Submitted:
27 June 2024
Posted:
29 June 2024
You are already at the latest version
Abstract
Keywords:
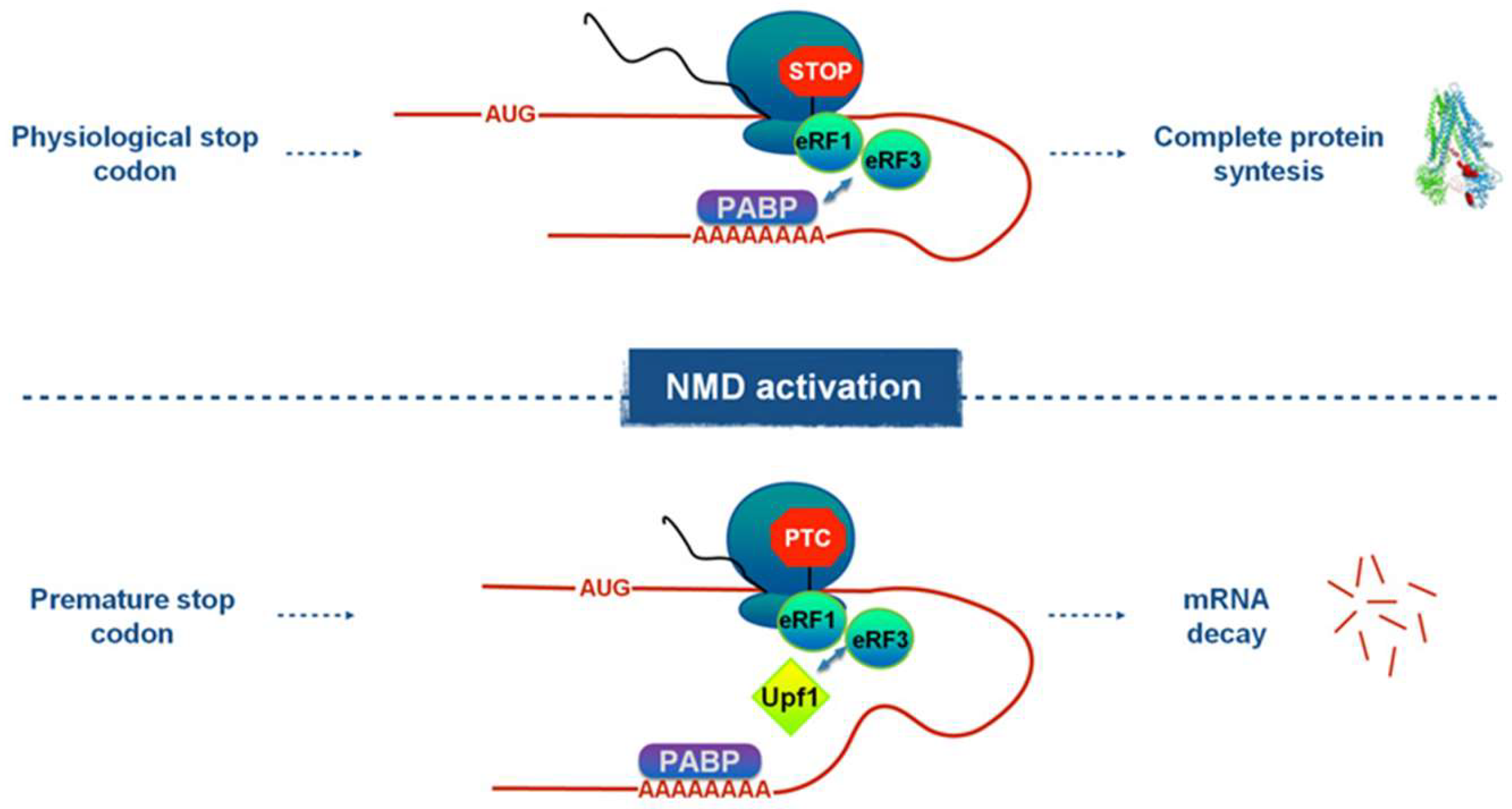
1. Introduction
2. Materials and Methods
2.1. Cell Cultures
2.2. Condition of Culture and TRIDs Treatment
2.3. Real-Time RT-PCR
2.4. Western Blotting
2.5. Immunofluorescence Microscopy
2.6. Functionality Assay: Quenching of Fluorescence of YFP Protein
2.7. Reduction of NMD Pathway Activity
3. Results
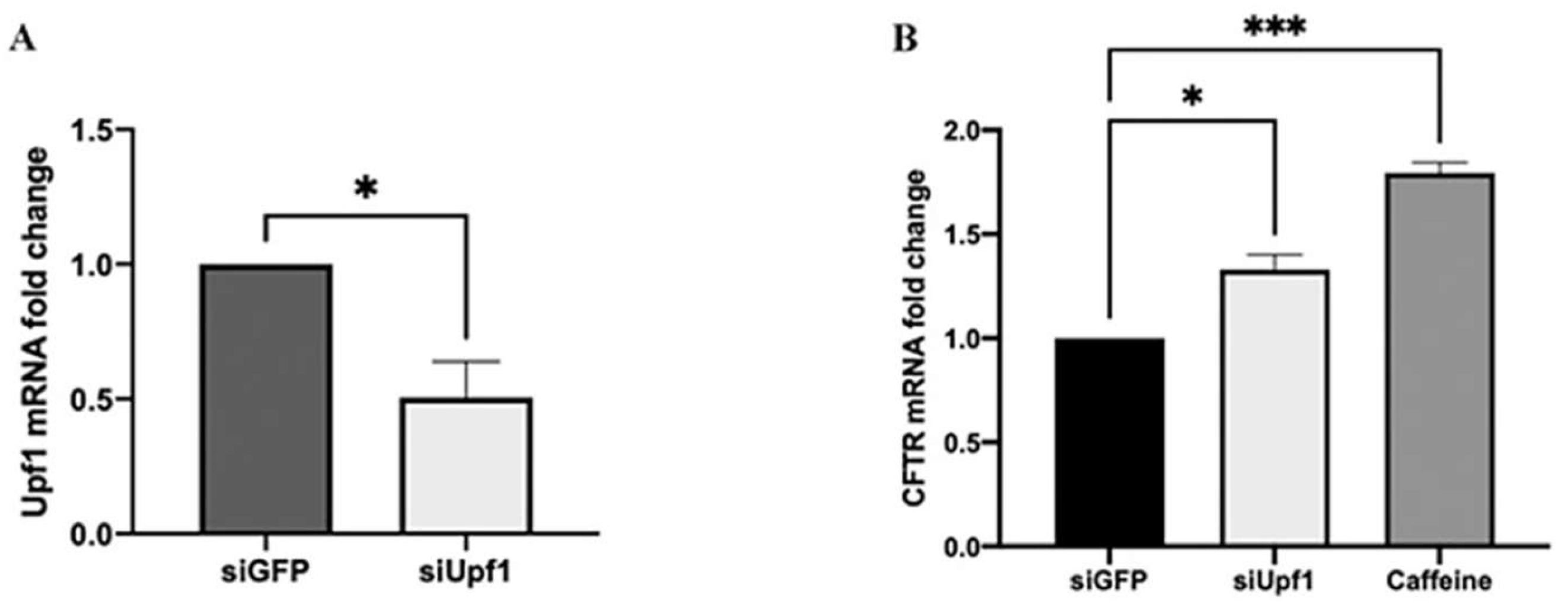
3.1. NMD Pathway Inhibition Increases CFTR mRNA Level Favoring TRIDs Readthrough Action
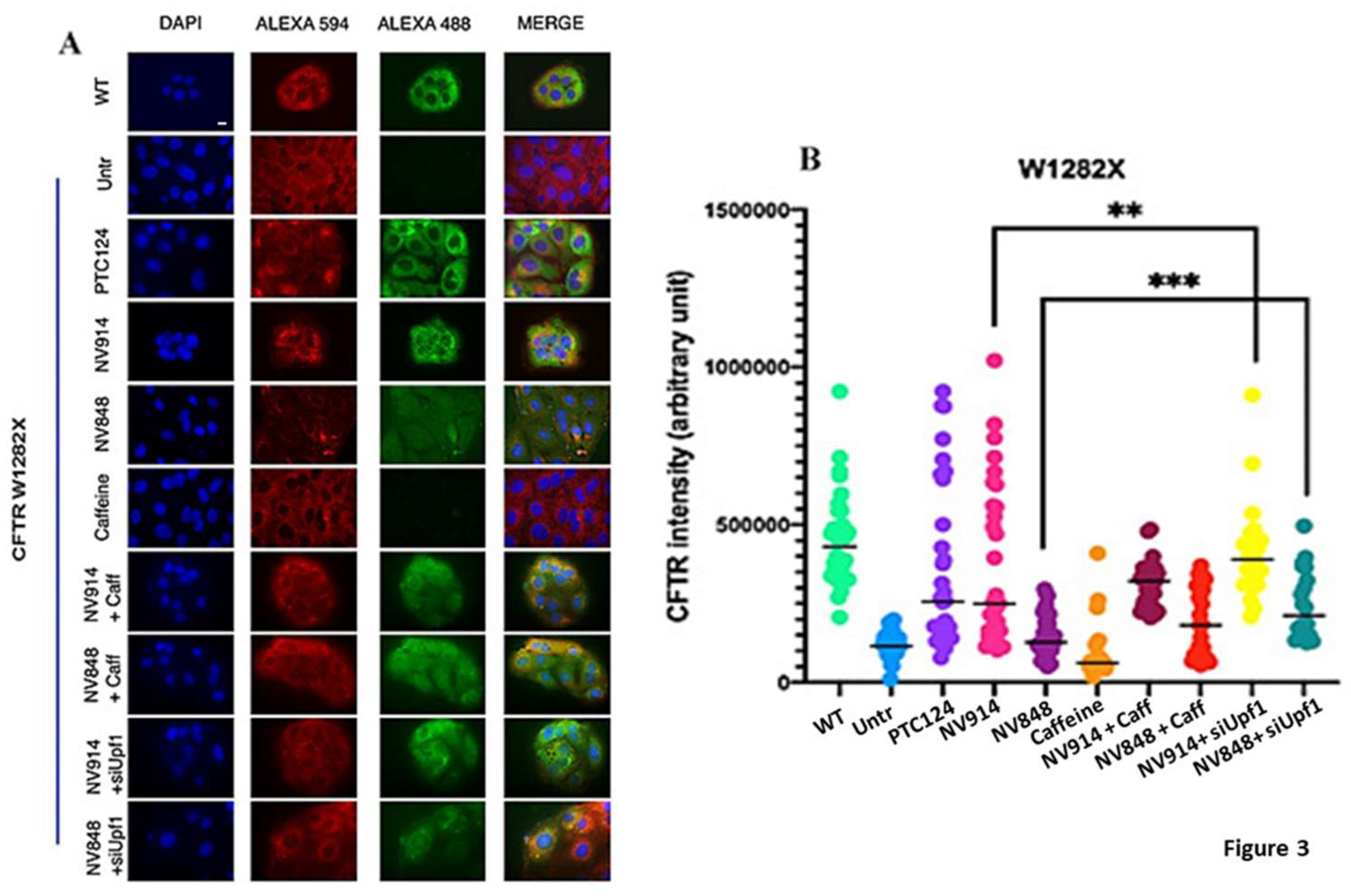
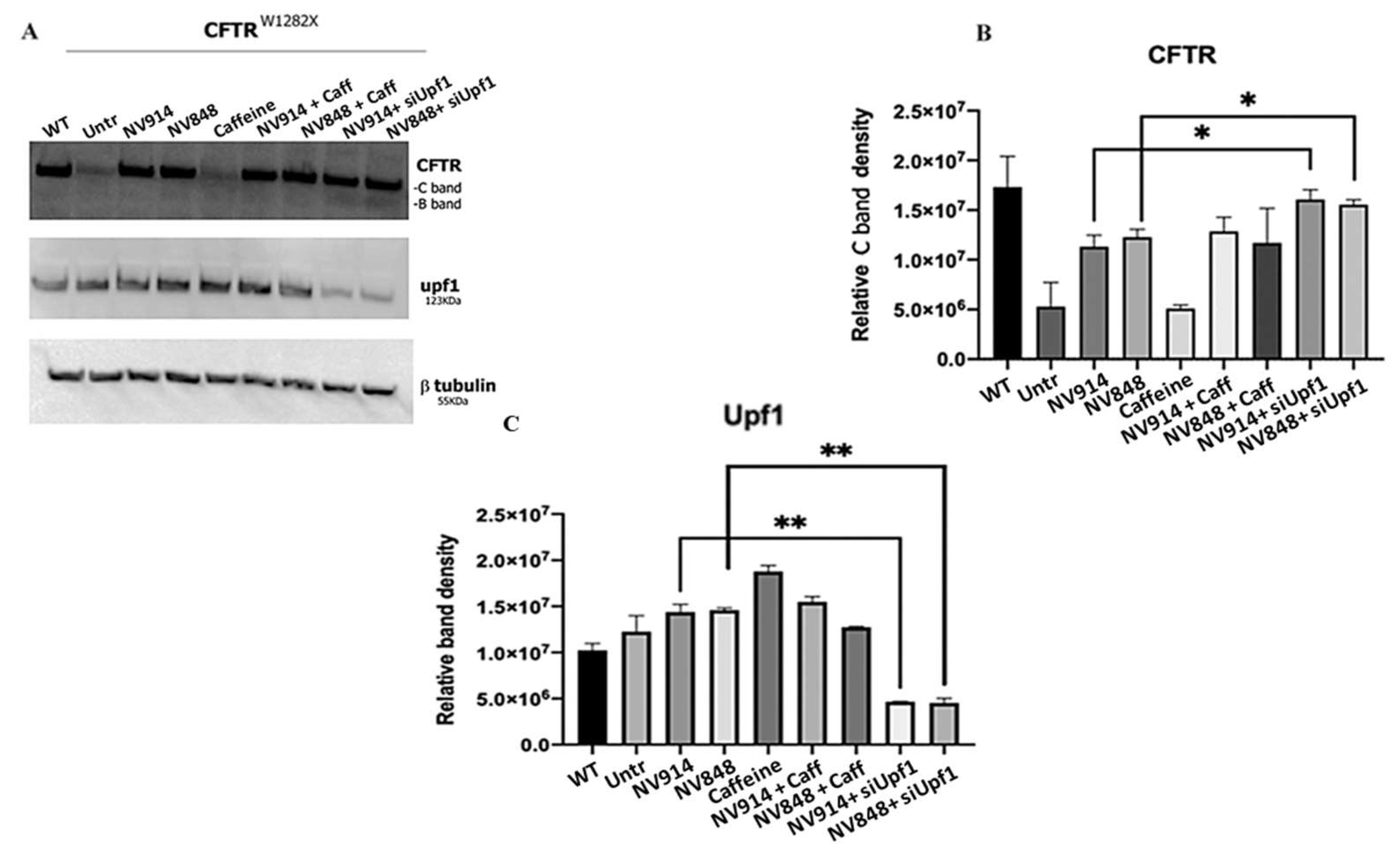
3.2. Combined Treatment of the NMD pathway Inhibition and Readthrough Induction with NV914 or NV848 TRID Molecules Increases CFTR Expression
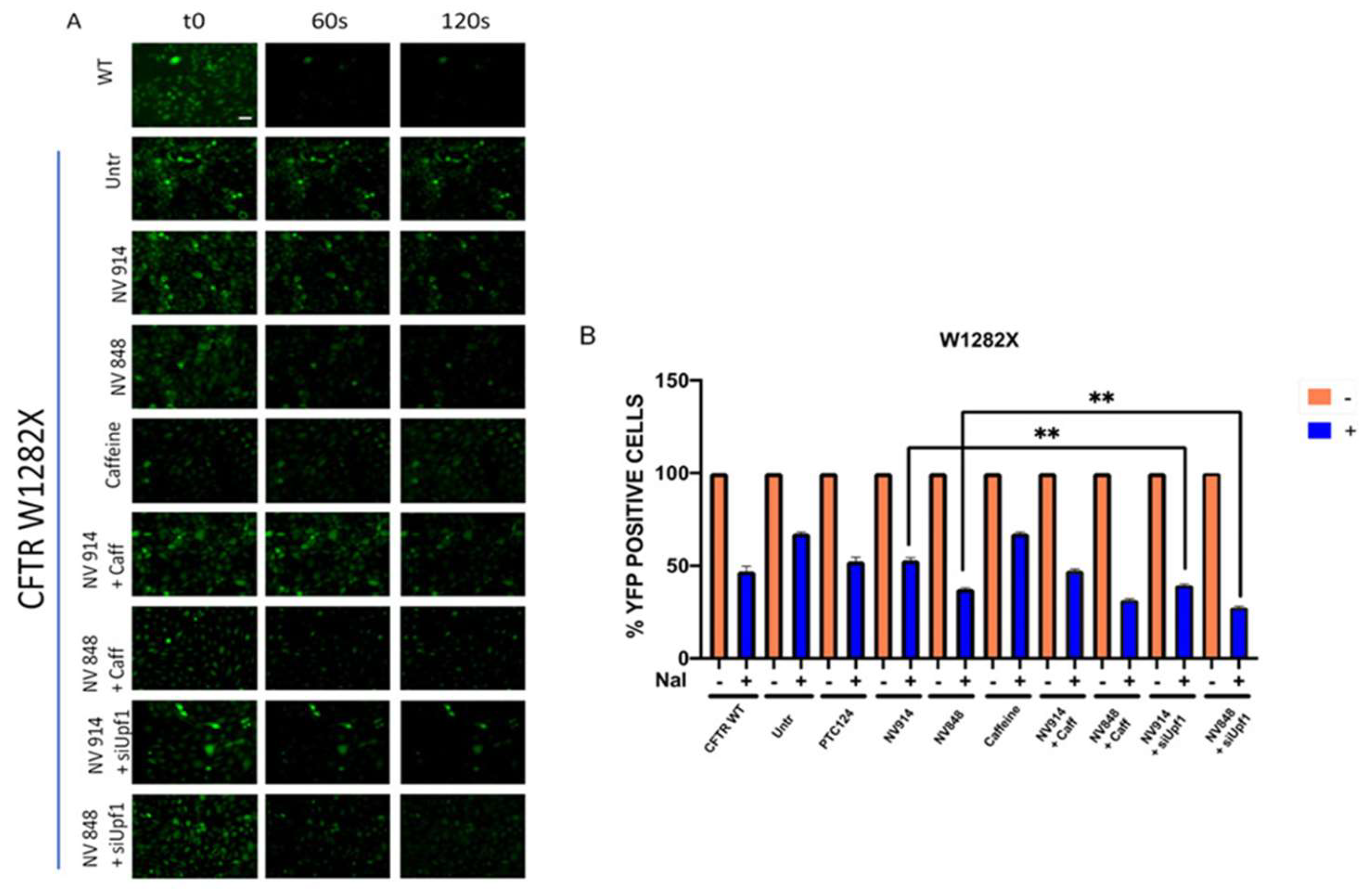
3.3. Functional Analysis of CFTR Channel after TRIDs Treatment
4. Discussion
Conclusions
5. Patents
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ardicli, D.; Haliloglu, G.; Alikasifoglu, M.; Topaloglu, H. (2019). Diagnostic Pathway to Nonsense Mutation Dystrophinopathy: A Tertiary-Center, Retrospective Experience. Neuropediatrics 2019, 50, 41–45. [CrossRef]
- Bongiorno R., Colombo M.P., Lecis D. (2021). Deciphering the nonsense-mediated mRNA decay pathway to identify cancer cell vulnerabilities for effective cancer therapy. J Exp Clin Cancer Res. 2021 Dec 1;40(1):376. PMID: 34852841; PMCID: PMC8638473. [CrossRef]
- Campofelice A., Lentini L., Di Leonardo A., Melfi R., Tutone M., Pace A. and Pibiri I. (2019). Strategies against Nonsense: Oxadiazoles as Translational Readthrough-Inducing Drugs (TRIDs). Int. J. Mol. Sci. 2019, 20, 3329;. [CrossRef]
- Chang, Y.F.; Imam, J. S.; Wilkinson, M. F. (2007). The Nonsense- Mediated Decay RNA Surveillance Pathway. Annu. Rev. Biochem. 2007, 76, 51−74. [CrossRef]
- Dabrowski M., Bukowy-Bieryllo Z., Zietkiewicz E. (2015). Translational readthrough potential of natural termination codons in eucaryotes--The impact of RNA sequence. RNA Biol; 12(9):950-8. PMID: 26176195; PMCID: PMC4615788. [CrossRef]
- Farinha, C.M.; Canato, S. (2017). From the endoplasmic reticulum to the plasma membrane: Mechanisms of CFTR folding and trafficking Cell. Mol. Life Sci. 2017, 74, 39–55. [CrossRef]
- Fernandes P., Miotto B., Saint-Ruf C., Said M., Barra V., Nähse V., Ravera S., Cappelli E., Naim V. FANCD2 modulates the mitochondrial stress response to prevent common fragile site instability. Commun Biol.2021 Jan 29;4(1):127.
- Fritz S. E., Ranganathan S., Wang C. D. , Hogg R. J. (2020). The RNA-binding protein PTBP1 promotes ATPase-dependent dissociation of the RNA helicase UPF1 to protect transcripts from nonsense-mediated mRNA decay, Journal of biological chemistry Volume 295, Issue 33, 14 August 2020, Pages 11613-11625. ISSN 0021-9258.
- Galietta L.J., Haggie P.M., Verkman A.S. (2001). Green fluorescent protein-based halide indicator s with improved chloride and iodide affinities. FEBS Letters, pp 220-24. [CrossRef]
- Hogg L. Goff. S., (2010). Upf1 senses 3′UTR length to potentiate mRNA decay. Cell 2010 Oct 29; 143 (3): 379-389.
- Ivanov, P. V.; Gehring, N. H.; Kunz, J. B.; Hentze, M. W.; Kulozik, A. E. (2008). Interactions between Upf1, ERFs, PABP and the Exon Junction Complex Suggest an Integrated Model for Mammalian NMD Pathways. EMBO J. 2008, 27 (5), 736−747. [CrossRef]
- Karousis, E. D.; Nasif, S.; Mühlemann, O. (2016). Nonsense-Mediated MRNA Decay: Novel Mechanistic Insights and Biological Impact. Wiley Interdiscip Rev RNA 2016, 7 (5), 661−682. doi: 10.1002/wrna.1357.
- Keeling K., Wang D., Dai Y., Murugesan S., Chenna B., Clark J., Belakhov V, Kandasamy J., Velu S., Baasov T., Bedwell DM. (2013). Attenuation of Nonsense- Mediated mRNA Decay Enhances In Vivo Nonsense Suppression, Plos One; 8(4):e60478. [CrossRef]
- Kerem E. (2020). ELX-02: an investigational read-through agent for the treatment of nonsense mutation-related genetic disease. Expert Opin Investig Drugs. Dec;29(12):1347-1354. Epub. 2020 Oct 12. PMID: 32972261. [CrossRef]
- Lentini L., Melfi R., Cancemi P., Pibiri I., Di Leonardo A. (2019). Caffeine boosts Ataluren's readthrough activity. Molecular Biology, Biochemistry, Volume 5, Issue 6 . [CrossRef]
- Lentini L., Melfi R., Di Leonardo A., Spinello A., Barone G., Pace A., Palumbo Piccionello A., Pibiri I. (2014). Toward a rationale for the PTC124 (Ataluren) promoted readthrough of premature stop codons: a computational approach and GFP-reporter cell-based assay. Mol Pharm. 2014 Mar 3;11(3):653-64. Epub 2014 Feb 7. PMID: 24483936; PMCID: PMC4167060. [CrossRef]
- Lou C.H., Dumdie J., Nonsense-Mediated RNA Decay Influences Human Embryonic Stem Cell Fate Stem Cell Reports 2016 Jun 14;6(6):844-857. [CrossRef]
- Loughran G, Chou M-Y, Ivanov IP, Jungreis I, Kellis M, Kiran AM, Baranov PV, Atkins JF. (2014). Evidence of efficient stop codon readthrough in four mammalian genes. Nucleic Acids Res, 42:8928-38. [CrossRef]
- Metze S., Herzog V.A, Ruepp M., and Mühlemann O. Comparison of EJC-enhanced and EJC-independent NMD in human cells reveals two partially redundant degradation pathways. RNA. 2013 Oct; 19(10): 1432–1448. [CrossRef]
- Neu-Yilik, Raimondeau E., et al.Dual function of UPF3B in early and late translation termination EMBO J, 2017 Oct 16;36(20):2968-2986. Epub 2017 Sep 12. [CrossRef]
- Peixeiro I, Inácio Â, Barbosa C, Silva AL, Liebhaber SA, Romão L. (2012). Interaction of PABPC1 with the translation initiation complex is critical to the NMD resistance of AUG-proximal nonsense mutations. Nucleic Acids Res. Feb;40(3):1160-73. Epub 2011 Oct 11. PMID: 21989405; PMCID: PMC3273812. [CrossRef]
- Pibiri I., Lentini L., Melfi R., Gallucci G., Pace A., Spinello A., Barone G., Di Leonardo A. (2015). Enhancement of premature stop codon readthrough in the CFTR gene by Ataluren (PTC124) derivatives. Eur J Med Chem. 2015 Aug 28;101:236-44. Epub 2015 Jun 21. PMID: 26142488. [CrossRef]
- Pibiri I., Lentini L., Melfi R., Tutone M., Baldassano S., Ricco Galluzzo P., Di Leonardo A., Pace A. (2018). Rescuing the CFTR protein function: Introducing 1,3,4-oxadiazoles as translational readthrough inducing drugs. Eur J Med Chem. 2018 Nov 5;159:126-142. Epub 2018 Sep 26. PMID: 30278331. [CrossRef]
- Pibiri I., Lentini L., Tutone M., Melfi R., Pace A., Di Leonardo A. (2016). Exploring the readthrough of nonsense mutations by non-acidic Ataluren analogues selected by ligand-based virtual screening. European Journal of Medicinal Chemistry. [CrossRef]
- Pibiri I., Melfi R., Tutone M., Di Leonardo A., Pace A. and Lentini L. (2020). Targeting Nonsense: Optimization of 1,2,4-Oxadiazole TRIDs to Rescue CFTR Expression and Functionality in Cystic Fibrosis Cell Model Systems Int. J. Mol. Sci. 2020, 21, 6420;. [CrossRef]
- Popp M.W., Maquat L.E. (2018). Nonsense-mediated mRNA Decay and Cancer. Curr Opin Genet Dev. 2018 Feb;48:44-50. Epub 2017 Nov 7. PMID: 29121514; PMCID: PMC5869107. [CrossRef]
- Pranke I., Golec A., Hinzpeter A., Edelman A., Sermet-Gaudelus I. (2019). Emerging Therapeutic Approaches for Cystic Fibrosis. From Gene Editing to Personalized Medicine. Front Pharmacol. 2019 Feb 27;10:121. PMID: 30873022; PMCID: PMC6400831. [CrossRef]
- Quon B.S., Rowe S.M. (2016). New and emerging targeted therapies for cystic. BMJ. Mar 30;352:i859. PMID: 27030675; PMCID: PMC4817245. [CrossRef]
- Rafeeq M.M., Murad H.A.S. (2017). Cystic fibrosis: current therapeutic targets and future approaches. J Transl Med. 2017 Apr 27;15(1):84. PMID: 28449677; PMCID: PMC5408469. [CrossRef]
- Richardson R., Smart M., Tracey-White D., Webster A., Moosajee M. (2017) Mechanism and evidence of nonsense suppression therapy for genetic eye disorders, Experimental Eye Research. [CrossRef]
- Roy B., Friesen W.J., Tomizawa Y., Leszyk J.D., Zhuo J., Johnson B., Dakka J., Trotta C.R., Xue X., Mutyam V., Keeling K.M., Mobley J.A., Rowe S.M., Bedwell D.M., Welch E.M., Jacobson A. (2016). Ataluren stimulates ribosomal selection of near-cognate tRNAs to promote nonsense suppression. Proc Natl Acad Sci U S A. Nov 1;113(44):12508-12513. Epub 2016 Oct 4. PMID: 27702906; PMCID: PMC5098639. [CrossRef]
- Salvatori F., Pappadà M., Breveglieri G., D'Aversa E., Finotti A., Lampronti I., Gambari R. and Borgatti M. (2018). UPF1 silenced cellular model systems for screening of read-through agents active on ß039 thalassemia point mutation. BMC biotechnology, 18(1), 28.
- Samanta A., Stingl K., Kohl S., Ries J., Linnert J., Nagel-Wolfrum K. (2019). Ataluren for the Treatment of Usher Syndrome 2A Caused by Nonsense Mutations. Int J Mol Sci.Dec 12;20(24). pii: E6274. [CrossRef]
- Sato H., Singer R.(2021) Cellular variability of nonsense-mediated mRNA decay Nat Commun 2021 Dec 10;12(1):7203.
- Sharma N., Evans T. A., Pellicore M. J., Davis E., Aksit, M. A., McCague A. F., Joynt A. T., Lu Z., Han S. T., Anzmann A. F., Lam A. N., Thaxton A., West N., Merlo C., Gottschalk L. B., Raraigh K. S., Sosnay P. R., Cotton C. U. and Cutting G. R. (2018). Capitalizing on the heterogeneous effects of CFTR nonsense and frameshift variants to inform therapeutic strategy for cystic fibrosis. PLoS genetics, 14(11), e1007723.
- Tutone M., Pibiri I., Lentini L., Pace A., Almerico A.M. (2019). Deciphering the Nonsense Readthrough Mechanism of Action of Ataluren: An in Silico Compared Study. ACS Med Chem Lett. 2019 Feb 7;10(4):522-527. PMID: 30996790; PMCID: PMC6466511. [CrossRef]
- Tutone M., Pibiri I., Perriera R., Campofelice A., Culletta G., Melfi R., Pace A., Almerico A.M., Lentini L. (2020). Pharmacophore-Based Design of New Chemical Scaffolds as Translational Readthrough-Inducing Drugs (TRIDs). ACS Med Chem Lett. 2020 Feb 18;11(5):747-753. PMID: 32435380; PMCID: PMC7236267. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



