Submitted:
20 August 2024
Posted:
22 August 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Case Series and Research Ethics
2.2. Clinical Data and Biological Samples
2.3. Genome-Wide Genotyping
2.4. Statistical Analyses
2.5. In Silico Functional Analyses
3. Results
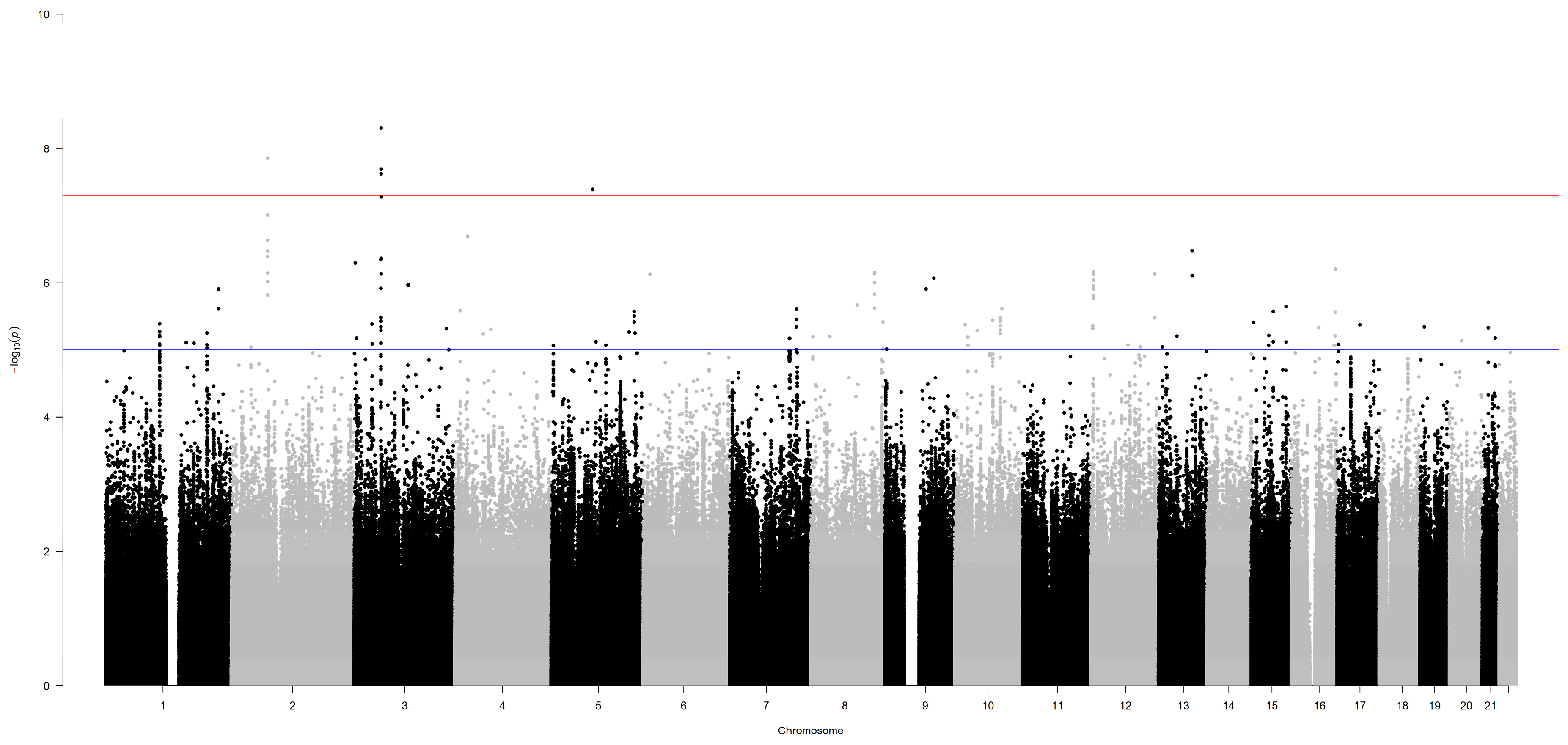
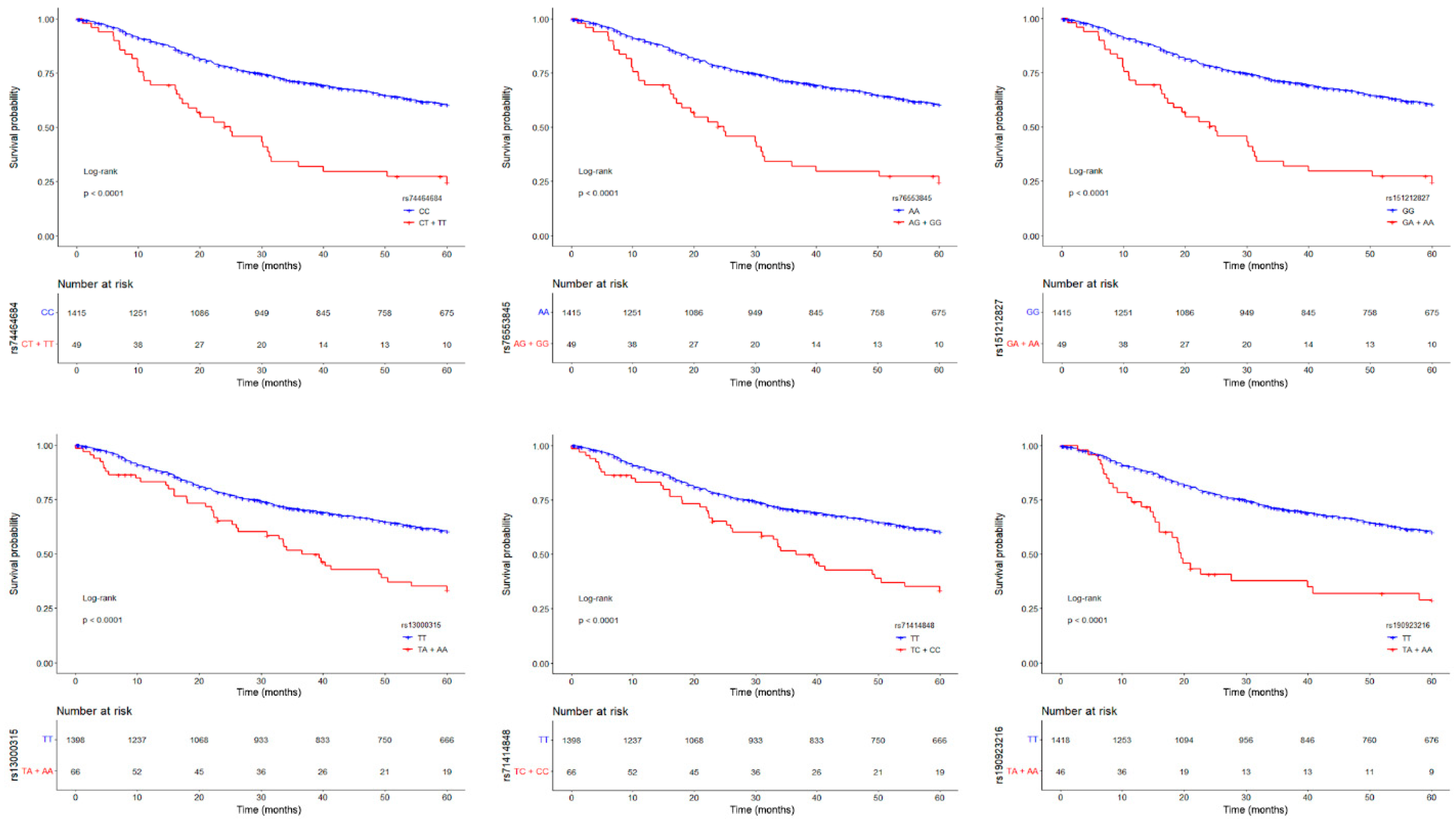
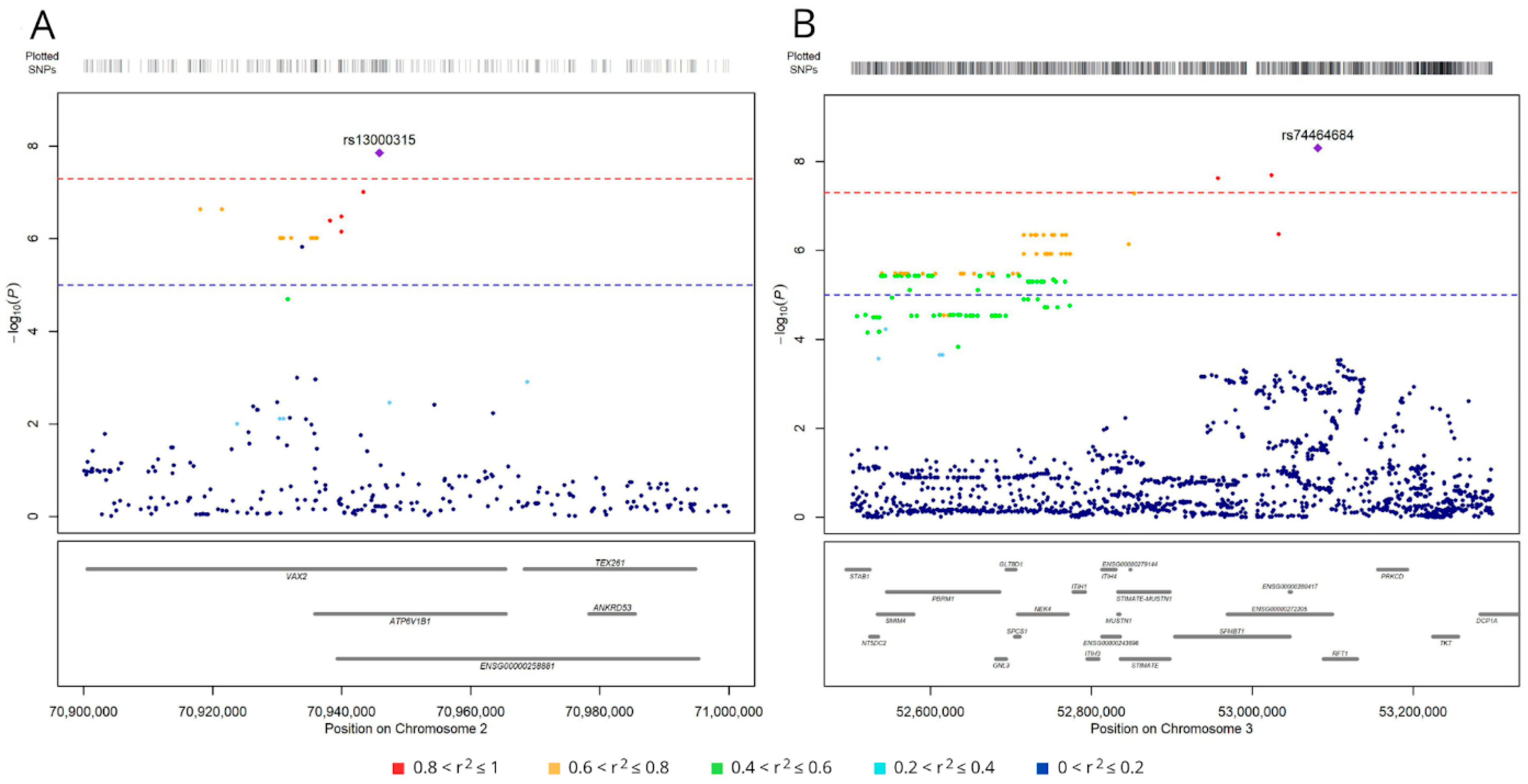
3.1. Germline Variants Associated with Overall Survival
3.2. SNPs Associated with Lung Adenocarcinoma Survival Have Regulatory Roles
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin 2021, 71, 209–249. [CrossRef]
- Siegel, R.L.; Miller, K.D.; Wagle, N.S.; Jemal, A. Cancer Statistics, 2023. CA Cancer J Clin 2023, 73, 17–48. [CrossRef]
- Potter, A.L.; Rosenstein, A.L.; Kiang, M. V; Shah, S.A.; Gaissert, H.A.; Chang, D.C.; Fintelmann, F.J.; Yang, C.-F.J. Association of Computed Tomography Screening with Lung Cancer Stage Shift and Survival in the United States: Quasi-Experimental Study. BMJ 2022, e069008. [CrossRef]
- Whitson, B.A.; Groth, S.S.; Duval, S.J.; Swanson, S.J.; Maddaus, M.A. Surgery for Early-Stage Non-Small Cell Lung Cancer: A Systematic Review of the Video-Assisted Thoracoscopic Surgery Versus Thoracotomy Approaches to Lobectomy. Ann Thorac Surg 2008, 86, 2008–2018. [CrossRef]
- Rami-Porta, R.; Call, S.; Dooms, C.; Obiols, C.; Sánchez, M.; Travis, W.D.; Vollmer, I. Lung Cancer Staging: A Concise Update. European Respiratory Journal 2018, 51, 1800190. [CrossRef]
- Howlader, N.; Forjaz, G.; Mooradian, M.J.; Meza, R.; Kong, C.Y.; Cronin, K.A.; Mariotto, A.B.; Lowy, D.R.; Feuer, E.J. The Effect of Advances in Lung-Cancer Treatment on Population Mortality. New England Journal of Medicine 2020, 383, 640–649. [CrossRef]
- Garinet, S.; Wang, P.; Mansuet-Lupo, A.; Fournel, L.; Wislez, M.; Blons, H. Updated Prognostic Factors in Localized NSCLC. Cancers (Basel) 2022, 14. [CrossRef]
- Chen, Z.; Fillmore, C.M.; Hammerman, P.S.; Kim, C.F.; Wong, K.-K. Non-Small-Cell Lung Cancers: A Heterogeneous Set of Diseases. Nat Rev Cancer 2014, 14, 535–546. [CrossRef]
- Chen, K.; Liu, H.; Liu, Z.; Luo, S.; Patz, E.F.; Moorman, P.G.; Su, L.; Shen, S.; Christiani, D.C.; Wei, Q. Genetic Variants in RUNX3 , AMD1 and MSRA in the Methionine Metabolic Pathway and Survival in Nonsmall Cell Lung Cancer Patients. Int J Cancer 2019, 145, 621–631. [CrossRef]
- Du, H.; Liu, L.; Liu, H.; Luo, S.; Patz, E.F.; Glass, C.; Su, L.; Du, M.; Christiani, D.C.; Wei, Q. Genetic Variants of DOCK2, EPHB1 and VAV2 in the Natural Killer Cell-Related Pathway Are Associated with Non-Small Cell Lung Cancer Survival. Am J Cancer Res 2021, 11, 2264–2277.
- Qian, D.; Liu, H.; Zhao, L.; Wang, X.; Luo, S.; Moorman, P.G.; Patz Jr, E.F.; Su, L.; Shen, S.; Christiani, D.C.; et al. Novel Genetic Variants in Genes of the Fc Gamma Receptor-Mediated Phagocytosis Pathway Predict Non-Small Cell Lung Cancer Survival. Transl Lung Cancer Res 2020, 9, 575–586. [CrossRef]
- Zhang, H.; Li, Y.; Guo, S.; Wang, Y.; Wang, H.; Lu, D.; Wang, J.; Jin, L.; Jiang, G.; Wu, J.; et al. Effect of ERCC2 Rs13181 and Rs1799793 Polymorphisms and Environmental Factors on the Prognosis of Patients with Lung Cancer. Am J Transl Res 2020, 12, 6941–6953.
- Pintarelli, G.; Cotroneo, C.E.; Noci, S.; Dugo, M.; Galvan, A.; Delli Carpini, S.; Citterio, L.; Manunta, P.; Incarbone, M.; Tosi, D.; et al. Genetic Susceptibility Variants for Lung Cancer: Replication Study and Assessment as Expression Quantitative Trait Loci. Sci Rep 2017, 7. [CrossRef]
- Du, H.; Mu, R.; Liu, L.; Liu, H.; Luo, S.; Patz, E.F.; Glass, C.; Su, L.; Du, M.; Christiani, D.C.; et al. Single Nucleotide Polymorphisms in FOXP1 and RORA of the Lymphocyte Activation-Related Pathway Affect Survival of Lung Cancer Patients. Transl Lung Cancer Res 2022, 11, 890–901. [CrossRef]
- Chen, A.S.; Liu, H.; Wu, Y.; Luo, S.; Patz, E.F.; Glass, C.; Su, L.; Du, M.; Christiani, D.C.; Wei, Q. Genetic Variants in DDO and PEX5L in Peroxisome-Related Pathways Predict Non-Small Cell Lung Cancer Survival. Mol Carcinog 2022, 61, 619–628. [CrossRef]
- Yang, S.; Tang, D.; Zhao, Y.C.; Liu, H.; Luo, S.; Stinchcombe, T.E.; Glass, C.; Su, L.; Shen, S.; Christiani, D.C.; et al. Potentially Functional Variants of ERAP1, PSMF1 and NCF2 in the MHC-I-Related Pathway Predict Non-Small Cell Lung Cancer Survival. Cancer Immunol Immunother 2021, 70, 2819–2833. [CrossRef]
- Galvan, A.; Colombo, F.; Frullanti, E.; Dassano, A.; Noci, S.; Wang, Y.; Eisen, T.; Matakidou, A.; Tomasello, L.; Vezzalini, M.; et al. Germline Polymorphisms and Survival of Lung Adenocarcinoma Patients: A Genome-Wide Study in Two European Patient Series. Int J Cancer 2015, 136. [CrossRef]
- Zhu, M.; Geng, L.; Shen, W.; Wang, Y.; Liu, J.; Cheng, Y.; Wang, C.; Dai, J.; Jin, G.; Hu, Z.; et al. Exome-Wide Association Study Identifies Low-Frequency Coding Variants in 2p23.2 and 7p11.2 Associated with Survival of Non–Small Cell Lung Cancer Patients. Journal of Thoracic Oncology 2017, 12, 644–656. [CrossRef]
- Dragani, T.A.; Muley, T.; Schneider, M.A.; Kobinger, S.; Eichhorn, M.; Winter, H.; Hoffmann, H.; Kriegsmann, M.; Noci, S.; Incarbone, M.; et al. Lung Adenocarcinoma Diagnosed at a Younger Age Is Associated with Advanced Stage, Female Sex, and Ever-Smoker Status, in Patients Treated with Lung Resection. Cancers (Basel) 2023, 15, 2395. [CrossRef]
- TNM Classification of Malignant Tumours.; Sobin, L., Christian, W., Eds.; UICC International Union Against Cancer, 2002;.
- TNM Classification of Malignant Tumours; Sobin LH, Gospodarowicz Mary K, Wittekind Christian, Eds.; 7th ed.; Wiley-Blackwell, 2009; ISBN 978-1-4443-3241-4.
- .
- Maspero, D.; Dassano, A.; Pintarelli, G.; Noci, S.; de Cecco, L.; Incarbone, M.; Tosi, D.; Santambrogio, L.; Dragani, T.A.; Colombo, F. Read-through Transcripts in Lung: Germline Genetic Regulation and Correlation with the Expression of Other Genes. Carcinogenesis 2020, 41. [CrossRef]
- Cotroneo, C.E.; Mangano, N.; Dragani, T.A.; Colombo, F. Lung Expression of Genes Putatively Involved in SARS-CoV-2 Infection Is Modulated in Cis by Germline Variants. European Journal of Human Genetics 2021, 29, s41431–s021.
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; De Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am J Hum Genet 2007, 81, 559–575. [CrossRef]
- Anderson, C.A.; Pettersson, F.H.; Clarke, G.M.; Cardon, L.R.; Morris, A.P.; Zondervan, K.T. Data Quality Control in Genetic Case-Control Association Studies. Nat Protoc 2010, 5, 1564–1573. [CrossRef]
- Das, S.; Forer, L.; Schönherr, S.; Sidore, C.; Locke, A.E.; Kwong, A.; Vrieze, S.I.; Chew, E.Y.; Levy, S.; McGue, M.; et al. Next-Generation Genotype Imputation Service and Methods. Nature Genetics 2016 48:10 2016, 48, 1284–1287. [CrossRef]
- Loh, P.R.; Danecek, P.; Palamara, P.F.; Fuchsberger, C.; Reshef, Y.A.; Finucane, H.K.; Schoenherr, S.; Forer, L.; McCarthy, S.; Abecasis, G.R.; et al. Reference-Based Phasing Using the Haplotype Reference Consortium Panel. Nature Genetics 2016 48:11 2016, 48, 1443–1448. [CrossRef]
- Fuchsberger, C.; Abecasis, G.R.; Hinds, D.A. Minimac2: Faster Genotype Imputation. Bioinformatics 2015, 31, 782–784. [CrossRef]
- Taliun, D.; Harris, D.N.; Kessler, M.D.; Carlson, J.; Szpiech, Z.A.; Torres, R.; Taliun, S.A.G.; Corvelo, A.; Gogarten, S.M.; Kang, H.M.; et al. Sequencing of 53,831 Diverse Genomes from the NHLBI TOPMed Program. Nature 2021 590:7845 2021, 590, 290–299. [CrossRef]
- Verlouw, J.A.M.; Clemens, E.; de Vries, J.H.; Zolk, O.; Verkerk, A.J.M.H.; am Zehnhoff-Dinnesen, A.; Medina-Gomez, C.; Lanvers-Kaminsky, C.; Rivadeneira, F.; Langer, T.; et al. A Comparison of Genotyping Arrays. European Journal of Human Genetics 2021 29:11 2021, 29, 1611–1624. [CrossRef]
- Chang, C.C.; Chow, C.C.; Tellier, L.C.A.M.; Vattikuti, S.; Purcell, S.M.; Lee, J.J. Second-Generation PLINK: Rising to the Challenge of Larger and Richer Datasets. Gigascience 2015, 4, 7. [CrossRef]
- Delaneau, O.; Marchini, J.; McVean, G.A.; Donnelly, P.; Lunter, G.; Marchini, J.L.; Myers, S.; Gupta-Hinch, A.; Iqbal, Z.; Mathieson, I.; et al. Integrating Sequence and Array Data to Create an Improved 1000 Genomes Project Haplotype Reference Panel. Nat Commun 2014, 5, 3934. [CrossRef]
- Clark, T.G.; Bradburn, M.J.; Love, S.B.; Altman, D.G. Survival Analysis Part I: Basic Concepts and First Analyses. British Journal of Cancer 2003 89:2 2003, 89, 232–238. [CrossRef]
- Terry M. Therneau and Patricia M. Grambsch Modeling Survival Data: Extending the Cox Model. , Springer-Verlag, New York, 2000. ; Springer-Verlag: New York, 2000;.
- Aulchenko, Y.S.; Ripke, S.; Isaacs, A.; van Duijn, C.M. GenABEL: An R Library for Genome-Wide Association Analysis. Bioinformatics 2007, 23, 1294–1296. [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society: Series B (Methodological) 1995, 57, 289–300. [CrossRef]
- Võsa, U.; Claringbould, A.; Westra, H.-J.; Bonder, M.J.; Deelen, P.; Zeng, B.; Kirsten, H.; Saha, A.; Kreuzhuber, R.; Yazar, S.; et al. Large-Scale Cis- and Trans-EQTL Analyses Identify Thousands of Genetic Loci and Polygenic Scores That Regulate Blood Gene Expression. Nat Genet 2021, 53, 1300–1310. [CrossRef]
- Gyorffy, B.; Surowiak, P.; Budczies, J.; Lánczky, A. Online Survival Analysis Software to Assess the Prognostic Value of Biomarkers Using Transcriptomic Data in Non-Small-Cell Lung Cancer. PLoS ONE 2013, 8. [CrossRef]
- Kratzer, T.B.; Bandi, P.; Freedman, N.D.; Smith, R.A.; Travis, W.D.; Jemal, A.; Siegel, R.L. Lung Cancer Statistics, 2023. Cancer 2024, 130, 1330–1348. [CrossRef]
- Kawaguchi, T.; Takada, M.; Kubo, A.; Matsumura, A.; Fukai, S.; Tamura, A.; Saito, R.; Maruyama, Y.; Kawahara, M.; Ignatius Ou, S.-H. Performance Status and Smoking Status Are Independent Favorable Prognostic Factors for Survival in Non-Small Cell Lung Cancer: A Comprehensive Analysis of 26,957 Patients with NSCLC. J Thorac Oncol 2010, 5, 620–630. [CrossRef]
- Sheikh, M.; Mukeriya, A.; Shangina, O.; Brennan, P.; Zaridze, D. Postdiagnosis Smoking Cessation and Reduced Risk for Lung Cancer Progression and Mortality : A Prospective Cohort Study. Ann Intern Med 2021, 174, 1232–1239. [CrossRef]
- Schulze, A.B.; Kuntze, A.; Schmidt, L.H.; Mohr, M.; Marra, A.; Hillejan, L.; Schulz, C.; Görlich, D.; Hartmann, W.; Bleckmann, A.; et al. High Expression of NT5DC2 Is a Negative Prognostic Marker in Pulmonary Adenocarcinoma. Cancers (Basel) 2022, 14. [CrossRef]
- Jin, X.; Liu, X.; Zhang, Z.; Xu, L. NT5DC2 Suppression Restrains Progression towards Metastasis of Non-Small-Cell Lung Cancer through Regulation P53 Signaling. Biochem Biophys Res Commun 2020, 533, 354–361. [CrossRef]
- Niu, C.; Qiu, W.; Li, X.; Li, H.; Zhou, J.; Zhu, H. Transketolase Serves as a Biomarker for Poor Prognosis in Human Lung Adenocarcinoma. J Cancer 2022, 13, 2584–2593. [CrossRef]
- Zhang, S.; Zeng, X.; Lin, S.; Liang, M.; Huang, H. Identification of Seven-Gene Marker to Predict the Survival of Patients with Lung Adenocarcinoma Using Integrated Multi-Omics Data Analysis. J Clin Lab Anal 2022, 36, e24190. [CrossRef]
| Characteristic | Total (n = 1479) |
Italian series (n = 1049) |
German series (n = 430) |
P | ||
|---|---|---|---|---|---|---|
| Age at surgery, years, median (range) | 65 (30–90) | 66 (30–85) | 63 (37–88) | < 0.001 a | ||
| Age group, years, n (%) | < 0.001 b | |||||
| < 55 | 227 (15.3) | 139 (13.3) | 88 (20.5) | |||
| 55-64 | 496 (33.5) | 347 (33.1) | 149 (34.7) | |||
| 65-74 | 563 (38.1) | 413 (39.3) | 150 (34.9) | |||
| ³ 75 | 193 (13.0) | 150 (14.3) | 43 (10.0) | |||
| Sex, n (%) | 0.004 b | |||||
| Male | 922 (62.4) | 679 (64.7) | 243 (56.5) | |||
| Female | 557 (37.6) | 370 (35.3) | 187 (43.5) | |||
| Smoking habit, n (%) | 0.67 b | |||||
| Never | 225 (15.3) | 160 (15.3) | 65 (15.1) | |||
| Ever | 1190 (80.4) | 826 (78.7) | 364 (84.7) | |||
| Missing | 64 (4.3) | 63 (6.0) | 1 (0.2) | |||
| Pathological stage, n (%) | < 0.001 b | |||||
| I | 778 (52.6) | 612 (58.7) | 166 (38.6) | |||
| II | 257 (17.4) | 174 (16.3) | 83 (19.3) | |||
| III | 348 (23.6) | 195 (18.5) | 153 (35.6) | |||
| IV | 82 (5.5) | 54 (5.2) | 28 (6.5) | |||
| Missing | 14 (0.95) | 14 (1.3) | 0 (0) | |||
| Decade of surgery, n (%) | ||||||
| Before 2000 | 221 (14.9) | 221 (21.1) | 0 (0) | 0.82 b | ||
| 2001–2010 | 558 (37.8) | 370 (35.3) | 188 (43.7) | |||
| After 2010 | 699 (47.2) | 458 (43.6) | 241 (56.0) | |||
| Missing | 1 (0.1) | 0 (0) | 1 (0.2) | |||
| Follow-up, months, median (IQR) | 54 (21–60 ) | 50 (20–60) | 60 (23–60) | 0.011 a | ||
| Survival status at 60 months, n (%) | 0.002 b | |||||
| Alive | 928 (62.7) | 685 (65.3) | 243 (56.5) | |||
| Dead | 551 (37.3) | 364 (34.7) | 187 (43.5) | |||
| Genotyping array, n (%) | < 0.001 b | |||||
| Infinium Omni2.5-8 | ||||||
| All tumors | 559 (37.7) | 559 (53.3) | 0 (0) | |||
| Stage I tumors * | 366 (65.5) | 366 (65.5) | 0 (0) | |||
| Axiom PMRA | ||||||
| All tumors | 920 (62.3) | 490 (46.7) | 430 (100) | |||
| Stage I tumors * | 412 (44.8) | 246 (50.2) | 166 (38.6) | |||
| Characteristic | Univariable analyses a | Multivariable analyses b | ||||
|---|---|---|---|---|---|---|
| HR (95% CI) | Cox P | HR (95% CI) | Cox P | |||
| Age, years | 1.00 (1.00–1.01) | 0.40 | 1.02 (1.01–1.03) | 2.93 x 10-5 | ||
| Age group, years | ||||||
| < 55 | 1.00 | 1.00 | ||||
| 55-64 | 0.84 (0.66–1.08) | 0.18 | 1.06 (0.82-1.37)* | 0.62* | ||
| 65-74 | 0.91 (0.71–1.16) | 0.45 | 1.33 (1.03-1.71)* | 0.028* | ||
| ³ 75 | 1.08 (0.80–1.46) | 0.62 | 1.82 (1.31-2.52)* | 2.98 × 10-4* | ||
| Sex | ||||||
| Male | 1.00 | 1.00 | ||||
| Female | 0.61 (0.51–0.74) | 1.88 x 10-7 | 0.66 (0.54–0.81) | 6.53 x 10-5 | ||
| Smoking habit | ||||||
| Never | 1.00 | 1.00 | ||||
| Ever | 1.25 (0.98–1.60) | 0.078 | 1.05 (0.81–1.37) | 0.72 | ||
| Pathological stage | ||||||
| I | 1.00 | 1.00 | ||||
| II | 2.06 (1.61–2.64) | 9.01 x 10-9 | 2.05 (1.60–2.65) | 2.38 x 10-8 | ||
| III | 4.04 (3.30–4.94) | < 2.00 x 10-16 | 4.14 (3.35–5.12) | < 2.00 x 10-16 | ||
| IV | 5.95 (4.45–7.97) | < 2.00 x 10-16 | 5.59 (4.12–7.57) | < 2.00 x 10-16 | ||
| Decade of surgery | ||||||
| Before 2000 | 1.00 | 1.00 | ||||
| 2001–2010 | 0.77 (0.62–0.95) | 0.017 | 0.68 (0.53–0.86) | 0.0014 | ||
| After 2010 | 0.45 (0.36–0.57) | 2.5 x 10-11 | 0.48 (0.37–0.63) | 8.62 x 10-8 | ||
| Country | ||||||
| Italy | 0.85 (0.71–1.01) | 0.063 | 0.96 (0.77–1.21) | 0.73 | ||
| Germany | 1.00 | 1.00 | ||||
| Genotyping array | ||||||
| Infinium Omni2.5-8 | 0.79 (0.66–0.94) | 0.0075 | 0.84 (0.68 –1.04) | 0.11 | ||
| Axiom PRMA | 1.00 | 1.00 | ||||
| Variable | HR (95% CI) | Cox P | |
|---|---|---|---|
| Age | 1.02 (1.01 – 1.03) | 2.8 x 10-6 | |
| Sex | |||
| Male | 1.00 | ||
| Female | 0.68 (0.56 – 0.82) | 8.0 x 10-5 | |
| Pathological stage | |||
| I | 1.00 | ||
| II | 2.00 (1.56 – 2.57) | 4.8 x 10-8 | |
| III | 4.34 (3.53 – 5.34) | < 2.0 x 10-16 | |
| IV | 6.24 (4.62 – 8.41) | < 2.0 x 10-16 | |
| Decade of surgery | |||
| Before 2000 | 1.00 | ||
| 2001 – 2010 | 0.70 (0.56 – 0.88) | 2.4 x 10-3 | |
| After 2010 | 0.49 (0.39 – 0.63) | 2.4 x 10-8 | |
| Genomic variant a | |||
| rs13000315 | 2.62 (1.92 – 3.56) | 9.6 x 10-10 | |
| rs151212827 | 2.32 (1.63 – 3.29) | 2.8 x 10-6 | |
| rs190923216 | 2.58 (1.75 – 3.79) | 1.5 x 10-6 | |
| SNP | Chr. | Minor allele | Major allele | Regulated gene | P-value | Z-score | FDR |
|---|---|---|---|---|---|---|---|
| rs13000315 | 2 | A | T | CLEC4F | 5.08 x 10-53 | 15.3 | 0 |
| A | T | NAGK | 4.30 x 10-10 | 6.24 | 0 | ||
| A | T | MCEE | 3.74 x 10-6 | 4.63 | 0.0100 | ||
| A | T | CD207 | 1.50 x 10-5 | 4.33 | 0.0387 | ||
| rs71414848 | 2 | C | T | CLEC4F | 3.16 x 10-53 | 15.4 | 0 |
| C | T | NAGK | 3.74 x 10-10 | 6.26 | 0 | ||
| C | T | MCEE | 4.81 x 10-6 | 4.57 | 0.0130 | ||
| C | T | CD207 | 1.88 x 10-5 | 4.28 | 0.0470 | ||
| rs74464684 | 3 | T | C | NT5DC2 | 6.54 x 10-49 | 14.7 | 0 |
| T | C | TKT | 8.97 x 10-16 | -8.04 | 0 | ||
| T | C | UQCC5 | 4.80 x 10-7 | -5.03 | 0.00147 | ||
| rs76553845 | 3 | G | A | NT5DC2 | 4.40 x 10-45 | 14.1 | 0 |
| G | A | TKT | 2.18 x 10-12 | -7.02 | 0 | ||
| G | A | UQCC5 | 4.12 x 10-7 | -5.06 | 0.00122 | ||
| rs151212827 | 3 | A | G | NT5DC2 | 1.57 x 10-47 | 14.5 | 0 |
| A | G | TKT | 1.08 x 10-11 | -6.80 | 0 | ||
| A | G | UQCC5 | 2.76 x 10-8 | -5.56 | 0.000121 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




