1. Introduction
Medical imaging (MI) is crucial in modern healthcare, providing essential information for diagnosing, treating, and monitoring diseases. Traditional image analysis relies on handcrafted features and expert knowledge, which can be time-consuming and prone to errors. Machine learning (ML) methods, such as Support Vector Machines (SVMs), decision trees, random forests, and logistic regression, have improved efficiency and reduced errors in tasks like image segmentation, object detection, and disease classification. These methods require manual feature selection and extraction. Recent advancements in deep learning (DL) have revolutionized medical image analysis by automatically learning and extracting hierarchical features from large volumes of medical image data [
1,
2,
3,
4,
5,
6,
7,
8,
9]. It has provided healthcare professionals with valuable insights for more accurate diagnoses and improved patient care.
Despite their exceptional performance, DL models in MI face challenges related to interpretability and transparency [
10,
11,
12]. Their opaque decision-making processes raise concerns about reliability in healthcare, where explanations for diagnostic decisions are crucial. Interpretability in artificial intelligence (AI)-driven healthcare models encourages trust and reliability by enabling practitioners to understand and verify model outputs. It ensures ethical and legal accountability by clarifying diagnostic and recommendation processes, which is crucial for compliance and ethical standards. Interpretability supports clinical decision-making by providing insights into AI recommendations, leading to better-informed patient care. Additionally, transparent models help identify biases and errors, enhancing fairness and accuracy.
Ongoing efforts to enhance the interpretability of DL models in MI involve developing techniques to clarify how these models reach conclusions [
13,
14,
15]. Researchers are creating explanations and visualizations to make the decision-making processes of these models more understandable and trustworthy for healthcare practitioners. This paper contributes to this area through several key aspects:
Comprehensive Survey: This paper offers a thorough survey of innovative approaches for interpreting and visualizing DL models in MI. It includes a wide array of techniques aimed at enhancing model transparency and trust.
Methodological Review: We provide an in-depth review of current methodologies, focusing on post-hoc visualization techniques such as perturbation-based, gradient-based, decomposition-based, trainable attention (TA)-based methods, and vision transformers (ViT). Each method’s effectiveness and applicability in MI are evaluated.
Clinical Relevance: This paper emphasizes the importance of interpretability techniques in clinical settings, showing how they lead to more reliable and actionable insights from DL models, thus supporting better decision-making in healthcare.
Future Directions: We outline future research directions in model interpretability and visualization, highlighting the need for more robust and scalable techniques that can manage the complexity of DL models while ensuring practical utility in medical applications.
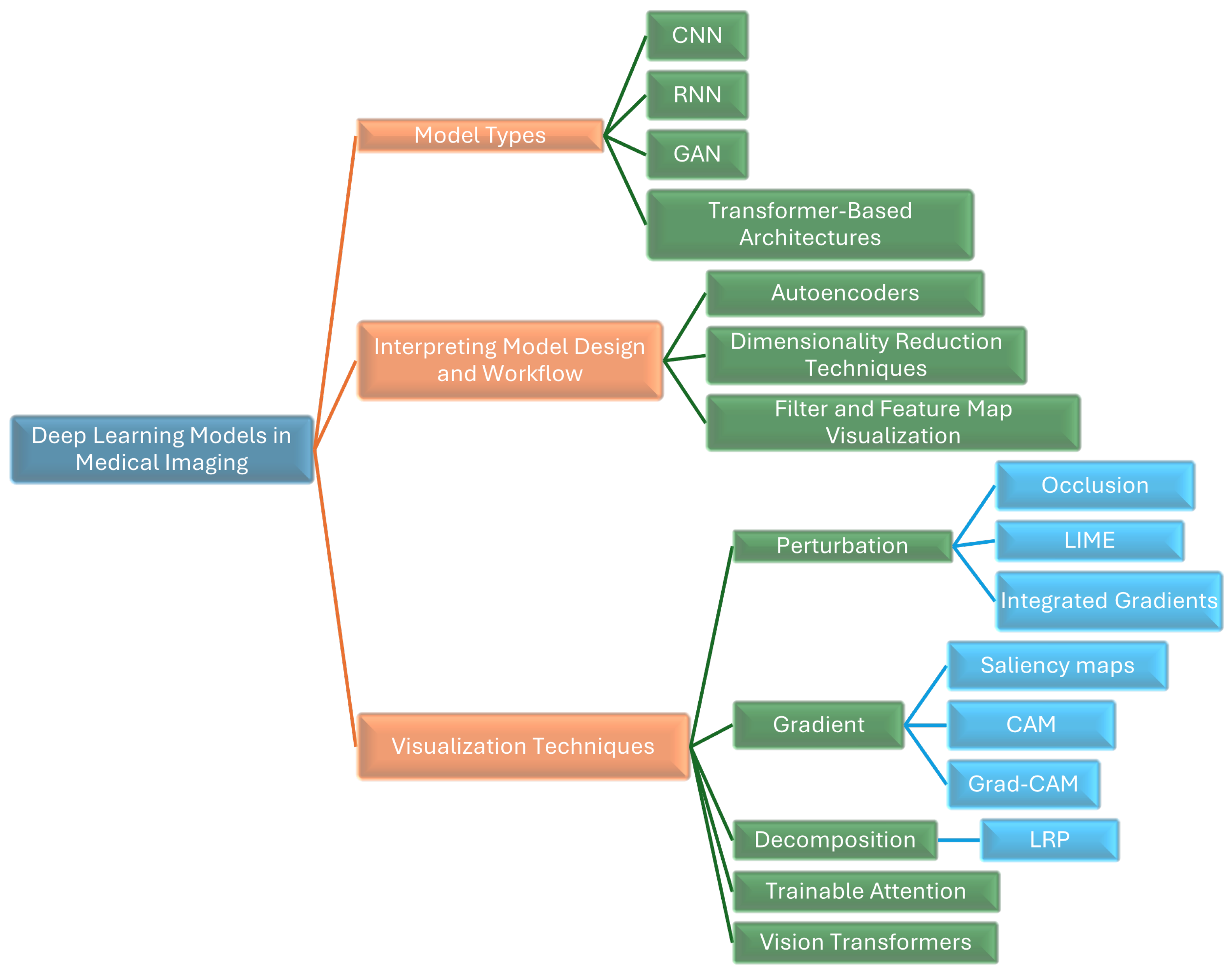
We provide a thorough survey of innovative approaches for interpreting and visualizing DL models in MI. As shown in
Figure 1, we illustrate various DL models and techniques used in MI, highlighting methods like CNNs, RNNs, GANs, transformer-based architectures, autoencoders, LIME, Grad-CAM, and Layer-Wise Relevance Propagation (LRP),attention-based methods, and vision transformers (ViTs).
The rest of this paper is divided into four sections, with multiple subsections within each of them.
Section 3 is focused on interpreting model design and workflow.
Section 4 Visualizing DL models in MI.
Section 5 presents an overview of post-hoc interpretation and Visualization Techniques. Comparison of Different Interpretation Methods discussed in
Section 6 and concludes the work with current challenges and future directions in
Section 8.
2. Research Methodology
A comprehensive review of explainable AI in medical image analysis was published by [
16,
17,
18,
19,
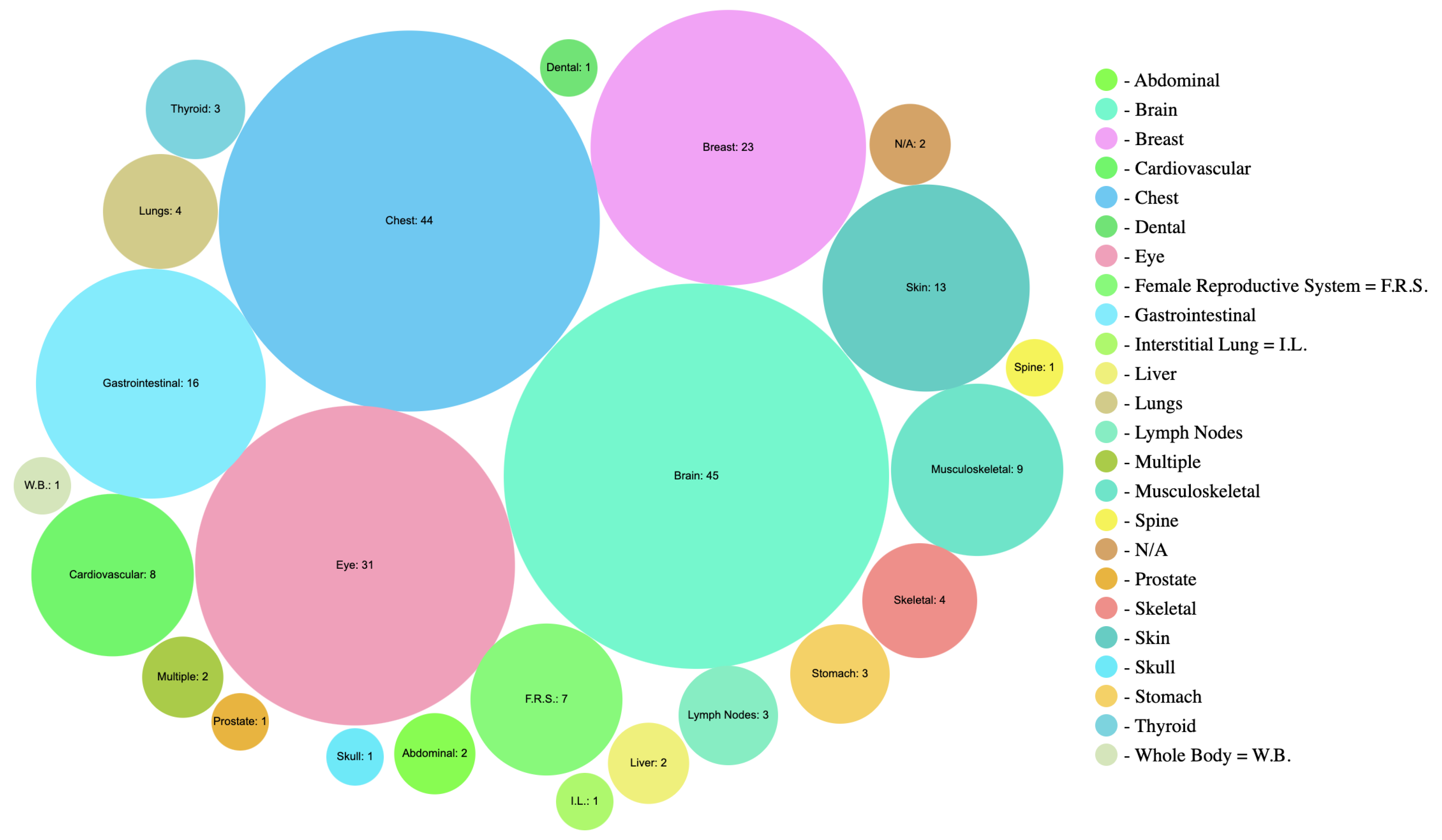
20]. While this review covers a broad range of topics, some critical areas, such as research on trainable attention (TA)-based methods, vision transformers, and their applications, have been overlooked. Our review aims to fill this gap by providing an extensive overview of various domains within medical imaging, addressing key aspects such as Domain, Task, Modality, Performance, and Technique.
This research employs the Systematic Literature Review (SLR) method, which involves several stages. The research questions guiding this study are:
What innovative methods exist for interpreting and visualizing deep learning models in medical imaging?
How effective are post-hoc visualization techniques (perturbation-based, gradient-based, decomposition-based, TA-based, and ViT) in improving model transparency?
What is the clinical relevance of interpretability techniques for actionable insights from deep learning models in healthcare?
What are the future research directions for model interpretability and visualization in medical applications?
The survey examines over 300 recent papers on explainable AI (XAI) in medical image analysis. Relevant contributions were identified using keywords like “deep learning", “convolutional neural networks", “medical imaging", “surveys", “interpretation", “visualization" and “review". Sources included ArXiv, Google Scholar, and Science Direct, focusing on titles. Studies without results on medical image data or using only standard neural networks with manually designed features were excluded. In cases of similar work, the most significant publications were selected.
The findings will be comprehensively presented, including a detailed description of the research methodology for replication. The literature search results, relevant articles, and their quality evaluations will be summarized in overview tables. Drawing from expertise in applying XAI techniques to medical image analysis, ongoing challenges, and future research directions will be discussed.
3. Interpreting Model Design and Workflow
Interpreting model design and workflow involves examining the hidden layers of convolutional neural networks (CNNs). This can be achieved through methods such as:
Autoencoders for Learning Latent Representations
Visualizing High-Dimensional Latent Data in a Two-Dimensional Space
Visualizing Filters and Activations in Feature Maps
3.1. Autoencoders for Learning Latent Representations
Autoencoders (AE) are DL models for unsupervised feature learning [
21], with applications in anomaly detection [
22], image compression [
23], and representation learning [
24]. They consist of an encoder creating latent representations and a decoder reconstructing images. Variants include variational autoencoders (VAE) and adversarial autoencoders (AAE). In medical imaging, AEs detect abnormalities by comparing input images with reconstructions and highlighting high reconstruction loss areas. For instance, VAE has reconstructed OCT retinal images to detect pathologies [
25], and AAE has localized brain lesions in MRI images [
26]. Convolutional AEs have detected nuclei in histopathology images by combining learned representations with thresholding [
27].
3.2. Visualizing High-Dimensional Latent Data in a Two-Dimensional Space
CNNs produce high-dimensional features, making visualization challenging. Dimensionality reduction techniques like PCA and tSNE simplify this data. PCA performs linear transformations, while tSNE [
28] uses nonlinear methods to map high-dimensional data to lower dimensions. tSNE is effective for visualizing patterns and clusters, such as in abdominal ultrasound and histopathology image classification. The constraint-based embedding technique [
29], using a divide-and-conquer algorithm to preserve k-nearest neighbors in 2D projections, has assessed deep belief networks separating brain MRI images of schizophrenic and healthy patients, though both tSNE and constraint-based embedding struggled with raw data separation.
3.3. Visualizing Filters and Activations in Feature Maps
A convolutional block extracts local features from input images through convolution filters, ReLU or GELU activations, and pooling layers. Filter visualization reveals CNN’s feature extraction capabilities, with initial layers capturing basic elements and later layers capturing intricate patterns. In medical imaging, filter visualization compares filters in CADe for 3D CT images [
5]. Larger filters offer more insights but require more memory. Feature map visualization, representing layer outputs after activation, highlights active features and can indicate training issues. It is used in tasks like skin lesion classification [
30], fetal facial plan recognition in ultrasound [
31], brain lesion segmentation in MRI [
2], and Alzheimer’s diagnosis with PET/MRI [
32].
4. Deep Learning Models in Medical Imaging
Convolutional Neural Networks (CNNs) are essential in DL for MI. CNNs adept at processing X-rays, CT scans, and MRIs through their hierarchical feature representations. Studies [
1,
2,
3] have demonstrated their effectiveness in various medical image analysis tasks. For instance, in segmentation tasks, CNNs excel at delineating organ boundaries or identifying anomalies within medical images, providing valuable insights for accurate diagnosis and treatment planning.
Recurrent Neural Networks (RNNs) excel in temporal modeling of dynamic imaging sequences, such as functional MRI or video-based imaging, by capturing temporal patterns [
33].
Generative Adversarial Networks (GANs) are valuable for image synthesis, data augmentation, and anomaly detection, generating synthetic images and learning normal patterns [
34,
35,
36].
4.1. Transformer-Based Architectures
Transformer-based architectures, including bidirectional encoder representations from transformers (BERT) [
37,
38], and generative pre-trained transformer (GPT) [
39], are emerging for tasks like disease prediction, image reconstruction, and capturing complex dependencies in medical images.
5. Interpretation and Visualization Techniques
In recent years, numerous explainable artificial intelligence (XAI) techniques have been developed to enhance the interpretability of DL models, particularly in MI. These techniques can be broadly categorized into Perturbation-Based, Gradient-Based, Decomposition, and Attention methods.
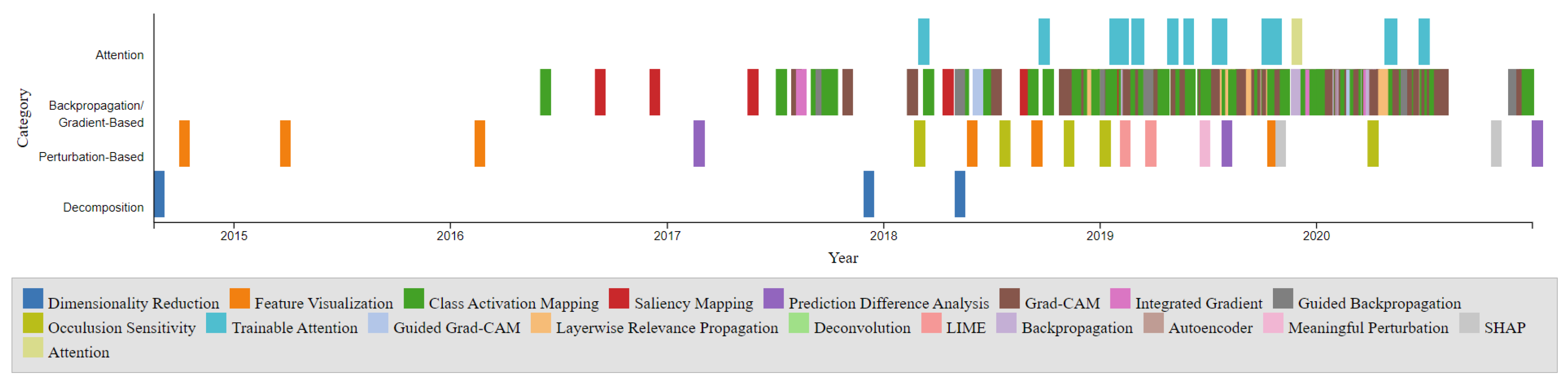
The timeline presented in
Figure 2 illustrates the development of these XAI techniques over the years. The points on the timeline are color-coordinated by respective categories, including Dimensionality Reduction, Feature Visualization, Class Activation Mapping, Saliency Mapping, Prediction Difference Analysis, Grad-CAM, Integrated Gradient, Guided Backpropagation, Occlusion Sensitivity, Trainable Attention, Guided Grad-CAM, Layerwise Relevance Propagation, Deconvolution, LIME, Backpropagation, Autoencoder, Meaningful Perturbation, SHAP, and Attention. Notably, the Gradient-Based category is the most densely populated, with CAM and Grad-CAM being among the most popular entries. The timeline also reveals a higher density of developments between 2017 and 2020. The comparison chart illustrates the visualization Techniques methods: Gradient-Based, Perturbation-Based, Decomposition-Based (LRP), and Trainable Attention Models in terms of model dependency, access to model parameters, and computational efficiency as shown in
Table 1.
5.1. Perturbation-Based Methods
Perturbation-based methods evaluate how input changes affect model outputs to determine feature importance. By altering specific image regions, these methods identify areas that significantly influence predictions, typically visualized with heatmaps. Techniques like Integrated Gradients (IG), Local Interpretable Model-agnostic Explanations (LIME), and Occlusion Sensitivity (OS) are used in various domains, including breast cancer detection, eye disease classification, and brain MRI analysis (see
Table 2). The table categorizes studies by domain, task, modality, performance, and technique. These methods also test model sensitivity to input variations, ensuring robust interpretations.
5.1.1. Occlusion
Zeiler and Fergus [
40] introduced an occlusion method to assess the impact on model output when parts of an image are obstructed. Kermany et al. [
41] utilized this method for interpreting optical coherence tomography images to diagnose retinal pathologies. A major limitation of occlusion is its high computational demand, as it requires inference for each occluded image region, increasing with image resolution and desired heatmaps.
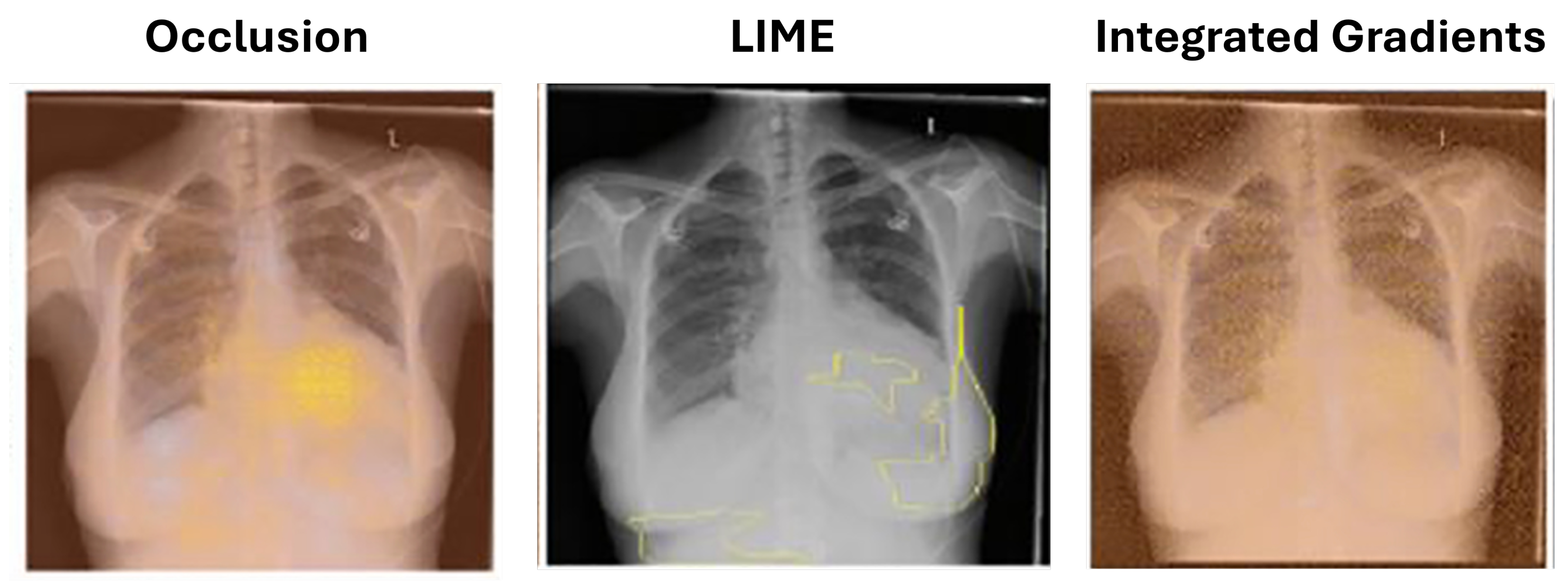
5.1.2. Local Interpretable Model-Agnostic Explanations (LIME)
Ribeiro et al. [
42] introduced local interpretable model-agnostic explanations (LIME) to identify superpixels (groups of connected pixels with similar intensities). Seah et al. [
43] applied LIME to identify congestive heart failure in chest radiographs. LIME offers an advantage over occlusion by preserving the altered image portions’ context, as they are not completely blocked as shown in
Figure 3.
5.1.3. Integrated Gradients
Sundararajan et al. [
44] introduced integrated gradients (IG) to measure pixel importance by computing gradients across images interpolated between the original and a baseline image with all non-values. Sayres et al. [
45] found that model-predicted grades and heatmaps improved the accuracy of diabetic retinopathy grading by readers.
Table 2.
Overview of Various Studies Using Perturbation-Based Methods in Medical Imaging
Table 2.
Overview of Various Studies Using Perturbation-Based Methods in Medical Imaging
| Domain |
Task |
Modality |
Performance |
Technique |
Citation |
| Breast |
Classification |
MRI |
N/A |
IG |
[46] |
| Eye |
Classification |
DR |
Accuracy: 95.5% |
IG |
[45] |
| Multiple |
Classification |
DR |
N/A |
IG |
[44] |
| Chest |
Detection |
X-ray |
Accuracy: 94.9%, AUC: 97.4% |
LIME |
[47] |
| Gastrointestinal |
Classification |
Endoscopy |
Accuracy: 97.9% |
LIME |
[48] |
| Brain |
Segmentation, Detection |
MRI |
ICC: 93.0% |
OS |
[49] |
| Brain |
Classification |
MRI |
Accuracy: 85.0% |
OS |
[50] |
| Breast |
Detection, Classification |
Histology |
Accuracy: 55.0% |
OS |
[51] |
| Eye, Chest |
Classification, Detection |
OCT, X-ray |
Eye Accuracy: 94.7%, Chest Accuracy: 92.8% |
OS |
[41] |
| Chest |
Classification |
X-ray |
AUC: 82.0% |
OS, IG, LIME |
[43] |
5.2. Gradient-Based Methods
Backpropagation, used for weight adjustment in neural network training, is also employed in model interpretation methods to compute gradients. Unlike training, these methods do not alter weights but use gradients to highlight important image areas.
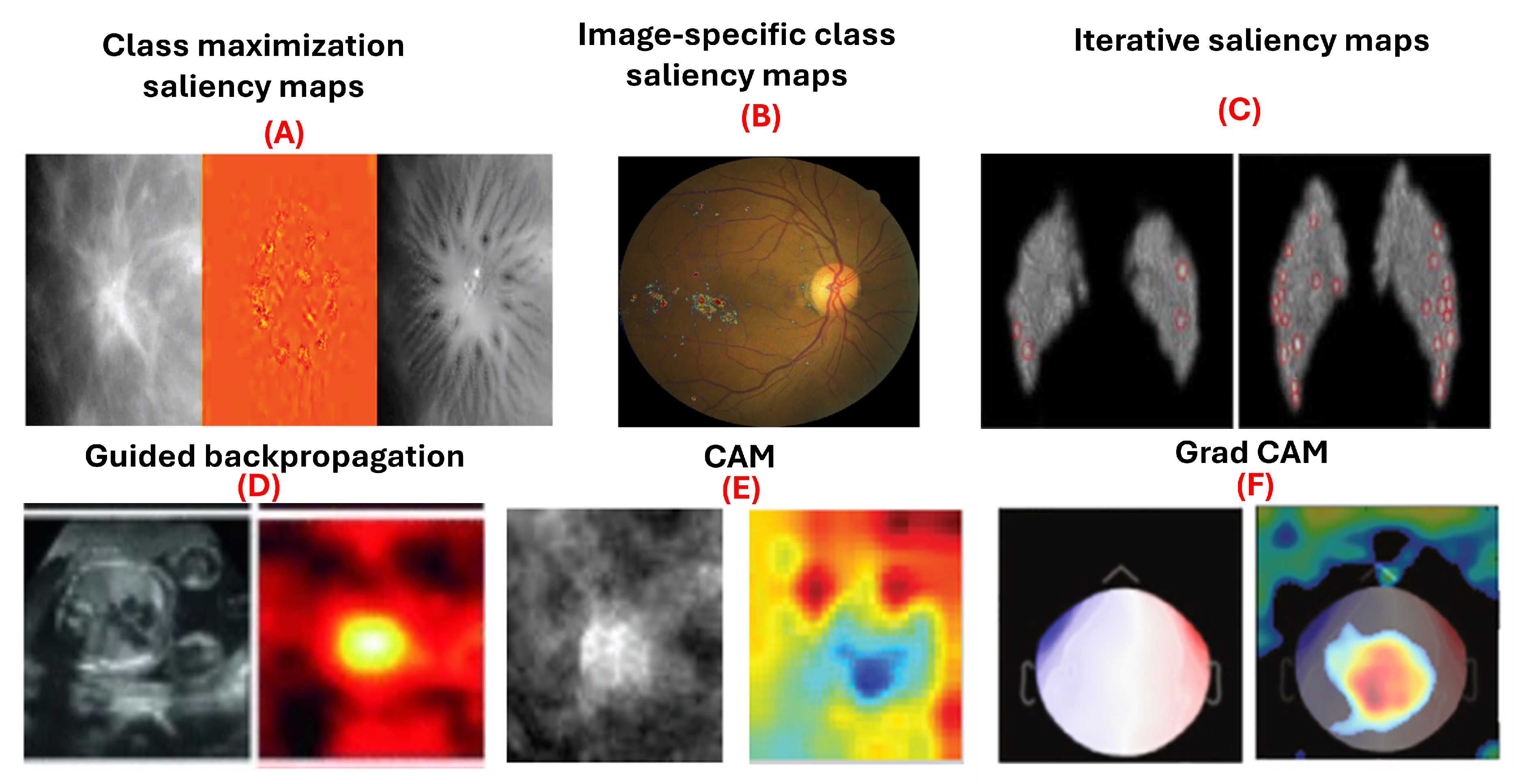
Figure 4.
Examples of gradient-based attribution methods for model interpretability. (A) Class maximization visualization of malignant and benign breast masses on mammograms [
9]. (B) Integrated gradients visualizing evidence of diabetic retinopathy on retinal fundus images [
45]. (C) Visualization of malignant and benign breast masses [
9]. (D) Guided backpropagation applied to ultrasound images for fetal heartbeat localization [
4]. (E) Differentiation between benign and malignant breast masses in mammograms [
6]. (F) Grad-CAM visualizations identifying discriminative regions in magnetoencephalography images for detecting eye-blink artifacts [
52].
Figure 4.
Examples of gradient-based attribution methods for model interpretability. (A) Class maximization visualization of malignant and benign breast masses on mammograms [
9]. (B) Integrated gradients visualizing evidence of diabetic retinopathy on retinal fundus images [
45]. (C) Visualization of malignant and benign breast masses [
9]. (D) Guided backpropagation applied to ultrasound images for fetal heartbeat localization [
4]. (E) Differentiation between benign and malignant breast masses in mammograms [
6]. (F) Grad-CAM visualizations identifying discriminative regions in magnetoencephalography images for detecting eye-blink artifacts [
52].
5.2.1. Saliency Maps
Saliency maps, introduced by Simonyan et al. [
53], use gradient information to explain how deep convolutional networks classify images. They are used in class maximization and image-specific class saliency maps. Class maximization generates an image maximizing activation for a class, as in:
Yi et al. [
9] applied this to visualize malignant and benign breast masses. Image-specific class saliency maps create heatmaps showing each pixel’s significance in classification, computed as:
Dubost et al. [
54] used these maps in a weakly supervised method for segmenting brain MRI structures. Saliency maps have also been utilized in diagnosing heart diseases in chest x-rays [
8], classifying breast masses in mammography [
55] with accuracy ranging from 85% to 92.9%, and identifying pediatric elbow fractures in x-rays [
56] with accuracy of 88.0% and area under curve (AUC) of 95.0%. Moreover, iterative saliency maps [
57] enhance less obvious image regions by generating a saliency map, inpainting prominent areas, and iterating the process until the image classification changes or a limit is reached. This approach, applied to retinal fundus images for diabetic retinopathy grading, demonstrated higher sensitivity compared to traditional saliency maps. However, saliency maps have limitations. They do not distinguish if a pixel supports or contradicts a class, and their effectiveness diminishes in binary classification.s.
5.2.2. Guided Backpropagation
Guided backpropagation, introduced by Springenberg et al. [
58], builds on the saliency map approach by Simonyan et al. [
53] and the deconvnet concept by Zeiler and Fergus [
40]. It improves gradient backpropagation through ReLU layers, where negative activations are set to zero during the forward pass. Guided backpropagation discards gradients where either the forward activation or the backward gradient is negative, producing heatmaps that highlight pixels positively contributing to the classification.
In 2017, Gao and Noble [
4] applied guided backpropagation to ultrasound images for fetal heartbeat localization. They found that the heatmaps remained consistent despite variations in the heartâs appearance, size, position, and contrast. Conversely, Böhle et al. [
59] discovered that guided backpropagation was less effective for visualizing Alzheimerâs disease in brain MRIs compared to other methods. Similarly, Dubost et al. [
60] achieved an intraclass correlation coefficient (ICC) of 93.0% in brain MRI detection using guided backpropagation. Wang et al. [
61] obtained an average accuracy of 93.7% in brain MRI classification with this technique. Gessert et al. [
62] reported an accuracy of 99.0% in cardiovascular classification using OCT images. Wickstrøm et al. [
63] achieved a 94.9% accuracy in gastrointestinal segmentation using endoscopy. Lastly, Jamaludin et al. [
64] reported an accuracy of 82.5% in musculoskeletal spine classification using MRI images with guided backpropagation.
5.2.3. Class Activation Maps (CAM)
Class Activation Mapping (CAM), introduced by Zhou et al. [
65], visualizes regions of an image most influential in a neural network’s classification decision. CAM is computed as a weighted sum of feature maps from the final convolutional layer, using weights from the fully connected layer following global average pooling [
66]. For a specific class
c and image
x:
This heatmap highlights regions most relevant for classification. CAM has been applied in various medical imaging applications, such as segmenting lung nodules in thoracic CT scans [
67] and differentiating between benign and malignant breast masses in mammograms [
6]. However, CAM’s effectiveness depends on the network architecture, requiring a global pooling (GAP) layer followed by a fully connected layer. While Zhou et al. [
68] originally used GAP, Oquab et al. [
69] demonstrated that global max pooling and log-sum-exponential pooling can also be used, with the latter yielding finer localization.
Table 3.
Performance metrics of various Medical Imaging tasks across different modalities using CAM
Table 3.
Performance metrics of various Medical Imaging tasks across different modalities using CAM
| Domain-Task |
Modality |
Performance |
Citation |
| Bladder Classification |
Histology |
Mean Accuracy: 69.9% |
[70] |
| Brain Classification |
MRI |
Accuracy: 86.7% |
[71,72] |
| Brain Detection |
MRI,PET,CT |
Accuracy: 90.2% - 95.3%, F1: 91.6% - 94.3% |
[73,74] |
| Breast Classification |
X-ray,Ultrasound, MRI |
Accuracy: 83.0% -89.0% |
[75,76,77,78] |
| Breast Detection |
X-ray, Ultrasound |
Mean AUC: 81.0%, AUC: Mt-Net 98.0%, Sn-Net 92.8%, Accuracy: 92.5% |
[79,80,81,82] |
| Chest Classification |
X-ray,CT |
Accuracy: 97.8% , Average AUC: 75.5%- 96.0% |
[83,84,85,86,87,88,89,90] |
| Chest Segmentation |
X-ray |
Accuracy: 95.8% |
[91] |
| Eye Classification |
Fundus Photography,OCT,CT |
F1-score: 95.0% , Precision: 93.0%, AUC: 88.0%- 99.0% |
[92,93,94,95,96] |
| Eye Detection |
Fundus Photography |
Accuracy: 73.2% -99.1%, AUC: 99.0% |
[97,98,99,100,101] |
| GI Classification |
Endoscopy |
Mean Accuracy: 93.2% |
[102,103,104,105] |
| Liver Classification, Segmentation |
Histology |
Mean Accuracy: 87.5% |
[106,107] |
| Musculoskeletal Classification |
MRI, X-ray |
Accuracy: 86.0%, AUC: 85.3% |
[108,109] |
| Skin Classification, Segmentation |
Dermatoscopy |
Accuracy: 83.6%, F1-score: 82.7% |
[110,111] |
| Skull Classification |
X-ray |
AUC: 88.0%- 93.0% |
[112] |
| Thyroid Classification |
Ultrasound |
Accuracy: 87.3% ± 0.0007, AUC: 90.1% ± 0.0007 |
[113] |
| Lymph Node Classification, Detection |
Histology |
Accuracy: 91.9%, AUC: 97.0% |
[114] |
| Various Classification |
CT, MRI, Ultrasound, X-ray, Fundoscopy |
F1-score: 98.0%, Model Accuracy: 98.0% |
[115,116] |
5.2.4. Grad-CAM
Grad-CAM, an extension of CAM by Selvaraju et al. [
117], broadens its application to any network architecture and output, including image segmentation and captioning. It bypasses the global pooling layer and weights feature maps directly with gradients calculated via backpropagation from a target class. The gradients of the output for class
c concerning feature maps
are averaged globally, multiplied by
, and passed through a ReLU activation to discard negative values:
Garg et al. employed grad-CAM visualizations to identify discriminative regions of magnetoencephalography images in the task of detecting eye-blink artifacts [
52]. The authors found that the regions of the eye highlighted by grad-CAM are the same regions that human experts rely on.
Table 4 summarizes the effectiveness of Grad-CAM across various medical imaging tasks, highlighting domains, tasks, modalities, and performance metrics.
5.3. Decomposition-Based Methods
Decomposition-based techniques for model interpretation focus on breaking down a modelâs prediction into a heatmap showing each pixelâs contribution to the final decision. These techniques, such as LRP, have been widely applied across different domains.
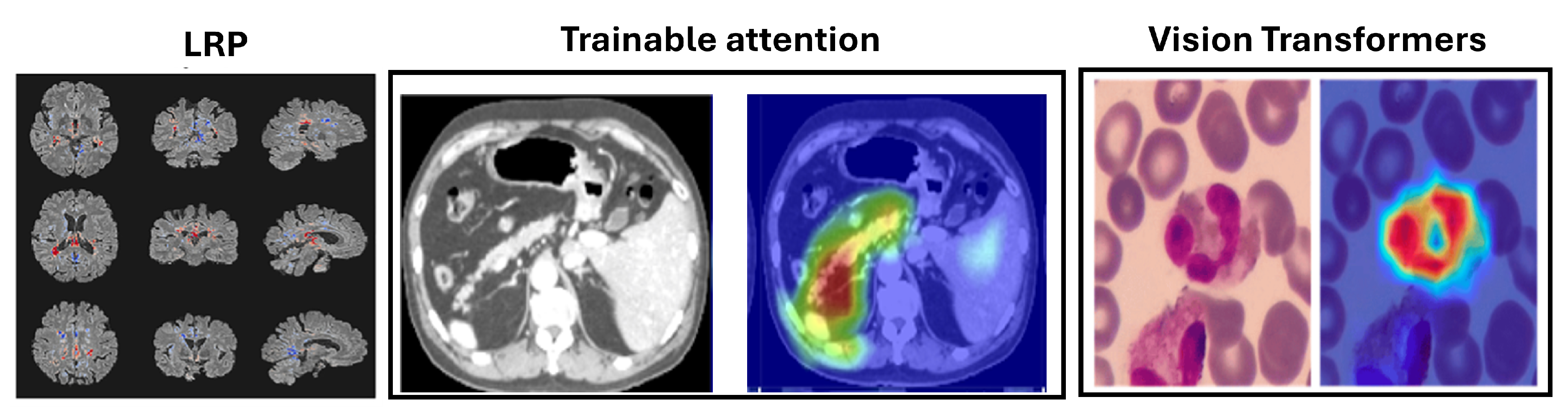
Figure 5.
Example uses of decomposition-based attribution methods for model interpretability to show the Layerwise relevance propagation for model interpretation in diagnosing multiple sclerosis on brain MRI (Eitel et al., 2019)[
160]. The Attention U-Net for organ segmentation in abdominal CT scans [
161] and the areas upon which the model correctly focuses its predictions on the test images in explainable Vision Transformers model [
162].
Figure 5.
Example uses of decomposition-based attribution methods for model interpretability to show the Layerwise relevance propagation for model interpretation in diagnosing multiple sclerosis on brain MRI (Eitel et al., 2019)[
160]. The Attention U-Net for organ segmentation in abdominal CT scans [
161] and the areas upon which the model correctly focuses its predictions on the test images in explainable Vision Transformers model [
162].
5.3.1. Layer-Wise Relevance Propagation (LRP)
Layer-Wise Relevance Propagation (LRP), introduced by Bach et al. in 2015 [
163], offers an alternative to gradient-based techniques like saliency mapping, guided backpropagation, and Grad-CAM. Instead of relying on gradients, LRP distributes the output of the final layer back through the network to calculate relevance scores for each neuron. This process is repeated recursively from the final layer to the input layer, generating a relevancy heatmap that can be overlaid on the input image. Further properties of LRP and details of its theoretical basis are given in (Montavon et al., 2017 [
164]), and a comparison of LRP to other interpretation methods can be found in ([
165,
166,
167]).
The relevance score
for a neuron
i in layer
l from a neuron
k in layer
is defined as:
The overall relevance score for neuron
i in layer
l is:
LRP has been applied in MI, such as diagnosing multiple sclerosis (MS) and Alzheimerâs disease (AD) using MRI scans. For MS, LRP heatmaps highlighted hyperintense lesions and affected brain areas [
160] as shown in
Figure 5, while for AD, they emphasized the hippocampal volume, a critical region for diagnosis [
168]. LRP has been found to provide clearer distinctions compared to gradient-based methods and has been used in frameworks like DeepLight for linking brain regions with cognitive states [
169].
5.4. Trainable Attention Models
Trainable Attention (TA) Mechanisms provide a dynamic approach to model interpretation by integrating attention modules into neural networks. Introduced for CNNs by Jetley et al. [
170], these soft attention modules generate attention maps that highlight important image parts. They compute compatibility scores between local and global features
using dot products and learned vectors
a:
The output
adjusts local features based on the attention map weights:
This method enhances signals from compatible features while reducing those from less compatible ones. Applications of attention mechanisms in medical imaging include the Attention U-Net for organ segmentation in abdominal CT scans [
161], fetal ultrasound classification, and breast mass segmentation in mammograms [
171]. Additionally, they have improved melanoma lesion classification [
172] and osteoarthritis grading in knee X-rays [
173]. Attention mechanisms are valued for their interpretability and performance enhancement, though optimal configurations are application-specific.
Table 5 provides an overview of various studies using TA models, highlighting the domains, tasks, modalities including MRI and histology, and performance metrics such as accuracy and F1-score, demonstrating the broad applicability and effectiveness of TA models in MI.
5.5. Vision Transformers
Vision Transformers (ViTs) have emerged as a prominent alternative to convolutional neural networks (CNNs) in medical imaging. Unlike CNNs, which use local receptive fields to capture spatial hierarchies, ViTs employ self-attention to model long-range dependencies and global context [
38,
184,
185]. By partitioning images into fixed-size patches treated as a sequence, ViTs utilize transformer encoder layers, effectively capturing complex anatomical structures and pathological patterns.
ViTs have shown superior performance in segmentation, classification, and detection tasks, achieving high accuracy in segmenting tumors and organs in MRI and CT scans, as reflected in Dice scores. Their interpretability is enhanced through attention maps, gradient-based methods, and occlusion sensitivity, which aid in visualizing model predictions. These advancements highlight ViTs’ potential to improve diagnostic accuracy and provide deeper insights into medical image analysis, as discussed in
Table 6. The areas upon which the model correctly focuses its predictions on the test image are presented in
Figure 5. The regions of focus identified by the ViT model exhibit significant overlap with the areas of White blood cells [
162].
6. Comparison of Different Interpretation Methods
6.1. Categorization by Visualization Technique
Visualization techniques in DL can be categorized based on their application and effectiveness.
Table 7 summarizes various visualization techniques used in DL for interpretability. It categorizes methods based on their tasks, body parts, modalities, accuracy, and evaluation metrics. This table highlights that techniques like CAM and Grad-CAM are effective for image classification and localization across different modalities such as X-ray and MRI, achieving high accuracy. LRP is noted for its accuracy in segmentation tasks, while IG is utilized for classification with notable AUC-Receiver Operating Characteristic (ROC) scores. Attention-based methods improve performance and interpretability by focusing on relevant regions, whereas perturbation-based methods assess model robustness. LIME provides model-agnostic explanations, and trainable attention models dynamically enhance feature focus.
6.2. Categorization by Body Parts, Modality, and Accuracy
This table provides a concise overview of the imaging techniques, their applications, accuracy, and specific considerations for different anatomical contexts.
Table 7 provides a concise overview of imaging techniques categorized by anatomical context (Body Parts). It lists various modalities such as MRI, X-ray, and ultrasound, and highlights specific techniques used for different body parts. For instance, CAM and Grad-CAM are prominent in brain imaging with high accuracy, while LRP and attention-based methods excel in breast imaging. The table also emphasizes the adaptation of methods to address challenges such as speckle noise in ultrasound imaging. The imaging techniques used such as CT, dermatoscopy, diabetic retinopathy (DR), endoscopy, fundus photography, histology, histopath, mammo, magnetoencephalography (MEG), MRI, OCT, PET, photography, ultrasound, and X-ray. Studies ranged from the years 2014 to 2020 with a majority coming from 2019 and 2020 as shown in
Figure 6.
6.3. Categorization by Task
This section organizes the techniques and their applications across different tasks, highlighting performance metrics and examples for clarity.
Table 8 organizes interpretability techniques according to their tasks, including classification, segmentation, and detection. It details the applications, performance metrics, and specific examples for each task. Techniques like CAM, Grad-CAM, and TA models are effective for classification tasks, providing high accuracy and AUC-ROC scores. LRP and Integrated Gradient are highlighted for segmentation tasks, with metrics like dice similarity coefficient (DSC) and intersection over union (IoU). Detection tasks benefit from methods such as saliency maps and CAM, with metrics including mean average precision (mAP) and sensitivity.
7. Current Challenges and Future Directions
7.1. Current Challenges
Despite significant advancements, several challenges remain in the interpretability and visualization of DL models in MI:
Scalability and Efficiency: Many interpretability methods, such as occlusion and perturbation-based techniques, are computationally intensive. This limits their scalability, especially with high-resolution medical images that require real-time analysis.
Clinical Integration: Translating interpretability techniques into clinical practice requires seamless integration with existing workflows and systems. This includes ensuring that the visualizations are intuitive for non-technical healthcare practitioners and that they provide actionable insights.
Robustness and Generalization: Interpretability methods must be robust across diverse patient populations and medical imaging modalities. Models trained on specific datasets might not generalize well to other contexts, leading to potential biases and inaccuracies in interpretations.
Standardization and Validation: There is a lack of standardized metrics and benchmarks for evaluating the effectiveness of interpretability methods. Rigorous validation in clinical settings is essential to establish the reliability and trustworthiness of these techniques.
Ethical and Legal Considerations: The opacity of deep learning models raises ethical and legal concerns, especially in healthcare where decisions can have critical consequences. Ensuring transparency, accountability, and fairness in AI-driven diagnostics is paramount.
7.2. Future Directions
To address these challenges, future research should focus on the following directions:
Development of Lightweight Methods: Creating computationally efficient interpretability techniques that can handle high-resolution images and deliver results in real-time is crucial. This includes optimizing existing methods and exploring new algorithmic approaches.
Enhanced Clinical Collaboration: Collaborative efforts between AI researchers, clinicians, and medical practitioners are needed to design interpretability methods that are clinically relevant and user-friendly. This could involve interactive visualization tools that allow clinicians to explore model outputs intuitively.
Robustness to Variability: Developing interpretability techniques that are robust to variations in imaging modalities, patient demographics, and clinical conditions is essential. This requires extensive training on diverse datasets and continuous validation across different settings.
Establishment of Standards: Creating standardized benchmarks and validation protocols for interpretability methods will help in objectively assessing their effectiveness and reliability. This includes developing common datasets and metrics for comparative evaluations.
Ethical Frameworks: Integrating ethical considerations into the design and deployment of interpretability methods is critical. This involves ensuring that models are transparent, explainable, and free from biases, as well as addressing privacy and data security concerns.
Hybrid Approaches: Combining different interpretability techniques, such as perturbation-based and gradient-based methods, can provide more comprehensive insights into model behavior. Hybrid approaches can leverage the strengths of various methods to enhance overall interpretability.
8. Conclusion
In conclusion, the integration of interpretability and visualization techniques into DL models for MI holds immense potential for advancing healthcare diagnostics and treatment planning. While significant progress has been made, challenges related to scalability, clinical integration, robustness, standardization, and ethical considerations persist. Addressing these challenges requires ongoing collaboration between AI researchers, clinicians, and healthcare practitioners. Future research should focus on developing efficient and clinically relevant interpretability methods, establishing standardized evaluation protocols, and ensuring ethical and transparent AI applications in healthcare. By overcoming these hurdles, we can enhance the trustworthiness, reliability, and clinical impact of DL models in MI, ultimately leading to better patient outcomes and more informed clinical decision-making.
References
- Hu, P.; Wu, F.; Peng, J.; Bao, Y.; Chen, F.; Kong, D. Automatic abdominal multi-organ segmentation using deep convolutional neural network and time-implicit level sets. International journal of computer assisted radiology and surgery 2017, 12, 399–411. [Google Scholar] [CrossRef] [PubMed]
- Kamnitsas, K.; Ledig, C.; Newcombe, V.F.; Simpson, J.P.; Kane, A.D.; Menon, D.K.; Rueckert, D.; Glocker, B. Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Medical image analysis 2017, 36, 61–78. [Google Scholar] [CrossRef]
- Roth, H.R.; Lu, L.; Farag, A.; Shin, H.C.; Liu, J.; Turkbey, E.B.; Summers, R.M. Deeporgan: Multi-level deep convolutional networks for automated pancreas segmentation. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference, Munich, Germany, 2015, Proceedings, Part I 18. Springer, 2015, October 5-9; pp. 556–564.
- Gao, Y.; Alison Noble, J. Detection and characterization of the fetal heartbeat in free-hand ultrasound sweeps with weakly-supervised two-streams convolutional networks. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention- MICCAI 2017: 20th International Conference, Quebec City, QC, Canada, 2017, Proceedings, Part II 20. Springer, 2017, September 11-13; pp. 305–313.
- Roth, H.R.; Lu, L.; Liu, J.; Yao, J.; Seff, A.; Cherry, K.; Kim, L.; Summers, R.M. Improving computer-aided detection using convolutional neural networks and random view aggregation. IEEE transactions on medical imaging 2015, 35, 1170–1181. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.T.; Lee, J.H.; Lee, H.; Ro, Y.M. Visually interpretable deep network for diagnosis of breast masses on mammograms. Physics in Medicine & Biology 2018, 63, 235025. [Google Scholar]
- Yang, X.; Do Yang, J.; Hwang, H.P.; Yu, H.C.; Ahn, S.; Kim, B.W.; You, H. Segmentation of liver and vessels from CT images and classification of liver segments for preoperative liver surgical planning in living donor liver transplantation. Computer methods and programs in biomedicine 2018, 158, 41–52. [Google Scholar] [CrossRef]
- Chen, X.; Shi, B. Deep mask for x-ray based heart disease classification. arXiv preprint, 2018. Available online: https://arxiv.org/abs/1808.08277.
- Yi, D.; Sawyer, R.L.; Cohn III, D.; Dunnmon, J.; Lam, C.; Xiao, X.; Rubin, D. Optimizing and visualizing deep learning for benign/malignant classification in breast tumors. arXiv preprint arXiv:1705.06362, arXiv:1705.06362 2017.
- Hengstler, M.; Enkel, E.; Duelli, S. Applied artificial intelligence and trustâThe case of autonomous vehicles and medical assistance devices. Technological Forecasting and Social Change 2016, 105, 105–120. [Google Scholar] [CrossRef]
- Nundy, S.; Montgomery, T.; Wachter, R.M. Promoting trust between patients and physicians in the era of artificial intelligence. Jama 2019, 322, 497–498. [Google Scholar] [CrossRef] [PubMed]
- Hosny, A.; Parmar, C.; Quackenbush, J.; Schwartz, L.H.; Aerts, H.J. Artificial intelligence in radiology. Nature Reviews Cancer 2018, 18, 500–510. [Google Scholar] [CrossRef] [PubMed]
- Jia, X.; Ren, L.; Cai, J. Clinical implementation of AI technologies will require interpretable AI models. Medical physics 2020, 1–4. [Google Scholar] [CrossRef]
- Reyes, M.; Meier, R.; Pereira, S.; Silva, C.A.; Dahlweid, F.M.; Tengg-Kobligk, H.v.; Summers, R.M.; Wiest, R. On the interpretability of artificial intelligence in radiology: challenges and opportunities. Radiology: artificial intelligence 2020, 2, e190043. [Google Scholar] [CrossRef]
- Gastounioti, A.; Kontos, D. Is it time to get rid of black boxes and cultivate trust in AI? Radiology: Artificial Intelligence 2020, 2, e200088. [Google Scholar] [CrossRef]
- Guo, R.; Wei, J.; Sun, L.; Yu, B.; Chang, G.; Liu, D.; Zhang, S.; Yao, Z.; Xu, M.; Bu, L. A survey on advancements in image-text multimodal models: From general techniques to biomedical implementations. Computers in Biology and Medicine, 2024; 108709. [Google Scholar]
- Rasool, N.; Bhat, J.I. Brain tumour detection using machine and deep learning: a systematic review. Multimedia Tools and Applications 2024, 1–54. [Google Scholar] [CrossRef]
- Huff, D.T.; Weisman, A.J.; Jeraj, R. Interpretation and visualization techniques for deep learning models in medical imaging. Physics in Medicine & Biology 2021, 66, 04TR01. [Google Scholar]
- Hohman, F.; Kahng, M.; Pienta, R.; Chau, D.H. Visual analytics in deep learning: An interrogative survey for the next frontiers. IEEE transactions on visualization and computer graphics 2018, 25, 2674–2693. [Google Scholar] [CrossRef] [PubMed]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; Van Der Laak, J.A.; Van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Medical image analysis 2017, 42, 60–88. [Google Scholar] [CrossRef] [PubMed]
- Vincent, P.; Larochelle, H.; Bengio, Y.; Manzagol, P.A. Extracting and composing robust features with denoising autoencoders. In Proceedings of the Proceedings of the 25th international conference on Machine learning, 2008, pp.; pp. 1096–1103.
- Kiran, B.R.; Thomas, D.M.; Parakkal, R. An overview of deep learning based methods for unsupervised and semi-supervised anomaly detection in videos. Journal of Imaging 2018, 4, 36. [Google Scholar] [CrossRef]
- Theis, L.; Shi, W.; Cunningham, A.; Huszár, F. Lossy image compression with compressive autoencoders. In Proceedings of the International conference on learning representations; 2022. [Google Scholar]
- Tschannen, M.; Bachem, O.; Lucic, M. Recent advances in autoencoder-based representation learning. arXiv preprint arXiv:1812.05069, arXiv:1812.05069 2018.
- Uzunova, H.; Ehrhardt, J.; Kepp, T.; Handels, H. Interpretable explanations of black box classifiers applied on medical images by meaningful perturbations using variational autoencoders. In Proceedings of the Medical Imaging 2019: Image Processing. SPIE, Vol. 10949; 2019; pp. 264–271. [Google Scholar]
- Chen, X.; You, S.; Tezcan, K.C.; Konukoglu, E. Unsupervised lesion detection via image restoration with a normative prior. Medical image analysis 2020, 64, 101713. [Google Scholar] [CrossRef] [PubMed]
- Hou, L.; Nguyen, V.; Kanevsky, A.B.; Samaras, D.; Kurc, T.M.; Zhao, T.; Gupta, R.R.; Gao, Y.; Chen, W.; Foran, D.; et al. Sparse autoencoder for unsupervised nucleus detection and representation in histopathology images. Pattern recognition 2019, 86, 188–200. [Google Scholar] [CrossRef] [PubMed]
- Van der Maaten, L.; Hinton, G. Visualizing data using t-SNE. Journal of machine learning research 2008, 9. [Google Scholar]
- Plis, S.M.; Hjelm, D.R.; Salakhutdinov, R.; Allen, E.A.; Bockholt, H.J.; Long, J.D.; Johnson, H.J.; Paulsen, J.S.; Turner, J.A.; Calhoun, V.D. Deep learning for neuroimaging: a validation study. Frontiers in neuroscience 2014, 8, 229. [Google Scholar] [CrossRef]
- Stoyanov, D.; Taylor, Z.; Kia, S.M.; Oguz, I.; Reyes, M.; Martel, A.; Maier-Hein, L.; Marquand, A.F.; Duchesnay, E.; Löfstedt, T.; et al. Understanding and interpreting machine learning in medical image computing applications; Springer, 2018.
- Yu, Z.; Tan, E.L.; Ni, D.; Qin, J.; Chen, S.; Li, S.; Lei, B.; Wang, T. A deep convolutional neural network-based framework for automatic fetal facial standard plane recognition. IEEE journal of biomedical and health informatics 2017, 22, 874–885. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Li, Z.; Zhang, B.; Du, H.; Wang, B.; Zhang, X. Multi-modal deep learning model for auxiliary diagnosis of Alzheimerâs disease. Neurocomputing 2019, 361, 185–195. [Google Scholar] [CrossRef]
- AlâAref, S.J.; Anchouche, K.; Singh, G.; Slomka, P.J.; Kolli, K.K.; Kumar, A.; Pandey, M.; Maliakal, G.; Van Rosendael, A.R.; Beecy, A.N.; et al. Clinical applications of machine learning in cardiovascular disease and its relevance to cardiac imaging. European heart journal 2019, 40, 1975–1986. [Google Scholar] [CrossRef] [PubMed]
- Nie, D.; Trullo, R.; Lian, J.; Petitjean, C.; Ruan, S.; Wang, Q.; Shen, D. Medical image synthesis with context-aware generative adversarial networks. In Proceedings of the Medical Image Computing and Computer Assisted Intervention- MICCAI 2017: 20th International Conference, Quebec City, QC, Canada, 2017, Proceedings, Part III 20. Springer, 2017, September 11-13; pp. 417–425.
- Frid-Adar, M.; Diamant, I.; Klang, E.; Amitai, M.; Goldberger, J.; Greenspan, H. GAN-based synthetic medical image augmentation for increased CNN performance in liver lesion classification. Neurocomputing 2018, 321, 321–331. [Google Scholar] [CrossRef]
- Yi, X.; Walia, E.; Babyn, P. Generative adversarial network in medical imaging: A review. Medical image analysis 2019, 58, 101552. [Google Scholar] [CrossRef] [PubMed]
- Devlin, J.; Chang, M.W.; Lee, K.; Toutanova, K. Bert: Pre-training of deep bidirectional transformers for language understanding. arXiv preprint arXiv:1810.04805, arXiv:1810.04805 2018.
- Al-Hammuri, K.; Gebali, F.; Kanan, A.; Chelvan, I.T. Vision transformer architecture and applications in digital health: a tutorial and survey. Visual computing for industry, biomedicine, and art 2023, 6, 14. [Google Scholar] [CrossRef] [PubMed]
- Lecler, A.; Duron, L.; Soyer, P. Revolutionizing radiology with GPT-based models: current applications, future possibilities and limitations of ChatGPT. Diagnostic and Interventional Imaging 2023, 104, 269–274. [Google Scholar] [CrossRef] [PubMed]
- Zeiler, M.D.; Fergus, R. Visualizing and understanding convolutional networks. In Proceedings of the Computer Vision–ECCV 2014: 13th European Conference, Zurich, Switzerland, 2014, Proceedings, Part I 13. Springer, 2014, September 6-12; pp. 818–833.
- Kermany, D.S.; Goldbaum, M.; Cai, W.; Valentim, C.C.; Liang, H.; Baxter, S.L.; McKeown, A.; Yang, G.; Wu, X.; Yan, F.; et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. cell 2018, 172, 1122–1131. [Google Scholar] [CrossRef]
- Ribeiro, M.T.; Singh, S.; Guestrin, C. " Explaining the predictions of any classifier. In Proceedings of the Proceedings of the 22nd ACM SIGKDD international conference on knowledge discovery and data mining, 2016, pp.; pp. 1135–1144.
- Seah, J.C.; Tang, J.S.; Kitchen, A.; Gaillard, F.; Dixon, A.F. Chest radiographs in congestive heart failure: visualizing neural network learning. Radiology 2019, 290, 514–522. [Google Scholar] [CrossRef]
- Sundararajan, M.; Taly, A.; Yan, Q. Axiomatic attribution for deep networks. In Proceedings of the International conference on machine learning. PMLR; 2017; pp. 3319–3328. [Google Scholar]
- Sayres, R.; Taly, A.; Rahimy, E.; Blumer, K.; Coz, D.; Hammel, N.; Webster, D.R. Using a deep learning algorithm and integrated gradients explanation to assist grading for diabetic retinopathy. Ophthalmology 2019, 126, 552–564. [Google Scholar] [CrossRef]
- Papanastasopoulos, Z.; Samala, R.K.; Chan, H.P.; Hadjiiski, L.; Paramagul, C.; Helvie, M.A.; Neal, C.H. Explainable AI for medical imaging: deep-learning CNN ensemble for classification of estrogen receptor status from breast MRI. In Proceedings of the Medical imaging 2020: Computer-aided diagnosis. SPIE, Vol. 11314; 2020; pp. 228–235. [Google Scholar]
- Rajaraman, S.; Candemir, S.; Thoma, G.; Antani, S. Visualizing and explaining deep learning predictions for pneumonia detection in pediatric chest radiographs. In Proceedings of the Medical Imaging 2019: Computer-Aided Diagnosis. SPIE, Vol. 10950; 2019; pp. 200–211. [Google Scholar]
- Malhi, A.; Kampik, T.; Pannu, H.; Madhikermi, M.; Främling, K. Explaining machine learning-based classifications of in-vivo gastral images. 2019 Digital Image Computing: Techniques and Applications (DICTA) 2019, 1–7. [Google Scholar]
- Dubost, F.; Adams, H.; Bortsova, G.; Ikram, M.A.; Niessen, W.; Vernooij, M.; De Bruijne, M. 3D regression neural network for the quantification of enlarged perivascular spaces in brain MRI. Medical image analysis 2019, 51, 89–100. [Google Scholar] [CrossRef] [PubMed]
- Shahamat, H.; Abadeh, M.S. Brain MRI analysis using a deep learning based evolutionary approach. Neural Networks 2020, 126, 218–234. [Google Scholar] [CrossRef] [PubMed]
- Gecer, B.; Aksoy, S.; Mercan, E.; Shapiro, L.G.; Weaver, D.L.; Elmore, J.G. Detection and classification of cancer in whole slide breast histopathology images using deep convolutional networks. Pattern recognition 2018, 84, 345–356. [Google Scholar] [CrossRef] [PubMed]
- Garg, P.; Davenport, E.; Murugesan, G.; Wagner, B.; Whitlow, C.; Maldjian, J.; Montillo, A. Using convolutional neural networks to automatically detect eye-blink artifacts in magnetoencephalography without resorting to electrooculography. In Proceedings of the Medical Image Computing and Computer Assisted Intervention- MICCAI 2017: 20th International Conference, Quebec City, QC, Canada, 2017, Proceedings, Part III 20. Springer, 2017, September 11-13; pp. 374–381.
- Simonyan, K.; Vedaldi, A.; Zisserman, A. Deep inside convolutional networks: Visualising image classification models and saliency maps. arXiv preprint, /: Available at: https, 1312. [Google Scholar]
- Dubost, F.; Bortsova, G.; Adams, H.; Ikram, A.; Niessen, W.J.; Vernooij, M.; De Bruijne, M. Gp-unet: Lesion detection from weak labels with a 3d regression network. In Proceedings of the Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), Cham, September 2017; pp. 214–221.
- Lévy, D.; Jain, A. Breast mass classification from mammograms using deep convolutional neural networks. arXiv preprint, /: Available at: https, 1612. [Google Scholar]
- Rayan, J.C.; Reddy, N.; Kan, J.H.; Zhang, W.; Annapragada, A. Binomial classification of pediatric elbow fractures using a deep learning multiview approach emulating radiologist decision making. Radiology: Artificial Intelligence 2019, 1, e180015. [Google Scholar] [CrossRef]
- Liefers, B.; González-Gonzalo, C.; Klaver, C.; van Ginneken, B.; Sánchez, C.I. Dense segmentation in selected dimensions: application to retinal optical coherence tomography. In Proceedings of the Proceedings of the International Conference on Medical Imaging with Deep Learning (MIDL).; pp. 2019337–346.
- Springenberg, J.T.; Dosovitskiy, A.; Brox, T.; Riedmiller, M. Striving for simplicity: The all convolutional net. arXiv preprint, /: Available at: https, 1412. [Google Scholar]
- Böhle, M.; Eitel, F.; Weygandt, M.; Ritter, K. Layer-wise relevance propagation for explaining deep neural network decisions in MRI-based Alzheimer’s disease classification. Frontiers in Aging Neuroscience 2019, 11, 456892. [Google Scholar]
- Dubost, F.; Yilmaz, P.; Adams, H.; Bortsova, G.; Ikram, M.A.; Niessen, W.; Vernooij, M.; de Bruijne, M. Enlarged perivascular spaces in brain MRI: automated quantification in four regions. Neuroimage 2019, 185, 534–544. [Google Scholar] [CrossRef]
- Wang, X.; Liang, X.; Jiang, Z.; Nguchu, B.A.; Zhou, Y.; Wang, Y.; Wang, H.; Li, Y.; Zhu, Y.; Wu, F.; et al. Decoding and mapping task states of the human brain via deep learning. Human brain mapping 2020, 41, 1505–1519. [Google Scholar] [CrossRef] [PubMed]
- Gessert, N.; Latus, S.; Abdelwahed, Y.S.; Leistner, D.M.; Lutz, M.; Schlaefer, A. Bioresorbable scaffold visualization in IVOCT images using CNNs and weakly supervised localization. In Proceedings of the Medical Imaging 2019: Image Processing. SPIE, Vol. 10949, March 2019; pp. 606–612. [Google Scholar]
- Wickstrøm, K.; Kampffmeyer, M.; Jenssen, R. Uncertainty and interpretability in convolutional neural networks for semantic segmentation of colorectal polyps. Medical Image Analysis 2020, 60, 101619. [Google Scholar] [CrossRef]
- Jamaludin, A.; Kadir, T.; Zisserman, A. SpineNet: Automated classification and evidence visualization in spinal MRIs. Medical Image Analysis 2017, 41, 63–73. [Google Scholar] [CrossRef]
- Zhou, B.; Khosla, A.; Lapedriza, A.; Oliva, A.; Torralba, A. Learning deep features for discriminative localization. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2016, pp.; pp. 2921–2929.
- Lin, M.; Chen, Q.; Yan, S. Network in network. arXiv preprint, /: Available at: https, 1312. [Google Scholar]
- Feng, X.; Lipton, Z.C.; Yang, J.; Small, S.A.; Provenzano, F.A.; Initiative, A.D.N.; Initiative, F.L.D.N. Estimating brain age based on a uniform healthy population with deep learning and structural magnetic resonance imaging. Neurobiology of Aging 2020, 91, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Zhou, B.; Wang, K.; Jiang, R.; Xu, M. Respond-CAM: Analyzing deep models for 3D imaging data by visualizations. In Proceedings of the Medical Image Computing and Computer Assisted InterventionâMICCAI 2018: 21st International Conference, Granada, Spain, 2018, Proceedings, Part I. Springer International Publishing, 2018, September 16-20; pp. 485–492.
- Oquab, M.; Bottou, L.; Laptev, I.; Sivic, J. Learning and transferring mid-level image representations using convolutional neural networks. In Proceedings of the Proceedings of the IEEE conference on computer vision and pattern recognition, 2014, pp.; pp. 1717–1724.
- Woerl, A.C.; Eckstein, M.; Geiger, J.; Wagner, D.C.; Daher, T.; Stenzel, P.; Fernandez, A.; Hartmann, A.; Wand, M.; Roth, W.; et al. Deep learning predicts molecular subtype of muscle-invasive bladder cancer from conventional histopathological slides. European urology 2020, 78, 256–264. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, A.; Sarkar, S.; Shah, A.; Gore, S.; Santosh, V.; Saini, J.; Ingalhalikar, M. Predictive and discriminative localization of IDH genotype in high grade gliomas using deep convolutional neural nets. In Proceedings of the 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019). IEEE; 2019; pp. 372–375. [Google Scholar]
- Shinde, S.; Prasad, S.; Saboo, Y.; Kaushick, R.; Saini, J.; Pal, P.K.; Ingalhalikar, M. Predictive markers for Parkinson’s disease using deep neural nets on neuromelanin sensitive MRI. NeuroImage: Clinical 2019, 22, 101748. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, S.; Aich, S.; Kim, H.C. Detection of Parkinsonâs disease from 3T T1 weighted MRI scans using 3D convolutional neural network. Diagnostics 2020, 10, 402. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.; Kim, Y.K.; Yoon, E.J.; Lee, J.Y.; Lee, D.S.; Initiative, A.D.N. Cognitive signature of brain FDG PET based on deep learning: domain transfer from Alzheimerâs disease to Parkinsonâs disease. European Journal of Nuclear Medicine and Molecular Imaging 2020, 47, 403–412. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Zhu, X.; Ding, M.; Zhang, X. Medical image classification using a light-weighted hybrid neural network based on PCANet and DenseNet. Ieee Access 2020, 8, 24697–24712. [Google Scholar] [CrossRef]
- Kim, C.; Kim, W.H.; Kim, H.J.; Kim, J. Weakly-supervised US breast tumor characterization and localization with a box convolution network. In Proceedings of the Medical Imaging 2020: Computer-Aided Diagnosis. SPIE, Vol. 11314; 2020; pp. 298–304. [Google Scholar]
- Luo, L.; Chen, H.; Wang, X.; Dou, Q.; Lin, H.; Zhou, J.; Li, G.; Heng, P.A. Deep angular embedding and feature correlation attention for breast MRI cancer analysis. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, 2019, Proceedings, Part IV 22. Springer, 2019, October 13–17; pp. 504–512.
- Yi, P.H.; Lin, A.; Wei, J.; Yu, A.C.; Sair, H.I.; Hui, F.K.; Hager, G.D.; Harvey, S.C. Deep-learning-based semantic labeling for 2D mammography and comparison of complexity for machine learning tasks. Journal of Digital Imaging 2019, 32, 565–570. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Nishikawa, R.M. Detecting mammographically occult cancer in women with dense breasts using deep convolutional neural network and Radon Cumulative Distribution Transform. Journal of Medical Imaging 2019, 6, 044502–044502. [Google Scholar] [CrossRef]
- Qi, X.; Zhang, L.; Chen, Y.; Pi, Y.; Chen, Y.; Lv, Q.; Yi, Z. Automated diagnosis of breast ultrasonography images using deep neural networks. Medical image analysis 2019, 52, 185–198. [Google Scholar] [CrossRef]
- Xi, P.; Guan, H.; Shu, C.; Borgeat, L.; Goubran, R. An integrated approach for medical abnormality detection using deep patch convolutional neural networks. The Visual Computer 2020, 36, 1869–1882. [Google Scholar] [CrossRef]
- Zhou, L.Q.; Wu, X.L.; Huang, S.Y.; Wu, G.G.; Ye, H.R.; Wei, Q.; Bao, L.Y.; Deng, Y.B.; Li, X.R.; Cui, X.W.; et al. Lymph node metastasis prediction from primary breast cancer US images using deep learning. Radiology 2020, 294, 19–28. [Google Scholar] [CrossRef] [PubMed]
- Dunnmon, J.A.; Yi, D.; Langlotz, C.P.; Ré, C.; Rubin, D.L.; Lungren, M.P. Assessment of convolutional neural networks for automated classification of chest radiographs. Radiology 2019, 290, 537–544. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Fu, D. Diagnose chest pathology in X-ray images by learning multi-attention convolutional neural network. In Proceedings of the 2019 IEEE 8th Joint International Information Technology and Artificial Intelligence Conference (ITAIC). IEEE; 2019; pp. 294–299. [Google Scholar]
- Khakzar, A.; Albarqouni, S.; Navab, N. Learning interpretable features via adversarially robust optimization. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, 2019, Proceedings, Part VI 22. Springer, 2019, October 13–17; pp. 793–800.
- Kumar, D.; Sankar, V.; Clausi, D.; Taylor, G.W.; Wong, A. Sisc: End-to-end interpretable discovery radiomics-driven lung cancer prediction via stacked interpretable sequencing cells. IEEE Access 2019, 7, 145444–145454. [Google Scholar] [CrossRef]
- Lei, Y.; Tian, Y.; Shan, H.; Zhang, J.; Wang, G.; Kalra, M.K. Shape and margin-aware lung nodule classification in low-dose CT images via soft activation mapping. Medical Image Analysis 2020, 60, 101628. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.X.; Tang, Y.B.; Peng, Y.; Yan, K.; Bagheri, M.; Redd, B.A.; Brandon, C.J.; Lu, Z.; Han, M.; Xiao, J.; et al. Automated abnormality classification of chest radiographs using deep convolutional neural networks. NPJ digital medicine 2020, 3, 70. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Zhang, X.; Huang, S. KGZNet: Knowledge-guided deep zoom neural networks for thoracic disease classification. In Proceedings of the 2019 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). IEEE; 2019; pp. 1396–1401. [Google Scholar]
- Yi, P.H.; Kim, T.K.; Yu, A.C.; Bennett, B.; Eng, J.; Lin, C.T. Can AI outperform a junior resident? Comparison of deep neural network to first-year radiology residents for identification of pneumothorax. Emergency Radiology 2020, 27, 367–375. [Google Scholar] [CrossRef]
- Liu, H.; Wang, L.; Nan, Y.; Jin, F.; Wang, Q.; Pu, J. SDFN: Segmentation-based deep fusion network for thoracic disease classification in chest X-ray images. Computerized Medical Imaging and Graphics 2019, 75, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.; Kasukurthi, N.; Pande, H. Deep learning for weak supervision of diabetic retinopathy abnormalities. In Proceedings of the 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019). IEEE; 2019; pp. 573–577. [Google Scholar]
- Liao, W.; Zou, B.; Zhao, R.; Chen, Y.; He, Z.; Zhou, M. Clinical interpretable deep learning model for glaucoma diagnosis. IEEE journal of biomedical and health informatics 2019, 24, 1405–1412. [Google Scholar] [CrossRef]
- Perdomo, O.; Rios, H.; Rodríguez, F.J.; Otálora, S.; Meriaudeau, F.; Müller, H.; González, F.A. Classification of diabetes-related retinal diseases using a deep learning approach in optical coherence tomography. Computer methods and programs in biomedicine 2019, 178, 181–189. [Google Scholar] [CrossRef]
- Shen, Y.; Sheng, B.; Fang, R.; Li, H.; Dai, L.; Stolte, S.; Qin, J.; Jia, W.; Shen, D. Domain-invariant interpretable fundus image quality assessment. Medical image analysis 2020, 61, 101654. [Google Scholar] [CrossRef]
- Wang, X.; Chen, H.; Ran, A.R.; Luo, L.; Chan, P.P.; Tham, C.C.; Chang, R.T.; Mannil, S.S.; Cheung, C.Y.; Heng, P.A. Towards multi-center glaucoma OCT image screening with semi-supervised joint structure and function multi-task learning. Medical Image Analysis 2020, 63, 101695. [Google Scholar] [CrossRef]
- Jiang, H.; Yang, K.; Gao, M.; Zhang, D.; Ma, H.; Qian, W. An interpretable ensemble deep learning model for diabetic retinopathy disease classification. In Proceedings of the 2019 41st annual international conference of the IEEE engineering in medicine and biology society (EMBC). IEEE; 2019; pp. 2045–2048. [Google Scholar]
- Tu, Z.; Gao, S.; Zhou, K.; Chen, X.; Fu, H.; Gu, Z.; Cheng, J.; Yu, Z.; Liu, J. SUNet: A lesion regularized model for simultaneous diabetic retinopathy and diabetic macular edema grading. In Proceedings of the 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI). IEEE; 2020; pp. 1378–1382. [Google Scholar]
- Kumar, D.; Taylor, G.W.; Wong, A. Discovery radiomics with CLEAR-DR: interpretable computer aided diagnosis of diabetic retinopathy. IEEE Access 2019, 7, 25891–25896. [Google Scholar] [CrossRef]
- Liu, C.; Han, X.; Li, Z.; Ha, J.; Peng, G.; Meng, W.; He, M. A self-adaptive deep learning method for automated eye laterality detection based on color fundus photography. Plos one 2019, 14, e0222025. [Google Scholar] [CrossRef]
- Narayanan, B.N.; Hardie, R.C.; De Silva, M.S.; Kueterman, N.K. Hybrid machine learning architecture for automated detection and grading of retinal images for diabetic retinopathy. Journal of Medical Imaging 2020, 7, 034501–034501. [Google Scholar] [CrossRef] [PubMed]
- Everson, M.; Herrera, L.G.P.; Li, W.; Luengo, I.M.; Ahmad, O.; Banks, M.; Magee, C.; Alzoubaidi, D.; Hsu, H.; Graham, D.; et al. Artificial intelligence for the real-time classification of intrapapillary capillary loop patterns in the endoscopic diagnosis of early oesophageal squamous cell carcinoma: A proof-of-concept study. United European gastroenterology journal 2019, 7, 297–306. [Google Scholar] [CrossRef]
- García-Peraza-Herrera, L.C.; Everson, M.; Lovat, L.; Wang, H.P.; Wang, W.L.; Haidry, R.; Stoyanov, D.; Ourselin, S.; Vercauteren, T. Intrapapillary capillary loop classification in magnification endoscopy: open dataset and baseline methodology. International journal of computer assisted radiology and surgery 2020, 15, 651–659. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Xing, Y.; Zhang, L.; Gao, H.; Zhang, H. Deep convolutional neural network for ulcer recognition in wireless capsule endoscopy: experimental feasibility and optimization. Computational and mathematical methods in medicine 2019, 2019, 7546215. [Google Scholar] [CrossRef]
- Yan, C.; Xu, J.; Xie, J.; Cai, C.; Lu, H. Prior-aware CNN with multi-task learning for colon images analysis. In Proceedings of the 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI). IEEE; 2020; pp. 254–257. [Google Scholar]
- Heinemann, F.; Birk, G.; Stierstorfer, B. Deep learning enables pathologist-like scoring of NASH models. Scientific reports 2019, 9, 18454. [Google Scholar] [CrossRef] [PubMed]
- Kiani, A.; Uyumazturk, B.; Rajpurkar, P.; Wang, A.; Gao, R.; Jones, E.; Yu, Y.; Langlotz, C.P.; Ball, R.L.; Montine, T.J.; et al. Impact of a deep learning assistant on the histopathologic classification of liver cancer. NPJ digital medicine 2020, 3, 23. [Google Scholar] [CrossRef]
- Chang, G.H.; Felson, D.T.; Qiu, S.; Guermazi, A.; Capellini, T.D.; Kolachalama, V.B. Assessment of knee pain from MR imaging using a convolutional Siamese network. European radiology 2020, 30, 3538–3548. [Google Scholar] [CrossRef]
- Yi, P.H.; Kim, T.K.; Wei, J.; Shin, J.; Hui, F.K.; Sair, H.I.; Hager, G.D.; Fritz, J. Automated semantic labeling of pediatric musculoskeletal radiographs using deep learning. Pediatric radiology 2019, 49, 1066–1070. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Zhuang, J.; Wang, R.; Zhang, J.; Zheng, W.S. Fusing metadata and dermoscopy images for skin disease diagnosis. In Proceedings of the 2020 IEEE 17th international symposium on biomedical imaging (ISBI). IEEE; 2020; pp. 1996–2000. [Google Scholar]
- Xie, Y.; Zhang, J.; Xia, Y.; Shen, C. A mutual bootstrapping model for automated skin lesion segmentation and classification. IEEE transactions on medical imaging 2020, 39, 2482–2493. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Lee, K.J.; Sunwoo, L.; Choi, D.; Nam, C.M.; Cho, J.; Kim, J.; Bae, Y.J.; Yoo, R.E.; Choi, B.S.; et al. Deep learning in diagnosis of maxillary sinusitis using conventional radiography. Investigative radiology 2019, 54, 7–15. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhang, L.; Zhu, M.; Qi, X.; Yi, Z. Automatic diagnosis for thyroid nodules in ultrasound images by deep neural networks. Medical image analysis 2020, 61, 101665. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Chung, A.C. Evidence localization for pathology images using weakly supervised learning. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, 2019, Proceedings, Part I 22. Springer, 2019, October 13–17; pp. 613–621.
- Kim, I.; Rajaraman, S.; Antani, S. Visual interpretation of convolutional neural network predictions in classifying medical image modalities. Diagnostics 2019, 9, 38. [Google Scholar] [CrossRef] [PubMed]
- Tang, C. Discovering Unknown Diseases with Explainable Automated Medical Imaging. In Proceedings of the Medical Image Understanding and Analysis: 24th Annual Conference, MIUA 2020, Oxford, UK, 2020, Proceedings 24. Springer, 2020, July 15-17; pp. 346–358.
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual explanations from deep networks via gradient-based localization. In Proceedings of the Proceedings of the IEEE International Conference on Computer Vision (ICCV), 2017, pp.; pp. 618–626.
- Hilbert, A.; Ramos, L.A.; van Os, H.J.; Olabarriaga, S.D.; Tolhuisen, M.L.; Wermer, M.J.; Marquering, H.A. Data-efficient deep learning of radiological image data for outcome prediction after endovascular treatment of patients with acute ischemic stroke. Computers in Biology and Medicine 2019, 115, 103516. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.H.; Ye, J.C. Understanding graph isomorphism network for rs-fMRI functional connectivity analysis. Frontiers in Neuroscience 2020, 14, 630. [Google Scholar] [CrossRef] [PubMed]
- Liao, L.; Zhang, X.; Zhao, F.; Lou, J.; Wang, L.; Xu, X.; Li, G. Multi-branch deformable convolutional neural network with label distribution learning for fetal brain age prediction. In Proceedings of the 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI). IEEE, April 2020; pp. 424–427. [Google Scholar]
- Natekar, P.; Kori, A.; Krishnamurthi, G. Demystifying brain tumor segmentation networks: interpretability and uncertainty analysis. Frontiers in Computational Neuroscience 2020, 14, 6. [Google Scholar] [CrossRef] [PubMed]
- Pereira, S.; Meier, R.; Alves, V.; Reyes, M.; Silva, C.A. Automatic brain tumor grading from MRI data using convolutional neural networks and quality assessment. In Proceedings of the Understanding and Interpreting Machine Learning in Medical Image Computing Applications: First International Workshops, MLCN 2018, DLF 2018, and iMIMIC 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, 2018, Proceedings 1. Springer International Publishing, 2018, September 16-20; pp. 106–114.
- Pominova, M.; Artemov, A.; Sharaev, M.; Kondrateva, E.; Bernstein, A.; Burnaev, E. Voxelwise 3D convolutional and recurrent neural networks for epilepsy and depression diagnostics from structural and functional MRI data. In Proceedings of the 2018 IEEE International Conference on Data Mining Workshops (ICDMW). IEEE, November 2018; pp. 299–307. [Google Scholar]
- Xie, B.; Lei, T.; Wang, N.; Cai, H.; Xian, J.; He, M.; Xie, H. Computer-aided diagnosis for fetal brain ultrasound images using deep convolutional neural networks. International Journal of Computer Assisted Radiology and Surgery 2020, 15, 1303–1312. [Google Scholar] [CrossRef]
- El Adoui, M.; Drisis, S.; Benjelloun, M. Multi-input deep learning architecture for predicting breast tumor response to chemotherapy using quantitative MR images. International Journal of Computer Assisted Radiology and Surgery 2020, 15, 1491–1500. [Google Scholar] [CrossRef]
- Obikane, S.; Aoki, Y. Weakly supervised domain adaptation with point supervision in histopathological image segmentation. In Proceedings of the Pattern Recognition: ACPR 2019 Workshops, Auckland, New Zealand, 2019, Proceedings 5. Springer Singapore, 2020, November 26; pp. 127–140.
- Candemir, S.; White, R.D.; Demirer, M.; Gupta, V.; Bigelow, M.T.; Prevedello, L.M.; Erdal, B.S. Automated coronary artery atherosclerosis detection and weakly supervised localization on coronary CT angiography with a deep 3-dimensional convolutional neural network. Computerized Medical Imaging and Graphics 2020, 83, 101721. [Google Scholar] [CrossRef] [PubMed]
- Cong, C.; Kato, Y.; Vasconcellos, H.D.; Lima, J.; Venkatesh, B. Automated stenosis detection and classification in X-ray angiography using deep neural network. In Proceedings of the 2019 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). IEEE, November 2019; pp. 1301–1308. [Google Scholar]
- Huo, Y.; Terry, J.G.; Wang, J.; Nath, V.; Bermudez, C.; Bao, S.; Landman, B.A. Coronary calcium detection using 3D attention identical dual deep network based on weakly supervised learning. In Proceedings of the Medical Imaging 2019: Image Processing. SPIE, Vol. 10949, March 2019; pp. 308–315. [Google Scholar]
- Patra, A.; Noble, J.A. Incremental learning of fetal heart anatomies using interpretable saliency maps. In Proceedings of the Medical Image Understanding and Analysis: 23rd Conference, MIUA 2019, Liverpool, UK, 2019, Proceedings 23. Springer International Publishing, 2020, July 24â26; pp. 129–141.
- Brunese, L.; Mercaldo, F.; Reginelli, A.; Santone, A. Explainable deep learning for pulmonary disease and coronavirus COVID-19 detection from X-rays. Computer Methods and Programs in Biomedicine 2020, 196, 105608. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Li, J.; Lu, G.; Zhang, D. Lesion location attention guided network for multi-label thoracic disease classification in chest X-rays. IEEE Journal of Biomedical and Health Informatics 2019, 24, 2016–2027. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Shang, L.; Ji, H.; Zhang, X. Deep learning features for lung adenocarcinoma classification with tissue pathology images. In Proceedings of the Neural Information Processing: 24th International Conference, ICONIP 2017, Guangzhou, China, 2017, Proceedings, Part IV. Springer International Publishing, 2017, Vol. 24, November 14â18; pp. 742–751.
- Hosny, A.; Parmar, C.; Coroller, T.P.; Grossmann, P.; Zeleznik, R.; Kumar, A.; Aerts, H.J. Deep learning for lung cancer prognostication: a retrospective multi-cohort radiomics study. PLoS Medicine 2018, 15, e1002711. [Google Scholar] [CrossRef] [PubMed]
- Humphries, S.M.; Notary, A.M.; Centeno, J.P.; Strand, M.J.; Crapo, J.D.; Silverman, E.K.; of COPD (COPDGene) Investigators, G.E. Deep learning enables automatic classification of emphysema pattern at CT. Radiology 2020, 294, 434–444. [Google Scholar] [CrossRef] [PubMed]
- Ko, H.; Chung, H.; Kang, W.S.; Kim, K.W.; Shin, Y.; Kang, S.J.; Lee, J. COVID-19 pneumonia diagnosis using a simple 2D deep learning framework with a single chest CT image: model development and validation. Journal of Medical Internet Research 2020, 22, e19569. [Google Scholar] [CrossRef] [PubMed]
- Mahmud, T.; Rahman, M.A.; Fattah, S.A. CovXNet: A multi-dilation convolutional neural network for automatic COVID-19 and other pneumonia detection from chest X-ray images with transferable multi-receptive feature optimization. Computers in Biology and Medicine 2020, 122, 103869. [Google Scholar] [CrossRef] [PubMed]
- Paul, R.; Schabath, M.; Gillies, R.; Hall, L.; Goldgof, D. Convolutional Neural Network ensembles for accurate lung nodule malignancy prediction 2 years in the future. Computers in Biology and Medicine 2020, 122, 103882. [Google Scholar] [CrossRef]
- Philbrick, K.A.; Yoshida, K.; Inoue, D.; Akkus, Z.; Kline, T.L.; Weston, A.D.; Erickson, B.J. What does deep learning see? Insights from a classifier trained to predict contrast enhancement phase from CT images. American Journal of Roentgenology 2018, 211, 1184–1193. [Google Scholar] [CrossRef]
- Qin, R.; Wang, Z.; Jiang, L.; Qiao, K.; Hai, J.; Chen, J.; Yan, B. FineâGrained Lung Cancer Classification from PET and CT Images Based on Multidimensional Attention Mechanism. Complexity 2020, 2020, 6153657. [Google Scholar] [CrossRef]
- Teramoto, A.; Yamada, A.; Kiriyama, Y.; Tsukamoto, T.; Yan, K.; Zhang, L.; Fujita, H. Automated classification of benign and malignant cells from lung cytological images using deep convolutional neural network. Informatics in Medicine Unlocked 2019, 16, 100205. [Google Scholar] [CrossRef]
- Xu, R.; Cong, Z.; Ye, X.; Hirano, Y.; Kido, S.; Gyobu, T.; Tomiyama, N. Pulmonary textures classification via a multi-scale attention network. IEEE Journal of Biomedical and Health Informatics 2019, 24, 2041–2052. [Google Scholar] [CrossRef]
- Vila-Blanco, N.; Carreira, M.J.; Varas-Quintana, P.; Balsa-Castro, C.; Tomas, I. Deep neural networks for chronological age estimation from OPG images. IEEE Transactions on Medical Imaging 2020, 39, 2374–2384. [Google Scholar] [CrossRef]
- Kim, M.; Han, J.C.; Hyun, S.H.; Janssens, O.; Van Hoecke, S.; Kee, C.; De Neve, W. Medinoid: computer-aided diagnosis and localization of glaucoma using deep learning. Applied Sciences 2019, 9, 3064. [Google Scholar] [CrossRef]
- Martins, J.; Cardoso, J.S.; Soares, F. Offline computer-aided diagnosis for Glaucoma detection using fundus images targeted at mobile devices. Computer Methods and Programs in Biomedicine 2020, 192, 105341. [Google Scholar] [CrossRef] [PubMed]
- Meng, Q.; Hashimoto, Y.; Satoh, S. How to extract more information with less burden: Fundus image classification and retinal disease localization with ophthalmologist intervention. IEEE Journal of Biomedical and Health Informatics 2020, 24, 3351–3361. [Google Scholar] [CrossRef]
- Wang, R.; Fan, D.; Lv, B.; Wang, M.; Zhou, Q.; Lv, C.; Xie, G.; Wang, L. OCT image quality evaluation based on deep and shallow features fusion network. In Proceedings of the 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI). IEEE; 2020; pp. 1561–1564. [Google Scholar]
- Zhang, R.; Tan, S.; Wang, R.; Manivannan, S.; Chen, J.; Lin, H.; Zheng, W.S. Biomarker localization by combining CNN classifier and generative adversarial network. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, 2019, Proceedings, Part I 22. Springer, 2019, October 13–17; pp. 209–217.
- Chen, X.; Lin, L.; Liang, D.; Hu, H.; Zhang, Q.; Iwamoto, Y.; Han, X.H.; Chen, Y.W.; Tong, R.; Wu, J. A dual-attention dilated residual network for liver lesion classification and localization on CT images. In Proceedings of the 2019 IEEE international conference on image processing (ICIP). IEEE; 2019; pp. 235–239. [Google Scholar]
- Itoh, H.; Lu, Z.; Mori, Y.; Misawa, M.; Oda, M.; Kudo, S.e.; Mori, K. Visualising decision-reasoning regions in computer-aided pathological pattern diagnosis of endoscytoscopic images based on CNN weights analysis. In Proceedings of the Medical Imaging 2020: Computer-Aided Diagnosis. SPIE, Vol. 11314; 2020; pp. 761–768. [Google Scholar]
- Korbar, B.; Olofson, A.M.; Miraflor, A.P.; Nicka, C.M.; Suriawinata, M.A.; Torresani, L.; Suriawinata, A.A.; Hassanpour, S. Looking under the hood: Deep neural network visualization to interpret whole-slide image analysis outcomes for colorectal polyps. In Proceedings of the Proceedings of the IEEE conference on computer vision and pattern recognition workshops, 2017, pp.; pp. 69–75.
- Kowsari, K.; Sali, R.; Ehsan, L.; Adorno, W.; Ali, A.; Moore, S.; Amadi, B.; Kelly, P.; Syed, S.; Brown, D. Hmic: Hierarchical medical image classification, a deep learning approach. Information 2020, 11, 318. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Cui, Y.; Shi, G.; Zhao, J.; Yang, X.; Qiang, Y.; Du, Q.; Ma, Y.; Kazihise, N.G.F. Multi-branch cross attention model for prediction of KRAS mutation in rectal cancer with t2-weighted MRI. Applied Intelligence 2020, 50, 2352–2369. [Google Scholar] [CrossRef]
- Cheng, C.T.; Ho, T.Y.; Lee, T.Y.; Chang, C.C.; Chou, C.C.; Chen, C.C.; Chung, I.; Liao, C.H.; et al. Application of a deep learning algorithm for detection and visualization of hip fractures on plain pelvic radiographs. European radiology 2019, 29, 5469–5477. [Google Scholar] [CrossRef] [PubMed]
- Gupta, V.; Demirer, M.; Bigelow, M.; Sarah, M.Y.; Joseph, S.Y.; Prevedello, L.M.; White, R.D.; Erdal, B.S. Using transfer learning and class activation maps supporting detection and localization of femoral fractures on anteroposterior radiographs. In Proceedings of the 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI). IEEE; 2020; pp. 1526–1529. [Google Scholar]
- Zhang, B.; Tan, J.; Cho, K.; Chang, G.; Deniz, C.M. Attention-based cnn for kl grade classification: Data from the osteoarthritis initiative. In Proceedings of the 2020 IEEE 17th international symposium on biomedical imaging (ISBI). IEEE; 2020; pp. 731–735. [Google Scholar]
- von Schacky, C.E.; Sohn, J.H.; Liu, F.; Ozhinsky, E.; Jungmann, P.M.; Nardo, L.; Posadzy, M.; Foreman, S.C.; Nevitt, M.C.; Link, T.M.; et al. Development and validation of a multitask deep learning model for severity grading of hip osteoarthritis features on radiographs. Radiology 2020, 295, 136–145. [Google Scholar] [CrossRef]
- Lee, J.H.; Ha, E.J.; Kim, D.; Jung, Y.J.; Heo, S.; Jang, Y.H.; An, S.H.; Lee, K. Application of deep learning to the diagnosis of cervical lymph node metastasis from thyroid cancer with CT: external validation and clinical utility for resident training. European radiology 2020, 30, 3066–3072. [Google Scholar] [CrossRef] [PubMed]
- Langner, T.; Wikström, J.; Bjerner, T.; Ahlström, H.; Kullberg, J. Identifying morphological indicators of aging with neural networks on large-scale whole-body MRI. IEEE transactions on medical imaging 2019, 39, 1430–1437. [Google Scholar] [CrossRef] [PubMed]
- Eitel, F.; Soehler, E.; Bellmann-Strobl, J.; Brandt, A.U.; Ruprecht, K.; Giess, R.M.; Kuchling, J.; Asseyer, S.; Weygandt, M.; Haynes, J.D.; et al. Uncovering convolutional neural network decisions for diagnosing multiple sclerosis on conventional MRI using layer-wise relevance propagation. NeuroImage: Clinical 2019, 24, 102003. [Google Scholar] [CrossRef] [PubMed]
- Schlemper, J.; Oktay, O.; Schaap, M.; Heinrich, M.; Kainz, B.; Glocker, B.; Rueckert, D. Attention gated networks: Learning to leverage salient regions in medical images. Medical image analysis 2019, 53, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Katar, O.; Yildirim, O. An explainable vision transformer model based white blood cells classification and localization. Diagnostics 2023, 13, 2459. [Google Scholar] [CrossRef] [PubMed]
- Bach, S.; Binder, A.; Montavon, G.; Klauschen, F.; Müller, K.R.; Samek, W. On pixel-wise explanations for non-linear classifier decisions by layer-wise relevance propagation. PloS one 2015, 10, e0130140. [Google Scholar] [CrossRef] [PubMed]
- Montavon, G.; Lapuschkin, S.; Binder, A.; Samek, W.; Müller, K.R. Explaining nonlinear classification decisions with deep taylor decomposition. Pattern recognition 2017, 65, 211–222. [Google Scholar] [CrossRef]
- Samek, W.; Binder, A.; Montavon, G.; Lapuschkin, S.; Müller, K.R. Evaluating the visualization of what a deep neural network has learned. IEEE transactions on neural networks and learning systems 2016, 28, 2660–2673. [Google Scholar] [CrossRef]
- Kohlbrenner, M.; Bauer, A.; Nakajima, S.; Binder, A.; Samek, W.; Lapuschkin, S. Towards best practice in explaining neural network decisions with LRP. In Proceedings of the 2020 International Joint Conference on Neural Networks (IJCNN). IEEE; 2020; pp. 1–7. [Google Scholar]
- Arquilla, K.; Gajera, I.D.; Darling, M.; Bhati, D.; Singh, A.; Guercio, A. Exploring Fine-Grained Feature Analysis for Bird Species Classification using Layer-wise Relevance Propagation. In Proceedings of the 2024 IEEE World AI IoT Congress (AIIoT). IEEE; 2024; pp. 625–631. [Google Scholar]
- Böhle, M.; Eitel, F.; Weygandt, M.; Ritter, K. Layer-wise relevance propagation for explaining deep neural network decisions in MRI-based Alzheimer’s disease classification. Frontiers in aging neuroscience 2019, 11, 456892. [Google Scholar] [CrossRef]
- Thomas, A.W.; Heekeren, H.R.; Müller, K.R.; Samek, W. Analyzing neuroimaging data through recurrent deep learning models. Frontiers in neuroscience 2019, 13, 1321. [Google Scholar] [CrossRef]
- Jetley, S.; Lord, N.A.; Lee, N.; Torr, P.H. Learn to pay attention. arXiv preprint arXiv:1804.02391, arXiv:1804.02391 2018.
- Li, S.; Dong, M.; Du, G.; Mu, X. Attention dense-u-net for automatic breast mass segmentation in digital mammogram. Ieee Access 2019, 7, 59037–59047. [Google Scholar] [CrossRef]
- Yan, Y.; Kawahara, J.; Hamarneh, G. Melanoma recognition via visual attention. In Proceedings of the Information Processing in Medical Imaging: 26th International Conference, IPMI 2019, Hong Kong, China, 2019, Proceedings 26. Springer, 2019, June 2–7; pp. 793–804.
- Górriz, M.; Antony, J.; McGuinness, K.; Giró-i Nieto, X.; OâConnor, N.E. Assessing knee OA severity with CNN attention-based end-to-end architectures. In Proceedings of the International conference on medical imaging with deep learning. PMLR; 2019; pp. 197–214. [Google Scholar]
- Dubost, F.; Adams, H.; Yilmaz, P.; Bortsova, G.; van Tulder, G.; Ikram, M.A.; Niessen, W.; Vernooij, M.W.; de Bruijne, M. Weakly supervised object detection with 2D and 3D regression neural networks. Medical image analysis 2020, 65, 101767. [Google Scholar] [CrossRef] [PubMed]
- Lian, C.; Liu, M.; Wang, L.; Shen, D. End-to-end dementia status prediction from brain mri using multi-task weakly-supervised attention network. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, 2019, Proceedings, Part IV 22. Springer, 2019, October 13–17; pp. 158–167.
- Wang, H.; Feng, J.; Zhang, Z.; Su, H.; Cui, L.; He, H.; Liu, L. Breast mass classification via deeply integrating the contextual information from multi-view data. Pattern Recognition 2018, 80, 42–52. [Google Scholar] [CrossRef]
- Li, L.; Xu, M.; Liu, H.; Li, Y.; Wang, X.; Jiang, L.; Wang, Z.; Fan, X.; Wang, N. A large-scale database and a CNN model for attention-based glaucoma detection. IEEE transactions on medical imaging 2019, 39, 413–424. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Kim, J.Y.; Kim, H.; Adhikari, S.P. Guided soft attention network for classification of breast cancer histopathology images. IEEE transactions on medical imaging 2019, 39, 1306–1315. [Google Scholar] [CrossRef] [PubMed]
- Pesce, E.; Withey, S.J.; Ypsilantis, P.P.; Bakewell, R.; Goh, V.; Montana, G. Learning to detect chest radiographs containing pulmonary lesions using visual attention networks. Medical image analysis 2019, 53, 26–38. [Google Scholar] [CrossRef] [PubMed]
- Singla, S.; Gong, M.; Ravanbakhsh, S.; Sciurba, F.; Poczos, B.; Batmanghelich, K.N. Subject2Vec: generative-discriminative approach from a set of image patches to a vector. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2018: 21st International Conference, Granada, Spain, 2018, Proceedings, Part I. Springer, 2018, September 16-20; pp. 502–510.
- Sun, J.; Darbehani, F.; Zaidi, M.; Wang, B. Saunet: Shape attentive u-net for interpretable medical image segmentation. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2020: 23rd International Conference, Lima, Peru, 2020, Proceedings, Part IV 23. Springer, 2020, October 4–8; pp. 797–806.
- Zhu, Z.; Ding, X.; Zhang, D.; Wang, L. Weakly-supervised balanced attention network for gastric pathology image localization and classification. In Proceedings of the 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI). IEEE; 2020; pp. 1–4. [Google Scholar]
- Barata, C.; Celebi, M.E.; Marques, J.S. Explainable skin lesion diagnosis using taxonomies. Pattern Recognition 2021, 110, 107413. [Google Scholar] [CrossRef]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv preprint arXiv:2010.11929, arXiv:2010.11929 2020.
- Srivastava, A.; Chandra, M.; Saha, A.; Saluja, S.; Bhati, D. Current Advances in Locality-Based and Feature-Based Transformers: A Review. In Proceedings of the International Conference on Data &, Information Sciences. Springer; 2023; pp. 321–335. [Google Scholar]
- Chen, J.; Lu, Y.; Yu, Q.; Luo, X.; Adeli, E.; Wang, Y.; Lu, L.; Yuille, A.L.; Zhou, Y. Transunet: Transformers make strong encoders for medical image segmentation. arXiv preprint arXiv:2102.04306, arXiv:2102.04306 2021.
- Karimi, D.; Vasylechko, S.D.; Gholipour, A. Convolution-free medical image segmentation using transformers. In Proceedings of the Medical image computing and computer assisted intervention–MICCAI 2021: 24th international conference, Strasbourg, France, 2021, proceedings, part I 24. Springer, 2021, September 27–October 1; pp. 78–88.
- Yun, B.; Wang, Y.; Chen, J.; Wang, H.; Shen, W.; Li, Q. Spectr: Spectral transformer for hyperspectral pathology image segmentation. arXiv preprint arXiv:2103.03604, arXiv:2103.03604 2021.
- Wenxuan, W.; Chen, C.; Meng, D.; Hong, Y.; Sen, Z.; Jiangyun, L. Transbts: Multimodal brain tumor segmentation using transformer. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Springer; 2021; pp. 109–119. [Google Scholar]
- Hatamizadeh, A.; Tang, Y.; Nath, V.; Yang, D.; Myronenko, A.; Landman, B.; Roth, H.R.; Xu, D. Unetr: Transformers for 3d medical image segmentation. In Proceedings of the Proceedings of the IEEE/CVF winter conference on applications of computer vision, 2022, pp.; pp. 574–584.
- Li, S.; Sui, X.; Luo, X.; Xu, X.; Liu, Y.; Goh, R. Medical image segmentation using squeeze-and-expansion transformers. arXiv preprint arXiv:2105.09511, arXiv:2105.09511 2021.
- Zhang, Y.; Higashita, R.; Fu, H.; Xu, Y.; Zhang, Y.; Liu, H.; Zhang, J.; Liu, J. A multi-branch hybrid transformer network for corneal endothelial cell segmentation. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, 2021, Proceedings, Part I 24. Springer, 2021, September 27–October 1; pp. 99–108.
- Lin, A.; Chen, B.; Xu, J.; Zhang, Z.; Lu, G.; Zhang, D. Ds-transunet: Dual swin transformer u-net for medical image segmentation. IEEE Transactions on Instrumentation and Measurement 2022, 71, 1–15. [Google Scholar] [CrossRef]
- Li, Y.; Cai, W.; Gao, Y.; Li, C.; Hu, X. More than encoder: Introducing transformer decoder to upsample. In Proceedings of the 2022 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). IEEE; 2022; pp. 1597–1602. [Google Scholar]
- Xu, G.; Zhang, X.; He, X.; Wu, X. Levit-unet: Make faster encoders with transformer for medical image segmentation. In Proceedings of the Chinese Conference on Pattern Recognition and Computer Vision (PRCV). Springer; 2023; pp. 42–53. [Google Scholar]
- Chang, Y.; Menghan, H.; Guangtao, Z.; Xiao-Ping, Z. Transclaw u-net: Claw u-net with transformers for medical image segmentation. arXiv preprint arXiv:2107.05188, arXiv:2107.05188 2021.
- Cao, H.; Wang, Y.; Chen, J.; Jiang, D.; Zhang, X.; Tian, Q.; Wang, M. Swin-unet: Unet-like pure transformer for medical image segmentation. In Proceedings of the European conference on computer vision. Springer; 2022; pp. 205–218. [Google Scholar]
- Petit, O.; Thome, N.; Rambour, C.; Themyr, L.; Collins, T.; Soler, L. U-net transformer: Self and cross attention for medical image segmentation. In Proceedings of the Machine Learning in Medical Imaging: 12th International Workshop, MLMI 2021, Held in Conjunction with MICCAI 2021, Strasbourg, France, 2021, Proceedings 12. Springer, 2021, September 27; pp. 267–276.
- Xie, Y.; Zhang, J.; Shen, C.; Xia, Y. Cotr: Efficiently bridging cnn and transformer for 3d medical image segmentation. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, 2021, Proceedings, Part III 24. Springer, 2021, September 27–October 1; pp. 171–180.
- Gao, Y.; Zhou, M.; Metaxas, D.N. UTNet: a hybrid transformer architecture for medical image segmentation. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, 2021, Proceedings, Part III 24. Springer, 2021, September 27–October 1; pp. 61–71.
- Chen, B.; Liu, Y.; Zhang, Z.; Lu, G.; Kong, A.W.K. Transattunet: Multi-level attention-guided u-net with transformer for medical image segmentation. IEEE Transactions on Emerging Topics in Computational Intelligence 2023. [Google Scholar] [CrossRef]
- Dong, B.; Wang, W.; Fan, D.P.; Li, J.; Fu, H.; Shao, L. Polyp-pvt: Polyp segmentation with pyramid vision transformers. arXiv preprint arXiv:2108.06932, arXiv:2108.06932 2021.
- Shen, Z.; Yang, H.; Zhang, Z.; Zheng, S. Automated kidney tumor segmentation with convolution and transformer network. In International Challenge on Kidney and Kidney Tumor Segmentation; Springer, 2021; pp. 1–12.
- Deng, K.; Meng, Y.; Gao, D.; Bridge, J.; Shen, Y.; Lip, G.; Zhao, Y.; Zheng, Y. Transbridge: A lightweight transformer for left ventricle segmentation in echocardiography. In Proceedings of the Simplifying Medical Ultrasound: Second International Workshop, ASMUS 2021, Held in Conjunction with MICCAI 2021, Strasbourg, France, 2021, Proceedings 2. Springer, 2021, September 27; pp. 63–72.
- Jia, Q.; Shu, H. Bitr-unet: a cnn-transformer combined network for mri brain tumor segmentation. In Proceedings of the International MICCAI Brainlesion Workshop. Springer; 2021; pp. 3–14. [Google Scholar]
- Hatamizadeh, A.; Nath, V.; Tang, Y.; Yang, D.; Roth, H.R.; Xu, D. Swin unetr: Swin transformers for semantic segmentation of brain tumors in mri images. In Proceedings of the International MICCAI brainlesion workshop. Springer; 2021; pp. 272–284. [Google Scholar]
- Li, Y.; Wang, S.; Wang, J.; Zeng, G.; Liu, W.; Zhang, Q.; Jin, Q.; Wang, Y. Gt u-net: A u-net like group transformer network for tooth root segmentation. In Proceedings of the Machine Learning in Medical Imaging: 12th International Workshop, MLMI 2021, Held in Conjunction with MICCAI 2021, Strasbourg, France, 2021, Proceedings 12. Springer, 2021, September 27; pp. 386–395.
- Gheflati, B.; Rivaz, H. Vision transformers for classification of breast ultrasound images. In Proceedings of the 2022 44th Annual International Conference of the IEEE Engineering in Medicine &, 2022, Biology Society (EMBC). IEEE; pp. 480–483.
- Zheng, Y.; Gindra, R.; Betke, M.; Beane, J.E.; Kolachalama, V.B. A deep learning based graph-transformer for whole slide image classification. medRxiv, 2021. [Google Scholar]
- Yu, S.; Ma, K.; Bi, Q.; Bian, C.; Ning, M.; He, N.; Li, Y.; Liu, H.; Zheng, Y. Mil-vt: Multiple instance learning enhanced vision transformer for fundus image classification. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, 2021, Proceedings, Part VIII 24. Springer, 2021, September 27–October 1; pp. 45–54.
- Sun, R.; Li, Y.; Zhang, T.; Mao, Z.; Wu, F.; Zhang, Y. Lesion-aware transformers for diabetic retinopathy grading. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2021, pp.; pp. 10938–10947.
- Perera, S.; Adhikari, S.; Yilmaz, A. Pocformer: A lightweight transformer architecture for detection of covid-19 using point of care ultrasound. In Proceedings of the 2021 IEEE international conference on image processing (ICIP). IEEE; 2021; pp. 195–199. [Google Scholar]
- Park, S.; Kim, G.; Kim, J.; Kim, B.; Ye, J.C. Federated split vision transformer for COVID-19 CXR diagnosis using task-agnostic training. arXiv preprint arXiv:2111.01338, arXiv:2111.01338 2021.
- Shome, D.; Kar, T.; Mohanty, S.N.; Tiwari, P.; Muhammad, K.; AlTameem, A.; Zhang, Y.; Saudagar, A.K.J. Covid-transformer: Interpretable covid-19 detection using vision transformer for healthcare. International Journal of Environmental Research and Public Health 2021, 18, 11086. [Google Scholar] [CrossRef]
- Liu, C.; Yin, Q. Automatic diagnosis of covid-19 using a tailored transformer-like network. In Proceedings of the Journal of Physics: Conference Series. IOP Publishing, Vol. 2010; 2021; p. 012175. [Google Scholar]
- Park, S.; Kim, G.; Oh, Y.; Seo, J.B.; Lee, S.M.; Kim, J.H.; Moon, S.; Lim, J.K.; Ye, J.C. Vision transformer for covid-19 cxr diagnosis using chest x-ray feature corpus. arXiv preprint arXiv:2103.07055, arXiv:2103.07055 2021.
- Gao, X.; Qian, Y.; Gao, A. COVID-VIT: Classification of COVID-19 from CT chest images based on vision transformer models. arXiv preprint arXiv:2107.01682, arXiv:2107.01682 2021.
- Mondal, A.K.; Bhattacharjee, A.; Singla, P.; Prathosh, A. xViTCOS: explainable vision transformer based COVID-19 screening using radiography. IEEE Journal of Translational Engineering in Health and Medicine 2021, 10, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.C.; Chen, G.L.; Wu, M.H. Visual transformer with statistical test for covid-19 classification. arXiv preprint arXiv:2107.05334, arXiv:2107.05334 2021.
- Zhang, L.; Wen, Y. A transformer-based framework for automatic COVID19 diagnosis in chest CTs. In Proceedings of the Proceedings of the IEEE/CVF international conference on computer vision, 2021, pp.; pp. 513–518.
- Ambita, A.A.E.; Boquio, E.N.V.; Naval Jr, P.C. Covit-gan: vision transformer forcovid-19 detection in ct scan imageswith self-attention gan forDataAugmentation. In Proceedings of the International Conference on Artificial Neural Networks. Springer; 2021; pp. 587–598. [Google Scholar]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).