Submitted:
05 August 2024
Posted:
06 August 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Preparation of Flowers
2.2. Screening and Isolation of LAB Strain from Flowers
2.3. Identification of Isolated LAB Strains from Flowers Using MALDI-TOF, PCR, and De-Replication
2.4. Screening for Exo-Cellular Polysaccharides Producers
2.5. Isolation and Purification of Exo-Cellular Polysaccharides
2.6. Determination of Monosaccharides by High Performance Liquid Chromatography (HPLC) Analysis
2.7. High-Throughput Screening for Texturing Strains in Plant-Base Drink
3. Results and Discussion
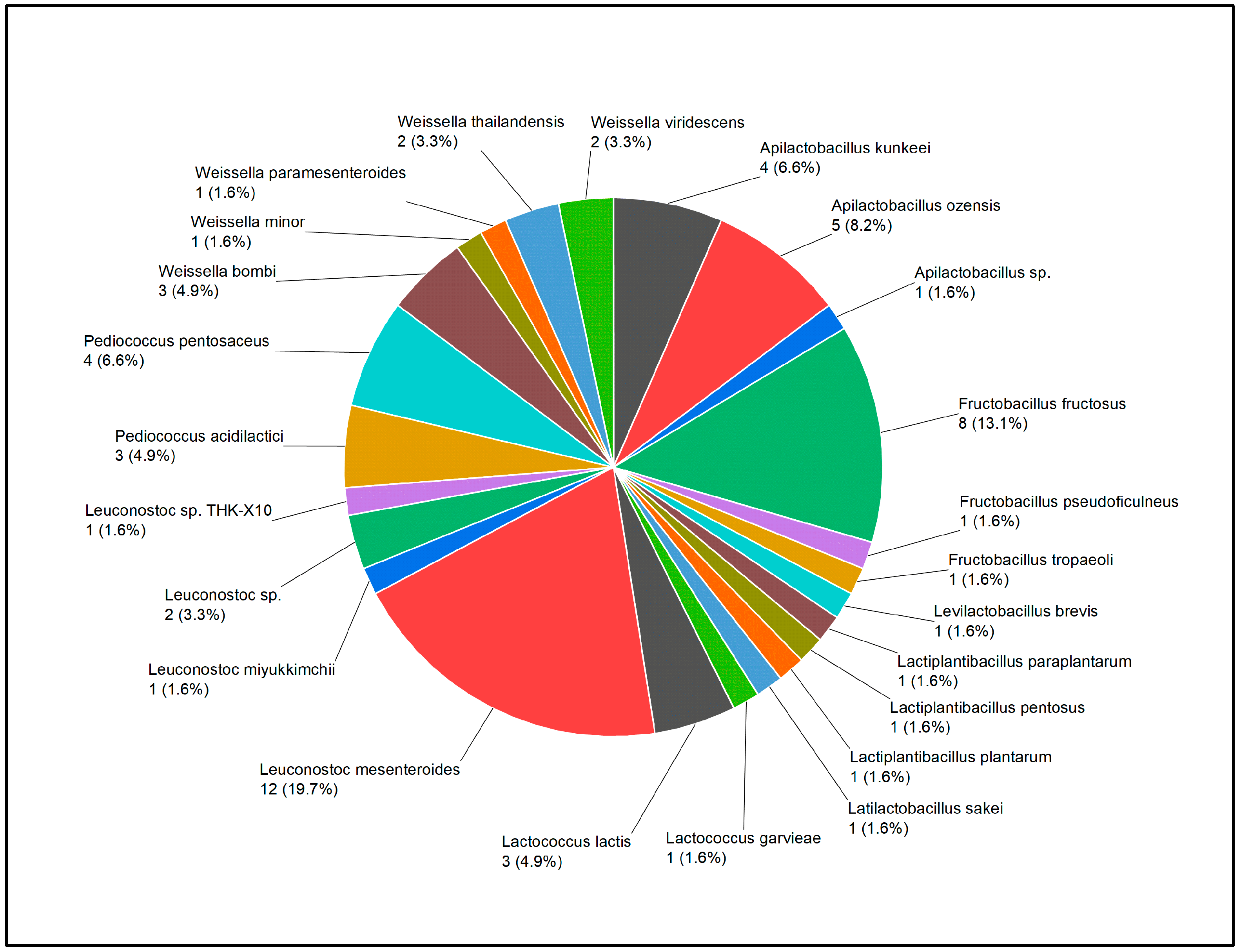
3.1. Diversity of plant-based LAB strains according to MALDI-TOF, PCR, and de-replication
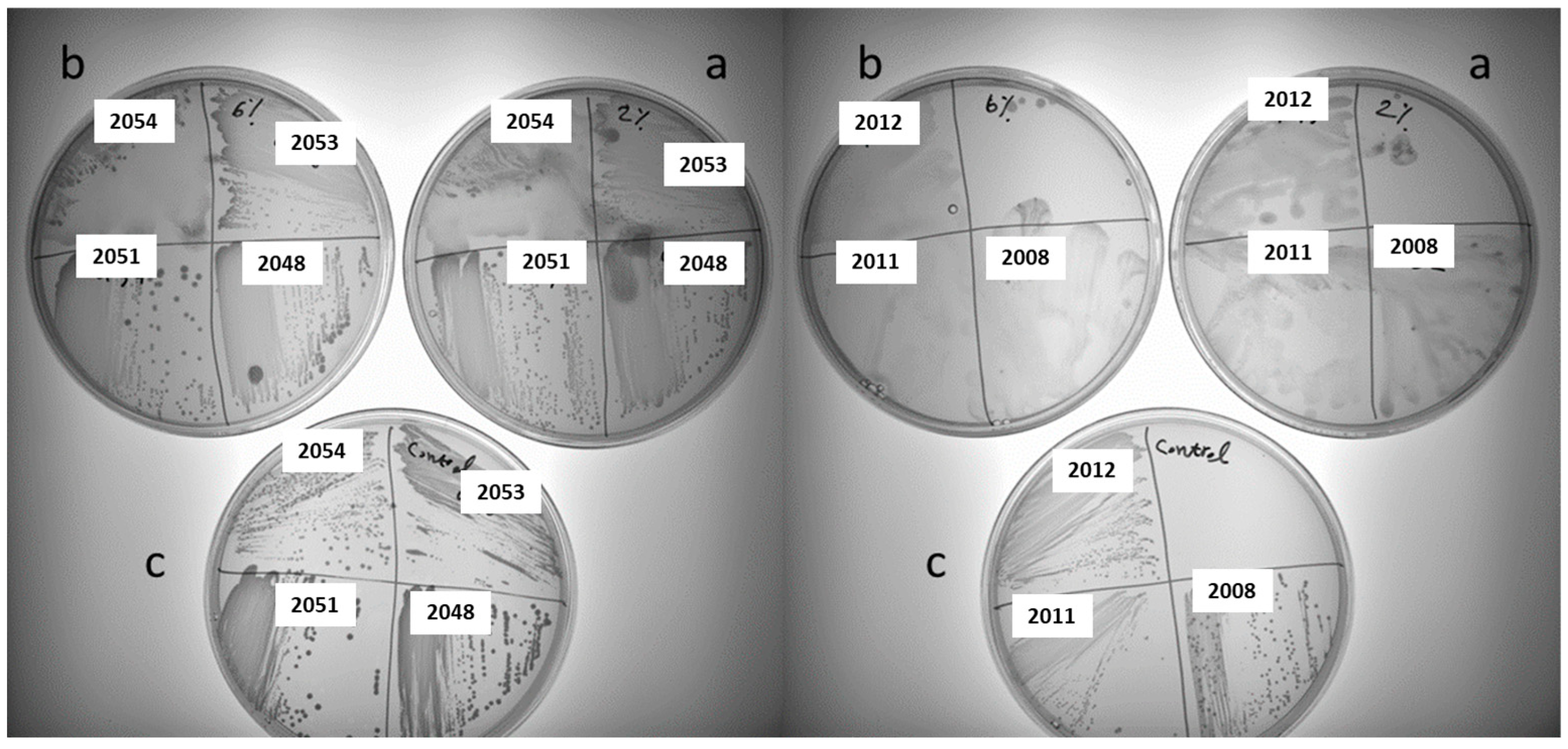
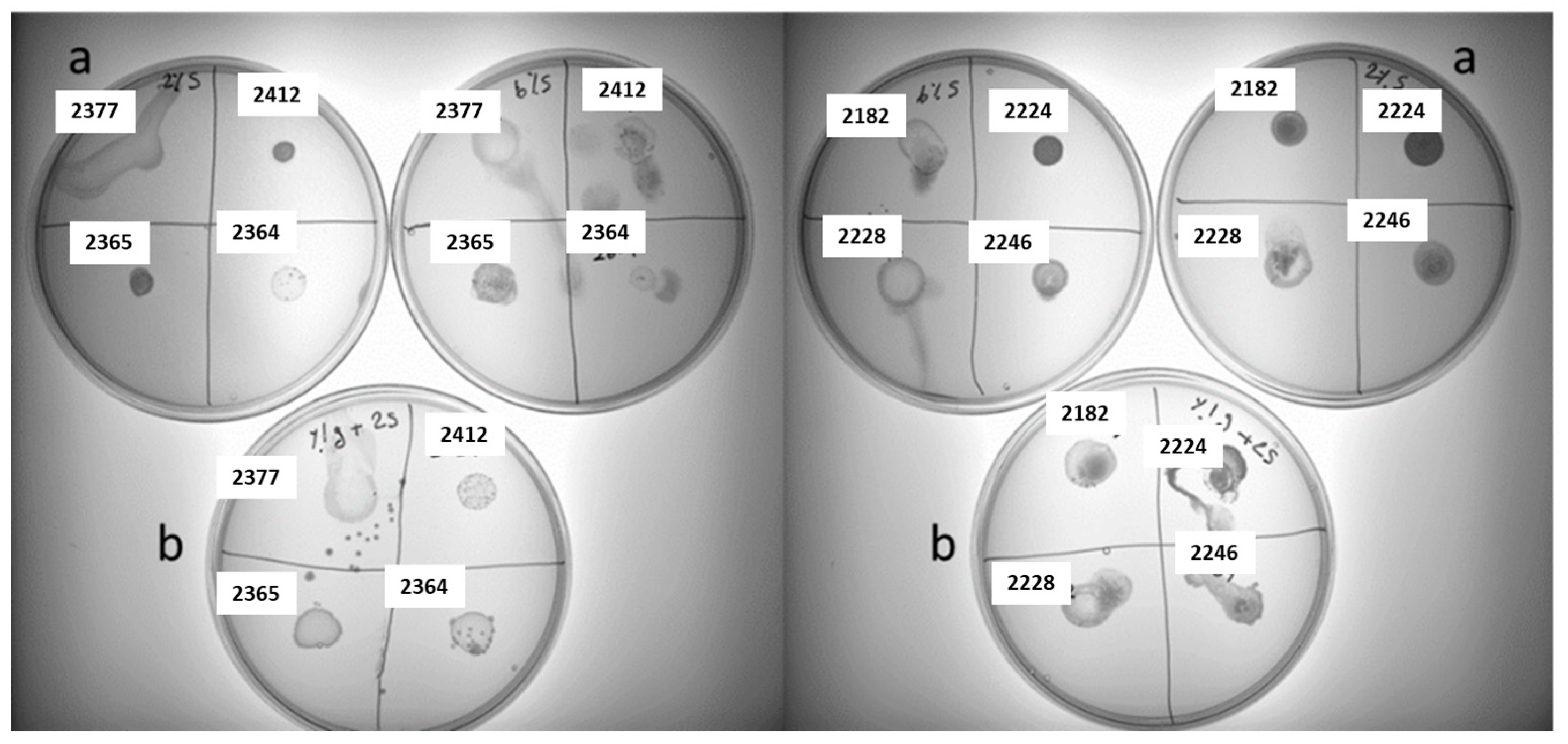
3.2. Screening for Polysaccharides Producing LAB Strains on Different Sucrose-Supplemented Media
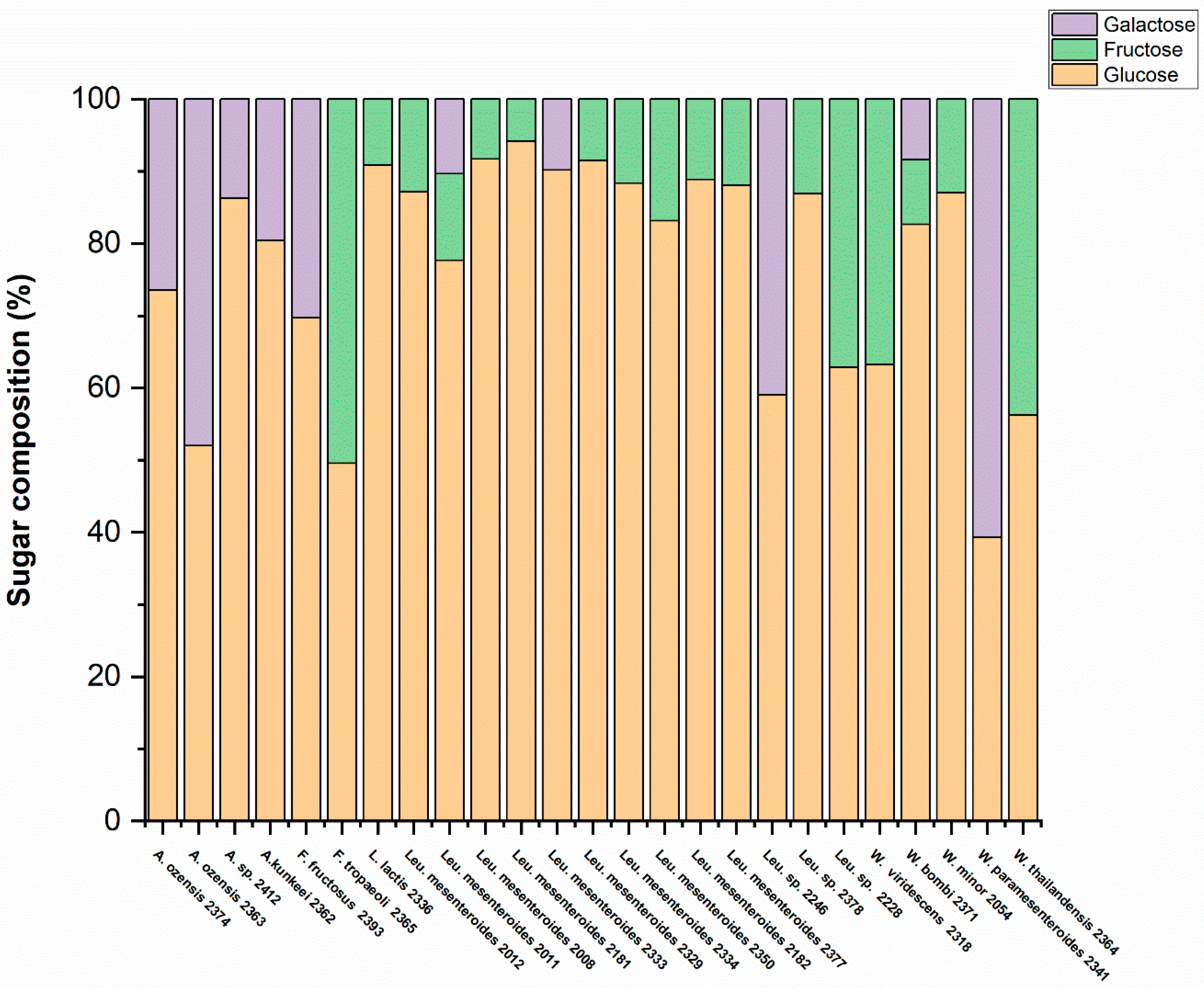
3.3. Determination of Monosaccharide Composition by HPLC Analysis
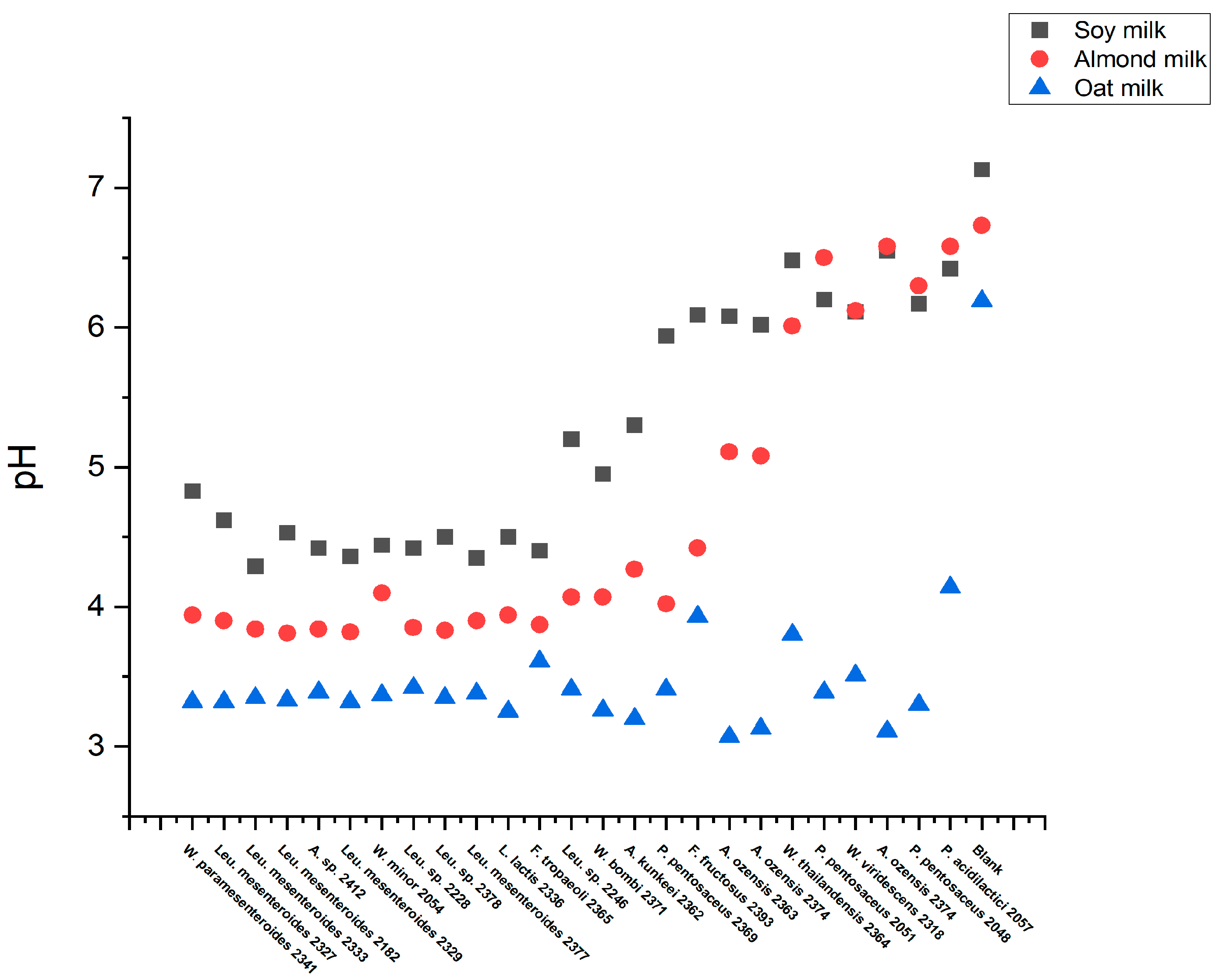
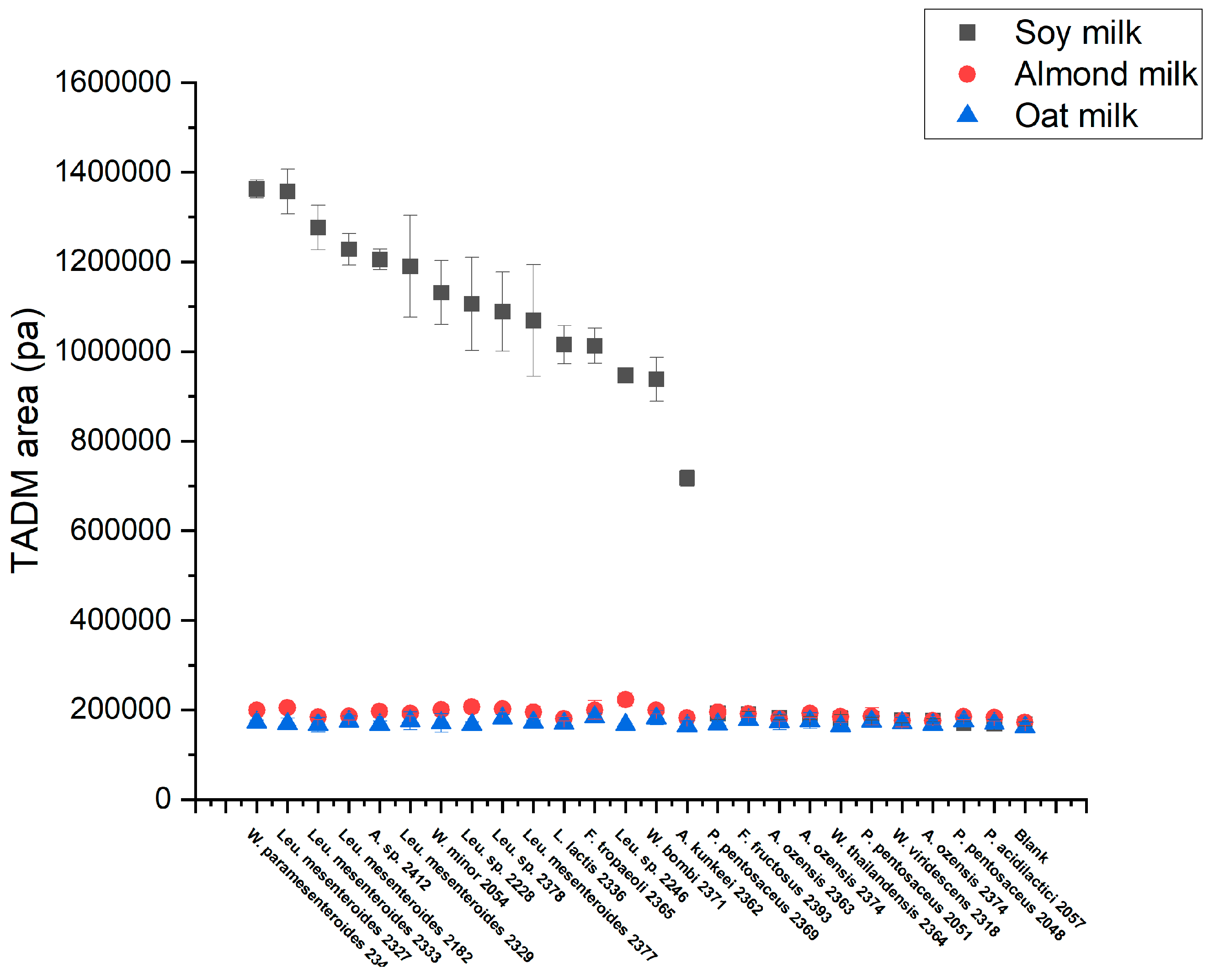
3.4. High-Throughput Screening for Texturing Strains
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lynch, K.M., A. Coffey, and E.K. Arendt, Exopolysaccharide producing lactic acid bacteria: Their techno-functional role and potential application in gluten-free bread products. Food research international 2018, 110, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Özpınar, F.B.; et al. , Physicochemical and structural characterisation of a branched dextran type exopolysaccharide (EPS) from Weissella confusa S6 isolated from fermented sausage (Sucuk). International Journal of Biological Macromolecules 2024, 130507. [Google Scholar] [CrossRef] [PubMed]
- Iosca, G.; et al. , Anti-Spoilage Activity and Exopolysaccharides Production by Selected Lactic Acid Bacteria. Foods 2022, 11, 1914. [Google Scholar] [CrossRef] [PubMed]
- Ruiz Rodríguez, L.G. , Mohamed, F., Bleckwedel, J., Medina, R., De Vuyst, L., Hebert, E.M. and Mozzi, F. Diversity and functional properties of lactic acid bacteria isolated from wild fruits and flowers present in Northern Argentina. Frontiers in microbiology 2019, 10, 1091. [Google Scholar] [CrossRef] [PubMed]
- Xiao, H.; et al. , Isolation and characterization of plant-based lactic acid bacteria from spontaneously fermented foods using a new modified medium. LWT 2023, 115695. [Google Scholar] [CrossRef]
- Wu, J.; et al. , Exopolysaccharides synthesized by lactic acid bacteria: Biosynthesis pathway, structure-function relationship, structural modification and applicability. Critical Reviews in Food Science and Nutrition 2023, 63, 7043–7064. [Google Scholar] [CrossRef] [PubMed]
- Verón, H.E.; et al. , Assessment of technological and functional features of Lactiplantibacillus and Fructobacillus strains isolated from Opuntia ficus-indica fruits. NFS Journal 2023, 31, 110–122. [Google Scholar] [CrossRef]
- Yalmanci, D., H. İspirli, and E. Dertli, Identification of Lactic Acid Bacteria (LAB) from pre-fermented liquids of selected cereals and legumes and characterization of their exopolysaccharides (EPS). Food Bioscience 2022, 50, 102014. [Google Scholar]
- Daba, G.M., M. O. Elnahas, and W.A. Elkhateeb, Contributions of exopolysaccharides from lactic acid bacteria as biotechnological tools in food, pharmaceutical, and medical applications. International Journal of Biological Macromolecules 2021, 173, 79–89. [Google Scholar] [CrossRef]
- Jurášková, D., S. C. Ribeiro, and C.C. Silva, Exopolysaccharides produced by lactic acid bacteria: From biosynthesis to health-promoting properties. Foods 2022, 11, 156. [Google Scholar] [CrossRef]
- Zeidan, A.A.; et al. , Polysaccharide production by lactic acid bacteria: From genes to industrial applications. FEMS microbiology reviews 2017, 41 (Suppl. 1), S168–S200. [Google Scholar] [CrossRef] [PubMed]
- Schmid, J., V. Sieber, and B. Rehm, Bacterial exopolysaccharides: Biosynthesis pathways and engineering strategies. Frontiers in microbiology 2015, 6, 496. [Google Scholar] [CrossRef] [PubMed]
- Kavitake, D., P. B. Devi, and P.H. Shetty, Overview of exopolysaccharides produced by Weissella genus–A review. International Journal of Biological Macromolecules 2020, 164, 2964–2973. [Google Scholar]
- Huang, W.; et al. , Evaluation of the fermentation potential of lactic acid bacteria isolated from herbs, fruits and vegetables as starter cultures in nut-based milk alternatives. Food Microbiology 2023, 112, 104243. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; et al. , Microbiological, physicochemical and rheological properties of fermented soymilk produced with exopolysaccharide (EPS) producing lactic acid bacteria strains. LWT-Food Science and Technology 2014, 57, 477–485. [Google Scholar] [CrossRef]
- Poulsen, V.K.; et al. , Screening for texturing Leuconostoc and genomics behind polysaccharide production. FEMS Microbiology Letters 2020, 367, fnaa179. [Google Scholar] [CrossRef] [PubMed]
- Molina, G.E.S.; et al. , Development of a novel lactic acid bacteria starter culture approach: From insect microbiome to plant-based fermentations. LWT 2022, 167, 113797. [Google Scholar] [CrossRef]
- Poulsen, V.K., P. Derkx, and G. Oregaard, High-throughput screening for texturing Lactococcus strains. FEMS microbiology letters 2019, 366, fnz001.
- Ruiz Rodríguez, L.G.; et al. , Diversity and functional properties of lactic acid bacteria isolated from wild fruits and flowers present in Northern Argentina. Frontiers in microbiology 2019, 10, 452267. [Google Scholar] [CrossRef] [PubMed]
- Saleh, G. , Isolation and characterization of unique fructophilic Lactic acid bacteria from different flower sources. Iraqi Journal of Agricultural Sciences 2020, 51, 508–518. [Google Scholar] [CrossRef]
- Anacarso, I.; et al. , Isolation and identification of lactic acid bacteria from plants and other vegetable matrices and microbial recombination with Enterococcus spp. Am. Res. Thoughts 2015, 1, 1503–1515. [Google Scholar]
- Terzić-Vidojević, A.; et al. , Diversity of non-starter lactic acid bacteria in autochthonous dairy products from Western Balkan Countries-technological and probiotic properties. Food Research International 2020, 136, 109494. [Google Scholar] [CrossRef] [PubMed]
- Iorizzo, M.; et al. , Inter-and intra-species diversity of lactic acid bacteria in Apis mellifera ligustica colonies. Microorganisms 2020, 8, 1578. [Google Scholar] [CrossRef] [PubMed]
- Aleklett, K., M. Hart, and A. Shade, The microbial ecology of flowers: An emerging frontier in phyllosphere research. Botany 2014, 92, 253–266. [Google Scholar] [CrossRef]
- Pimentel, T.C.; et al. , Understanding the potential of fruits, flowers, and ethnic beverages as valuable sources of techno-functional and probiotics strains: Current scenario and main challenges. Trends in Food Science & Technology 2021, 114, 25–59. [Google Scholar]
- Ruas-Madiedo, P., N. Salazar, and C.G. de los Reyes-Gavilán, Exopolysaccharides produced by lactic acid bacteria in food and probiotic applications, in Microbial glycobiology. 2010, Elsevier. p. 885-902.
- Fuso, A.; et al. , Feeding lactic acid bacteria with different sugars: Effect on exopolysaccharides (EPS) production and their molecular characteristics. Foods 2023, 12, 215. [Google Scholar] [CrossRef] [PubMed]
- Paulo, E.M.; et al. , Método alternativo de triagem de bactérias láticas produtoras de exopolissacarídeos com confirmação rápida. Food Science and Technology 2012, 32, 710–714. [Google Scholar] [CrossRef]
- Subramanian, S.B.; et al. , Extracellular polymeric substances (EPS) producing bacterial strains of municipal wastewater sludge: Isolation, molecular identification, EPS characterization and performance for sludge settling and dewatering. Water research 2010, 44, 2253–2266. [Google Scholar] [CrossRef] [PubMed]
- van Hijum, S.A.; et al. , Structure-function relationships of glucansucrase and fructansucrase enzymes from lactic acid bacteria. Microbiology and molecular biology reviews 2006, 70, 157–176. [Google Scholar] [CrossRef] [PubMed]
- Görke, B. and J. Stülke, Carbon catabolite repression in bacteria: Many ways to make the most out of nutrients. Nature Reviews Microbiology 2008, 6, 613–624. [Google Scholar] [CrossRef]
- Gangoiti, J., T. Pijning, and L. Dijkhuizen, Biotechnological potential of novel glycoside hydrolase family 70 enzymes synthesizing α-glucans from starch and sucrose. Biotechnology advances 2018, 36, 196–207. [Google Scholar] [CrossRef]
- Angelin, J. and M. Kavitha, Exopolysaccharides from probiotic bacteria and their health potential. International Journal of Biological Macromolecules 2020, 162, 853–865. [Google Scholar] [PubMed]
- Ayyash, M.; et al. , Physicochemical, bioactive and rheological properties of an exopolysaccharide produced by a probiotic Pediococcus pentosaceus M41. Carbohydrate polymers 2020, 229, 115462. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, M.T.; et al. , Characterisation of dextran AP-27 produced by bee pollen isolate Lactobacillus kunkeei AP-27. Process Biochemistry 2023, 129, 22–29. [Google Scholar] [CrossRef]
- Poulsen, V.K.; et al. , Versatile Lactococcus lactis strains improve texture in both fermented milk and soybean matrices. FEMS Microbiology Letters 2022, 369, fnac117. [Google Scholar] [CrossRef]
- Erkus, O.; et al. , Multifactorial diversity sustains microbial community stability. The ISME journal 2013, 7, 2126–2136. [Google Scholar] [CrossRef]

| Oat, Isola Bio | Almond, Ecomil | Soy, Naturli | |
|---|---|---|---|
| Energy | 215 kJ / 51 kcal | 134.00 kJ / 32.00 kcal | 153kJ/37 kcal |
| Fat | 1.0 g | 2.10 g | 2.1 g |
| Carbohydrate | 9.0 g | 3.30 g | 0.6 g |
| -Here sugar | 4.2 g | <0.30 g | 0.6 g |
| Protein | 1.0 g | 1.00 g | 3.7 g |
| Salt | 0.08 g | 0.14 g | 0.09 g |
| No. | Species | NFICC ID | City | Flower |
|---|---|---|---|---|
| 1 | Apilactobacillus kunkeei | 2373 | Odense | Verbascum |
| 2 | Apilactobacillus kunkeei | 2324 | Copenhagen | Rosa |
| 3 | Apilactobacillus kunkeei | 2359 | Aarhus | Fallopia |
| 4 | Apilactobacillus kunkeei | 2362 | Silkeborg | Rubus |
| 5 | Apilactobacillus ozensis | 2370 | Odense | Potentilla |
| 6 | Apilactobacillus ozensis | 2374 | Odense | Jacobaea |
| 7 | Apilactobacillus ozensis | 2363 | Silkeborg | Rubus |
| 8 | Apilactobacillus ozensis | 2366 | Silkeborg | Lavandula |
| 9 | Apilactobacillus ozensis | 2368 | Odense | Jacobaea |
| 10 | Apilactobacillus sp. | 2412 | Silkeborg | Lavandula |
| 11 | Fructobacillus fructosus | 2323 | Copenhagen | Rosa |
| 12 | Fructobacillus fructosus | 2321 | Copenhagen | Geranium |
| 13 | Fructobacillus fructosus | 2361 | Silkeborg | Rubus |
| 14 | Fructobacillus fructosus | 2347 | Copenhagen | Rosa |
| 15 | Fructobacillus fructosus | 2325 | Copenhagen | Rubus |
| 16 | Fructobacillus fructosus | 2319 | Copenhagen | Syringa |
| 17 | Fructobacillus fructosus | 2376 | Odense | Convolvulus |
| 18 | Fructobacillus fructosus | 2393 | Silkeborg | Lavandula |
| 19 | Fructobacillus pseudoficulneus | 2222 | Aarhus | Betonica |
| 20 | Fructobacillus tropaeoli | 2365 | Silkeborg | Lavandula |
| 21 | Levilactobacillus brevis | 2055 | Aarhus | Eschscholzia |
| 22 | Lactiplantibacillus paraplantarum | 2184 | Odense | Cirsium |
| 23 | Lactiplantibacillus pentosus | 2185 | Odense | Cirsium |
| 24 | Lactiplantibacillus plantarum | 2183 | Odense | Cirsium |
| 25 | Latilactobacillus sakei | 2317 | Copenhagen | Stellaria |
| 26 | Lactococcus garvieae | 2331 | Copenhagen | Hyacinthus |
| 27 | Lactococcus lactis | 2358 | Aarhus | Erythranthe |
| 28 | Lactococcus lactis | 2332 | Copenhagen | Hyacinthus |
| 29 | Lactococcus lactis | 2336 | Copenhagen | Trifolium |
| 30 | Leuconostoc mesenteroides | 2012 | Copenhagen | Forsythia |
| 31 | Leuconostoc mesenteroides | 2011 | Copenhagen | Prunus |
| 32 | Leuconostoc mesenteroides | 2008 | Copenhagen | Aubrieta |
| 33 | Leuconostoc mesenteroides | 2181 | Aarhus | Telekia |
| 34 | Leuconostoc mesenteroides | 2333 | Copenhagen | Prunus |
| 35 | Leuconostoc mesenteroides | 2343 | Copenhagen | Trifolium |
| 36 | Leuconostoc mesenteroides | 2329 | Copenhagen | Ranunculus |
| 37 | Leuconostoc mesenteroides | 2327 | Copenhagen | Hottonia |
| 38 | Leuconostoc mesenteroides | 2334 | Copenhagen | Trifolium |
| 39 | Leuconostoc mesenteroides | 2350 | Copenhagen | Bellis |
| 40 | Leuconostoc mesenteroides | 2182 | Aarhus | Telekia |
| 41 | Leuconostoc mesenteroides | 2377 | Aarhus | Campanula |
| 42 | Leuconostoc miyukkimchii | 2224 | Copenhagen | Rosa |
| 43 | Leuconostoc sp. | 2228 | Aarhus | Dasiphora |
| 44 | Leuconostoc sp. | 2378 | Aarhus | Black mullein |
| 45 | Leuconostoc sp. THK-X10 | 2246 | Copenhagen | Rosa |
| 46 | Pediococcus acidilactici | 2053 | Aarhus | Betonica |
| 47 | Pediococcus acidilactici | 2057 | Aarhus | Fuchsia |
| 48 | Pediococcus acidilactici | 2357 | Aarhus | Erythranthe |
| 49 | Pediococcus pentosaceus | 2051 | Aarhus | Hydrangea |
| 50 | Pediococcus pentosaceus | 2048 | Aarhus | Dasiphora |
| 51 | Pediococcus pentosaceus | 2379 | Aarhus | Teucrium |
| 52 | Pediococcus pentosaceus | 2369 | Odense | Artemisia |
| 53 | Weissella bombi | 2356 | Aarhus | Erythranthe |
| 54 | Weissella bombi | 2346 | Aarhus | Teucrium |
| 55 | Weissella bombi | 2371 | Odense | Agastache |
| 56 | Weissella minor | 2054 | Aarhus | Betonica |
| 57 | Weissella paramesenteroides | 2341 | Copenhagen | Trifolium |
| 58 | Weissella thailandensis | 2056 | Odense | Convolvulus |
| 59 | Weissella thailandensis | 2364 | Silkeborg | Lavandula |
| 60 | Weissella viridescens | 2320 | Copenhagen | Cardamine |
| 61 | Weissella viridescens | 2318 | Copenhagen | Cotoneaster |
| NFICC codes | Species | 2% Sucrose | 2% Sucrose and 1% Glucose | ||
|---|---|---|---|---|---|
| Slimy | Ropy | Slimy | Ropy | ||
| 2324 | Apilactobacillus kunkeei | - | + | + | + |
| 2362 | Apilactobacillus kunkeei | + | + | ++ | - |
| 2370 | Apilactobacillus ozensis | ++ | + | - | + |
| 2374 | Apilactobacillus ozensis | +++ | - | ++ | - |
| 2363 | Apilactobacillus ozensis | - | + | - | + |
| 2366 | Apilactobacillus ozensis | - | +++ | - | ++ |
| 2347 | Fructobacillus fructosus | + | - | - | + |
| 2185 | Lactobacillus pentosus | +- | - | - | - |
| 2183 | Lactiplantibacillus plantarum | +- | - | - | - |
| 2331 | Lactococcus garvieae | +- | + | +- | + |
| 2336 | Lactococcus lactis | + | ++ | ++ | + |
| 2012 | Leuconostoc mesenteroides | - | + | ++ | +- |
| 2011 | Leuconostoc mesenteroides | +- | +- | +- | + |
| 2008 | Leuconostoc mesenteroides | - | + | + | + |
| 2181 | Leuconostoc mesenteroides | +- | - | ++ | - |
| 2333 | Leuconostoc mesenteroides | + | + | ++ | + |
| 2329 | Leuconostoc mesenteroides | + | - | + | - |
| 2327 | Leuconostoc mesenteroides | + | - | + | - |
| 2334 | Leuconostoc mesenteroides | +- | - | + | - |
| 2350 | Leuconostoc mesenteroides | - | +- | - | - |
| 2182 | Leuconostoc mesenteroides | + | - | ++ | - |
| 2224 | Leuconostoc miyukkimchii | - | + | + | ++ |
| 2228 | Leuconostoc sp. | + | + | + | + |
| 2246 | Leuconostoc sp. THK-X10 | + | - | + | + |
| 2057 | Pediococcus acidilactici | - | +++ | - | +++ |
| 2051 | Pediococcus pentosaceus | + | - | - | - |
| 2048 | Pediococcus pentosaceus | + | + | - | - |
| 2369 | Pediococcus pentosaceus | - | +++ | - | +++ |
| 2379 | Pediococcus pentosaceus | - | + | - | + |
| 2371 | Weissella bombi | ++ | - | ++ | - |
| 2054 | Weissella minor | + | + | ++ | + |
| 2341 | Weissella paramesenteroides | + | + | + | + |
| 2056 | Weissella thailandensis | + | - | ++ | - |
| 2318 | Weissella viridescens | + | - | - | +- |
| 2393 | Fructobacillus fructosus | + | - | - | - |
| 2378 | Leuconostoc sp. | + | - | ++ | + |
| 2377 | Leuconostoc mesenteroides | + | - | + | + |
| 2412 | Apilactobacillus sp. | + | - | - | - |
| 2365 | Fructobacillus tropaeoli | + | - | - | - |
| 2364 | Weissella thailandensis | + | - | - | ++ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




