Submitted:
21 August 2024
Posted:
21 August 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Methods
Plant Material and DNA Extraction
Bisulfite Sequencing
Histone Modification and Non-Coding RNA Analysis
Functional Validation of Differentially Methylated Regions (DMRs)
Expanded Analysis of Environmental Stress Response
Data Analysis
Differential Methylation Analysis
Results
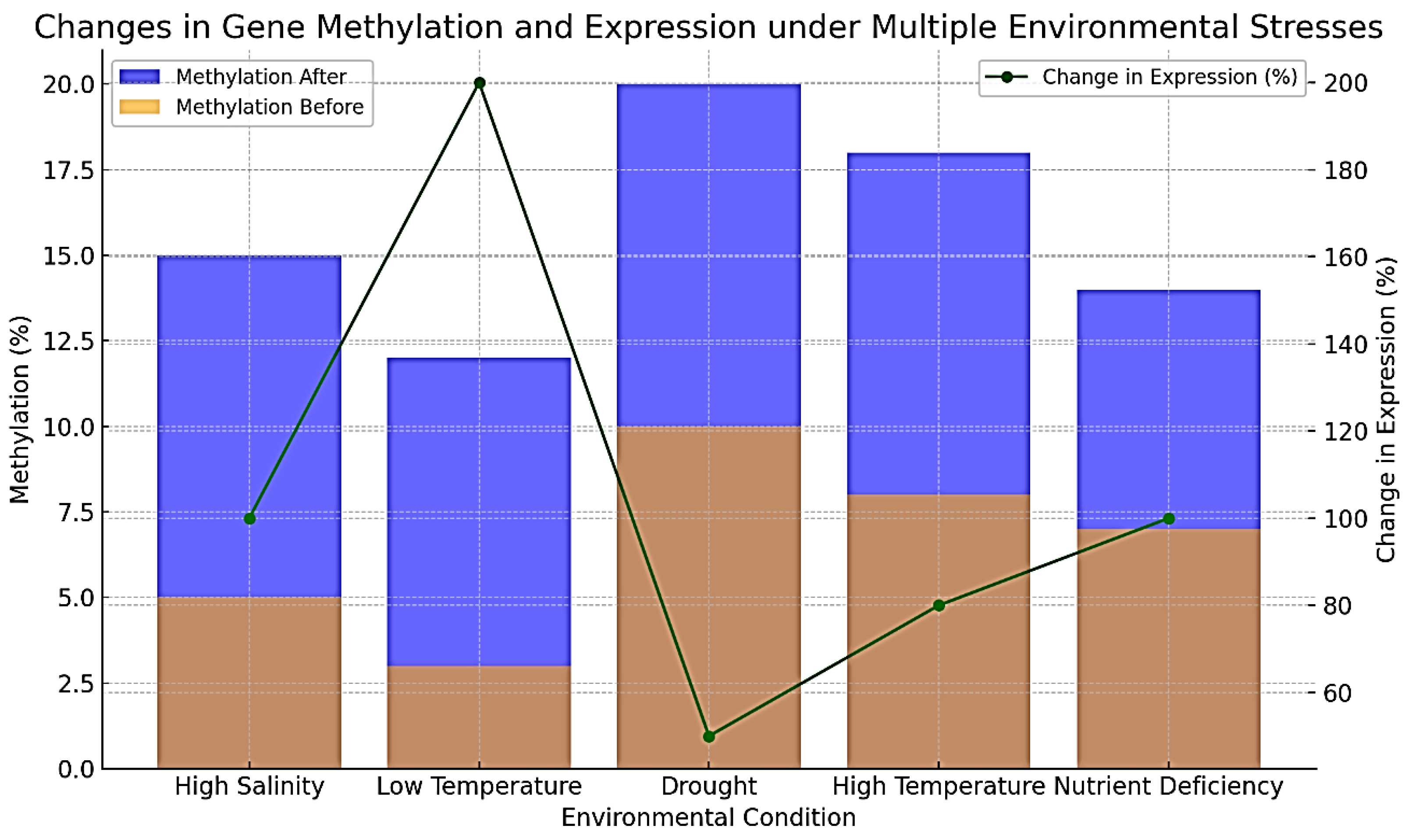
Environmental Stress Response
Discussion
Future Directions
Conclusion
References
- Bradshaw, A. D. (1965). Evolutionary significance of phenotypic plasticity in plants. Advances in Genetics, 13, 115-155. [CrossRef]
- Hancock, J. F. (2012). Plant Evolution and the Origin of Crop Species. CABI. [CrossRef]
- Rubio, J., et al. (2013). Genetic and phenotypic characterization of a core collection of chickpea (Cicer arietinum L.) cultivars. BMC Plant Biology, 13, 169. [CrossRef]
- Varshney, R. K., et al. (2013). Integrated physical, genetic and genome map of chickpea (Cicer arietinum L.). The Plant Journal, 75(5), 715-729. [CrossRef]
- Kumar, J., et al. (2018). Role of genetics and genomics in enhancing legume productivity and stress tolerance. Advances in Genetics, 97, 111-153. [CrossRef]
- Jiang, C., et al. (2016). Epigenetics and genome evolution. Genetics, 203(2), 375-387. [CrossRef]
- Springer, N. M. (2013). Epigenetics and crop improvement. Trends in Genetics, 29(4), 242-250. [CrossRef]
- Law, J. A., & Jacobsen, S. E. (2010). Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nature Reviews Genetics, 11(3), 204-220. [CrossRef]
- Zhang, H., et al. (2018). Genome-wide analysis of DNA methylation and transcriptional repression in Arabidopsis. The Plant Cell, 30(3), 537-558. [CrossRef]
- Finnegan, E. J., et al. (1996). Epialleles as a source of variation in plant development. Plant Molecular Biology, 32(1-2), 97-107. [CrossRef]
- Kumar, S., et al. (2017). Epigenetic regulation of abiotic stress tolerance in plants. Advances in Agronomy, 145, 1-65. [CrossRef]
- Schmitz, R. J., et al. (2013). Epigenome-wide inheritance of cytosine methylation variants in a recombinant inbred population. Genome Research, 23(10), 1663-1674. [CrossRef]
- Zhang, X., et al. (2013). DNA methylation profiles reveal novel targets for methylation-based gene regulation in Arabidopsis. PLoS Genetics, 9(1), e1003410. [CrossRef]
- Li, Q., et al. (2014). Systematic identification of regulatory elements in Arabidopsis thaliana by sequence genome-wide association. Nature Genetics, 46(8), 822-829. [CrossRef]
- Gupta, P. K., et al. (2012). Next-generation sequencing and its applications: elucidation of genetic variation in plants. Indian Journal of Genetics and Plant Breeding, 72(3), 269-284. [CrossRef]
- Eichten, S. R., & Springer, N. M. (2016). Minimal evidence for consistent changes in maize DNA methylation patterns following environmental stress. Frontiers in Plant Science, 7, 308. [CrossRef]
- Upadhyaya, H. D., et al. (2011). Genomic tools and germplasm diversity for chickpea improvement. Plant Genetic Resources, 9(1), 45-58. [CrossRef]
- Doyle, J. J., & Doyle, J. L. (1987). A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin, 19(1), 11-15.
- Murray, M. G., & Thompson, W. F. (1980). Rapid isolation of high molecular weight plant DNA. Nucleic Acids Research, 8(19), 4321-4325. [CrossRef]
- Saghai-Maroof, M. A., et al. (1984). Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proceedings of the National Academy of Sciences, 81(24), 8014-8018. [CrossRef]
- Clarke, J. D. (2009). Cetyltrimethyl ammonium bromide (CTAB) DNA miniprep for plant DNA isolation. Cold Spring Harbor Protocols, 2009(3), pdb.prot5177. [CrossRef]
- Porebski, S., et al. (1997). Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Molecular Biology Reporter, 15(1), 8-15. [CrossRef]
- Zhang, J., & Stewart, J. M. (2000). Economical and rapid method for extracting cotton genomic DNA. The Journal of Cotton Science, 4(3), 193-201.
- Clark, S. J., et al. (1994). High sensitivity mapping of methylated cytosines. Nucleic Acids Research, 22(15), 2990-2997. [CrossRef]
- Frommer, M., et al. (1992). A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proceedings of the National Academy of Sciences, 89(5), 1827-1831. [CrossRef]
- Rakyan, V. K., et al. (2004). DNA methylation profiling of the human major histocompatibility complex: a pilot study for the human epigenome project. PLoS Biology, 2(12), e405. [CrossRef]
- Gomez-Diaz, E., et al. (2012). The dual epigenomic and transcriptomic maps reveal an active role of DNA methylation in the regulation of a complex invertebrate behavior. BMC Genomics, 13, 152. [CrossRef]
- Molaro, A., et al. (2011). Sperm methylation profiles reveal features of epigenetic inheritance and evolution in primates. Cell, 146(6), 1029-1041. [CrossRef]
- Laird, P. W. (2010). Principles and challenges of genome-wide DNA methylation analysis. Nature Reviews Genetics, 11(3), 191-203. [CrossRef]
- Meyer, M., & Kircher, M. (2010). Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harbor Protocols, 2010(6), pdb.prot5448. [CrossRef]
- Booth, M. J., et al. (2012). Quantitative sequencing of 5-formylcytosine in DNA at single-base resolution. Nature Chemistry, 4(12), 766-774. [CrossRef]
- Krueger, F., et al. (2012). DNA methylome analysis using short bisulfite sequencing data. Nature Methods, 9(2), 145-151. [CrossRef]
- Cokus, S. J., et al. (2008). Shotgun bisulfite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature, 452(7184), 215-219. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




