 |
| Methods | Datasets | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | ||||
| DBSCAN HDBSCAN DPC DPCSA DGB SWCG-DGB |
8 9 9 10 3 9 |
1 2 2 2 1 2 |
9 6 7 7 6 7 |
29 28 31 31 29 31 |
7 8 10 10 8 10 |
23 18 19 18 12 20 |
|||
| Method | Datasets | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | ||||
| DBSCAN HDBSCAN DPC DPCSA DGB SWCG-DGB |
0.336 0.044 1.215 2.459 0.464 0.356 |
0.860 0.015 0.621 0.596 0.923 0.513 |
0.559 0.011 0.372 0.548 0.487 0.361 |
1.347 0.033 1.952 2.686 1.487 1.457 |
0.871 0.020 0.413 0.661 0.423 0.302 |
3.012 0.049 1.114 2.577 0.590 0.352 |
|||
| Method | Datasets | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | ||||
| DBSCAN HDBSCAN DPC DPCSA DGB SWCG-DGB |
0.792 0.951 0.954 0.801 0.798 0.972 |
0.756 0.910 0.918 0.721 0.756 0.942 |
0.795 0.936 0.970 0.820 0.812 0.976 |
0.730 0.960 0.976 0.919 0.852 0.961 |
0.786 0.971 0.972 0.716 0.721 0.983 |
0.798 0.906 0.922 0.738 0.797 0.936 |
|||
| Method | Datasets | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | ||||
| DBSCAN HDBSCAN DPC DPCSA DGB SWCG-DGB |
0.641 0.743 0.891 0.750 0.727 0.904 |
0.632 0.801 0.910 0.603 0.689 0.915 |
0.755 0.819 0.922 0.729 0.734 0.939 |
0.623 0.805 0.908 0.835 0.780 0.942 |
0.704 0.901 0.855 0.461 0.627 0.913 |
0.720 0.602 0.896 0.590 0.705 0.901 |
|||
| Method | Datasets | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | ||||
| DBSCAN HDBSCAN DPC DPCSA DGB SWCG-DGB |
0.716 0.832 0.841 0.765 0.742 0.932 |
0.702 0.815 0.816 0.678 0.713 0.928 |
0.811 0.875 0.972 0.790 0.789 0.953 |
0.718 0.855 0.953 0.862 0.832 0.957 |
0.723 0.940 0.912 0.524 0.664 0.952 |
0.755 0.723 0.811 0.622 0.765 0.933 |
|||
| Methods | QNC | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Dataset 7 (Iris) |
Dataset 8 (Seeds) | Dataset 9 (USPS) | ||||||||
| DBSCAN HDBSCAN DPC DPCSA DGB SWCG-DGB |
2 2 3 2 3 3 |
2 3 3 3 3 3 |
15 12 9 11 10 10 |
|||||||
| Methods | RI | ARI | NMI | |
|---|---|---|---|---|
| Dataset 7 (Iris) | ||||
| DBSCAN | 0.751 | 0.682 | 0.740 | |
| HDBSCAN | 0.776 | 0.568 | 0.733 | |
| DPC | 0.935 | 0.893 | 0.917 | |
| DPCSA | 0.892 | 0.886 | 0.870 | |
| DGB | 0.955 | 0.902 | 0.919 | |
| SWCG-DGB | 0.962 | 0.931 | 0.948 | |
| Dataset 8 (Seeds) | ||||
| DBSCAN | 0.782 | 0.632 | 0.704 | |
| HDBSCAN | 0.753 | 0.413 | 0.531 | |
| DPC | 0.921 | 0.910 | 0.933 | |
| DPCSA | 0.746 | 0.703 | 0.693 | |
| DGB | 0.923 | 0.905 | 0.910 | |
| SWCG-DGB | 0.934 | 0.887 | 0.895 | |
| Dataset 9 (USPS) | ||||
| DBSCAN | 0.675 | 0.596 | 0,638 | |
| HDBSCAN | 0.973 | 0.868 | 0.882 | |
| DPC | 0.927 | 0.903 | 0.916 | |
| DPCSA | 0.851 | 0.783 | 0.843 | |
| DGB | 0.945 | 0.901 | 0.916 | |
| SWCG-DGB | 0.984 | 0.925 | 0.938 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




