Submitted:
28 July 2024
Posted:
30 July 2024
You are already at the latest version
Abstract
Keywords:
Introduction
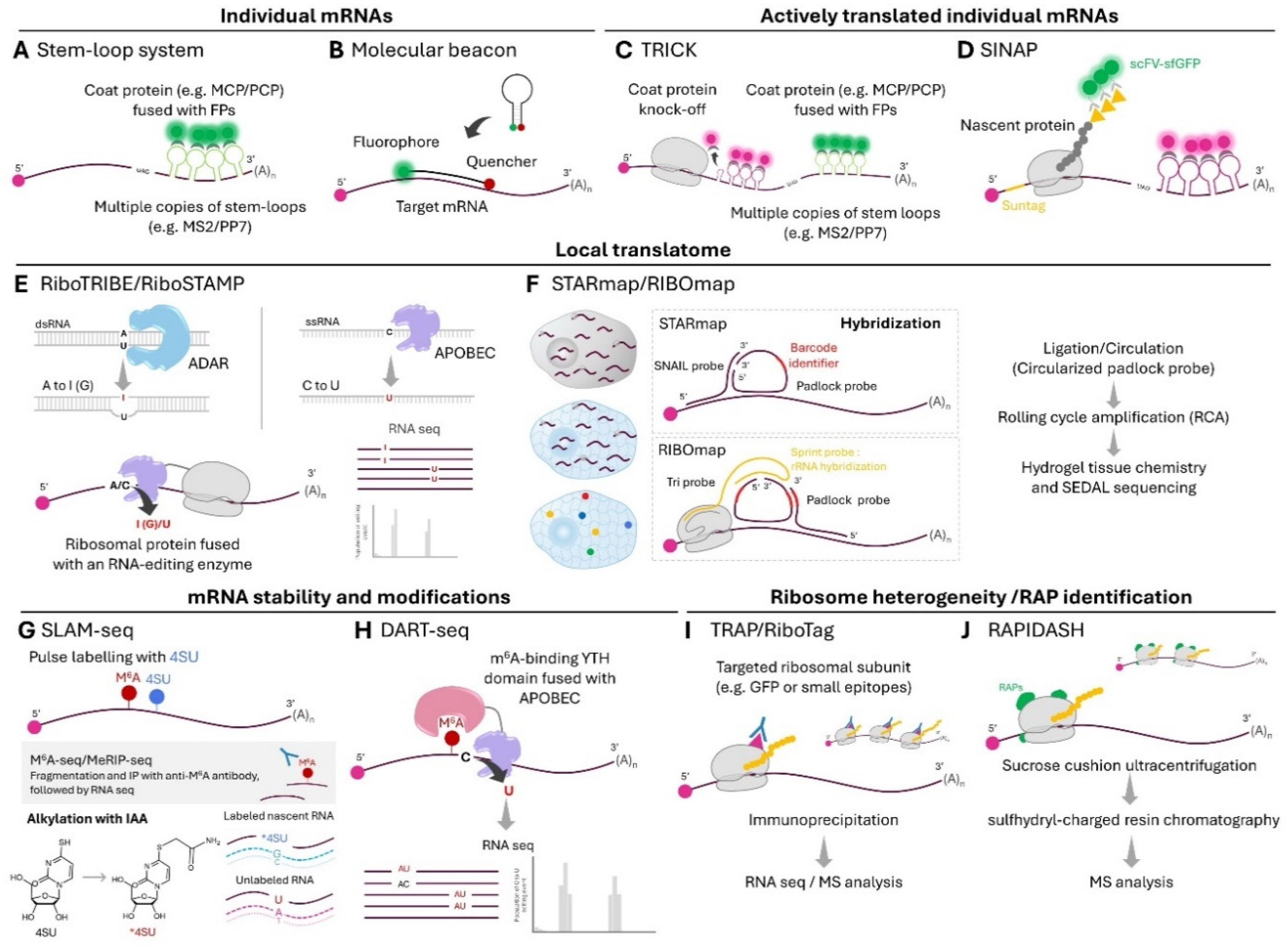
Imaging Actively Translated Individual mRNAs
Mapping Local Translatomes
Mapping Global Turnover and Associated Regulatory Elements of Localized mRNAs
Local Translation of Ribosomal Proteins and Local Remodeling of Ribosomes
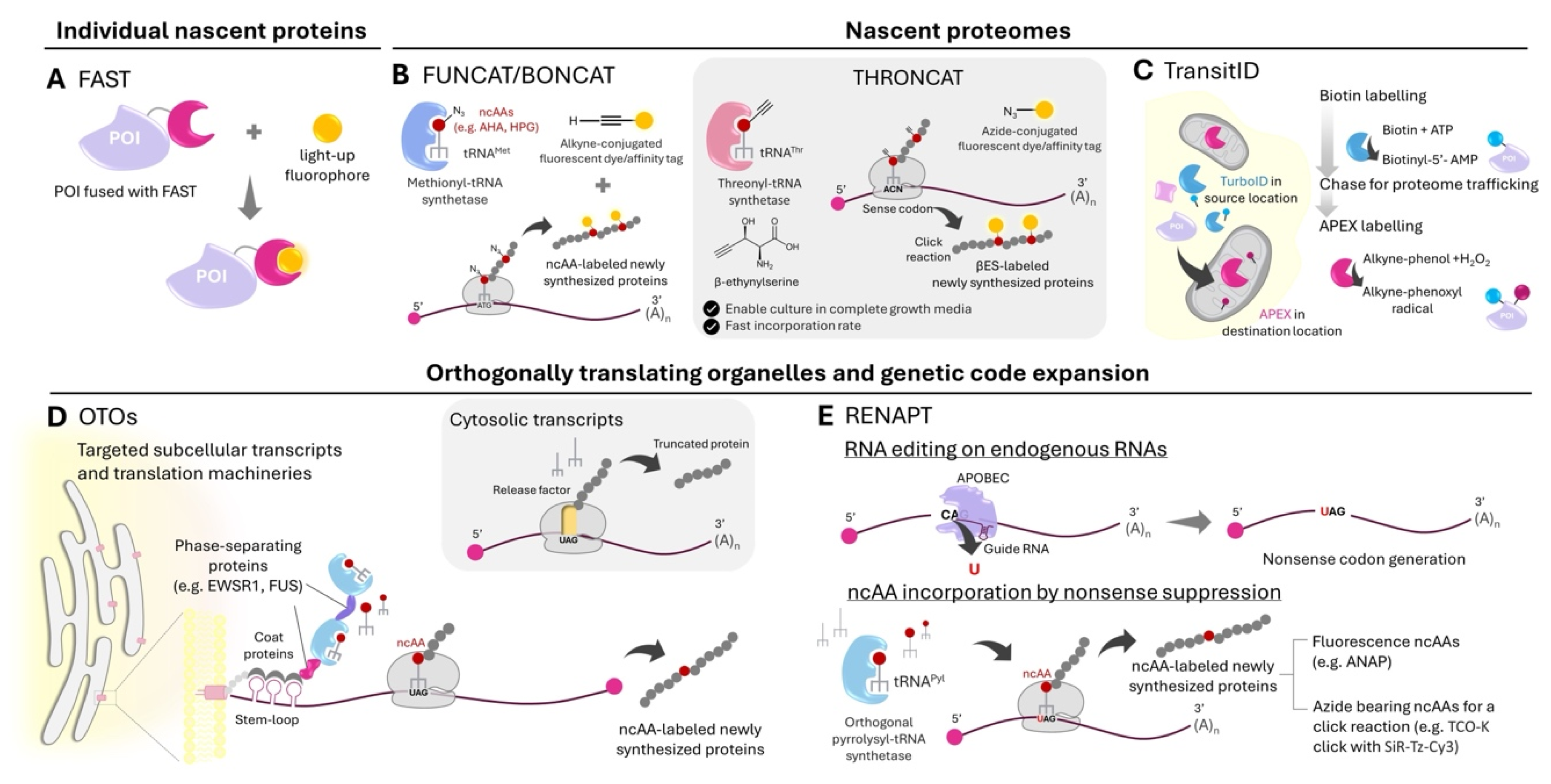
Imaging Newly Synthesized Individual Proteins
Mapping Local Nascent Proteomes and Imaging Nascent Proteins
Controlling and Monitoring Local Translation via Orthogonally Translating Organelles and Genetic Code Expansion
Controlling Translation via mRNA-Specific Ribosomes in Eukaryotes
Outlook of Local Translation Imaging and Controlling Tools for Mammalian Synthetic Biology
Acknowledgements
Conflict of Interest Statement
References and Recommended Reading
- Papers of particular interest, published within the period of review (2021-2024), have been highlighted as: * of special interest ** of outstanding interest.
- 1. Liao G, Mingle L, Van De Water L, Liu G: Control of cell migration through mRNA localization and local translation. Wiley Interdiscip Rev RNA 2015, 6:1-15.
- 2. Wang DO, Martin KC, Zukin RS: Spatially restricting gene expression by local translation at synapses. Trends Neurosci 2010, 33:173-182.
- 3. Medioni C, Mowry K, Besse F: Principles and roles of mRNA localization in animal development. Development 2012, 139:3263-3276.
- 4. Mountford PS, Smith AG: Internal ribosome entry sites and dicistronic RNAs in mammalian transgenesis. Trends Genet 1995, 11:179-184.
- 5. de Felipe P, Hughes LE, Ryan MD, Brown JD: Co-translational, intraribosomal cleavage of polypeptides by the foot-and-mouth disease virus 2A peptide. J Biol Chem 2003, 278:11441-11448.
- 6. Orr MW, Mao Y, Storz G, Qian S-B: Alternative ORFs and small ORFs: shedding light on the dark proteome. Nucleic Acids Research 2019, 48:1029-1042.
- 7. Shandell MA, Tan Z, Cornish VW: Genetic Code Expansion: A Brief History and Perspective. Biochemistry 2021, 60:3455-3469.
- 8. de la Torre D, Chin JW: Reprogramming the genetic code. Nat Rev Genet 2021, 22:169-184.
- 9. Liu CC, Schultz PG: Adding New Chemistries to the Genetic Code. Annual Review of Biochemistry 2010, 79:413-444.
- 10. Anzalone AV, Lin AJ, Zairis S, Rabadan R, Cornish VW: Reprogramming eukaryotic translation with ligand-responsive synthetic RNA switches. Nature Methods 2016, 13:453-458.
- 11. Holt CE, Martin KC, Schuman EM: Local translation in neurons: visualization and function. Nature Structural & Molecular Biology 2019, 26:557-566.
- 12. Taylor AM, Wu J, Tai HC, Schuman EM: Axonal translation of β-catenin regulates synaptic vesicle dynamics. J Neurosci 2013, 33:5584-5589.
- 13. Cagnetta R, Frese CK, Shigeoka T, Krijgsveld J, Holt CE: Rapid Cue-Specific Remodeling of the Nascent Axonal Proteome. Neuron 2018, 99:29-46.e24.
- 14. Aakalu G, Smith WB, Nguyen N, Jiang C, Schuman EM: Dynamic visualization of local protein synthesis in hippocampal neurons. Neuron 2001, 30:489-502.
- 15. Cajigas IJ, Tushev G, Will TJ, tom Dieck S, Fuerst N, Schuman EM: The local transcriptome in the synaptic neuropil revealed by deep sequencing and high-resolution imaging. Neuron 2012, 74:453-466.
- 16. Perez JD, Dieck ST, Alvarez-Castelao B, Tushev G, Chan IC, Schuman EM: Subcellular sequencing of single neurons reveals the dendritic transcriptome of GABAergic interneurons. Elife 2021, 10.
- 17. Glock C, Biever A, Tushev G, Nassim-Assir B, Kao A, Bartnik I, Tom Dieck S, Schuman EM: The translatome of neuronal cell bodies, dendrites, and axons. Proc Natl Acad Sci U S A 2021, 118.
- *18. Biever A, Glock C, Tushev G, Ciirdaeva E, Dalmay T, Langer JD, Schuman EM: Monosomes actively translate synaptic mRNAs in neuronal processes. Science 2020, 367.
- Polysome profiling of microdissected regions to determine association of axonal, dendritic, and somatic transcripts with monosomes or polysomes. Discovery that monosomes actively translate synaptic mRNAs in neuronal processes.
- **19. Fusco CM, Desch K, Dörrbaum AR, Wang M, Staab A, Chan ICW, Vail E, Villeri V, Langer JD, Schuman EM: Neuronal ribosomes exhibit dynamic and context-dependent exchange of ribosomal proteins. Nature Communications 2021, 12:6127.
- Local synthesis of ribosomal proteins and local ribosome remodeling in dendrites. Ribosomal protein incorporation into ribosomes is regulated in part by oxidative stress.
- **20. Shigeoka T, Koppers M, Wong HH, Lin JQ, Cagnetta R, Dwivedy A, de Freitas Nascimento J, van Tartwijk FW, Ströhl F, Cioni JM, et al.: On-Site Ribosome Remodeling by Locally Synthesized Ribosomal Proteins in Axons. Cell Rep 2019, 29:3605-3619.e3610.
- Local synthesis of ribosomal proteins and local ribosome remodeling in axons, which are needed for the maintenance of axon branching in vivo. Identification of a sequence motif for axonal ribosomal protein translation.
- 21. Clifton NE, Lin JQ, Holt CE, O’Donovan MC, Mill J: Enrichment of the Local Synaptic Translatome for Genetic Risk Associated With Schizophrenia and Autism Spectrum Disorder. Biol Psychiatry 2024, 95:888-895.
- 22. Hale CR, Sawicka K, Mora K, Fak JJ, Kang JJ, Cutrim P, Cialowicz K, Carroll TS, Darnell RB: FMRP regulates mRNAs encoding distinct functions in the cell body and dendrites of CA1 pyramidal neurons. Elife 2021, 10.
- 23. Meservey LM, Topkar VV, Fu MM: mRNA Transport and Local Translation in Glia. Trends Cell Biol 2021, 31:419-423.
- 24. Berkovits BD, Mayr C: Alternative 3′ UTRs act as scaffolds to regulate membrane protein localization. Nature 2015, 522:363-367.
- 25. Fasken MB, Corbett AH: Mechanisms of nuclear mRNA quality control. RNA Biology 2009, 6:237-241.
- 26. Bertrand E, Chartrand P, Schaefer M, Shenoy SM, Singer RH, Long RM: Localization of ASH1 mRNA particles in living yeast. Mol Cell 1998, 2:437-445.
- 27. Chao JA, Patskovsky Y, Almo SC, Singer RH: Structural basis for the coevolution of a viral RNA–protein complex. Nature Structural & Molecular Biology 2008, 15:103-105.
- 28. Le P, Ahmed N, Yeo GW: Illuminating RNA biology through imaging. Nature Cell Biology 2022, 24:815-824.
- 29. Yin P, Kuang S, Nie Z: Fluorescent RNA Tags for In Situ RNA Imaging in Living Cells. Analysis & Sensing 2023, 3:e202200090.
- 30. Das S, Vera M, Gandin V, Singer RH, Tutucci E: Intracellular mRNA transport and localized translation. Nat Rev Mol Cell Biol 2021, 22:483-504.
- 31. Tutucci E, Vera M, Biswas J, Garcia J, Parker R, Singer RH: An improved MS2 system for accurate reporting of the mRNA life cycle. Nature Methods 2018, 15:81-89.
- 32. Li W, Maekiniemi A, Sato H, Osman C, Singer RH: An improved imaging system that corrects MS2-induced RNA destabilization. Nature Methods 2022, 19:1558-1562.
- 33. Ohishi H, Shimada S, Uchino S, Li J, Sato Y, Shintani M, Owada H, Ohkawa Y, Pertsinidis A, Yamamoto T, et al.: STREAMING-tag system reveals spatiotemporal relationships between transcriptional regulatory factors and transcriptional activity. Nature Communications 2022, 13:7672.
- 34. Tyagi S, Kramer FR: Molecular Beacons: Probes that Fluoresce upon Hybridization. Nature Biotechnology 1996, 14:303-308.
- 35. Donlin-Asp PG, Polisseni C, Klimek R, Heckel A, Schuman EM: Differential regulation of local mRNA dynamics and translation following long-term potentiation and depression. Proceedings of the National Academy of Sciences 2021, 118:e2017578118.
- 36. Abudayyeh OO, Gootenberg JS, Essletzbichler P, Han S, Joung J, Belanto JJ, Verdine V, Cox DBT, Kellner MJ, Regev A, et al.: RNA targeting with CRISPR-Cas13. Nature 2017, 550:280-284.
- 37. Han Y, Branon TC, Martell JD, Boassa D, Shechner D, Ellisman MH, Ting A: Directed Evolution of Split APEX2 Peroxidase. ACS Chemical Biology 2019, 14:619-635.
- 38. Halstead JM, Lionnet T, Wilbertz JH, Wippich F, Ephrussi A, Singer RH, Chao JA: Translation. An RNA biosensor for imaging the first round of translation from single cells to living animals. Science 2015, 347:1367-1671.
- 39. Tanenbaum ME, Gilbert LA, Qi LS, Weissman JS, Vale RD: A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 2014, 159:635-646.
- 40. Wang C, Han B, Zhou R, Zhuang X: Real-Time Imaging of Translation on Single mRNA Transcripts in Live Cells. Cell 2016, 165:990-1001.
- 41. Wu B, Eliscovich C, Yoon YJ, Singer RH: Translation dynamics of single mRNAs in live cells and neurons. Science 2016, 352:1430-1435.
- 42. Yan X, Hoek TA, Vale RD, Tanenbaum ME: Dynamics of Translation of Single mRNA Molecules In Vivo. Cell 2016, 165:976-989.
- 43. Mateju D, Eichenberger B, Voigt F, Eglinger J, Roth G, Chao JA: Single-Molecule Imaging Reveals Translation of mRNAs Localized to Stress Granules. Cell 2020, 183:1801-1812.e1813.
- 44. Ingolia NT, Ghaemmaghami S, Newman JR, Weissman JS: Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 2009, 324:218-223.
- 45. Jan CH, Williams CC, Weissman JS: Principles of ER cotranslational translocation revealed by proximity-specific ribosome profiling. Science 2014, 346:1257521.
- 46. Williams CC, Jan CH, Weissman JS: Targeting and plasticity of mitochondrial proteins revealed by proximity-specific ribosome profiling. Science 2014, 346:748-751.
- 47. Xu W, Abruzzi K, Rosbash M: RiboTRIBE: Monitoring Translation with ADAR-meditated RNA Editing. bioRxiv 2021:2021.2006.2020.449184.
- 48. Brannan KW, Chaim IA, Marina RJ, Yee BA, Kofman ER, Lorenz DA, Jagannatha P, Dong KD, Madrigal AA, Underwood JG, et al.: Robust single-cell discovery of RNA targets of RNA-binding proteins and ribosomes. Nat Methods 2021, 18:507-519.
- **49. Zeng H, Huang J, Ren J, Wang CK, Tang Z, Zhou H, Zhou Y, Shi H, Aditham A, Sui X, et al.: Spatially resolved single-cell translatomics at molecular resolution. Science 2023, 380:eadd3067.
- A massively multiplexed platform to assess translation, based on clever probe design that captures only ribosome-bound mRNAs of interests and barcode-based detection with in situ sequencing.
- 50. Zeng H, Huang J, Zhou H, Meilandt WJ, Dejanovic B, Zhou Y, Bohlen CJ, Lee S-H, Ren J, Liu A, et al.: Integrative in situ mapping of single-cell transcriptional states and tissue histopathology in a mouse model of Alzheimer’s disease. Nature Neuroscience 2023, 26:430-446.
- 51. Wang X, Allen WE, Wright MA, Sylwestrak EL, Samusik N, Vesuna S, Evans K, Liu C, Ramakrishnan C, Liu J, et al.: Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 2018, 361:eaat5691.
- 52. Kislauskis EH, Zhu X, Singer RH: Sequences responsible for intracellular localization of beta-actin messenger RNA also affect cell phenotype. J Cell Biol 1994, 127:441-451.
- *53. Loedige I, Baranovskii A, Mendonsa S, Dantsuji S, Popitsch N, Breimann L, Zerna N, Cherepanov V, Milek M, Ameres S, et al.: mRNA stability and m(6)A are major determinants of subcellular mRNA localization in neurons. Mol Cell 2023, 83:2709-2725.e2710.
- The paper used SLAM-seq to characterize expression dynamics and global stability of RNAs, and found correlation between mRNA stability and their propensity to trafficking to distal neuronal processes.
- 54. Chen CYA, Ezzeddine N, Shyu AB: Chapter 17 Messenger RNA Half-Life Measurements in Mammalian Cells. In Methods in Enzymology. Edited by: Academic Press; 2008:335-357. vol 448.].
- 55. Herzog VA, Reichholf B, Neumann T, Rescheneder P, Bhat P, Burkard TR, Wlotzka W, von Haeseler A, Zuber J, Ameres SL: Thiol-linked alkylation of RNA to assess expression dynamics. Nat Methods 2017, 14:1198-1204.
- 56. Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, Cesarkas K, Jacob-Hirsch J, Amariglio N, Kupiec M, et al.: Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 2012, 485:201-206.
- 57. Meyer KD: DART-seq: an antibody-free method for global m6A detection. Nature Methods 2019, 16:1275-1280.
- 58. Poon MM, Choi SH, Jamieson CA, Geschwind DH, Martin KC: Identification of process-localized mRNAs from cultured rodent hippocampal neurons. J Neurosci 2006, 26:13390-13399.
- 59. Yang YM, Karbstein K: The chaperone Tsr2 regulates Rps26 release and reincorporation from mature ribosomes to enable a reversible, ribosome-mediated response to stress. Sci Adv 2022, 8:eabl4386.
- 60. Shi Z, Fujii K, Kovary KM, Genuth NR, Röst HL, Teruel MN, Barna M: Heterogeneous Ribosomes Preferentially Translate Distinct Subpools of mRNAs Genome-wide. Mol Cell 2017, 67:71-83.e77.
- 61. Ferretti MB, Ghalei H, Ward EA, Potts EL, Karbstein K: Rps26 directs mRNA-specific translation by recognition of Kozak sequence elements. Nat Struct Mol Biol 2017, 24:700-707.
- 62. Xue S, Tian S, Fujii K, Kladwang W, Das R, Barna M: RNA regulons in Hox 5′ UTRs confer ribosome specificity to gene regulation. Nature 2015, 517:33-38.
- 63. Majzoub K, Hafirassou ML, Meignin C, Goto A, Marzi S, Fedorova A, Verdier Y, Vinh J, Hoffmann JA, Martin F, et al.: RACK1 controls IRES-mediated translation of viruses. Cell 2014, 159:1086-1095.
- 64. Heiman M, Kulicke R, Fenster RJ, Greengard P, Heintz N: Cell type–specific mRNA purification by translating ribosome affinity purification (TRAP). Nature Protocols 2014, 9:1282-1291.
- 65. Simsek D, Tiu GC, Flynn RA, Byeon GW, Leppek K, Xu AF, Chang HY, Barna M: The Mammalian Ribo-interactome Reveals Ribosome Functional Diversity and Heterogeneity. Cell 2017, 169:1051-1065.e1018.
- 66. Ferretti MB, Karbstein K: Does functional specialization of ribosomes really exist? Rna 2019, 25:521-538.
- *67. Susanto TT, Hung V, Levine AG, Kerr CH, Yoo Y, Chen Y, Oses-Prieto JA, Fromm L, Fujii K, Wernig M, et al.: RAPIDASH: A tag-free enrichment of ribosome-associated proteins reveals compositional dynamics in embryonic tissues and stimulated macrophages. bioRxiv 2023:2023.2012.2007.570613.
- An enabling technique for the isolation of native ribosomes while preserving weak interactions of ribosome-associated proteins.
- 68. Georgeson J, Schwartz S: The ribosome epitranscriptome: inert-or a platform for functional plasticity? Rna 2021, 27:1293-1301.
- 69. Wang DO, Kim SM, Zhao Y, Hwang H, Miura SK, Sossin WS, Martin KC: Synapse- and stimulus-specific local translation during long-term neuronal plasticity. Science 2009, 324:1536-1540.
- 70. Wong HH, Lin JQ, Ströhl F, Roque CG, Cioni JM, Cagnetta R, Turner-Bridger B, Laine RF, Harris WA, Kaminski CF, et al.: RNA Docking and Local Translation Regulate Site-Specific Axon Remodeling In Vivo. Neuron 2017, 95:852-868.e858.
- 71. Ando R, Shimozono S, Ago H, Takagi M, Sugiyama M, Kurokawa H, Hirano M, Niino Y, Ueno G, Ishidate F, et al.: StayGold variants for molecular fusion and membrane-targeting applications. Nature Methods 2023.
- 72. Ivorra-Molla E, Akhuli D, McAndrew MBL, Scott W, Kumar L, Palani S, Mishima M, Crow A, Balasubramanian MK: A monomeric StayGold fluorescent protein. Nature Biotechnology 2023.
- 73. Zhang H, Lesnov GD, Subach OM, Zhang W, Kuzmicheva TP, Vlaskina AV, Samygina VR, Chen L, Ye X, Nikolaeva AY, et al.: Bright and stable monomeric green fluorescent protein derived from StayGold. Nature Methods 2024.
- 74. Zhang D, Chen Z, Du Z, Bao B, Su N, Chen X, Ge Y, Lin Q, Yang L, Hua Y, et al.: Design of a palette of SNAP-tag mimics of fluorescent proteins and their use as cell reporters. Cell Discovery 2023, 9:56.
- 75. Frei MS, Tarnawski M, Roberti MJ, Koch B, Hiblot J, Johnsson K: Engineered HaloTag variants for fluorescence lifetime multiplexing. Nature Methods 2022, 19:65-70.
- 76. Gautier A: Fluorescence-Activating and Absorption-Shifting Tags for Advanced Imaging and Biosensing. Accounts of Chemical Research 2022, 55:3125-3135.
- 77. Plamont M-A, Billon-Denis E, Maurin S, Gauron C, Pimenta FM, Specht CG, Shi J, Quérard J, Pan B, Rossignol J, et al.: Small fluorescence-activating and absorption-shifting tag for tunable protein imaging in vivo. Proceedings of the National Academy of Sciences 2016, 113:497-502.
- *78. Benaissa H, Ounoughi K, Aujard I, Fischer E, Goïame R, Nguyen J, Tebo AG, Li C, Le Saux T, Bertolin G, et al.: Engineering of a fluorescent chemogenetic reporter with tunable color for advanced live-cell imaging. Nature Communications 2021, 12:6989.
- The paper described the development of promiscuous FAST variants which can bind to different fluorogenic chromophores covering the visible spectrum.
- 79. Kang H, Schuman EM: A Requirement for Local Protein Synthesis in Neurotrophin-Induced Hippocampal Synaptic Plasticity. Science 1996, 273:1402-1406.
- 80. Martin KC, Casadio A, Zhu H, E Y, Rose JC, Chen M, Bailey CH, Kandel ER: Synapse-Specific, Long-Term Facilitation of Aplysia Sensory to Motor Synapses: A Function for Local Protein Synthesis in Memory Storage. Cell 1997, 91:927-938.
- 81. Mineev KS, Goncharuk SA, Goncharuk MV, Povarova NV, Sokolov AI, Baleeva NS, Smirnov AY, Myasnyanko IN, Ruchkin DA, Bukhdruker S, et al.: NanoFAST: structure-based design of a small fluorogen-activating protein with only 98 amino acids. Chemical Science 2021, 12:6719-6725.
- 82. Tang Q, Chen X: Nascent Proteomics: Chemical Tools for Monitoring Newly Synthesized Proteins. Angewandte Chemie International Edition 2023, 62:e202305866.
- 83. Stone SE, Glenn WS, Hamblin GD, Tirrell DA: Cell-selective proteomics for biological discovery. Current Opinion in Chemical Biology 2017, 36:50-57.
- 84. Beatty KE, Liu JC, Xie F, Dieterich DC, Schuman EM, Wang Q, Tirrell DA: Fluorescence Visualization of Newly Synthesized Proteins in Mammalian Cells. Angewandte Chemie International Edition 2006, 45:7364-7367.
- 85. Dieterich DC, Link AJ, Graumann J, Tirrell DA, Schuman EM: Selective identification of newly synthesized proteins in mammalian cells using bioorthogonal noncanonical amino acid tagging (BONCAT). Proceedings of the National Academy of Sciences 2006, 103:9482-9487.
- 86. Sun C, Nold A, Fusco CM, Rangaraju V, Tchumatchenko T, Heilemann M, Schuman EM: The prevalence and specificity of local protein synthesis during neuronal synaptic plasticity. Science Advances 2021, 7:eabj0790.
- *87. Alvarez-Castelao B, Schanzenbächer CT, Hanus C, Glock C, tom Dieck S, Dörrbaum AR, Bartnik I, Nassim-Assir B, Ciirdaeva E, Mueller A, et al.: Cell-type-specific metabolic labeling of nascent proteomes in vivo. Nature Biotechnology 2017, 35:1196-1201.
- Cell-type specific metabolic labeling of nascent proteomes using a genetically targetable mutant methionyl-tRNA synthetase.
- *88. Ignacio BJ, Dijkstra J, Mora N, Slot EFJ, van Weijsten MJ, Storkebaum E, Vermeulen M, Bonger KM: THRONCAT: metabolic labeling of newly synthesized proteins using a bioorthogonal threonine analog. Nature Communications 2023, 14:3367.
- A new metabolic labeling approach for newly synthesized proteins based on a threonine analog.
- 89. Elliott TS, Townsley FM, Bianco A, Ernst RJ, Sachdeva A, Elsässer SJ, Davis L, Lang K, Pisa R, Greiss S, et al.: Proteome labeling and protein identification in specific tissues and at specific developmental stages in an animal. Nat Biotechnol 2014, 32:465-472.
- 90. tom Dieck S, Kochen L, Hanus C, Heumüller M, Bartnik I, Nassim-Assir B, Merk K, Mosler T, Garg S, Bunse S, et al.: Direct visualization of newly synthesized target proteins in situ. Nat Methods 2015, 12:411-414.
- *91. Qin W, Cheah JS, Xu C, Messing J, Freibaum BD, Boeynaems S, Taylor JP, Udeshi ND, Carr SA, Ting AY: Dynamic mapping of proteome trafficking within and between living cells by TransitID. Cell 2023, 186:3307-3324.e3330.
- Orthogonal proximity labeling approaches allow intracellular and intercellular protein trafficking studies.
- 92. Reinkemeier CD, Girona GE, Lemke EA: Designer membraneless organelles enable codon reassignment of selected mRNAs in eukaryotes. Science 2019, 363:eaaw2644.
- **93. Reinkemeier CD, Lemke EA: Dual film-like organelles enable spatial separation of orthogonal eukaryotic translation. Cell 2021, 184:4886-4903.e4821.
- Phase separation of mRNAs and aminoacyl-tRNA synthetases at the membranes of organelles enables orthogonal local translation of the mRNAs.
- 94. Reinkemeier CD, Lemke EA: Condensed, Microtubule-coating Thin Organelles for Orthogonal Translation in Mammalian Cells. Journal of Molecular Biology 2022, 434:167454.
- *95. Hao M, Ling X, Sun Y, Wang X, Li W, Chang L, Zeng Z, Shi X, Niu M, Chen L, et al.: Tracking endogenous proteins based on RNA editing-mediated genetic code expansion. Nat Chem Biol 2024.
- The combination of RNA-targeted base editing with genetic code expansion to incorporate non-canonical amino acids to proteins synthesized from endogenous transcripts.
- 96. Sappakhaw K, Jantarug K, Slavoff SA, Israsena N, Uttamapinant C: A Genetic Code Expansion-Derived Molecular Beacon for the Detection of Intracellular Amyloid-β Peptide Generation. Angew Chem Int Ed Engl 2021, 60:3934-3939.
- 97. Uttamapinant C, Howe JD, Lang K, Beránek V, Davis L, Mahesh M, Barry NP, Chin JW: Genetic Code Expansion Enables Live-Cell and Super-Resolution Imaging of Site-Specifically Labeled Cellular Proteins. Journal of the American Chemical Society 2015, 137:4602-4605.
- 98. Aphicho K, Kittipanukul N, Uttamapinant C: Visualizing the complexity of proteins in living cells with genetic code expansion. Curr Opin Chem Biol 2022, 66:102108.
- 99. Lampkin BJ, Kritzer JA: Engineered fluorogenic HaloTag ligands for turn-on labelling in live cells††Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d3cc05536a. Chemical Communications 2023, 60:200-203.
- 100. Martin A, Rivera-Fuentes P: A general strategy to develop fluorogenic polymethine dyes for bioimaging. Nature Chemistry 2024, 16:28-35.
- 101. Rackham O, Chin JW: A network of orthogonal ribosome·mRNA pairs. Nature Chemical Biology 2005, 1:159-166.
- 102. Fried SD, Schmied WH, Uttamapinant C, Chin JW: Ribosome Subunit Stapling for Orthogonal Translation in E. coli. Angew Chem Int Ed Engl 2015, 54:12791-12794.
- 103. Schmied WH, Tnimov Z, Uttamapinant C, Rae CD, Fried SD, Chin JW: Controlling orthogonal ribosome subunit interactions enables evolution of new function. Nature 2018, 564:444-448.
- 104. Orelle C, Carlson ED, Szal T, Florin T, Jewett MC, Mankin AS: Protein synthesis by ribosomes with tethered subunits. Nature 2015, 524:119-124.
- 105. Carlson ED, d’Aquino AE, Kim DS, Fulk EM, Hoang K, Szal T, Mankin AS, Jewett MC: Engineered ribosomes with tethered subunits for expanding biological function. Nat Commun 2019, 10:3920.
- 106. Adaligil E, Song A, Cunningham CN, Fairbrother WJ: Ribosomal Synthesis of Macrocyclic Peptides with Linear γ(4)- and β-Hydroxy-γ(4)-amino Acids. ACS Chem Biol 2021, 16:1325-1331.
- 107. Chan AI, Sawant MS, Burdick DJ, Tom J, Song A, Cunningham CN: Evaluating Translational Efficiency of Noncanonical Amino Acids to Inform the Design of Druglike Peptide Libraries. ACS Chem Biol 2023, 18:81-90.
- **108. Ferretti MB, Barre JL, Karbstein K: Translational Reprogramming Provides a Blueprint for Cellular Adaptation. Cell Chem Biol 2018, 25:1372-1379.e1373.
- Translational reprogramming in yeast through modulation of ribosomal protein compositions.
- 109. Shakiba N, Jones RD, Weiss R, Del Vecchio D: Context-aware synthetic biology by controller design: Engineering the mammalian cell. Cell Syst 2021, 12:561-592.
- 110. Huang Y, Liu T: Therapeutic applications of genetic code expansion. Synthetic and Systems Biotechnology 2018, 3:150-158.
- 111. Davis L, Radman I, Goutou A, Tynan A, Baxter K, Xi Z, O’Shea JM, Chin JW, Greiss S: Precise optical control of gene expression in C elegans using improved genetic code expansion and Cre recombinase. eLife 2021, 10:e67075.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



