Submitted:
24 August 2024
Posted:
26 August 2024
You are already at the latest version
Abstract
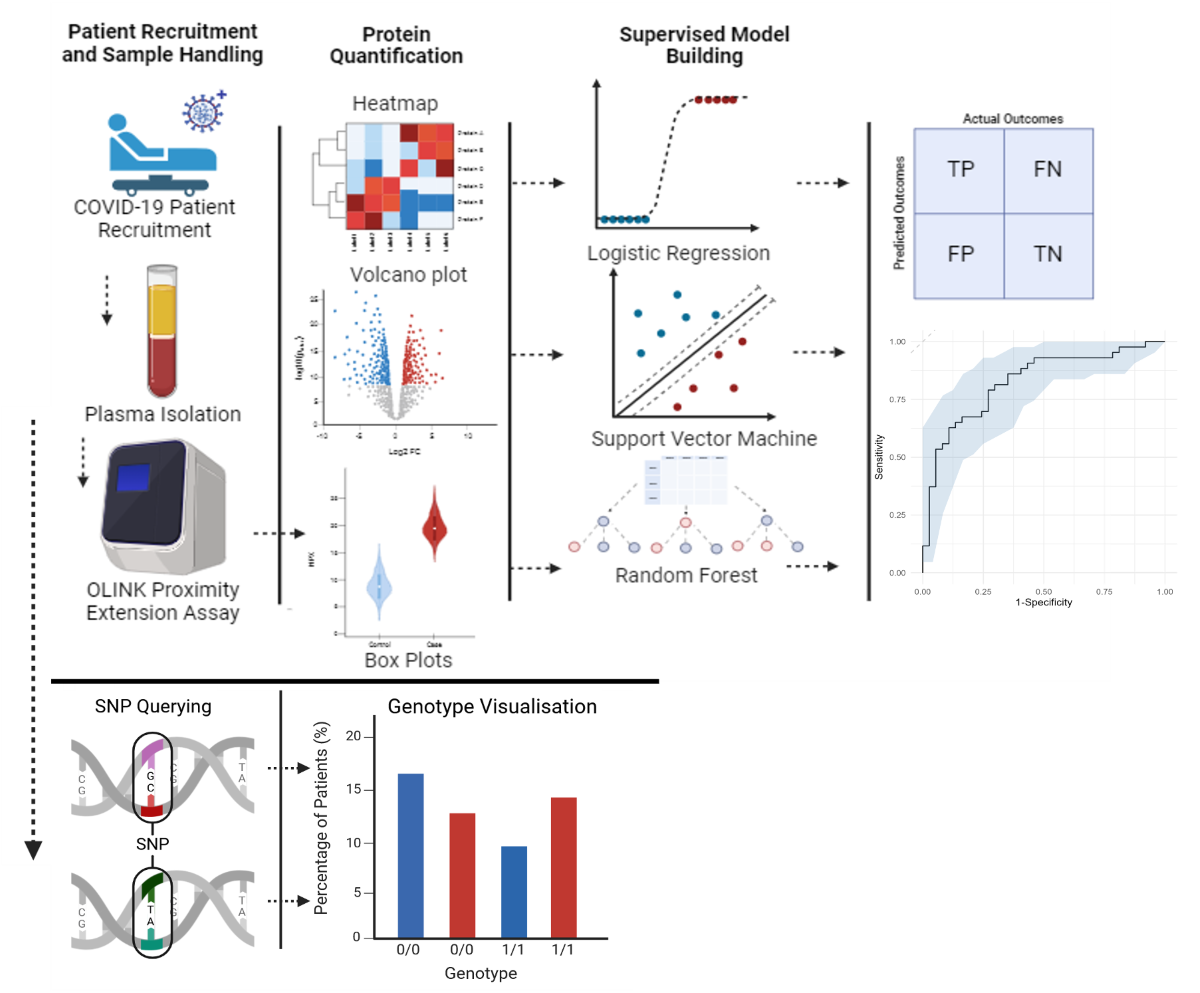
Keywords:
1. Introduction
2. Materials and Methods
2.1. Patient Recruitment
2.2.1. OLINK Plasma Protein Quantification
2.2.2. DNA Isolation
2.3. Genome Analysis
2.4. Statistical Testing
2.5. Differential Regulation Analysis
2.6. Proteomic Separation
2.7. Pathway Analysis
2.8. Machine Learning
2.9. Protein Association Analysis
3. Results
3.1. Demographics of COVID-19 Cohort
| Table 1. Base Demographics for Proteomic Analysis. Base statistics for n=400 COVID-19 patients who had their plasma proteins quantified. Table provides labelling for Age, Gender, and Severity across hospitalisation status. |
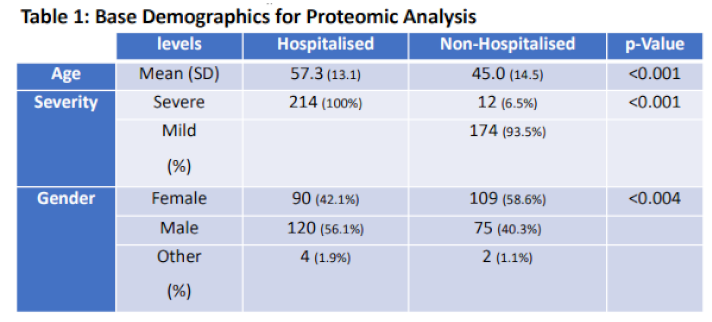 |
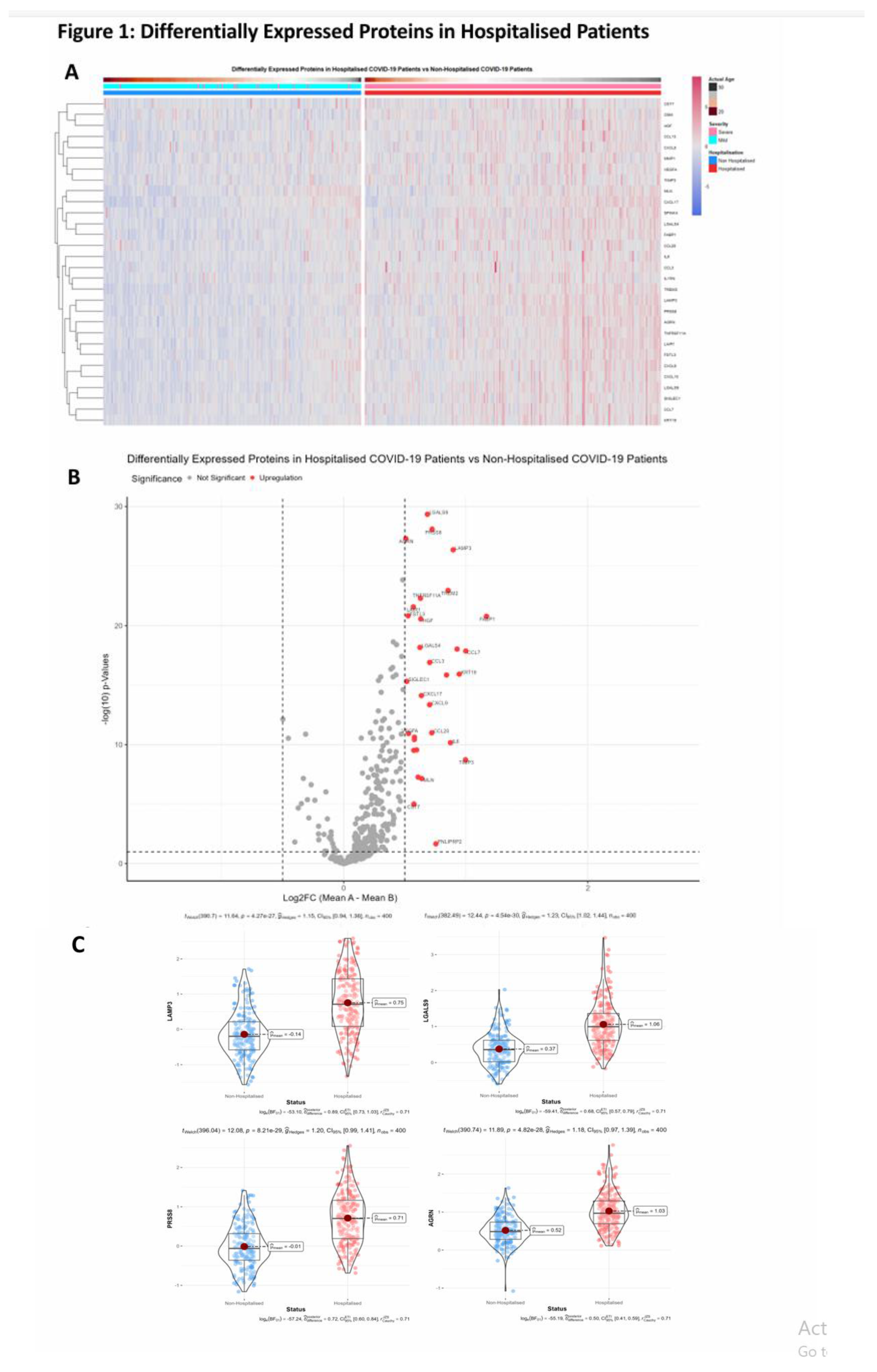
3.2. Differentially Expressed Proteins in Hospitalised Patients
| Table 2. Statistical Validation. Statistical data retrieved from testing NPX protein regulation between hospitalised COVID-19 patients and non-hospitalised COVID-19 patients, values for the top 10 most significant proteins identified. |
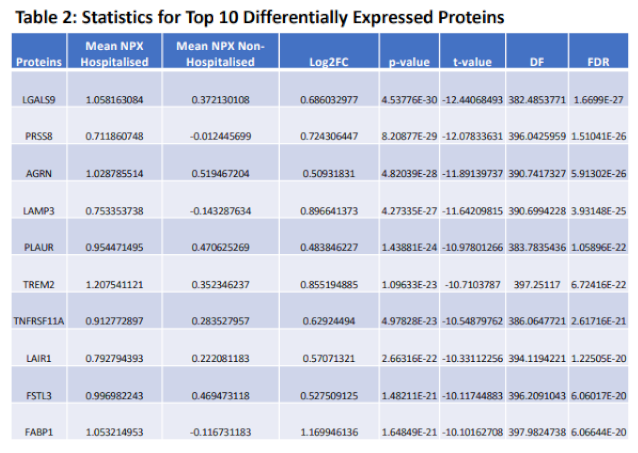 |
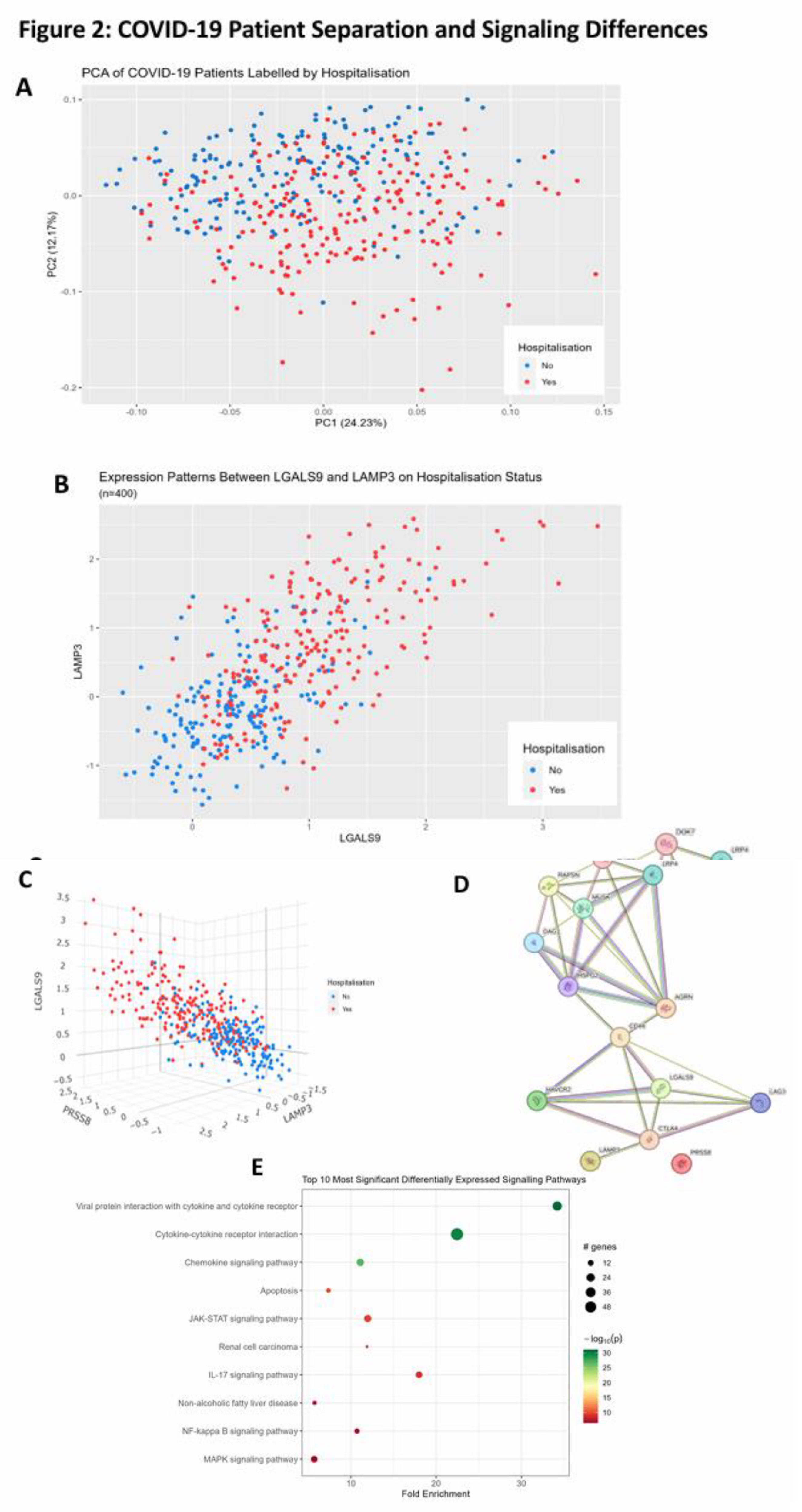
3.3. COVID-19 Patient Separation and Differential Signaling
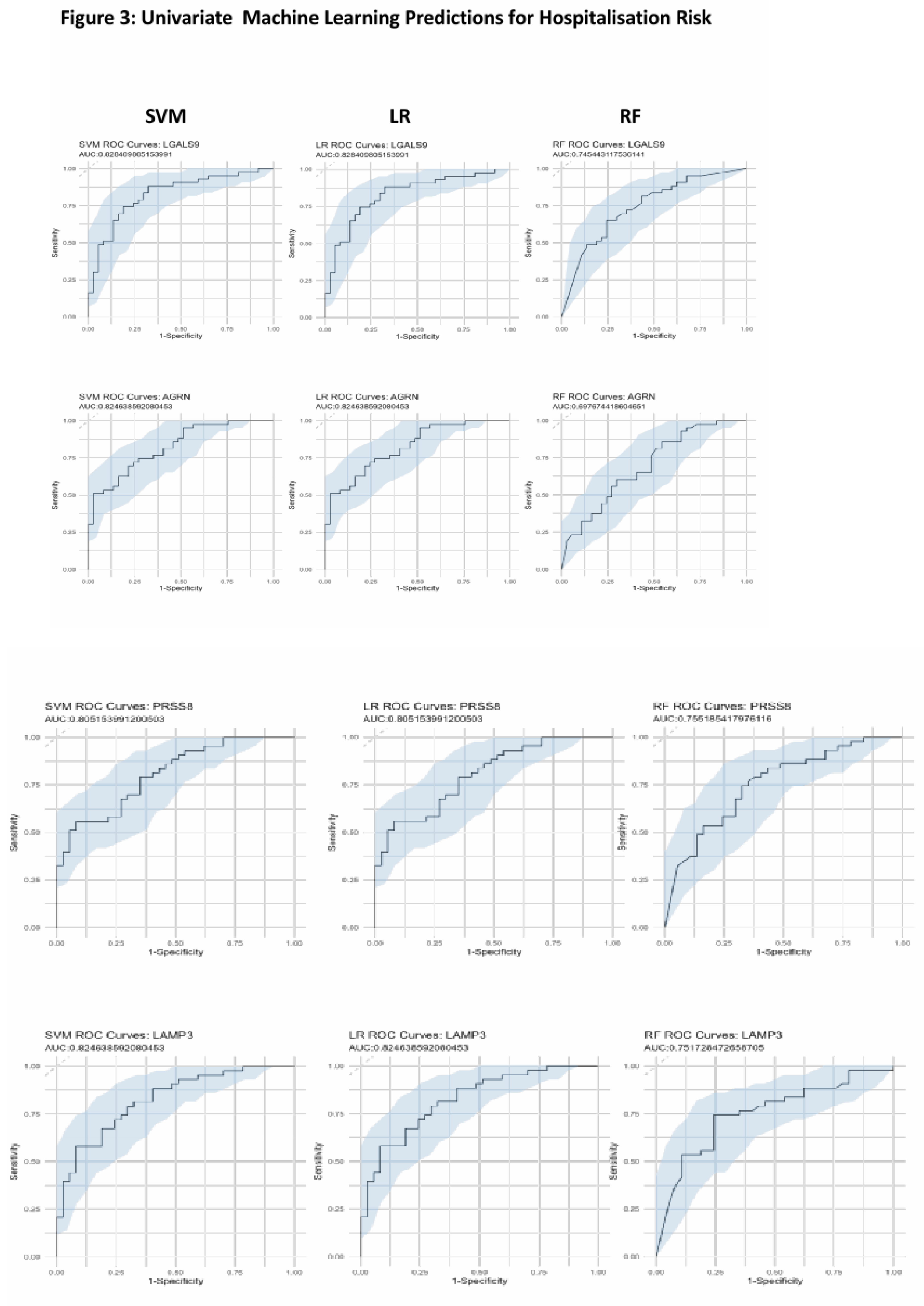
3.4. Univariate Machine Learning Predictions for Hospitalisation Risk
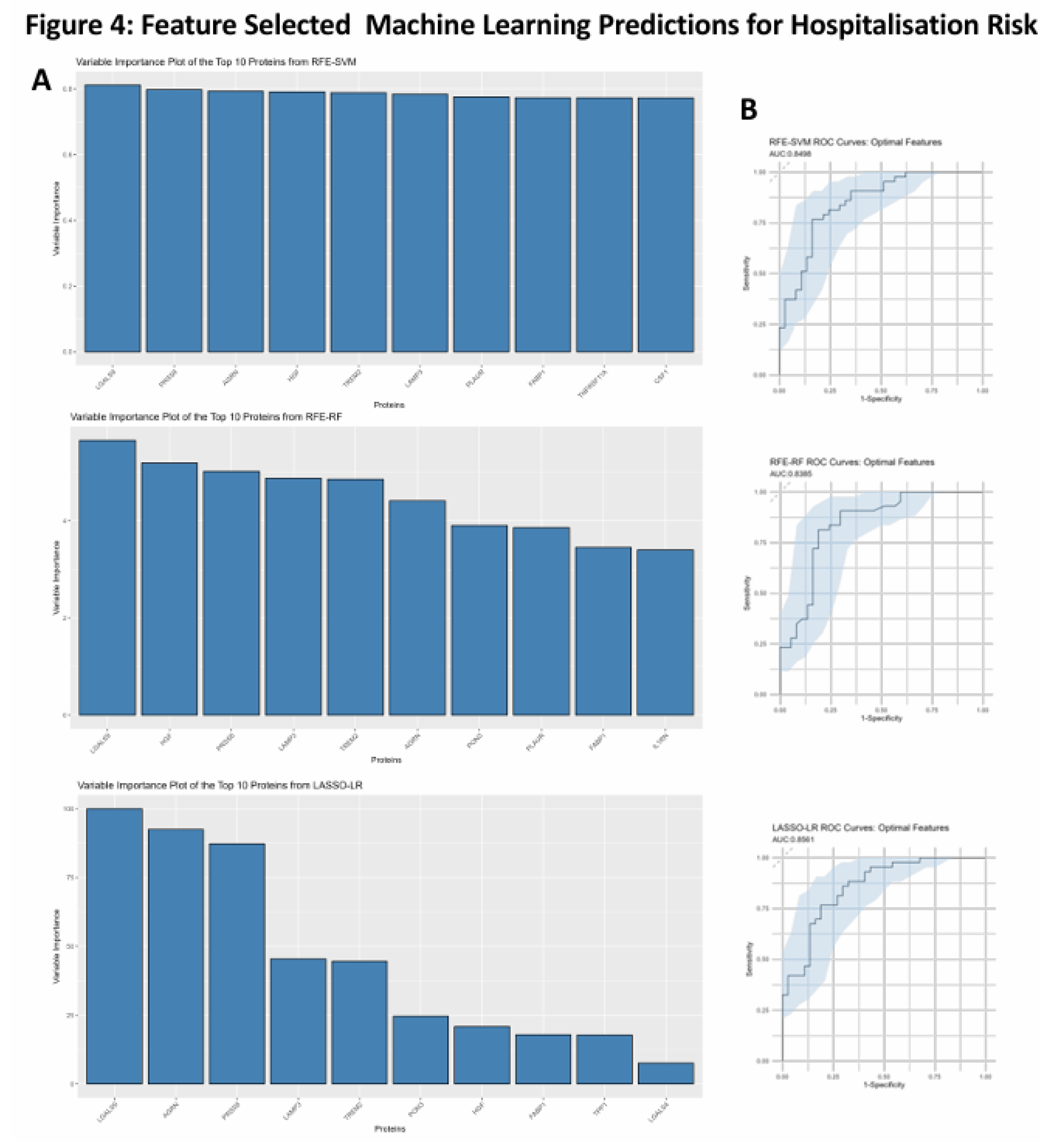
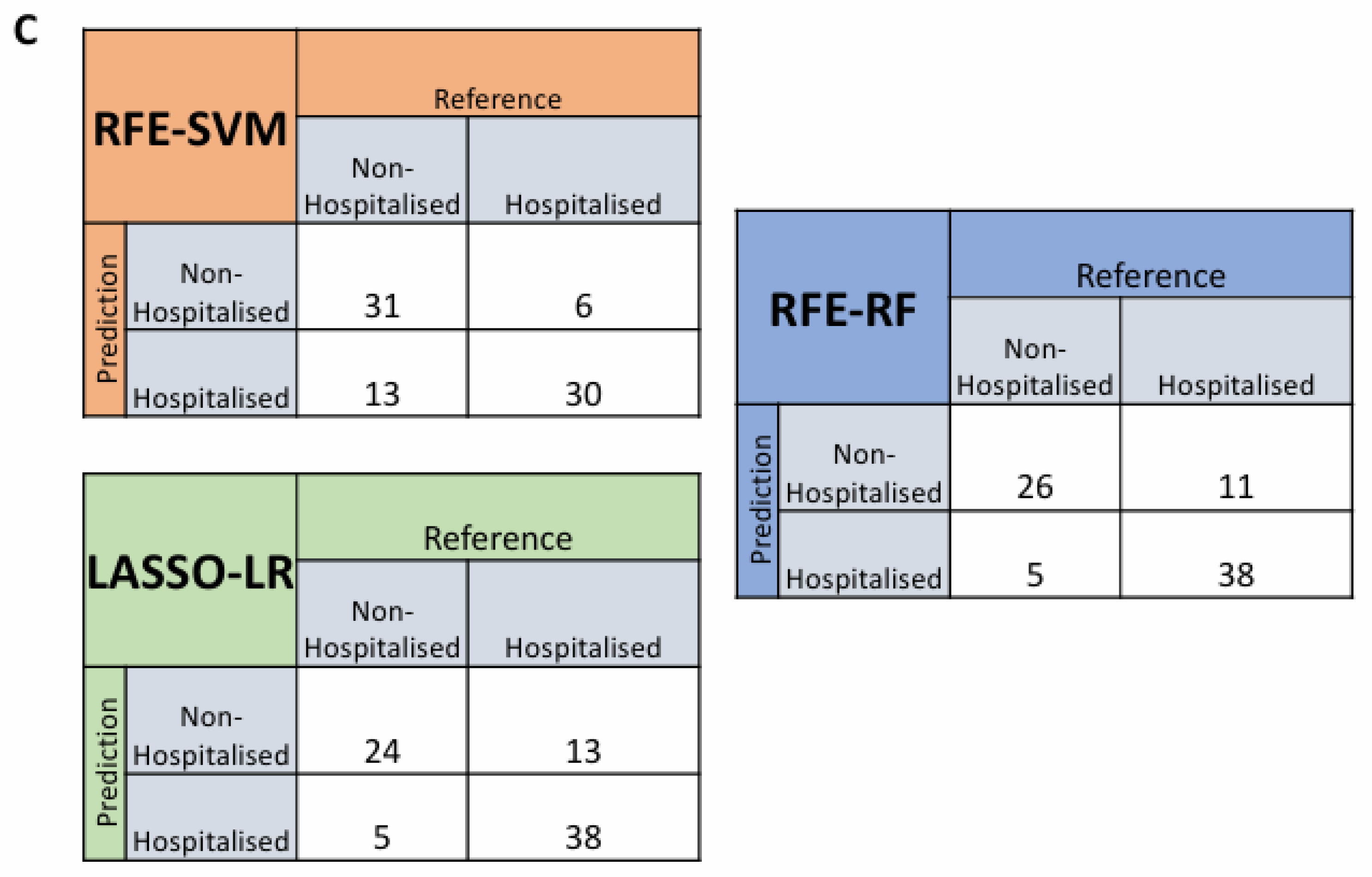
3.5. Feature Selected Machine Learning Predictions for Hospitalisation Risk
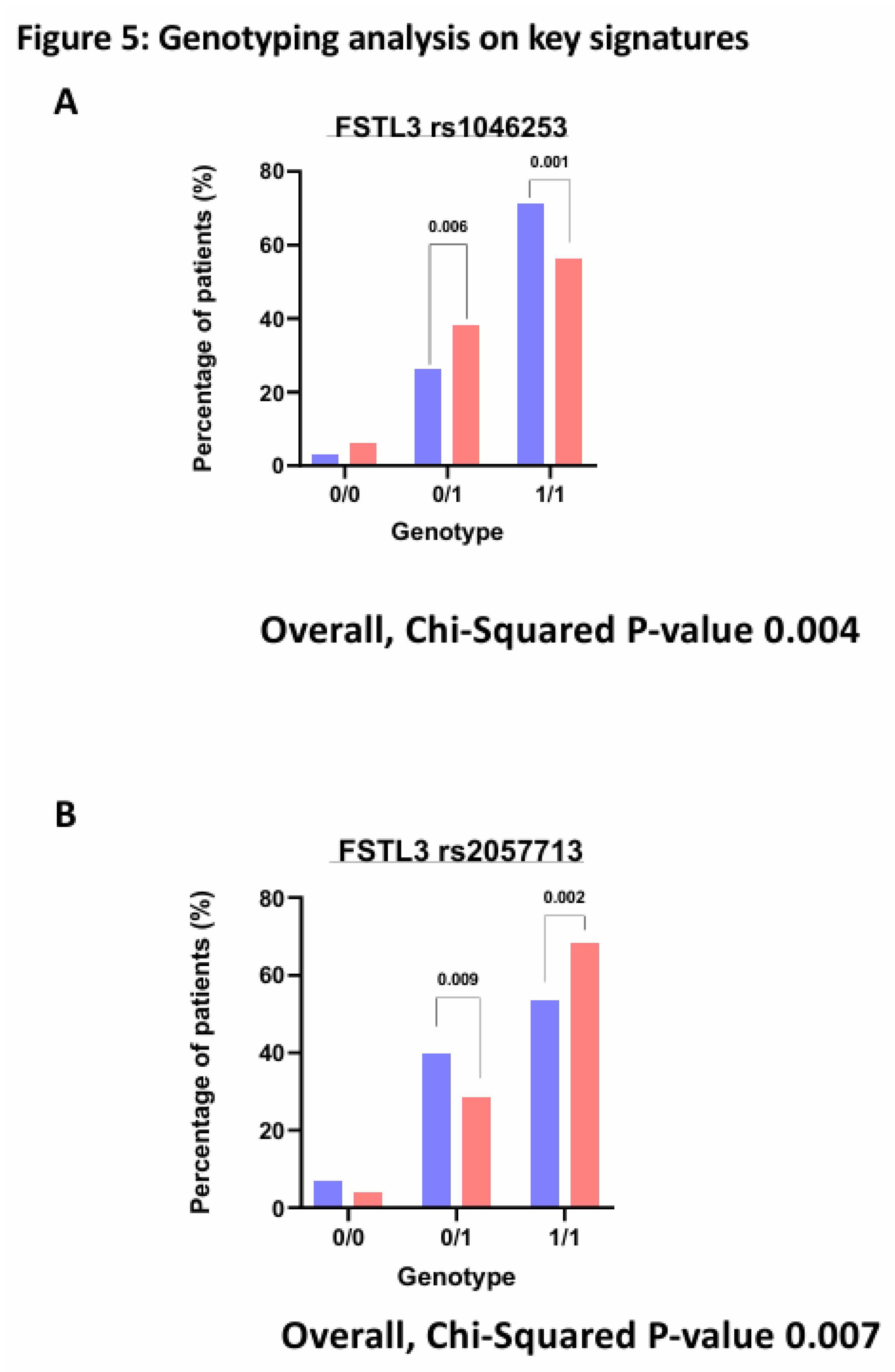
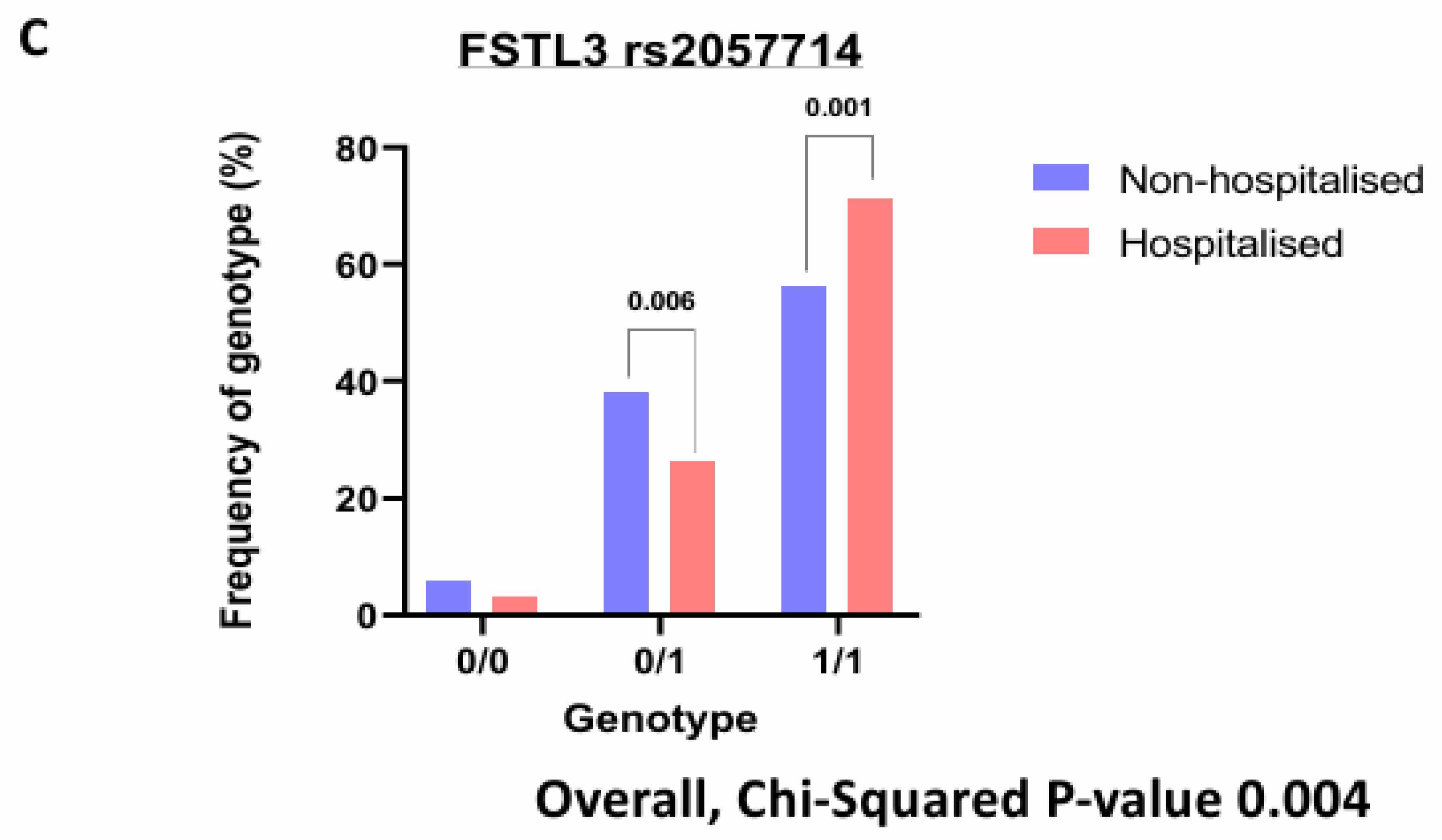
3.6. SNPs on Genes of Interest
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- World Health Organization, “World Health Organization: WHO Coronavirus (COVID-19) Dashboard.
- L. E. van Eijk et al., “COVID-19: immunopathology, pathophysiological mechanisms, and treatment options.,” J Pathol, vol. 254, no. 4, pp. 307–331, Jul. 2021. [CrossRef]
- S. Akira, S. Uematsu, and O. Takeuchi, “Pathogen recognition and innate immunity.,” Cell, vol. 124, no. 4, pp. 783–801, Feb. 2006. [CrossRef]
- Y. Tang, J. Liu, D. Zhang, Z. Xu, J. Ji, and C. Wen, “Cytokine Storm in COVID-19: The Current Evidence and Treatment Strategies.,” Front Immunol, vol. 11, p. 1708, 2020. [CrossRef]
- Y.-R. Guo et al., “The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak - an update on the status.,” Mil Med Res, vol. 7, no. 1, p. 11, Mar. 2020. [CrossRef]
- T. Beaney et al., “Trends and associated factors for Covid-19 hospitalisation and fatality risk in 2.3 million adults in England,” Nat Commun, vol. 13, no. 1, p. 2356, Apr. 2022. [CrossRef]
- R. S. Khedar et al., “Biomarkers and outcomes in hospitalised patients with COVID-19: a prospective registry,” BMJ Open, vol. 12, no. 12, p. e067430, Dec. 2022. [CrossRef]
- A. English et al., “Genomic, Proteomic and Phenotypic Biomarkers of COVID-19 Severity: Protocol for a Retrospective Observational Study (Preprint),” JMIR Res Protoc, Jul. 2023. [CrossRef]
- D. Yazici et al., “Disrupted epithelial permeability as a predictor of severe COVID-19 development.,” Allergy, Jul. 2023. [CrossRef]
- Y. Lv, X. Ma, Y. Ma, Y. Du, and J. Feng, “A new emerging target in cancer immunotherapy: Galectin-9 (LGALS9).,” Genes Dis, vol. 10, no. 6, pp. 2366–2382, Nov. 2023. [CrossRef]
- W. A. Teft, M. G. Kirchhof, and J. Madrenas, “A MOLECULAR PERSPECTIVE OF CTLA-4 FUNCTION,” Annu Rev Immunol, vol. 24, no. 1, pp. 65–97, Apr. 2006. [CrossRef]
- B. de Saint-Vis et al., “A Novel Lysosome-Associated Membrane Glycoprotein, DC-LAMP, Induced upon DC Maturation, Is Transiently Expressed in MHC Class II Compartment,” Immunity, vol. 9, no. 3, pp. 325–336, Sep. 1998. [CrossRef]
- H. Zhang et al., “AGRN promotes lung adenocarcinoma progression by activating Notch signaling pathway and acts as a therapeutic target,” Pharmacol Res, vol. 194, p. 106819, Aug. 2023. [CrossRef]
- L. Klaric et al., “Mendelian randomisation identifies alternative splicing of the FAS death receptor as a mediator of severe COVID-19.,” medRxiv, Apr. 2021. [CrossRef]
- A. J. A. Groffen, C. A. F. Buskens, T. H. van Kuppevelt, J. H. Veerkamp, L. A. H. Monnens, and L. P. W. J. van den Heuvel, “Primary structure and high expression of human agrin in basement membranes of adult lung and kidney,” Eur J Biochem, vol. 254, no. 1, pp. 123–128, May 1998. [CrossRef]
- A. Bauer et al., “Proteomics reveals antiviral host response and NETosis during acute COVID-19 in high-risk patients,” Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease, vol. 1869, no. 2, p. 166592, Feb. 2023. [CrossRef]
- N. Bozorgmehr et al., “Galectin-9, a Player in Cytokine Release Syndrome and a Surrogate Diagnostic Biomarker in SARS-CoV-2 Infection,” mBio, vol. 12, no. 3, Jun. 2021. [CrossRef]
- D. Szklarczyk et al., “The STRING database in 2023: protein–protein association networks and functional enrichment analyses for any sequenced genome of interest,” Nucleic Acids Res, vol. 51, no. D1, pp. D638–D646, Jan. 2023. [CrossRef]
- A. Funaro, G. C. Spagnoli, M. Momo, W. Knapp, and F. Malavasi, “Stimulation of T cells via CD44 requires leukocyte-function-associated antigen interactions and interleukin-2 production,” Hum Immunol, vol. 40, no. 4, pp. 267–278, Aug. 1994. [CrossRef]
- L. Yang et al., “COVID-19: immunopathogenesis and Immunotherapeutics,” Signal Transduct Target Ther, vol. 5, no. 1, p. 128, Jul. 2020. [CrossRef]
- J. D. Roh et al., “Plasma Proteomics of COVID-19–Associated Cardiovascular Complications,” JACC Basic Transl Sci, vol. 7, no. 5, pp. 425–441, May 2022. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).



