Submitted:
12 August 2024
Posted:
16 August 2024
You are already at the latest version
Abstract
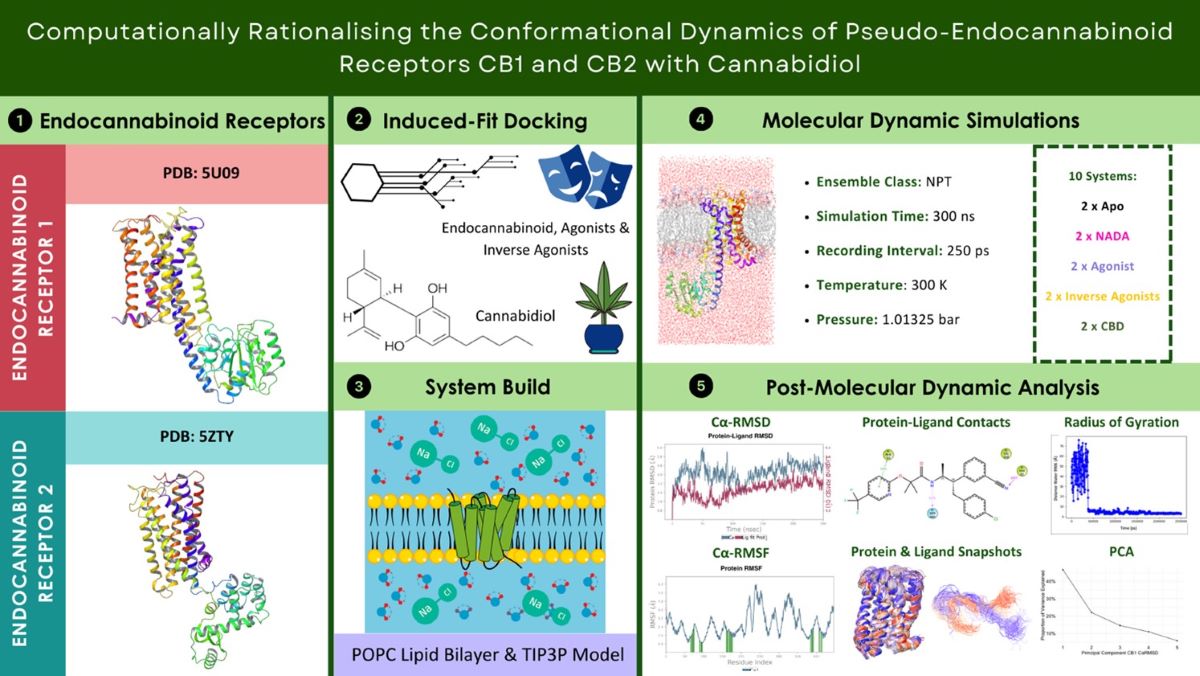
Keywords:
1. Introduction
2. Materials and Methods
3. Results
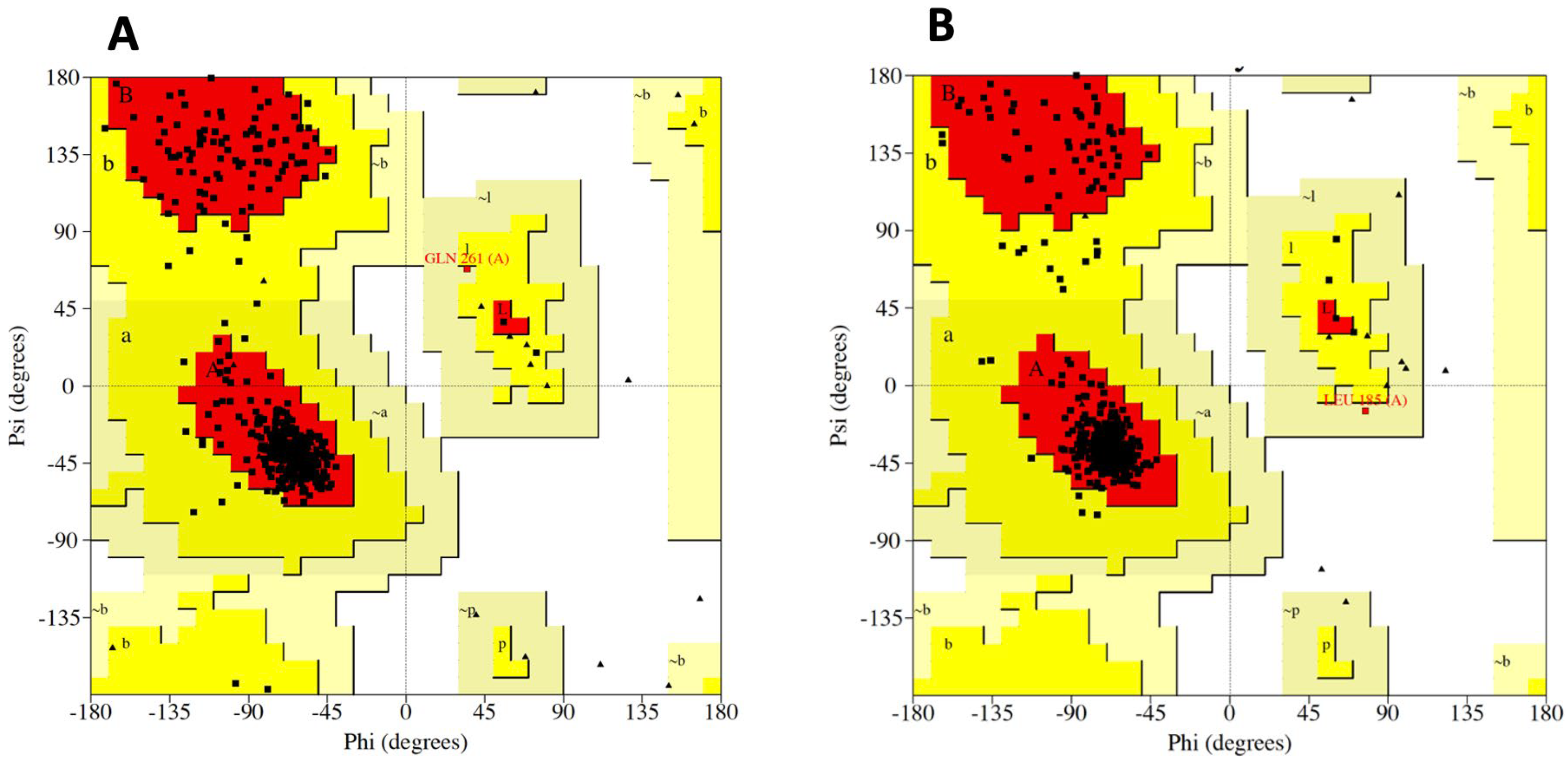
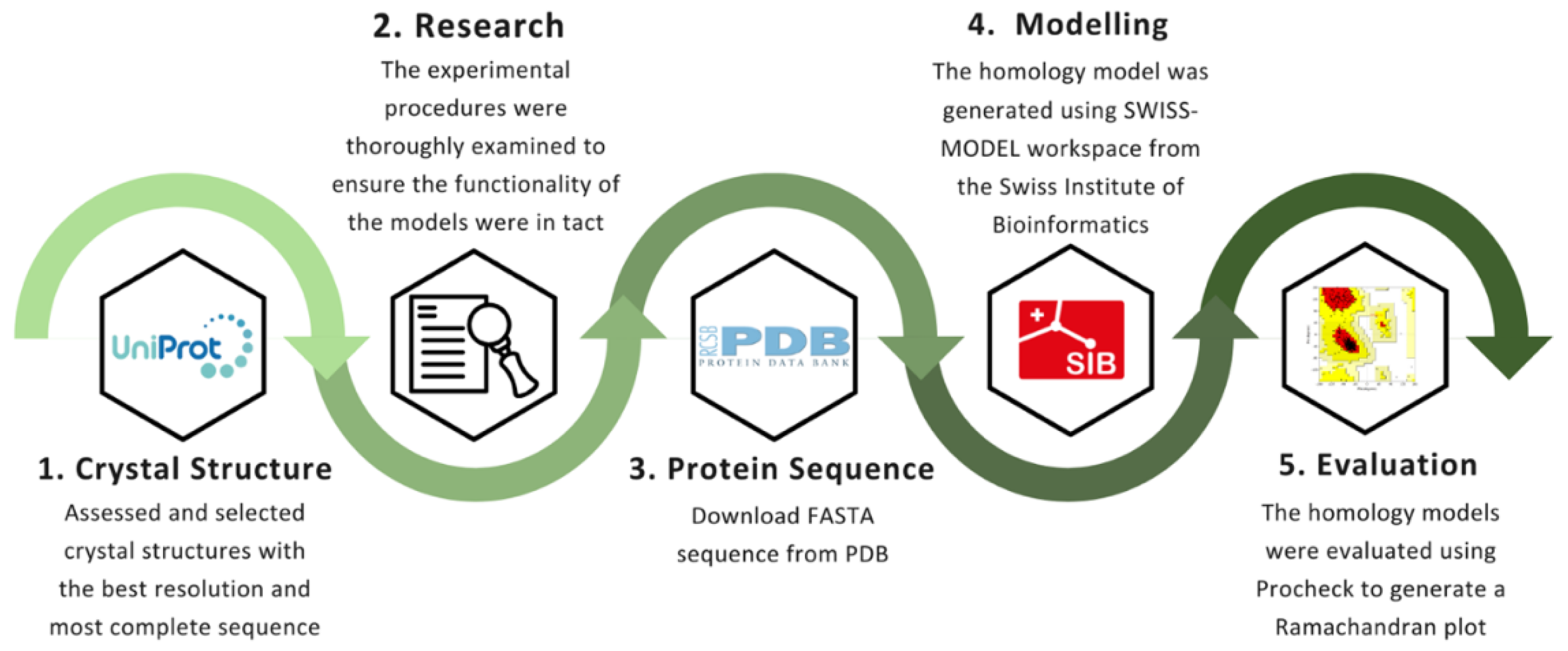
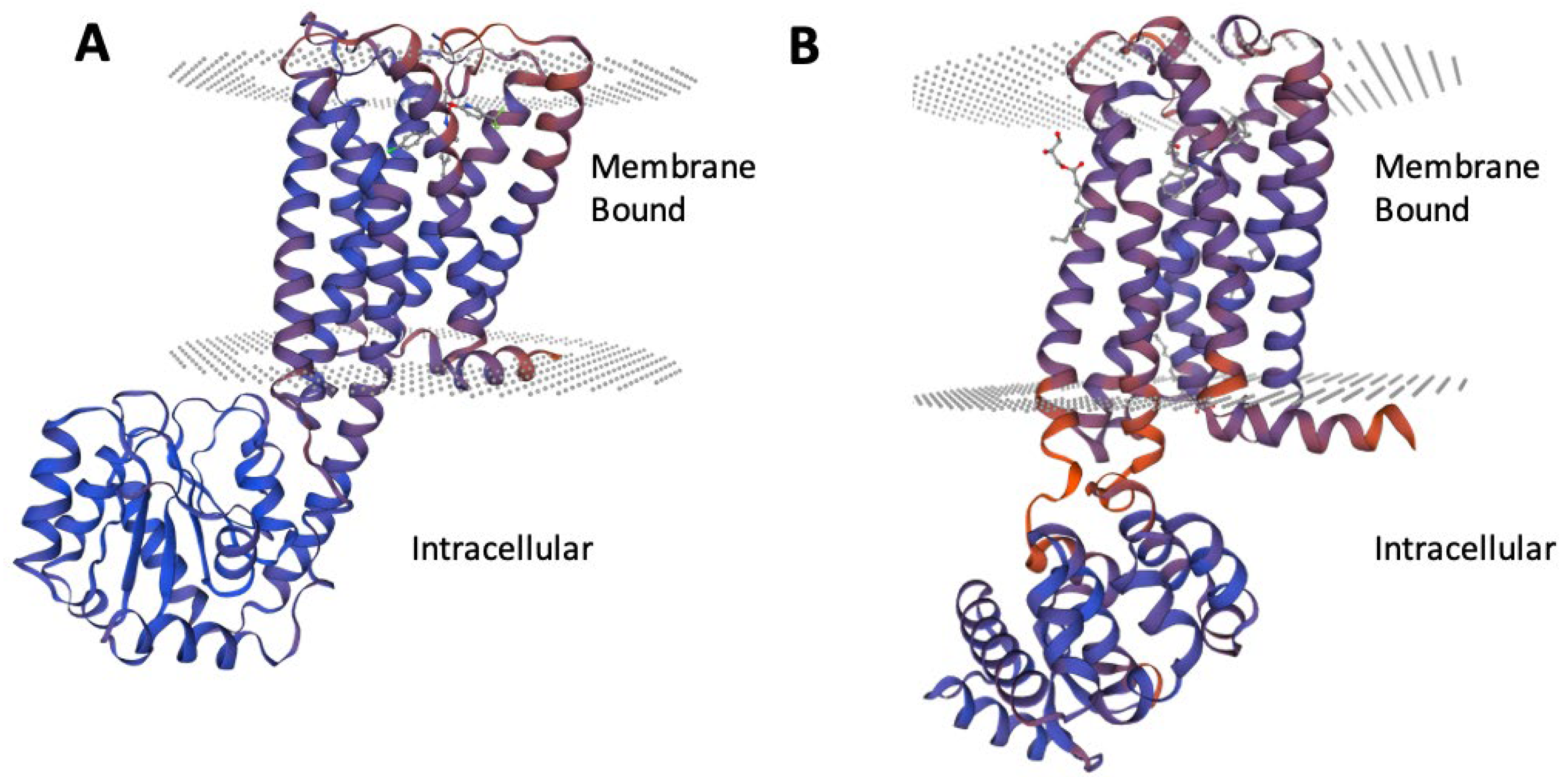
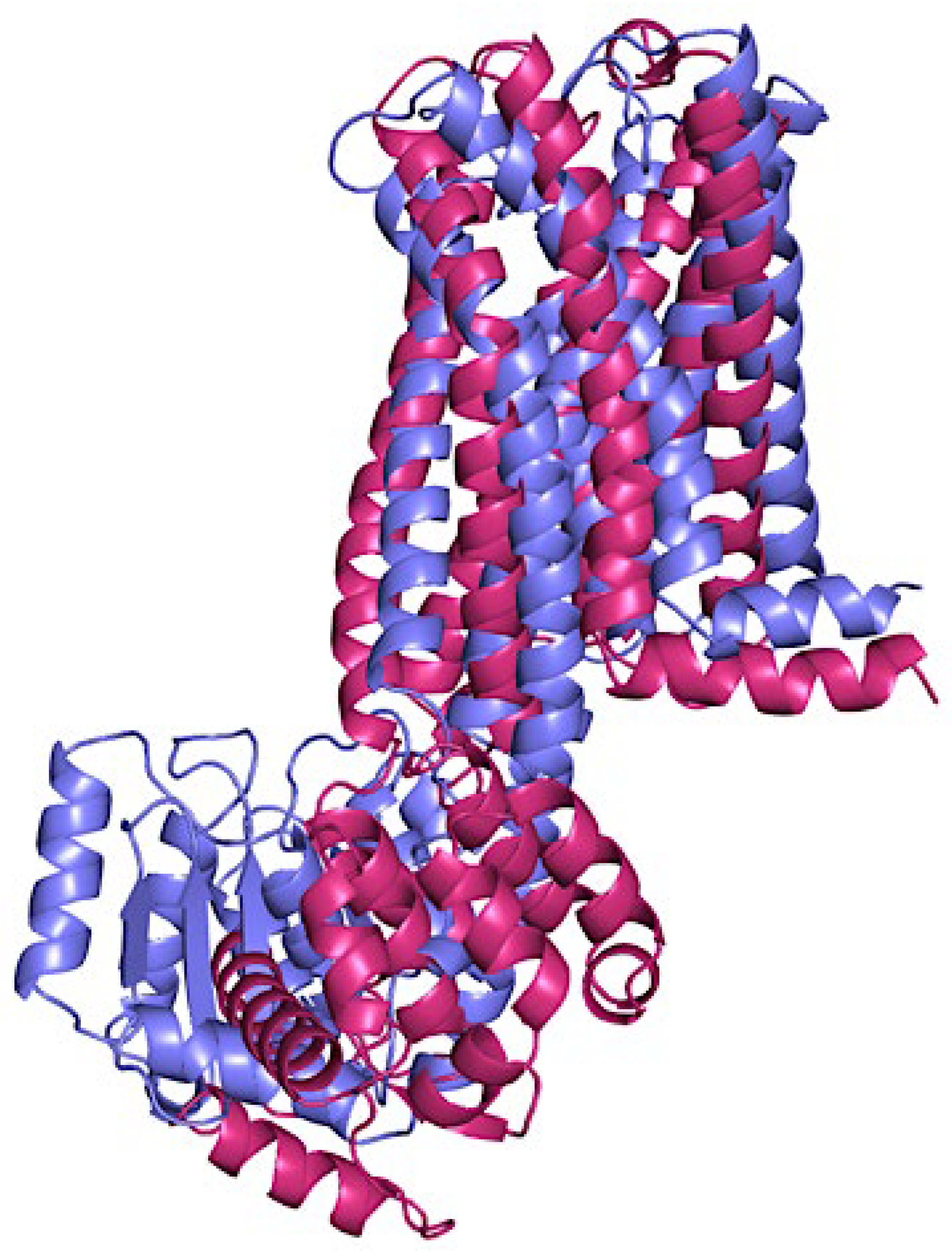
3.1. Crystal Selection and Homology Modelling
| Plot Statistics | Endocannabinoid Receptor 1 | Endocannabinoid Receptor 2 | ||
|---|---|---|---|---|
| Residues in most favoured regions | 409 | 94.2 % | 385 | 94.4 % |
| Residues in additional allowed regions | 24 | 5.5 % | 22 | 5.4 % |
| Residues in generously allowed regions | 1 | 0.2 % | 1 | 0.2 % |
| Residues in disallowed regions | 0 | 0.0 % | 0 | 0.0 % |
| Number of residues present in the structure | 478 | - | 449 | - |
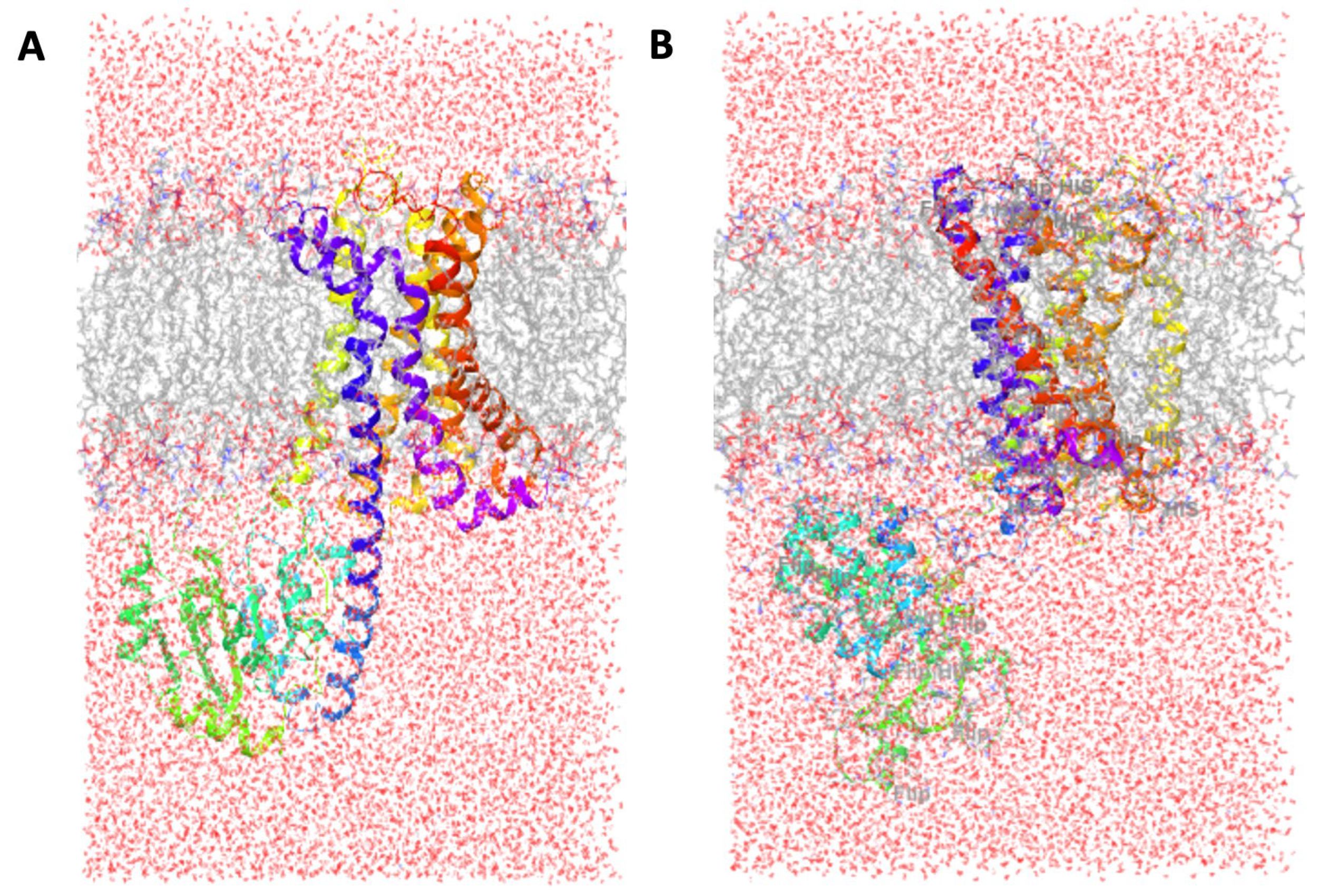
3.2. Solvation and Membrane System
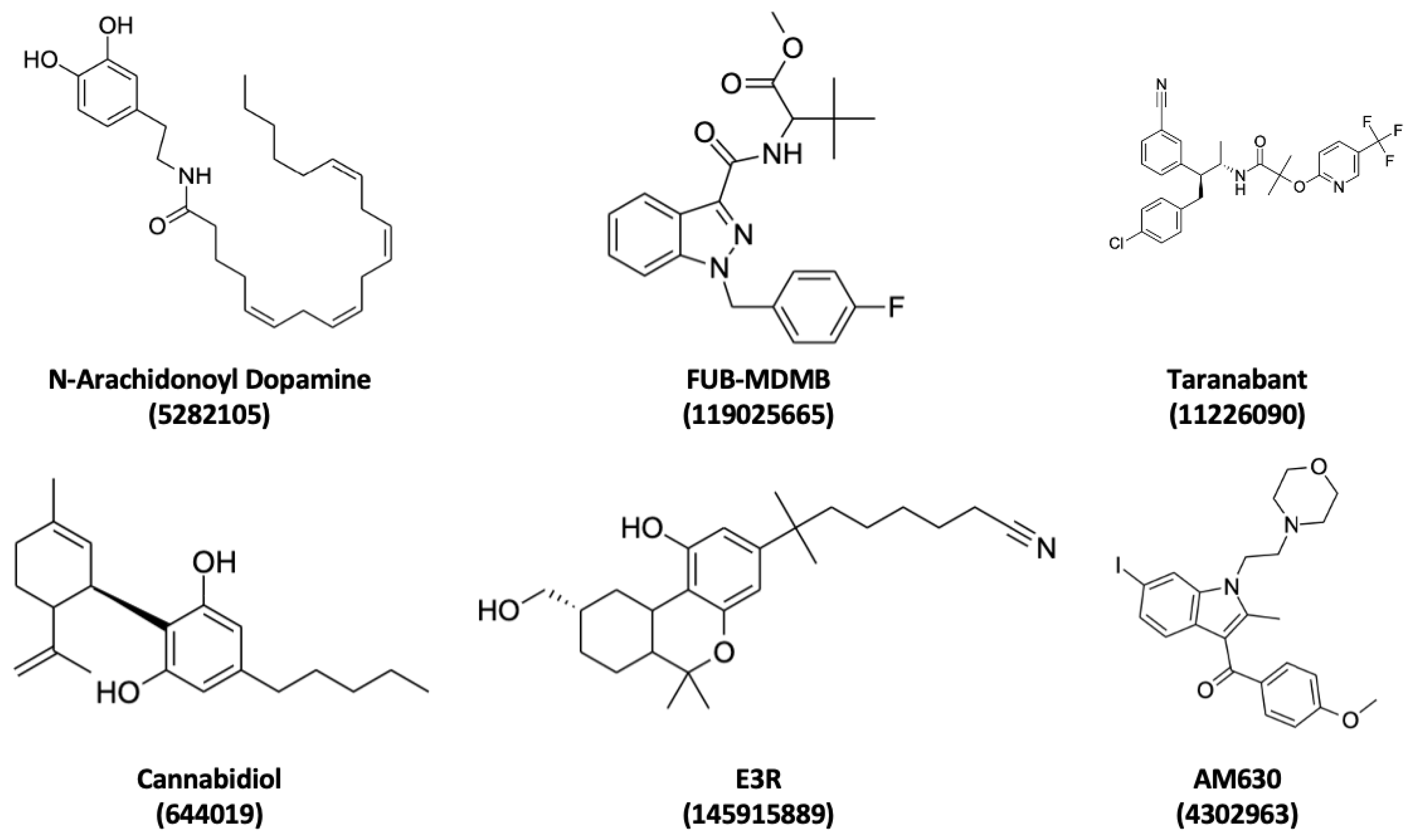
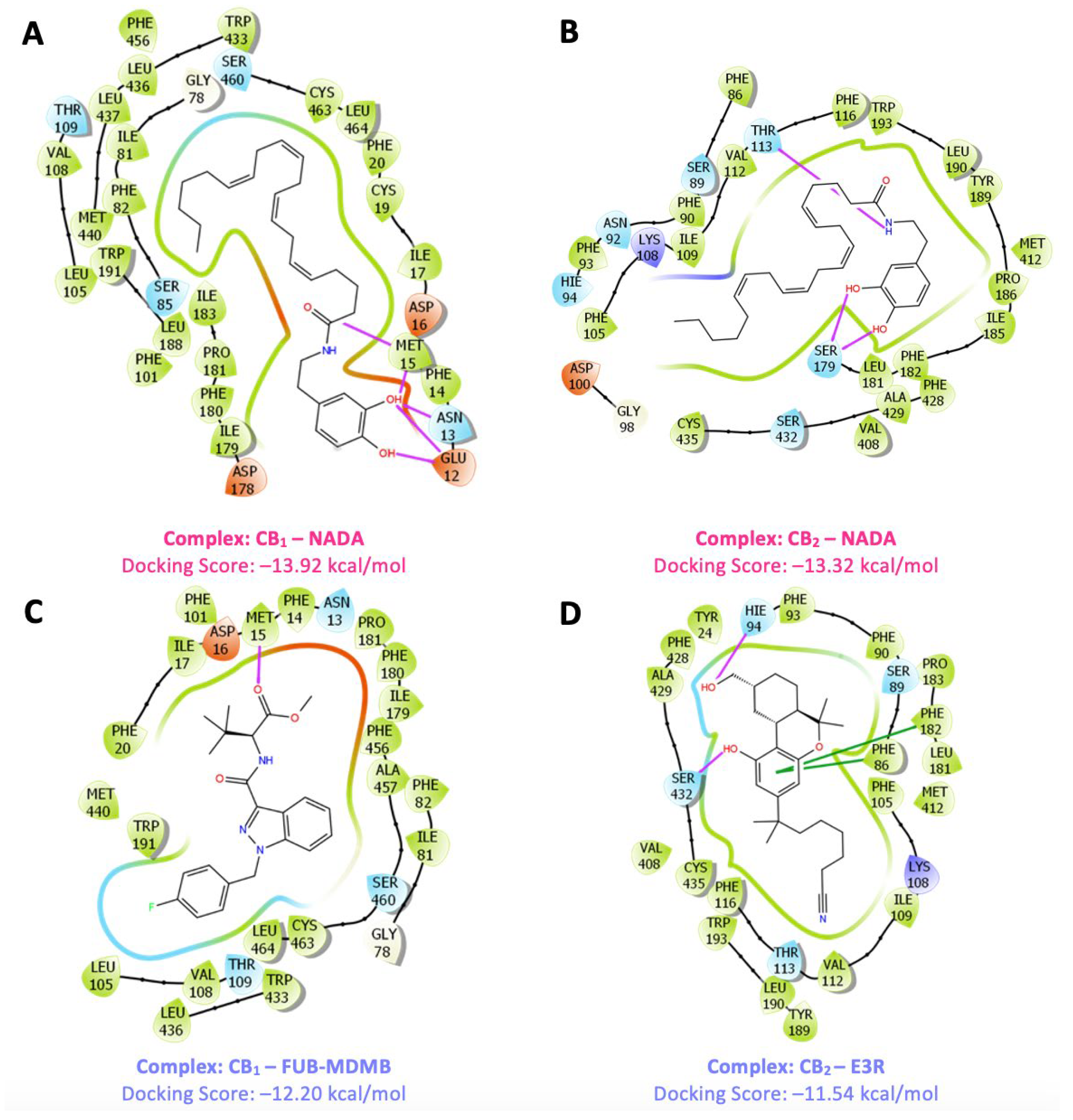
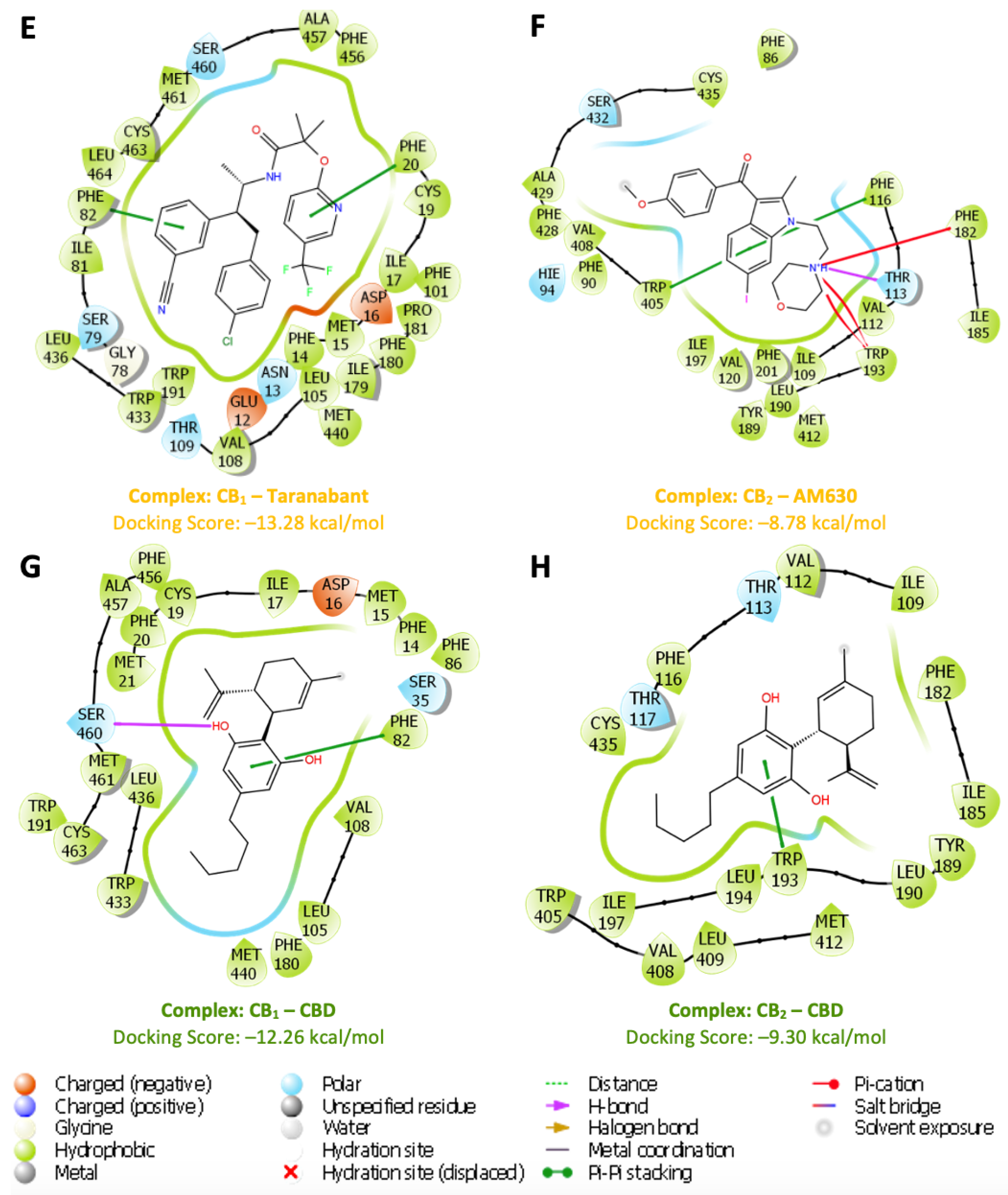
3.3. Ligand Interaction
3.4. Energy Properties
| Statistical Parameters | Apo | NADA | FUB-MDMB | Taranabant | CBD |
|---|---|---|---|---|---|
| Simulation Period (ns) | 250.00 | 250.00 | 250.00 | 250.00 | 250.00 |
| Degrees of Freedom | 141853 | 161211 | 153736 | 164349 | 146648 |
| Number of Atoms | 65690 | 74517 | 71224 | 76178 | 68022 |
| Average Total Energy (kcal/mol) | –116151.80 | –125664.51 | –124343.60 | –123603.39 | –121447.47 |
| Average Potential Energy (kcal/mol) | –158158.88 | –173394.12 | –169867.49 | –172266.69 | –164876.64 |
| Temperature (K) | 298.04 | 297.97 | 298.02 | 298.00 | 298.05 |
| Pressure (bar) | 0.87 | 1.47 | 1.25 | 1.29 | 1.30 |
| Volume (Å3) | 638265.35 | 723028.60 | 692012.22 | 739565.77 | 661813.67 |
| Slope (ps-1) | -0.001 | 0.000 | 0.000 | -0.001 | 0.000 |
| Statistical Parameters | Apo | NADA | E3R | AM630 | CBD |
|---|---|---|---|---|---|
| Simulation Period (ns) | 250.00 | 250.00 | 250.00 | 250.00 | 250.00 |
| Degrees of Freedom | 150195 | 154383 | 153622 | 140182 | 144228 |
| Number of Atoms | 69961 | 71701 | 71400 | 65151 | 66969 |
| Average Total Energy (kcal/mol) | –128263.10 | –126944.28 | –127483.56 | –118506.84 | –119839.14 |
| Average Potential Energy (kcal/mol) | –172746.90 | –172660.86 | –172976.50 | –160023.83 | –162550.75 |
| Temperature (K) | 298.08 | 298.03 | 298.04 | 298.07 | 298.05 |
| Pressure (bar) | 1.36 | 0.70 | 0.75 | 1.30 | 0.86 |
| Volume (Å3) | 682471.69 | 698028.68 | 695374.07 | 634755.01 | 651847.95 |
| Slope (ps-1) | 0.000 | 0.000 | 0.000 | -0.001 | -0.001 |
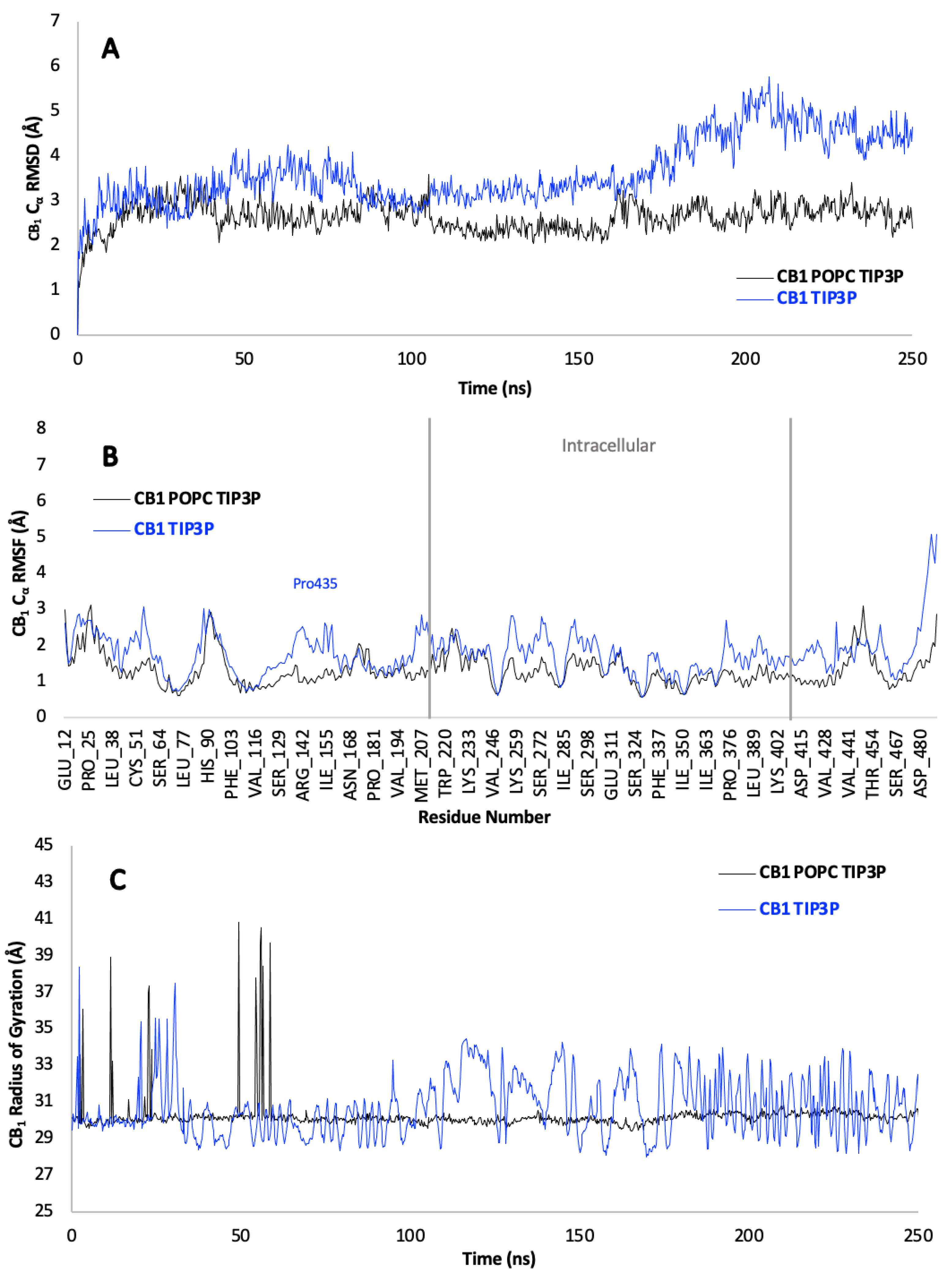
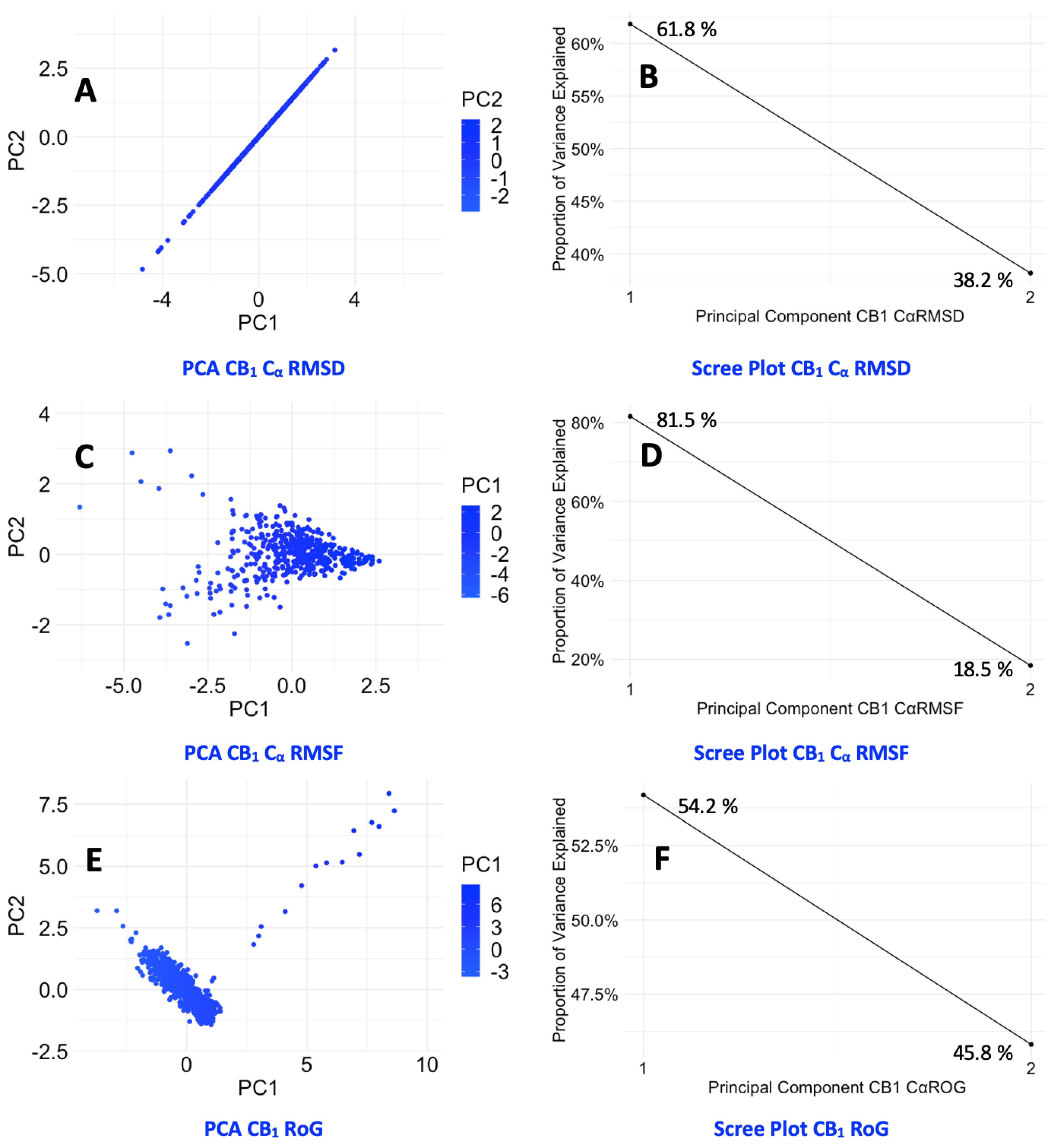
3.5. Protein Backbone Cα RMSD
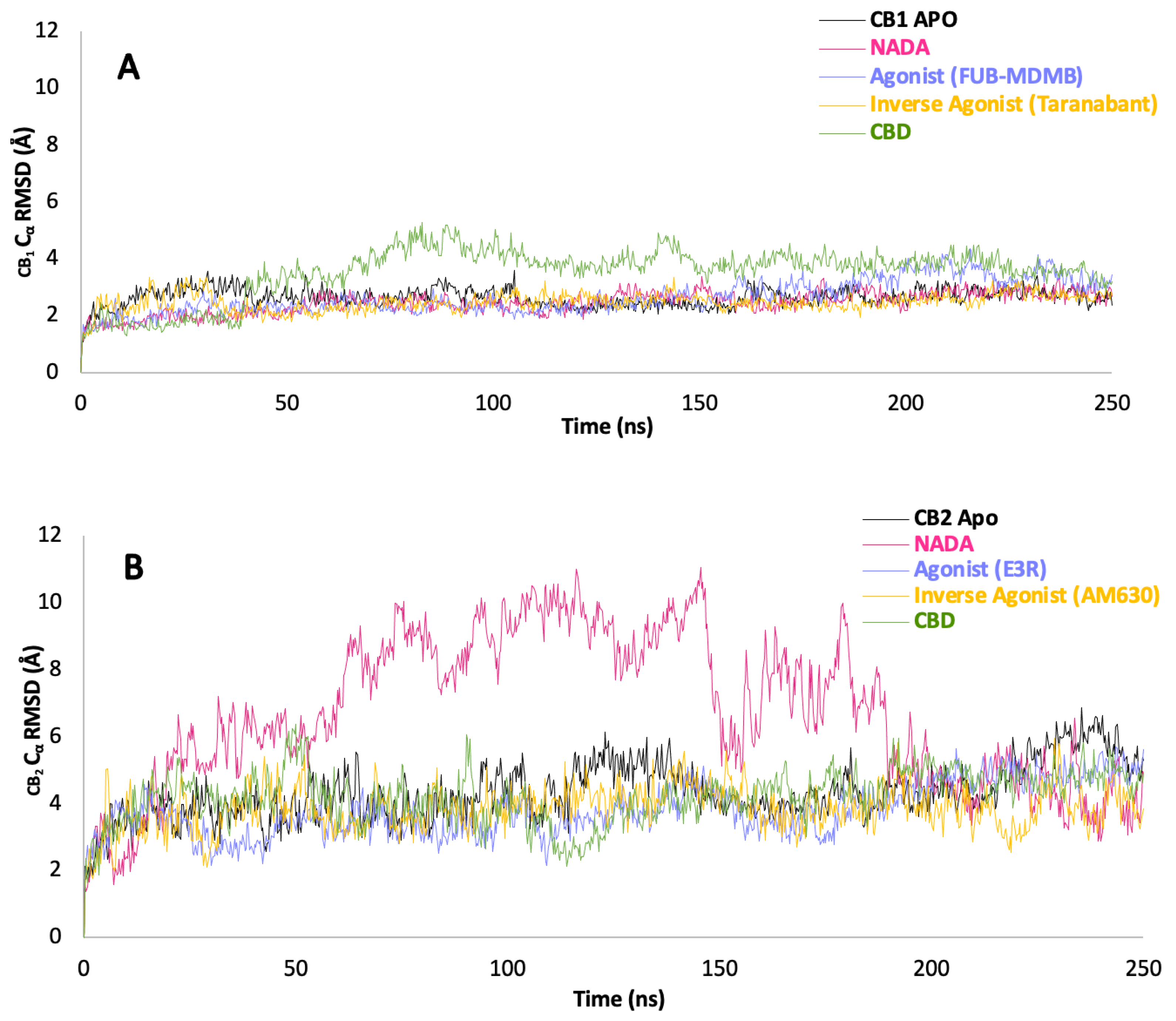
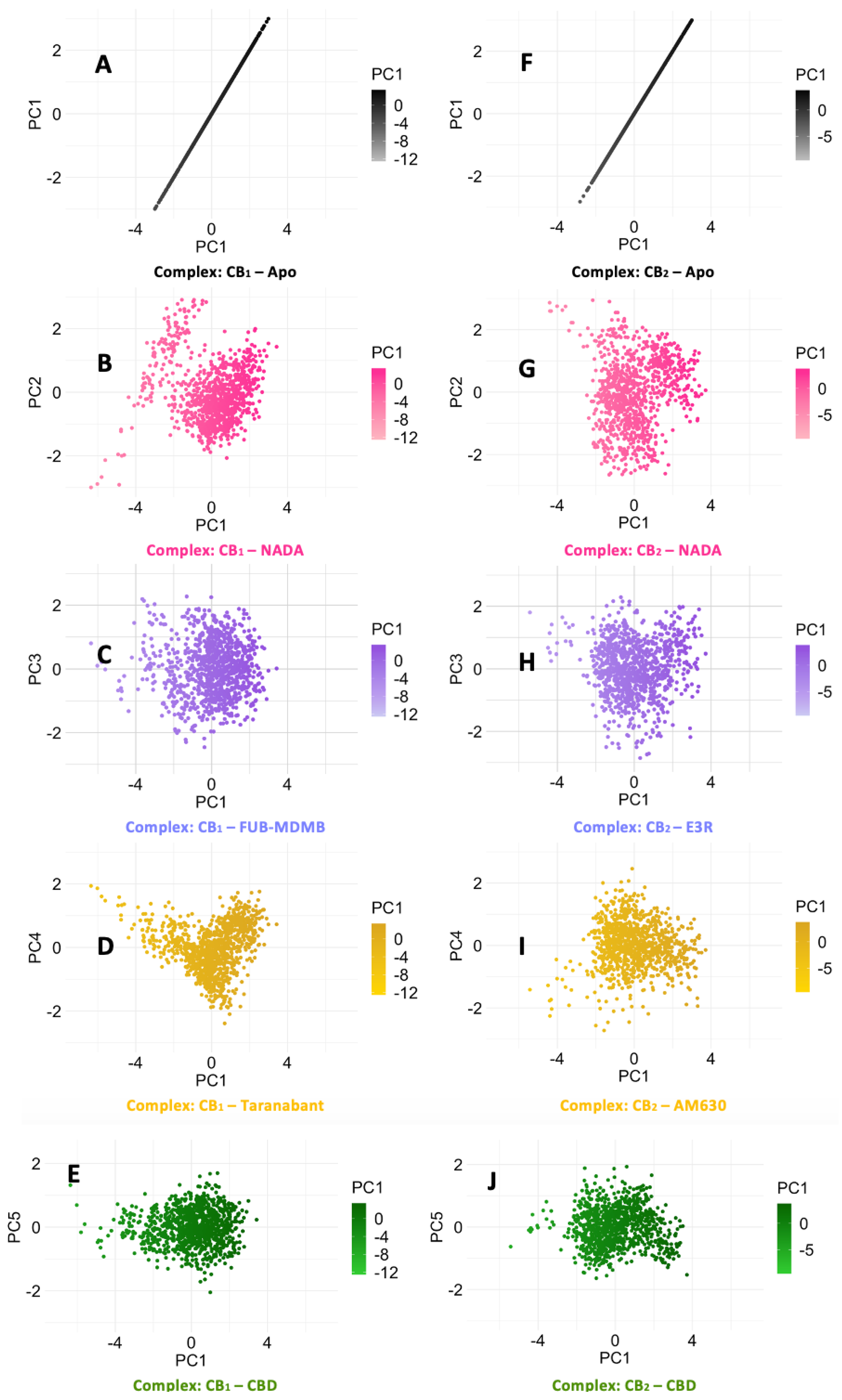
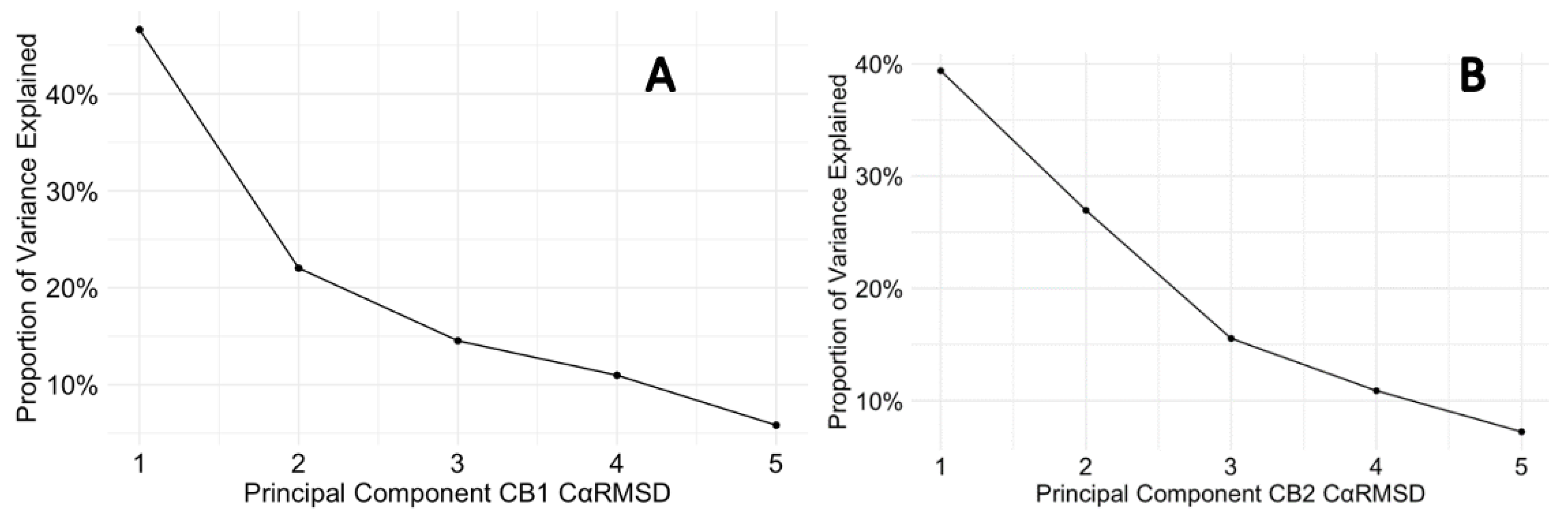
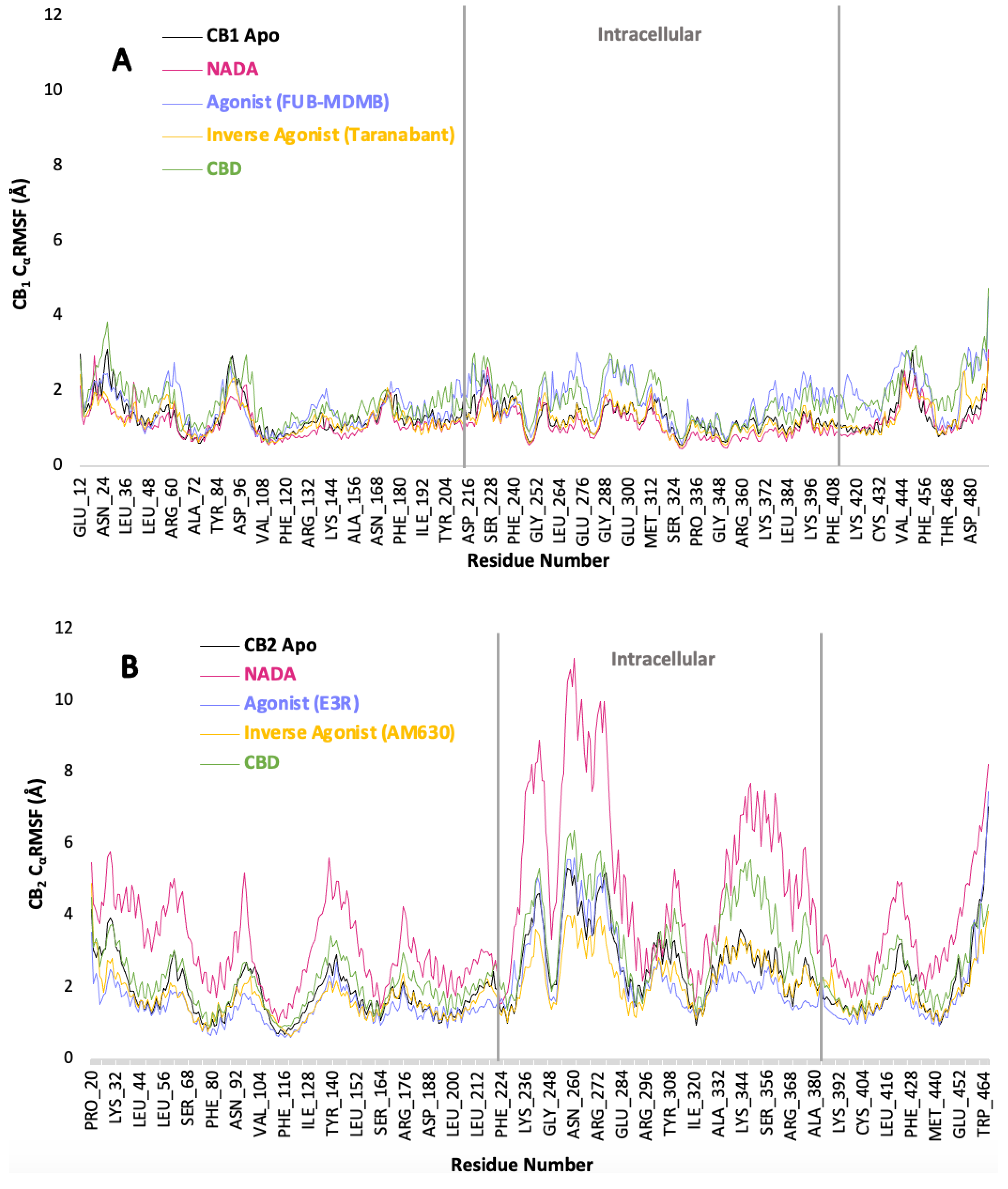
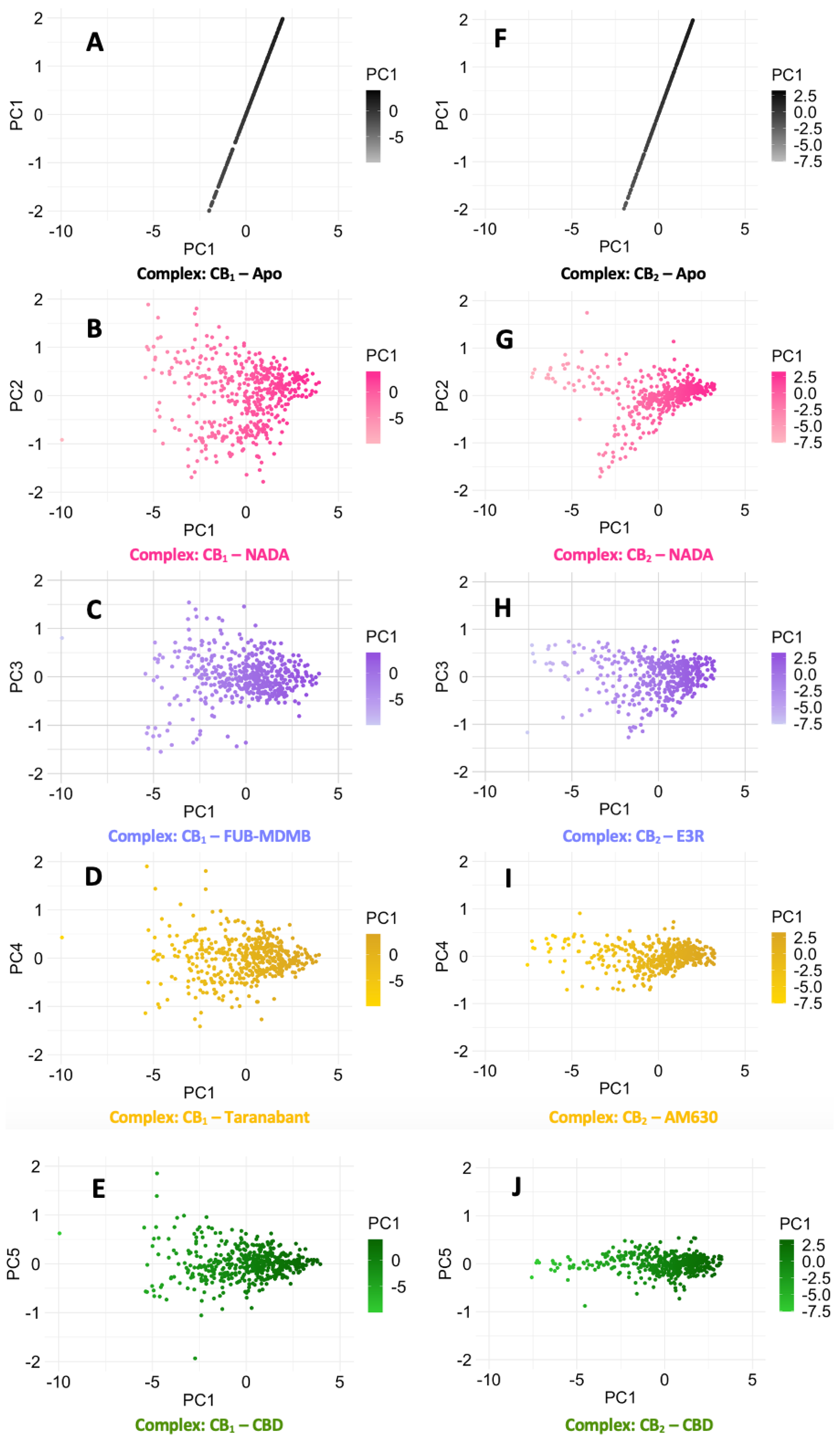
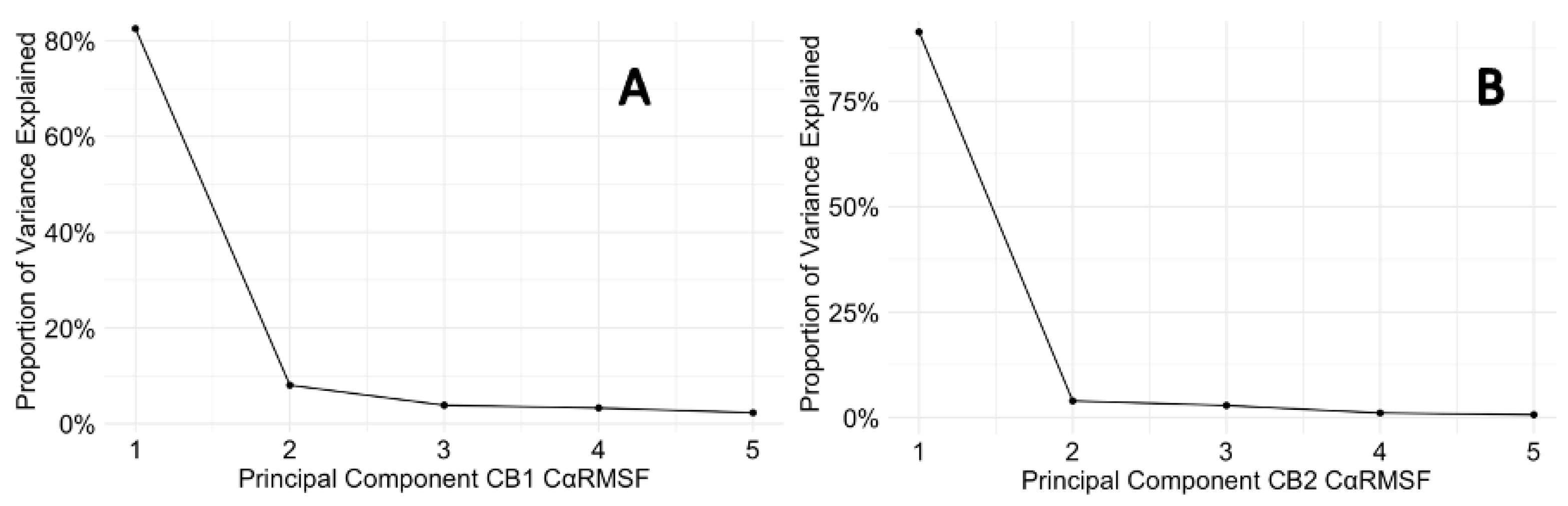
3.6. Protein Backbone Cα RMSF
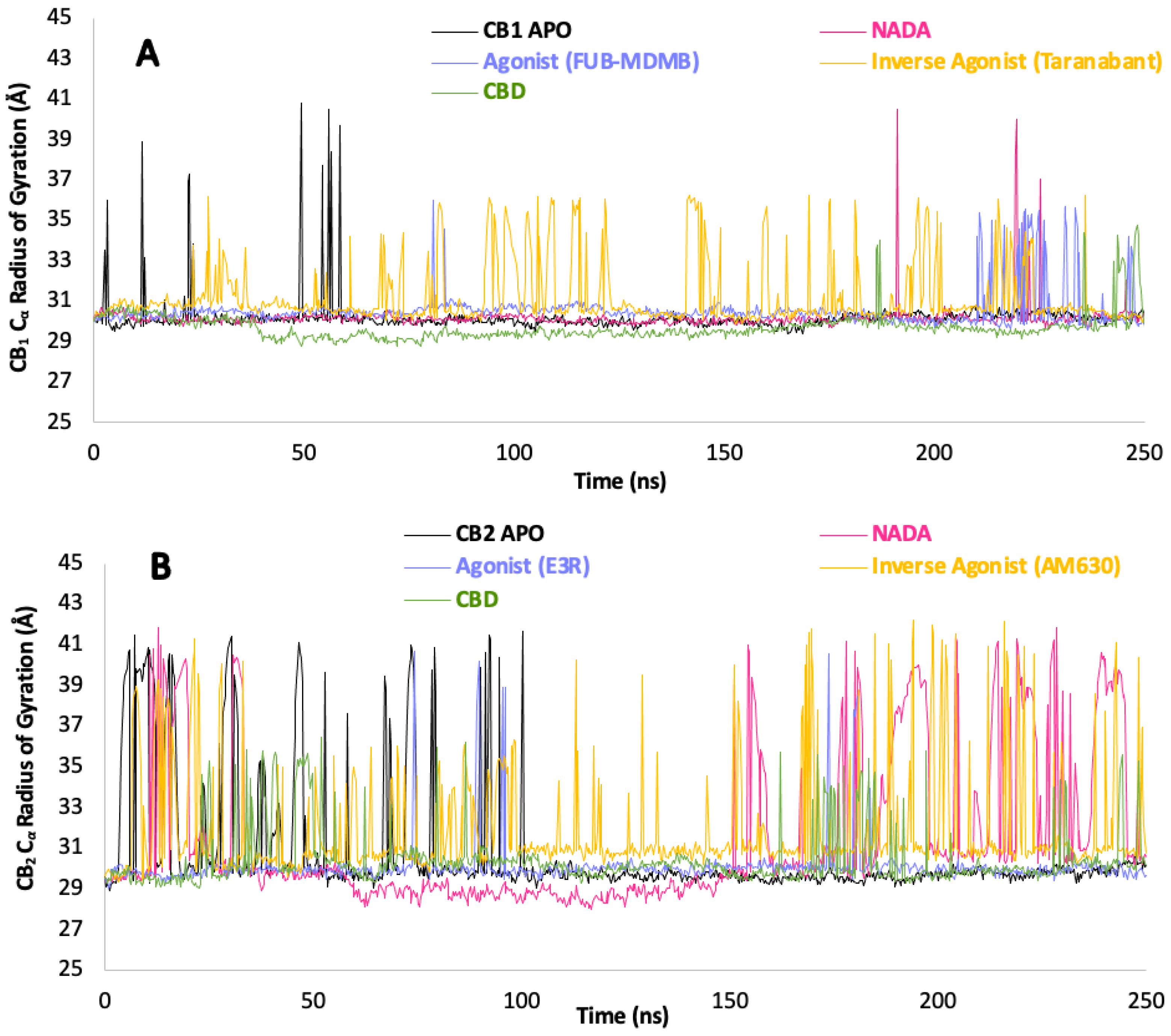
3.7. Protein Backbone Cα Radius of Gyration
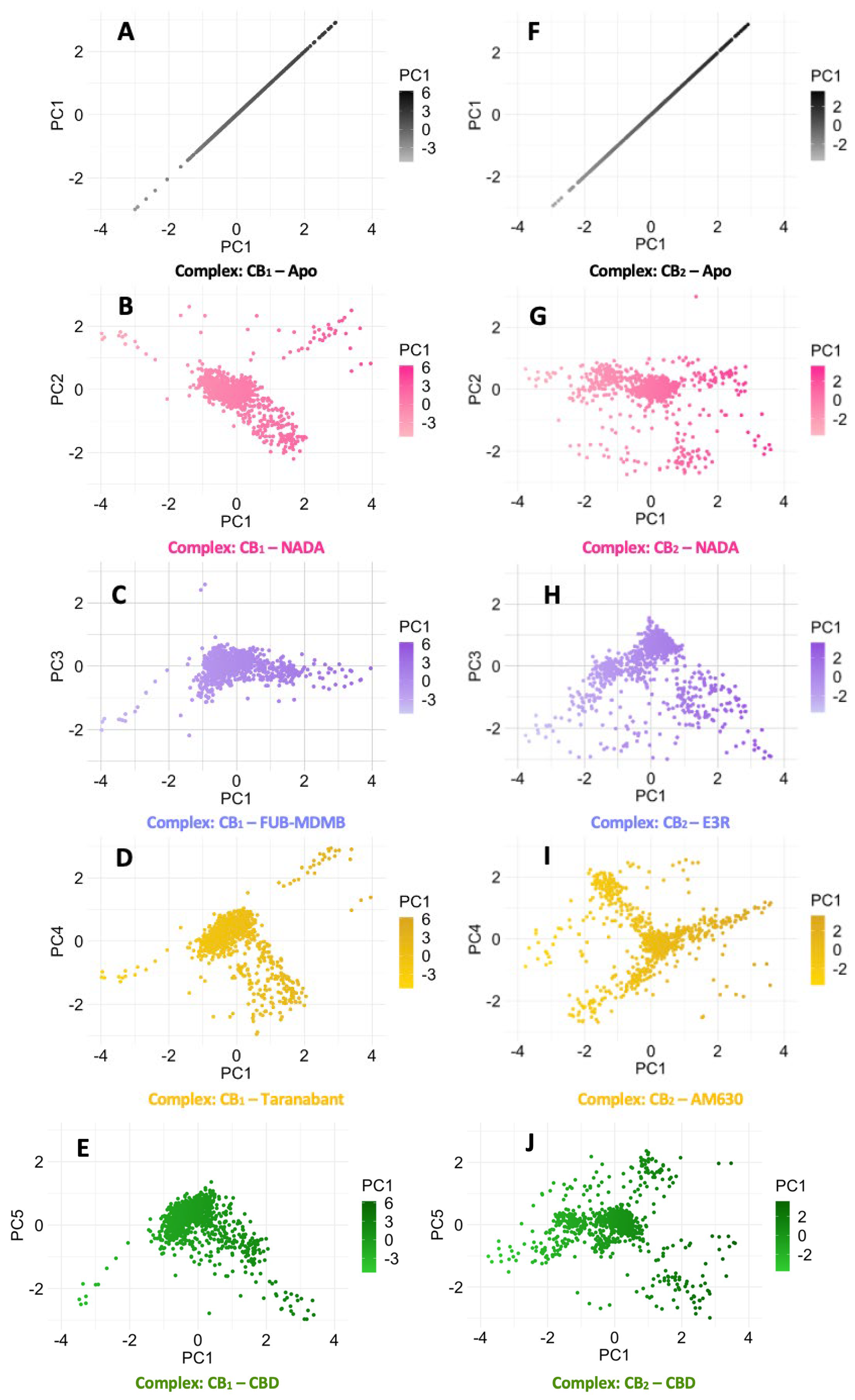
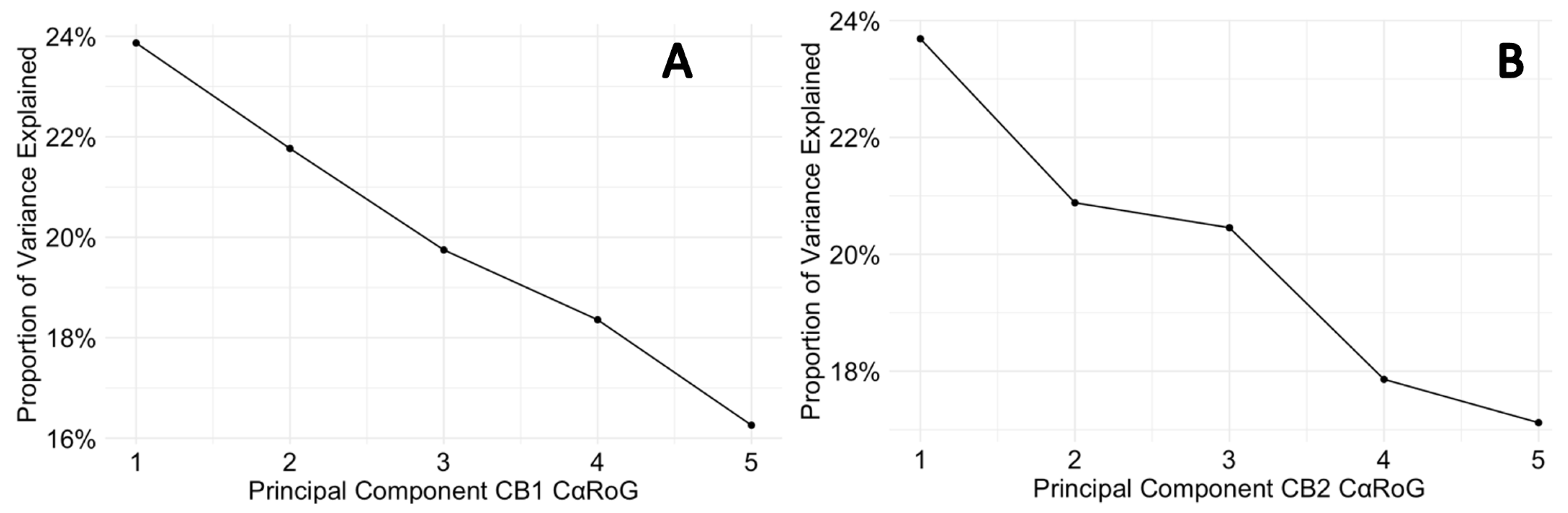
3.8. Simulation Trajectory Snapshots
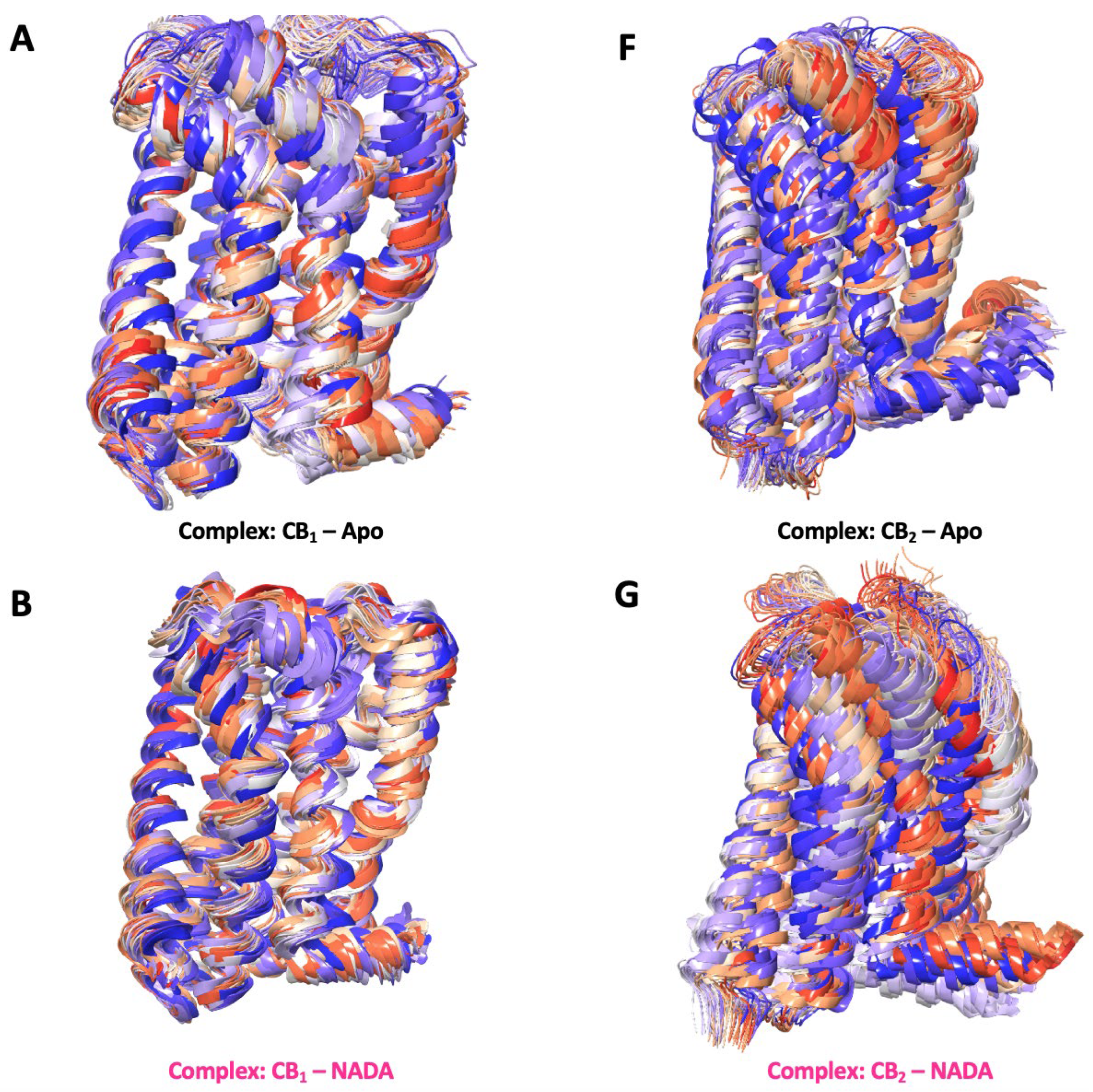
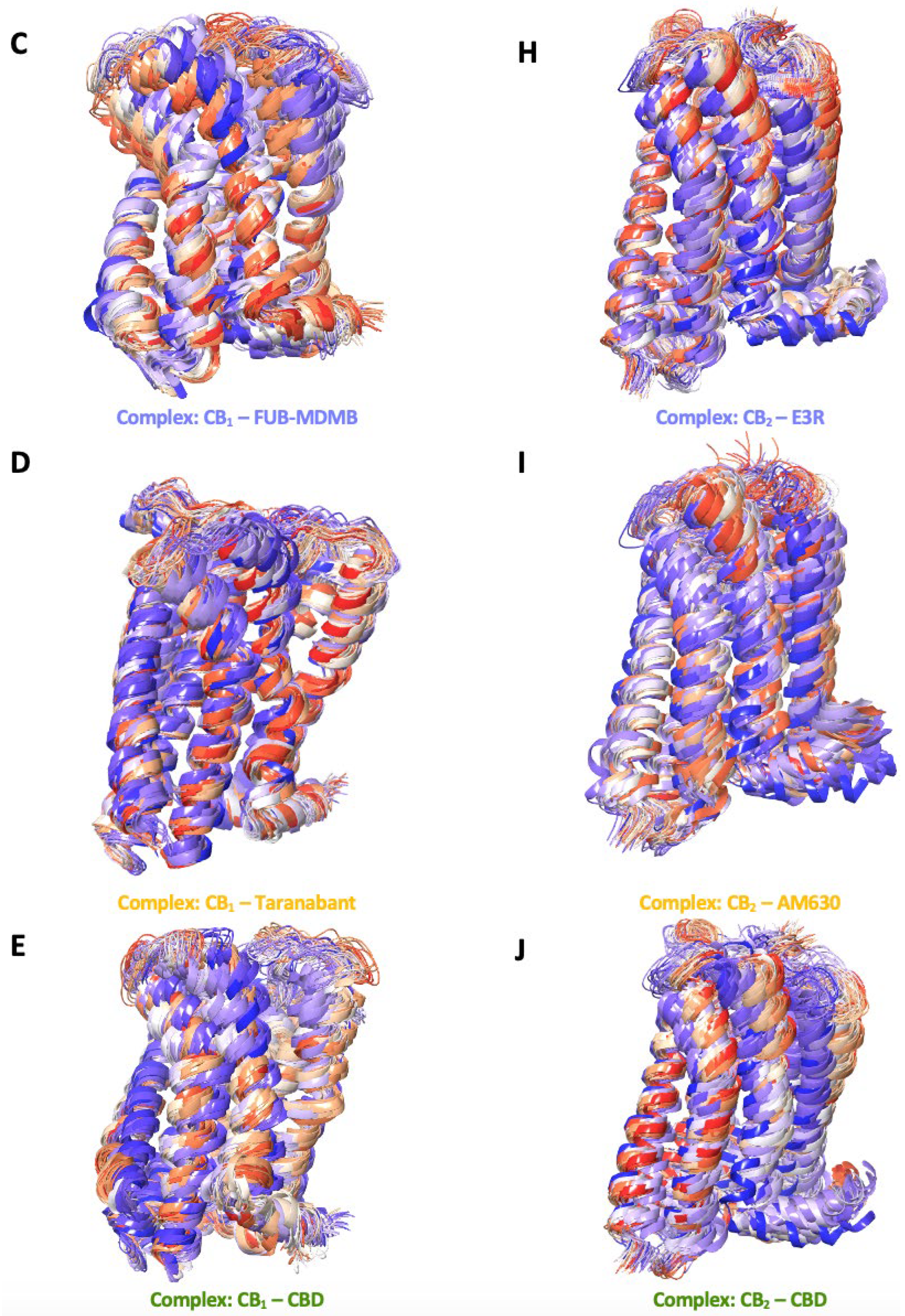
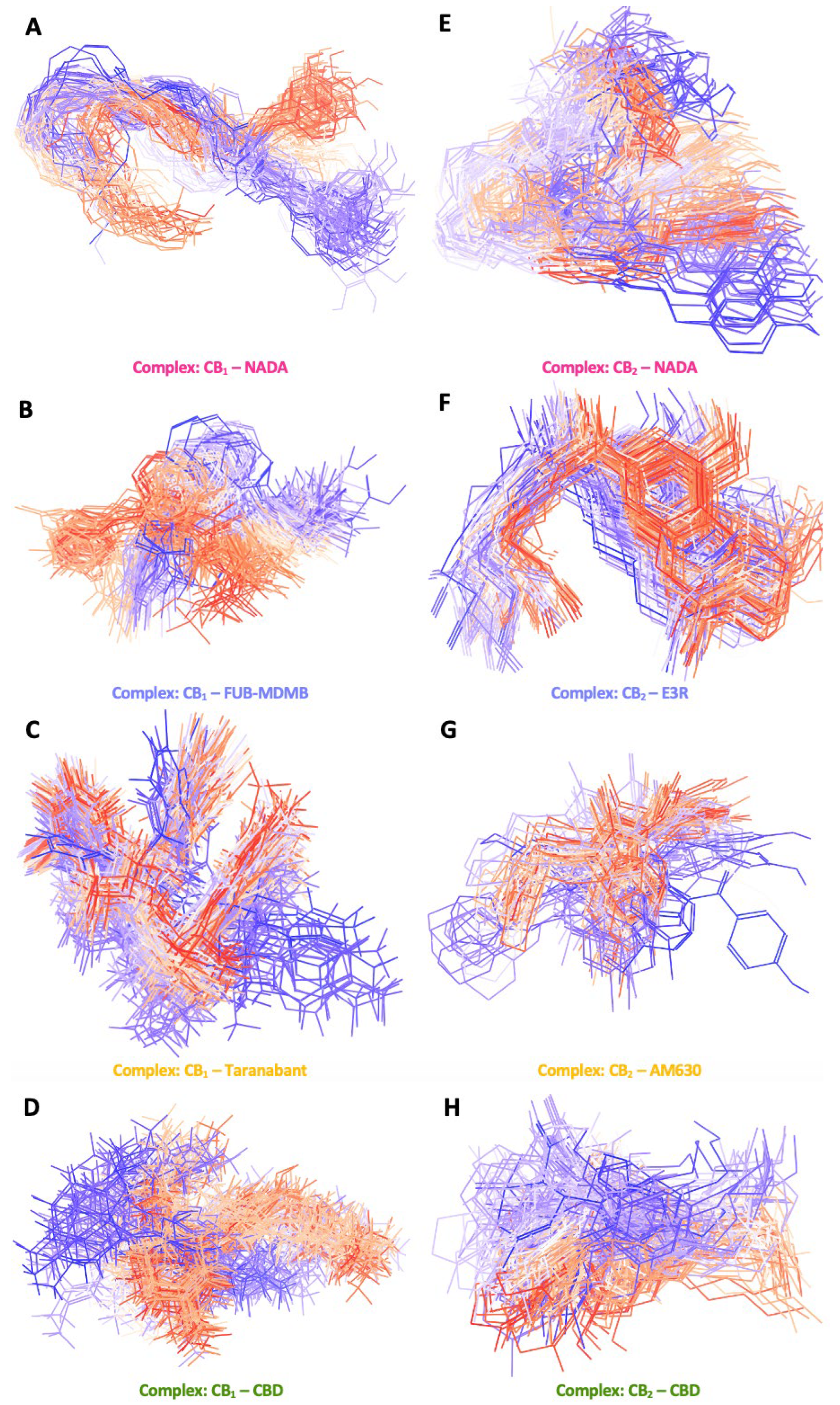
3.9. Ligand RMSD and Simulation Protein-Ligand Interactions
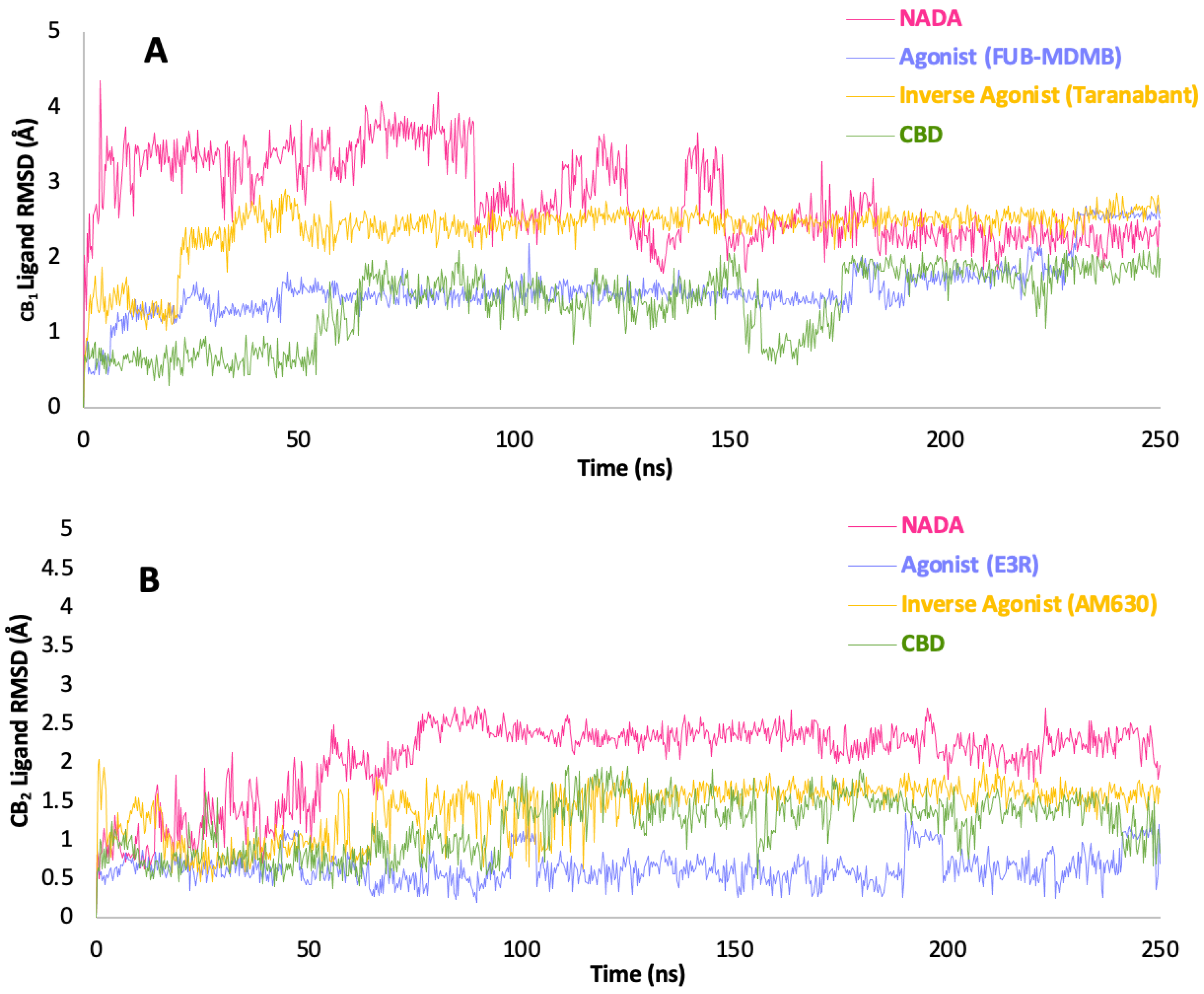
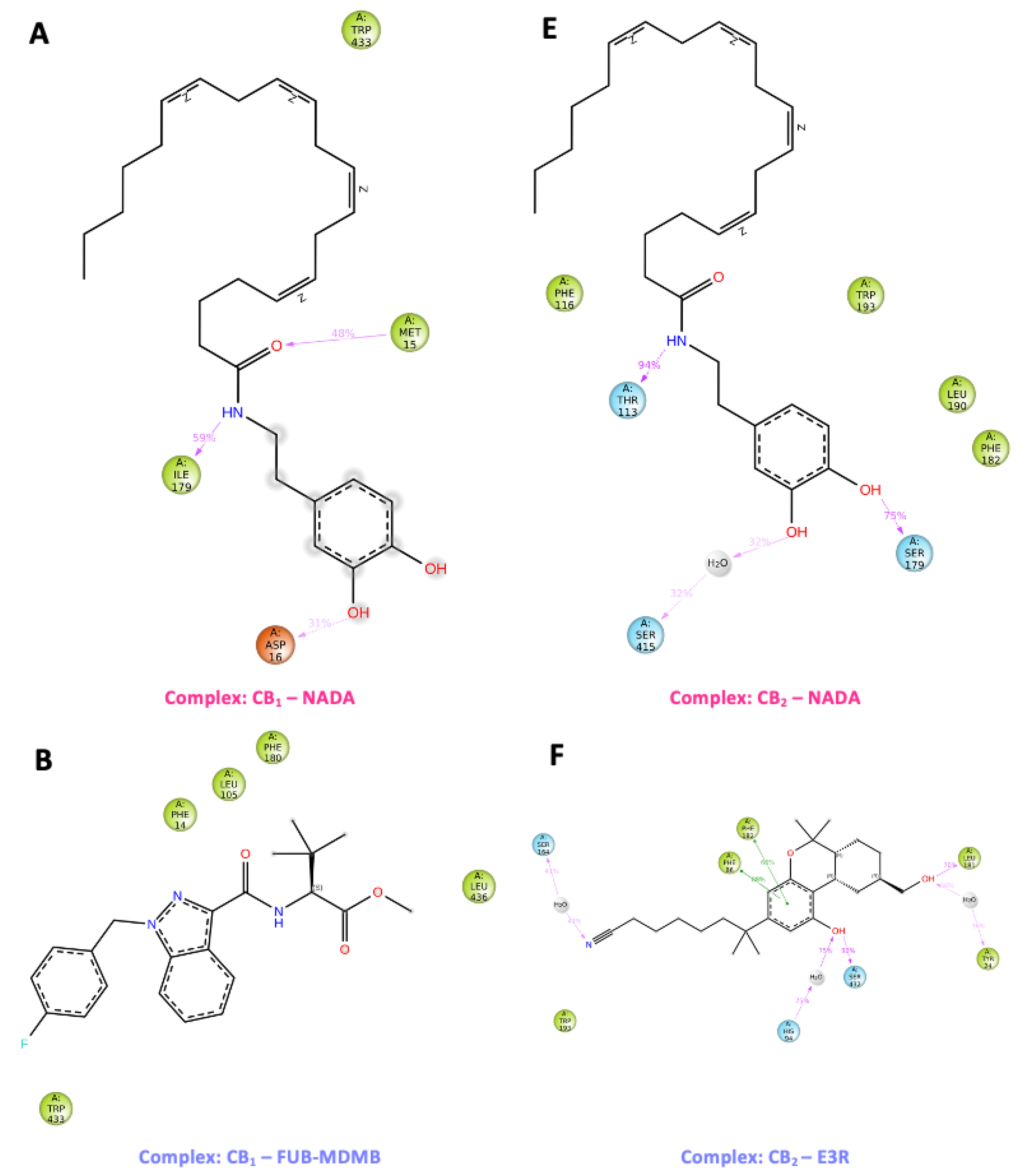
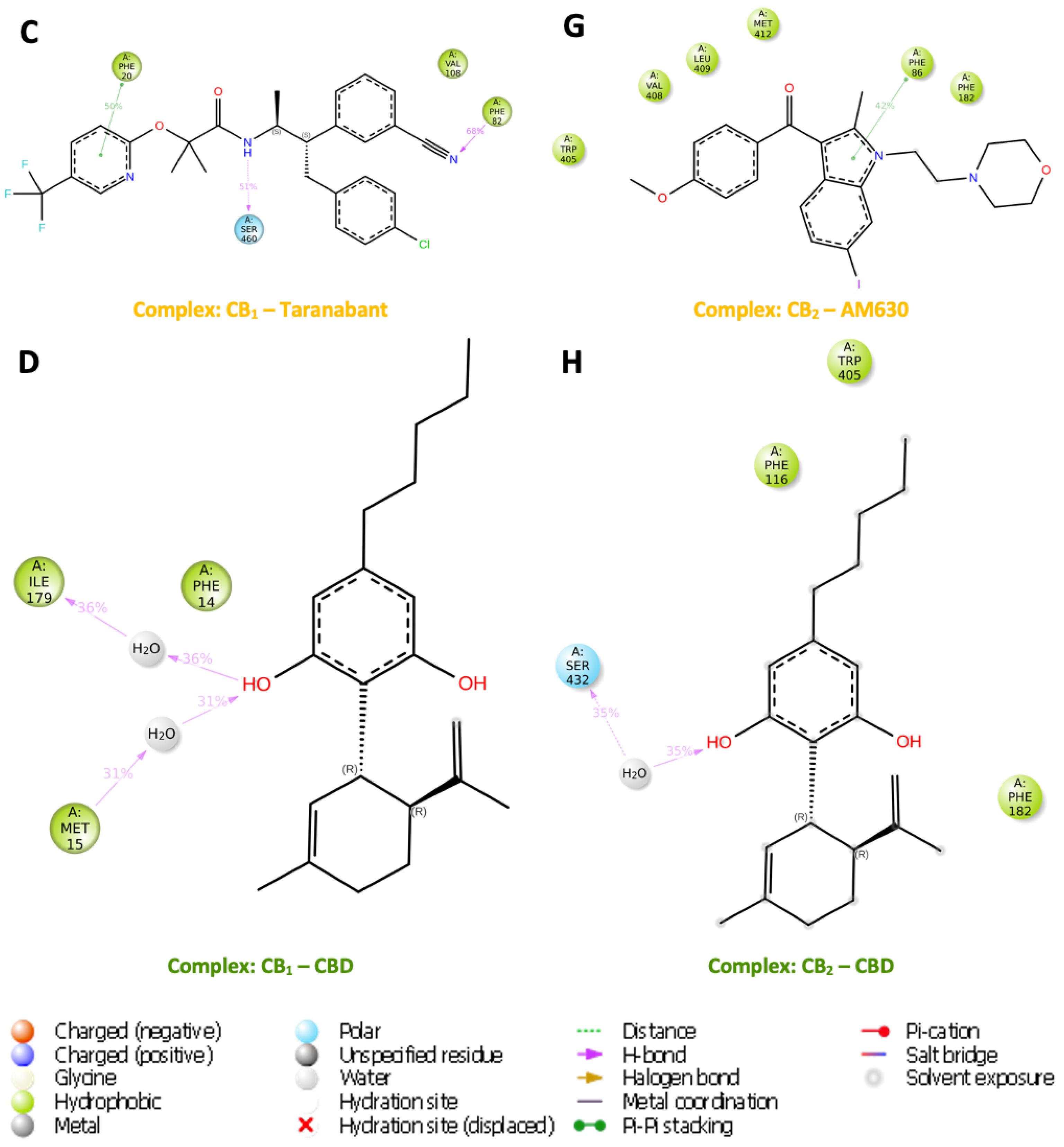
3.10. Water Analysis
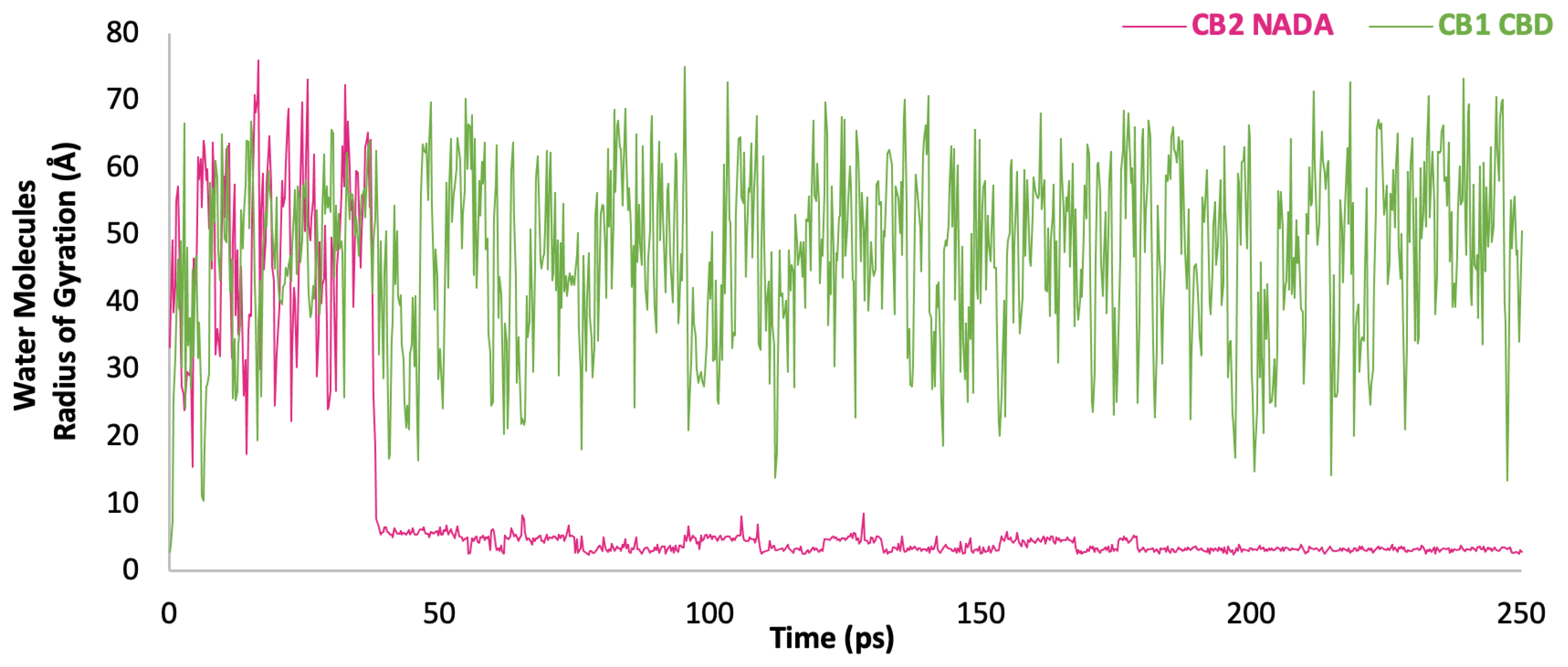
4. Discussion
5. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
Appendix A

References
- UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Research. 2023, 51, D523–D31. [CrossRef]
- Herkenham, M.; Lynn, A.B.; Johnson, M.R.; Melvin, L.S.; de Costa, B.R.; Rice, K.C. Characterization and localization of cannabinoid receptors in rat brain: a quantitative in vitro autoradiographic study. Journal of Neuroscience 1991, 11, 563–83. [Google Scholar] [CrossRef]
- Mackie, K. Cannabinoid receptors: where they are and what they do. Journal of neuroendocrinology. 2008, 20, 10–4. [Google Scholar] [CrossRef] [PubMed]
- Kano, M.; Ohno-Shosaku, T.; Hashimotodani, Y.; Uchigashima, M.; Watanabe, M. Endocannabinoid-mediated control of synaptic transmission. Physiological reviews. 2009.
- Marsicano, G.; Lutz, B. Neuromodulatory functions of the endocannabinoid system. Journal of endocrinological investigation. 2006, 29, 27. [Google Scholar]
- Turcotte, C.; Blanchet, M.-R.; Laviolette, M.; Flamand, N. The CB 2 receptor and its role as a regulator of inflammation. Cellular and Molecular Life Sciences 2016, 73, 4449–70. [Google Scholar] [CrossRef] [PubMed]
- Scuderi, C.; Filippis, D.D.; Iuvone, T.; Blasio, A.; Steardo, A.; Esposito, G. Cannabidiol in medicine: a review of its therapeutic potential in CNS disorders. Phytotherapy Research: An International Journal Devoted to Pharmacological and Toxicological Evaluation of Natural Product Derivatives. 2009, 23, 597–602. [Google Scholar] [CrossRef] [PubMed]
- Slomski, A. THC for Chronic Pain. JAMA. 2018, 320, 1631. [Google Scholar] [CrossRef]
- White, C.M. A review of human studies assessing cannabidiol's (CBD) therapeutic actions and potential. The Journal of Clinical Pharmacology. 2019, 59, 923–34. [Google Scholar] [CrossRef]
- Haubrich, J. Conformational dynamics and pharmacology in transmembrane dimeric receptors; Université Montpellier, 2021. [Google Scholar]
- Vaidehi, N. Dynamics and flexibility of G-protein-coupled receptor conformations and their relevance to drug design. Drug discovery today. 2010, 15, 951–7. [Google Scholar] [CrossRef]
- Eldeeb, K.; Leone-Kabler, S.; Howlett, A.C. CB1 cannabinoid receptor-mediated increases in cyclic AMP accumulation are correlated with reduced Gi/o function. Journal of basic and clinical physiology and pharmacology 2016, 27, 311–22. [Google Scholar] [CrossRef]
- Hilger, D. The role of structural dynamics in GPCR-mediated signaling. The FEBS journal 2021, 288, 2461–89. [Google Scholar] [CrossRef]
- Haschek-Hock, W.M.; Rousseaux, C.G.; Wallig, M.A.; Bolon, B. Haschek and Rousseaux's Handbook of Toxicologic Pathology; Principles and Practice of Toxicologic Pathology; Academic press, 2021; Volume 1. [Google Scholar]
- Mechoulam, R.; Fride, E.; Di Marzo, V. Endocannabinoids. European journal of pharmacology. 1998, 359, 1–18. [CrossRef] [PubMed]
- Adams, R.; Hunt, M.; Clark, J. Structure of cannabidiol, a product isolated from the marihuana extract of Minnesota wild hemp. I. Journal of the American chemical society 1940, 62, 196–200. [Google Scholar] [CrossRef]
- Adams, R.; Pease, D.; Cain, C.; Clark, J. Structure of cannabidiol. VI. Isomerization of cannabidiol to tetrahydrocannabinol, a physiologically active product. Conversion of cannabidiol to cannabinol1. Journal of the American Chemical Society 1940, 62, 2402–5. [Google Scholar] [CrossRef]
- Holt, A.K.; Poklis, J.L.; Peace, M.R. ∆ 8-THC, THC-O Acetates and CBD-di-O Acetate: Emerging Synthetic Cannabinoids Found in Commercially Sold Plant Material and Gummy Edibles. Journal of Analytical Toxicology 2022, 46, 940–8. [Google Scholar] [CrossRef]
- Gromiha, M.M. Protein bioinformatics: from sequence to function; Academic Press, 2010. [Google Scholar]
- Zhao, H.; Caflisch, A. Molecular dynamics in drug design. European journal of medicinal chemistry. 2015, 91, 4–14. [Google Scholar]
- Manandhar, A.; Haron, M.H.; Klein, M.L.; Elokely, K. Understanding the Dynamics of the Structural States of Cannabinoid Receptors and the Role of Different Modulators. Life 2022, 12, 2137. [Google Scholar] [CrossRef]
- Hua, T., Vemuri, K., Pu, M., Qu, L., Han, G.W., Wu, Y., et al. Crystal structure of the human cannabinoid receptor CB1. Cell 2016, 167, 750–62.e14. [CrossRef]
- Li, X., Hua, T., Vemuri, K., Ho, J.-H., Wu, Y., Wu, L., et al. Crystal structure of the human cannabinoid receptor CB2. Cell 2019, 176, 459–67.e13. [CrossRef]
- Aminpour, M., Montemagno, C., Tuszynski, J.A. An overviewof molecular modeling for drug discovery with specific illustrative examples of applications. Molecules 2019, 24, 1693. [CrossRef] [PubMed]
- Nadendla, R.R. Molecular modeling: A powerful tool for drug design and molecular docking. Resonance. 2004, 9, 51–60. [Google Scholar] [CrossRef]
- Karplus, M., Kuriyan, J. Molecular dynamics and protein function. Proceedings of the National Academy of Sciences 2005, 102, 6679–85. [CrossRef]
- Berman, H.M., Westbrook, J., Feng, Z., Gilliland, G., Bhat, T.N., Weissig, H., et al. The protein data bank. The protein data bank. Nucleic acids research 2000, 28, 235–42. [CrossRef] [PubMed]
- McCammon, J.A., Gelin, B.R., Karplus, M. Dynamics of folded proteins. Nature 1977, 267, 585–90. [CrossRef]
- Hua, T., Li, X., Wu, L., Iliopoulos-Tsoutsouvas, C., Wang, Y., Wu, M., et al. Activation and signaling mechanism revealed by cannabinoid receptor-Gi complex structures. Cell 2020, 180, 655–65 e18. [CrossRef]
- Shao, Z., Yin, J., Chapman, K., Grzemska, M., Clark, L., Wang, J., Rosenbaum, D.M. High-resolution crystal structure of the human CB1 cannabinoid receptor. Nature 2016, 540, 602–6. [CrossRef]
- Waterhouse, A., Bertoni, M., Bienert, S., Studer, G., Tauriello, G., Gumienny, R., et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic acids research 2018, 46, W296–W303. [CrossRef] [PubMed]
- Madhavi Sastry, G., Adzhigirey, M., Day, T., Annabhimoju, R., Sherman, W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. Journal of computer-aided molecular design 2013, 27, 221–34. [CrossRef]
- Jacobson, M.P., Friesner, R.A., Xiang, Z., Honig, B. On the role of the crystal environment in determining protein side-chain conformations. Journal of molecular biology 2002, 320, 597–608. [CrossRef]
- Jacobson, M.P., Pincus, D.L., Rapp, C.S., Day, T.J., Honig, B., Shaw, D.E., Friesner, R.A. A hierarchical approach to all-atom protein loop prediction. Proteins: Structure, Function, and Bioinformatics. 2004, 55, 351–67. [CrossRef] [PubMed]
- Jorgensen, W.L., Maxwell, D.S., Tirado-Rives, J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. Journal of the American Chemical Society 1996, 118, 11225–36. [CrossRef]
- Bochevarov, A.D., Watson, M.A., Greenwood, J.R., Philipp, D.M. Multiconformation, density functional theory-based p K a prediction in application to large, flexible organic molecules with diverse functional groups. Journal of chemical theory and computation 2016, 12, 6001–19. [CrossRef]
- Johnston, R.C., Yao, K., Kaplan, Z., Chelliah, M., Leswing, K., Seekins, S., et al. Epik: p K a and Protonation State Prediction through Machine Learning. Journal of Chemical Theory and Computation 2023, 19, 2380–8. [CrossRef]
- Shelley, J.C., Cholleti, A., Frye, L.L., Greenwood, J.R., Timlin, M.R., Uchimaya, M. Epik: a software program for pK a prediction and protonation state generation for drug-like molecules. Journal of computer-aided molecular design 2007, 21, 681–91. [CrossRef] [PubMed]
- Farid, R., Day, T., Friesner, R.A., Pearlstein, R.A. New insights about HERG blockade obtained from protein modeling, potential energy mapping, and docking studies. Bioorganic & medicinal chemistry 2006, 14, 3160–73.
- Friesner, R.A., Banks, J.L., Murphy, R.B., Halgren, T.A., Klicic, J.J., Mainz, D.T., et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. Journal of medicinal chemistry 2004, 47, 1739–49. [CrossRef] [PubMed]
- Friesner, R.A., Murphy, R.B., Repasky, M.P., Frye, L.L., Greenwood, J.R., Halgren, T.A., et al. Extra precision glide: Docking and scoring incorporating a model of hydrophobic enclosure for protein− ligand complexes. Journal of medicinal chemistry 2006, 49, 6177–96. [CrossRef]
- Halgren, T.A., Murphy, R.B., Friesner, R.A., Beard, H.S., Frye, L.L., Pollard, W.T., Banks, J.L. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. Journal of medicinal chemistry 2004, 47, 1750–9. [CrossRef] [PubMed]
- Sherman, W., Beard, H.S., Farid, R. Use of an induced fit receptor structure in virtual screening. Chemical biology & drug design 2006, 67, 83–4.
- Sherman, W., Day, T., Jacobson, M.P., Friesner, R.A., Farid, R. Novel procedure for modeling ligand/receptor induced fit effects. Journal of medicinal chemistry 2006, 49, 534–53. [CrossRef]
- Yang, Y., Yao, K., Repasky, M.P., Leswing, K., Abel, R., Shoichet, B.K., Jerome, S.V. Efficient exploration of chemical space with docking and deep learning. Journal of Chemical Theory and Computation 2021, 17, 7106–19. [CrossRef]
- Bowers, K.J., Chow, E., Xu, H., Dror, R.O., Eastwood, M.P., Gregersen, B.A., et al. Scalable algorithms for molecular dynamics simulations on commodity clusters. In Proceedings of the 2006 ACM/IEEE Conference on Supercomputing; 2006; p. 84-es.
- RStudio, R.T. Integrated development environment for R. RStudio, PBC: Boston, MA, USA, 2020. [Google Scholar]
- Tang, Y., Horikoshi, M., Li, W. ggfortify: unified interface to visualize statistical results of popular R packages. . R J. 2016, 8, 474.
- Aboytes-Ojeda, M., Castillo-Villar, K.K., Yu, T.-h.E., Boyer, C.N., English, B.C., Larson, J.A., et al. A principal component analysis in switchgrass chemical composition. Energies 2016, 9, 913. [CrossRef]
- Team, R.D.C. R: A language and environment for statistical computing. (No Title). 2010. [Google Scholar]
- Kleywegt, G.J., Jones, T.A. Phi/psi-chology: Ramachandran revisited. Structure. 1996, 4, 1395–400. [PubMed]
- Schrodinger, L. The PyMOL molecular graphics system. Version. 2015, 1, 8. [Google Scholar]
- Leach, A.R. Molecular modelling: principles and applications. Pearson education, 2001. [Google Scholar]
- Nickels, J.D., Katsaras, J. Water and lipid bilayers. Membrane hydration: The role of water in the structure and function of biological membranes. 2015, 45–67.
- Kumar, K.K., Shalev-Benami, M., Robertson, M.J., Hu, H., Banister, S.D., Hollingsworth, S.A., et al. Structure of a signaling cannabinoid receptor 1-G protein complex. Cell 2019, 176, 448–58 e12. [CrossRef] [PubMed]
- Wojtalla, A., Herweck, F., Granzow, M., Klein, S., Trebicka, J., Huss, S., et al. The endocannabinoid N-arachidonoyl dopamine (NADA) selectively induces oxidative stress-mediated cell death in hepatic stellate cells but not in hepatocytes. American Journal of Physiology-Gastrointestinal and Liver Physiology 2012, 302, G873–G87. [CrossRef]
- Ibsen, M.S., Connor, M., Glass, M. Cannabinoid CB1 and CB2 receptor signaling and bias. Cannabis and cannabinoid research 2017, 2, 48–60. [CrossRef] [PubMed]
- Bradshaw, H.B., Rimmerman, N., Krey, J.F., Walker, J.M. Sex and hormonal cycle differences in rat brain levels of pain-related cannabimimetic lipid mediators. American Journal of Physiology-Regulatory, Integrative and Comparative Physiology 2006, 291, R349–R58. [CrossRef] [PubMed]
- Levitt, M. The birth of computational structural biology. Nature structural biology. 2001, 8, 392–3. [CrossRef]
- Jiang, W., Lacroix, J., Luo, Y.L. Importance of molecular dynamics equilibrium protocol on protein-lipid interaction near channel pore. Biophysical Reports 2022, 2. [CrossRef] [PubMed]
- Jorgensen, W.L. The many roles of computation in drug discovery. Science. 2004, 303, 1813–8. [Google Scholar] [CrossRef]
- Aviz-Amador, A., Contreras-Puentes, N., Mercado-Camargo, J. Virtual screening using docking and molecular dynamics of cannabinoid analogs against CB1 and CB2 receptors. Computational Biology and Chemistry 2021, 95, 107590. [CrossRef]
- Di Marzo, V., Piscitelli, F. The endocannabinoid system and its modulation by phytocannabinoids. Neurotherapeutics 2015, 12, 692–8. [CrossRef]
- Russo, E.B., Hohmann, A.G. Role of cannabinoids in pain management. In Comprehensive Treatment of Chronic Pain by Medical, Interventional, and Integrative Approaches: The AMERICAN ACADEMY OF PAIN MEDICINE Textbook on Patient Management; Springer, 2012; pp. 181–97.
- Pertwee, R. The diverse CB1 and CB2 receptor pharmacology of three plant cannabinoids: Δ9-tetrahydrocannabinol, cannabidiol and Δ9-tetrahydrocannabivarin. British journal of pharmacology. 2008, 153, 199–215. [Google Scholar] [CrossRef]
- Altschul, S.F., Gish, W., Miller, W., Myers, E.W., Lipman, D.J. Basic local alignment search tool. Journal of molecular biology 1990, 215, 403–10. [CrossRef]
- Chen, V.B., Arendall, W.B., Headd, J.J., Keedy, D.A., Immormino, R.M., Kapral, G.J., et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallographica Section D: Biological Crystallography 2010, 66, 12–21. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




