Submitted:
09 August 2024
Posted:
12 August 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Methodological Development
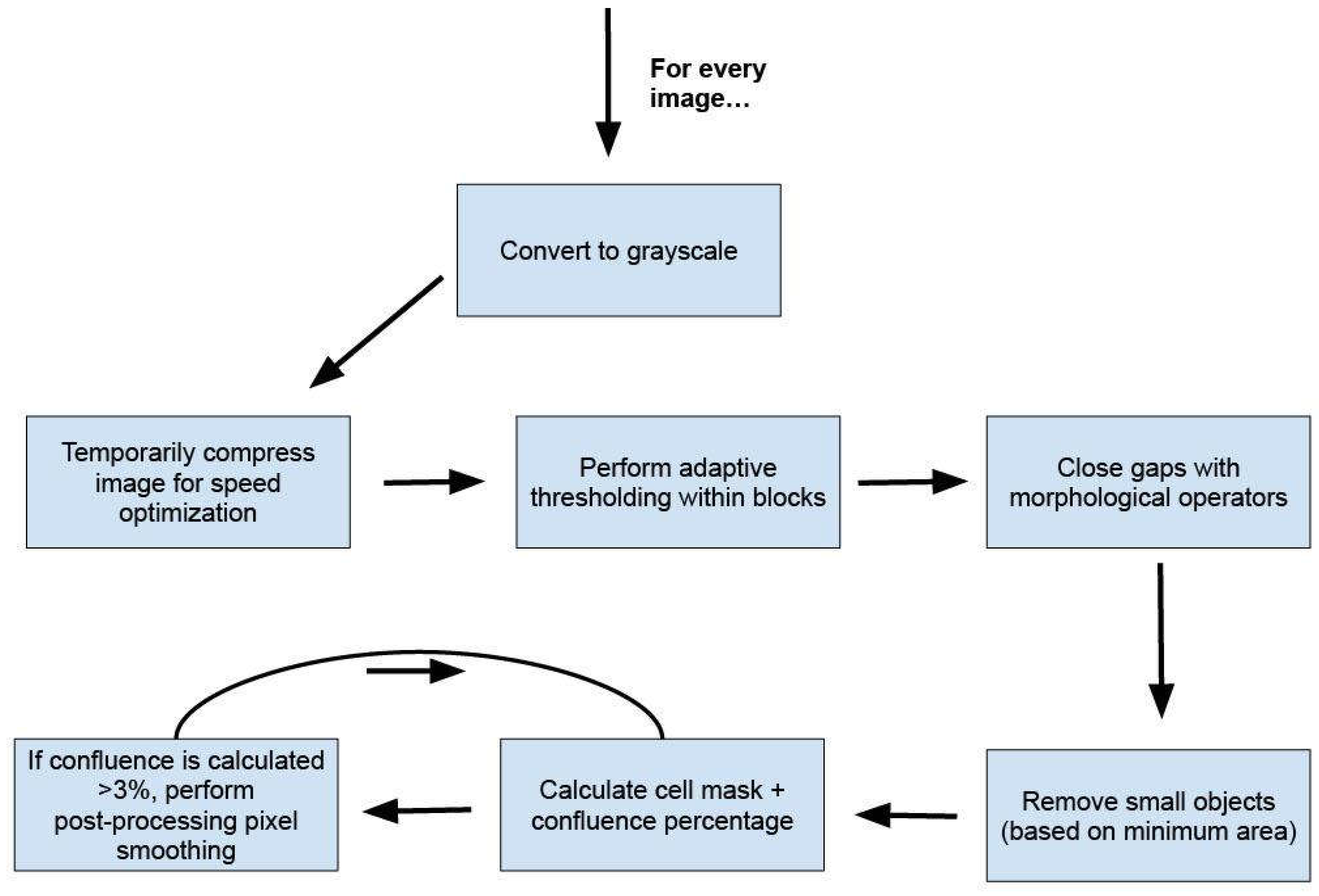
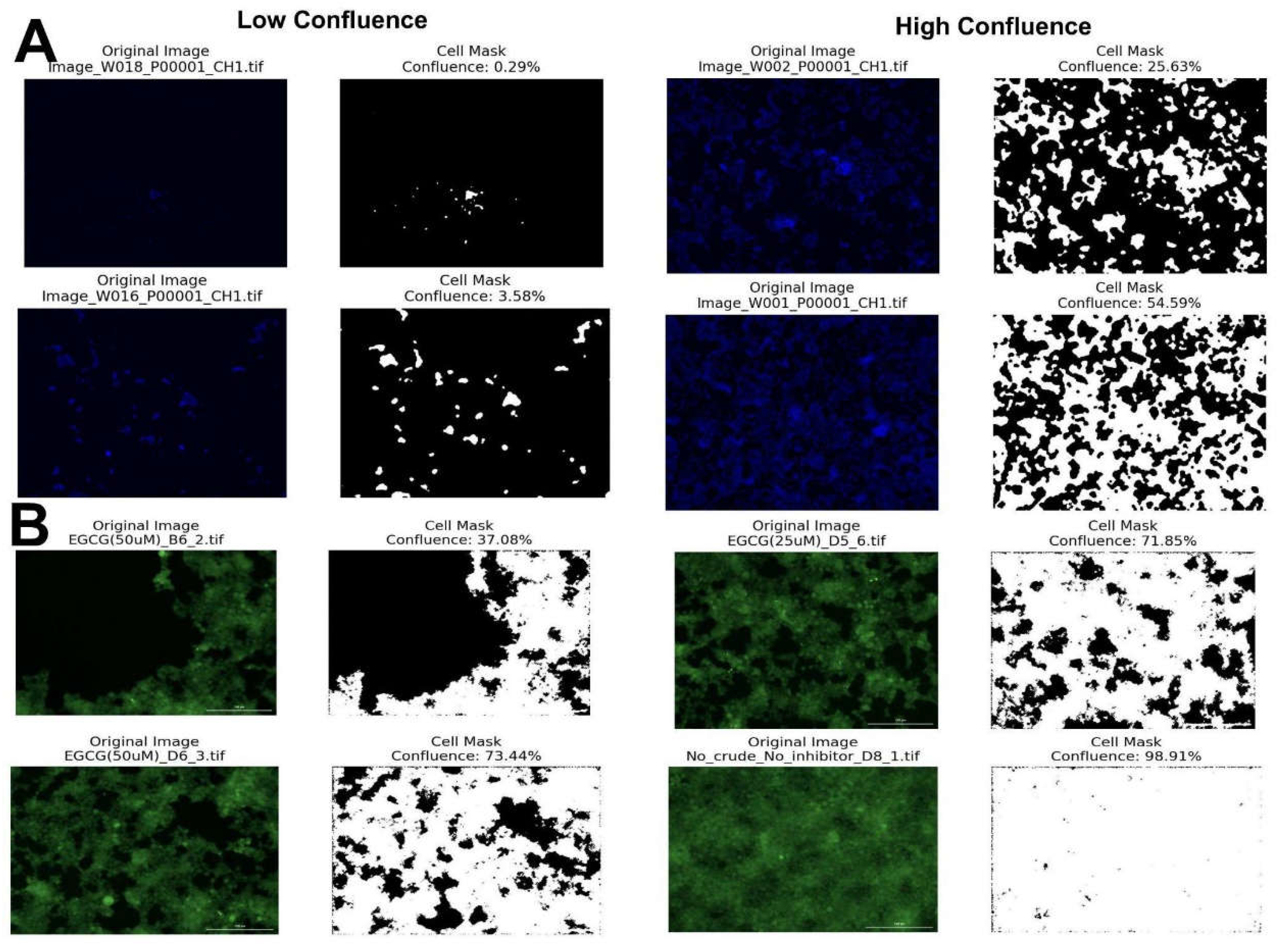
Results
Discussion
Conclusions
Instructions
-
Download Python:Go to python.org and download the appropriate version of Python3.
-
Install/Check Installation of Pip:Run one of the following commands in your Command-Line Interface (CLI):
- OS: python -m ensurepip --upgrade
- Windows: py -m ensurepip --upgrade
-
Downloading Script:Download the ConfluenceAnalysisProgram.txt file linked in supplementary material, change the file extension to .py, and place it in the desired file directory.
-
Running the Script:Launch your preferred CLI such as Terminal for OS, or Command Prompt for Windows. Within your CLI, navigate to the file directory containing ConfluenceAnalysisProgram.py. Then, run the script.
-
File Provision:The program will check that all necessary external libraries are installed (and install them if not). Then, it will prompt you to enter a FOLDER with images of type .jpg, .jpeg, .png, .bmp, .tiff, .tif, or .gif.
-
Pop-up Preview Window:After supplying the path to the relevant folder, a pop-up window will emerge displaying an image within the folder on the left, and the cell mask on the right. To the left are four adjustable parameters that should help you align the cell mask with perceived cell presence in the original image. They are each described below:
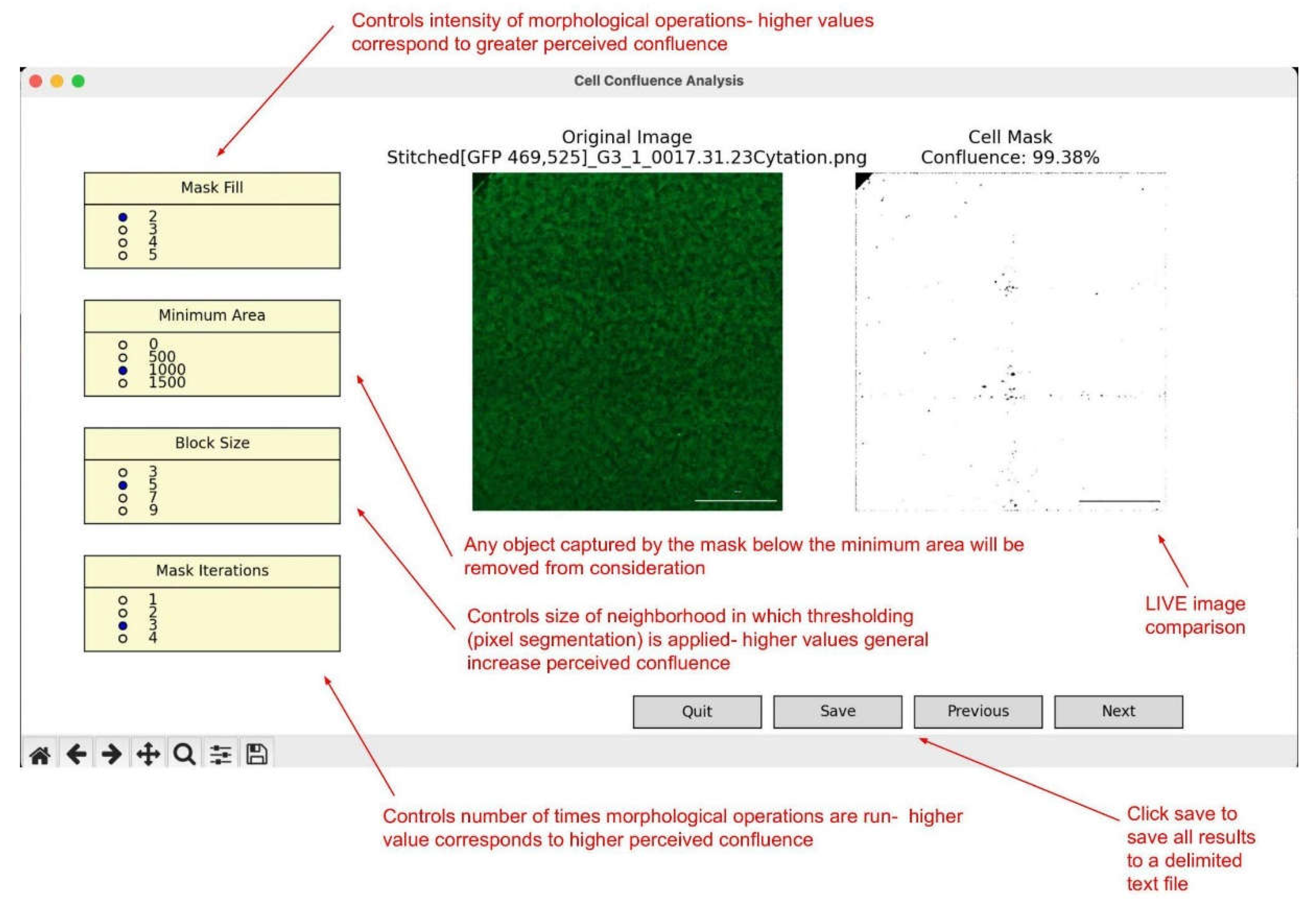
Saving Results
Supplementary Materials
Acknowledgments
References
- Busschots, S.; O’Toole, S.; O’Leary, J.J.; Stordal, B. Non-invasive and non-destructive measurements of confluence in cultured adherent cell lines. MethodsX 2015, 2, 8–13. [Google Scholar] [CrossRef]
- Lobo, V.; Shcherbinina, E.; Westholm, J.O.; et al. Integrative transcriptomic and proteomic profiling of the effects of cell confluency on gene expression. Sci Data 2024, 11, 617. [Google Scholar] [CrossRef] [PubMed]
- Hur, S.S.; del Álamo, J.C.; Park, J.S.; Chien, S. Roles of cell confluency and fluid shear in 3-dimensional intracellular forces in endothelial cells. Proceedings of the National Academy of Sciences 2012, 109, 11110–11115. [Google Scholar] [CrossRef] [PubMed]
- Shamhan, M.; Idris, A.S.; Toha, S.F.; Daud, M.F.; Idris, I.M.; Malik, H. An automated approach for fibroblast cell confluency characterisation and sample handling using AIoT for bio-research and bio-manufacturing. Cogent Engineering 2023, 10. [Google Scholar] [CrossRef]
- CellProfiler. Retrieved from https://cellprofiler.org/.
- Malik, H.; Idris, A.S.; Toha, S.F.; Mohd Idris, I.; Daud, M.F.; Azmi, N.L. A review of open-source image analysis tools for mammalian cell culture: algorithms, features and implementations. PeerJ Computer Science 2023, 9, e1364. [Google Scholar] [CrossRef]
- Logos Biosystems. Monitoring confluency of adherent cells in multi-well plates using the CELENA® X High Content Imaging System. 2020. Retrieved from https://logosbio.com/.
- Farrell, C.J.; Cicalese, S.M.; Davis, H.B.; Dogdas, B.; Shah, T.; Culp, T.; Hoang, V.M. Cell confluency analysis on microcarriers by micro-flow imaging. Cytotechnology 2016, 68, 2469–2478. [Google Scholar] [CrossRef] [PubMed]
- Forero, M.G.; Hidalgo, A. Image Processing Methods for Automatic Cell Counting In Vivo or In Situ Using 3D Confocal Microscopy. InTech 2011. [Google Scholar] [CrossRef]
- Chiu, C.-H.; Leu, J.-D.; Lin, T.-T.; Su, P.-H.; Li, W.-C.; Lee, Y.-J.; Cheng, D.-C. Systematic quantification of cell confluence in human normal oral fibroblasts. Applied Sciences 2020, 10, 9146. [Google Scholar] [CrossRef]
- Freshney, R.I. Culture of Animal Cells: A Manual of Basic Technique and Specialized Applications. Wiley-Blackwell, 2010. [Google Scholar]
- Xue, Z.; Zeng, J.; Li, Y.; Meng, B.; Gong, X.; Zhao, Y.; Dai, X. Proteomics reveals that cell density could affect the efficacy of drug treatment. Biochemistry and Biophysics Reports 2023, 33, 101403. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



