Submitted:
08 June 2024
Posted:
11 June 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Preparation of crude extract and active compound
2.2. Cell culture
2.3. Cell viability assay
2.4. RNA extraction and gene expression analysis
2.5. Measurement of the MMP-9 production
2.6. LC-MS/MS analysis
2.6.1. Preparation of protein sample
2.6.2. Proteomic data analysis
2.7. Molecular docking analysis
2.8. Statistical analysis
3. Results
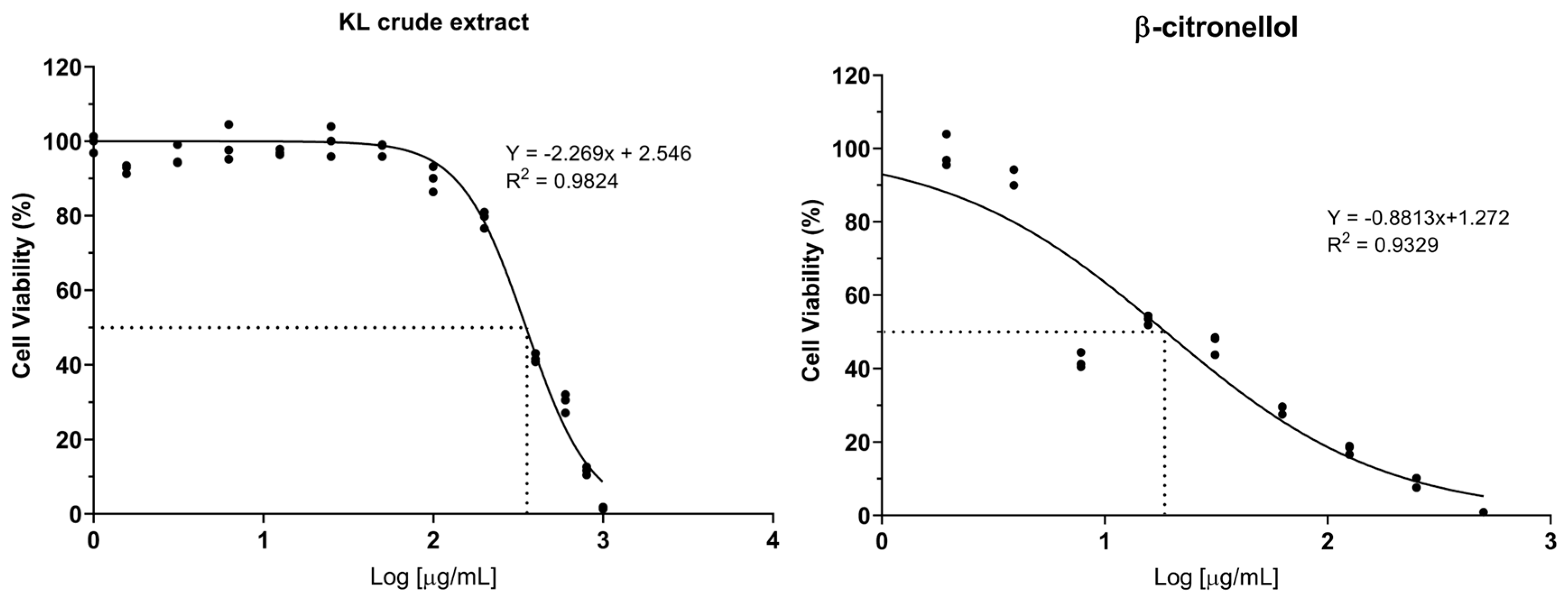
3.1. Cytotoxicity of crude extract and active compound on the LX-2 cell line
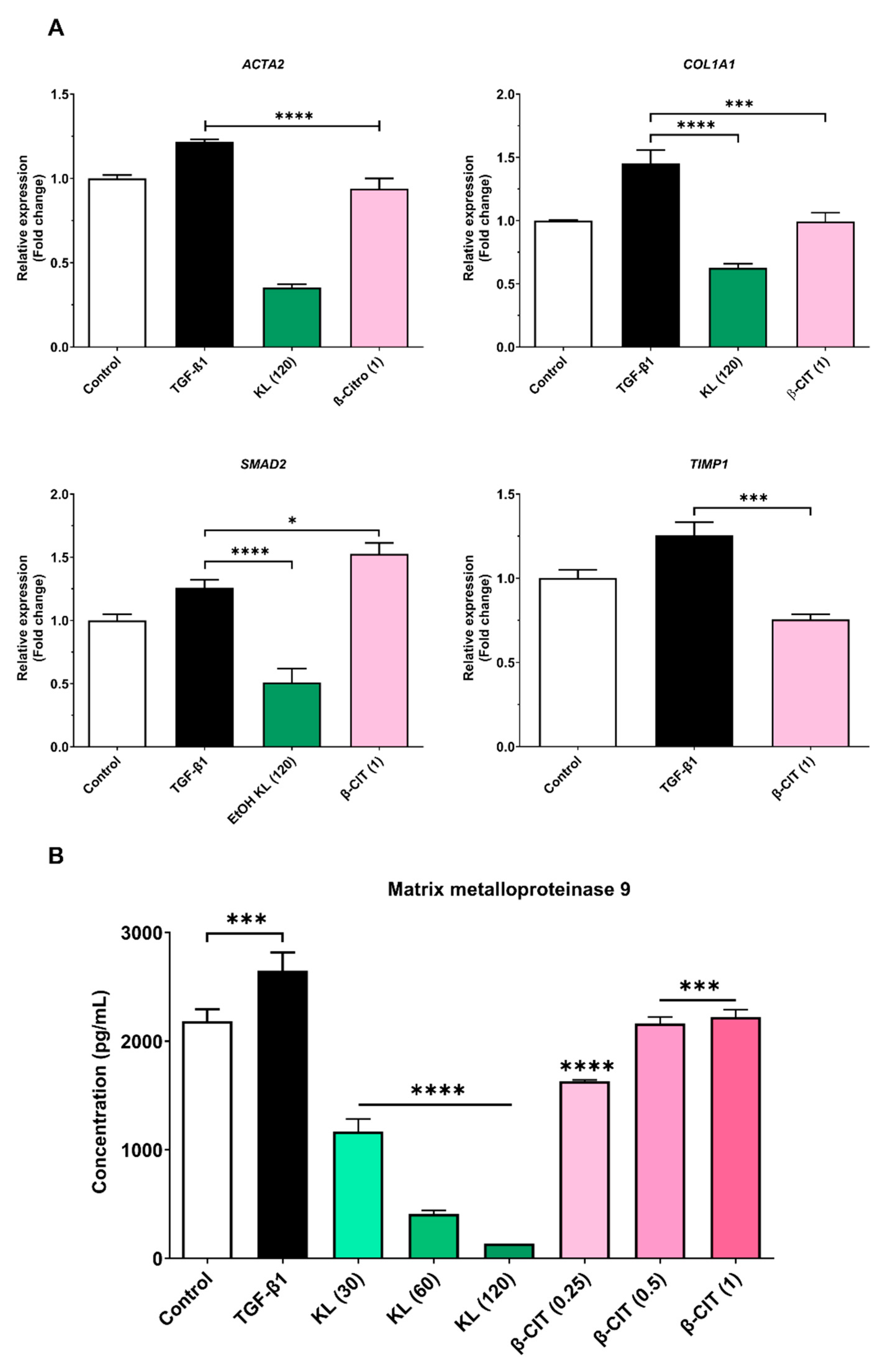
3.2. The inhibition of activated HSCs genes expression and MMP-9 production
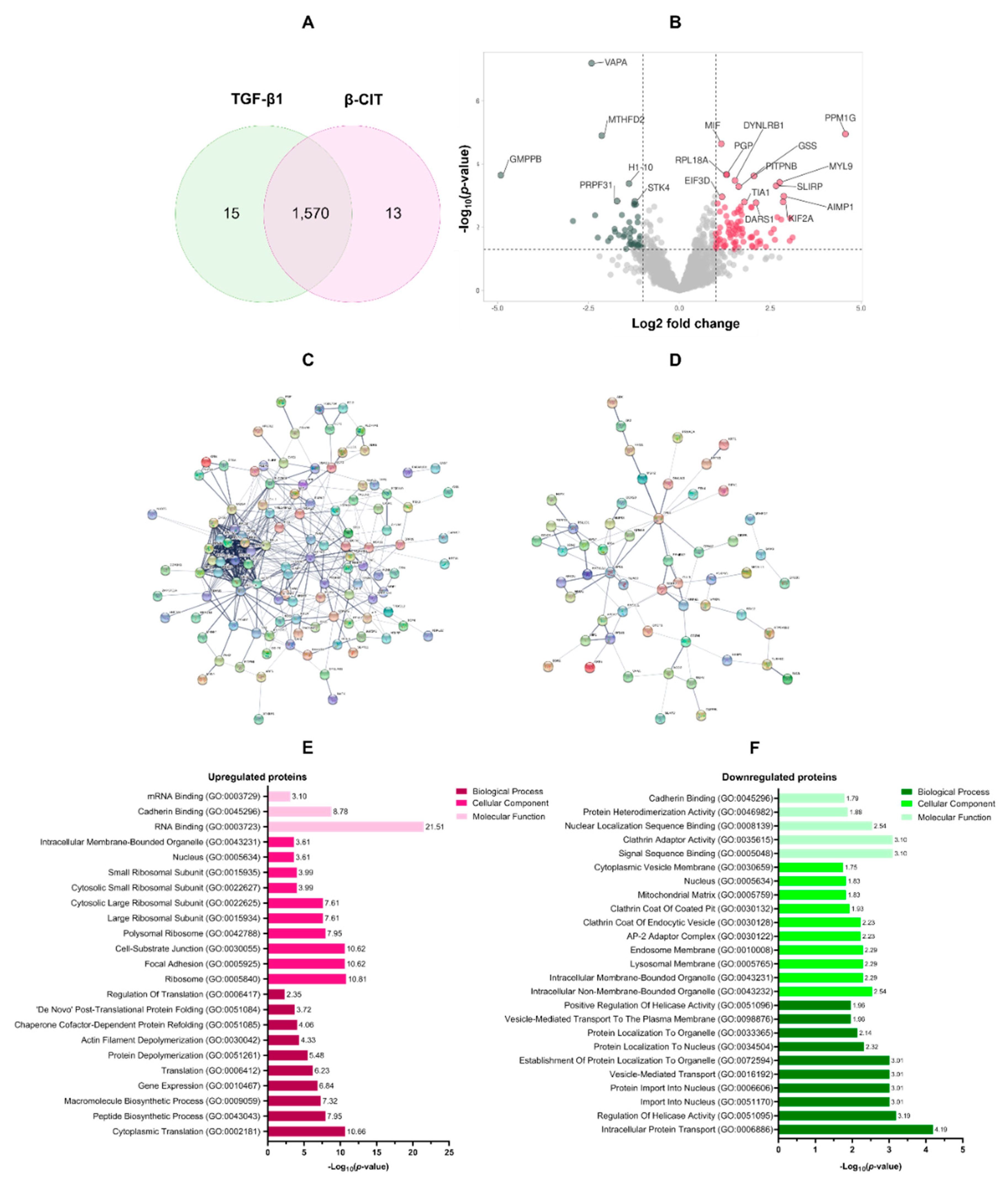
3.3. Proteomic analysis of the effect of β-citronellol on LX-2 cell
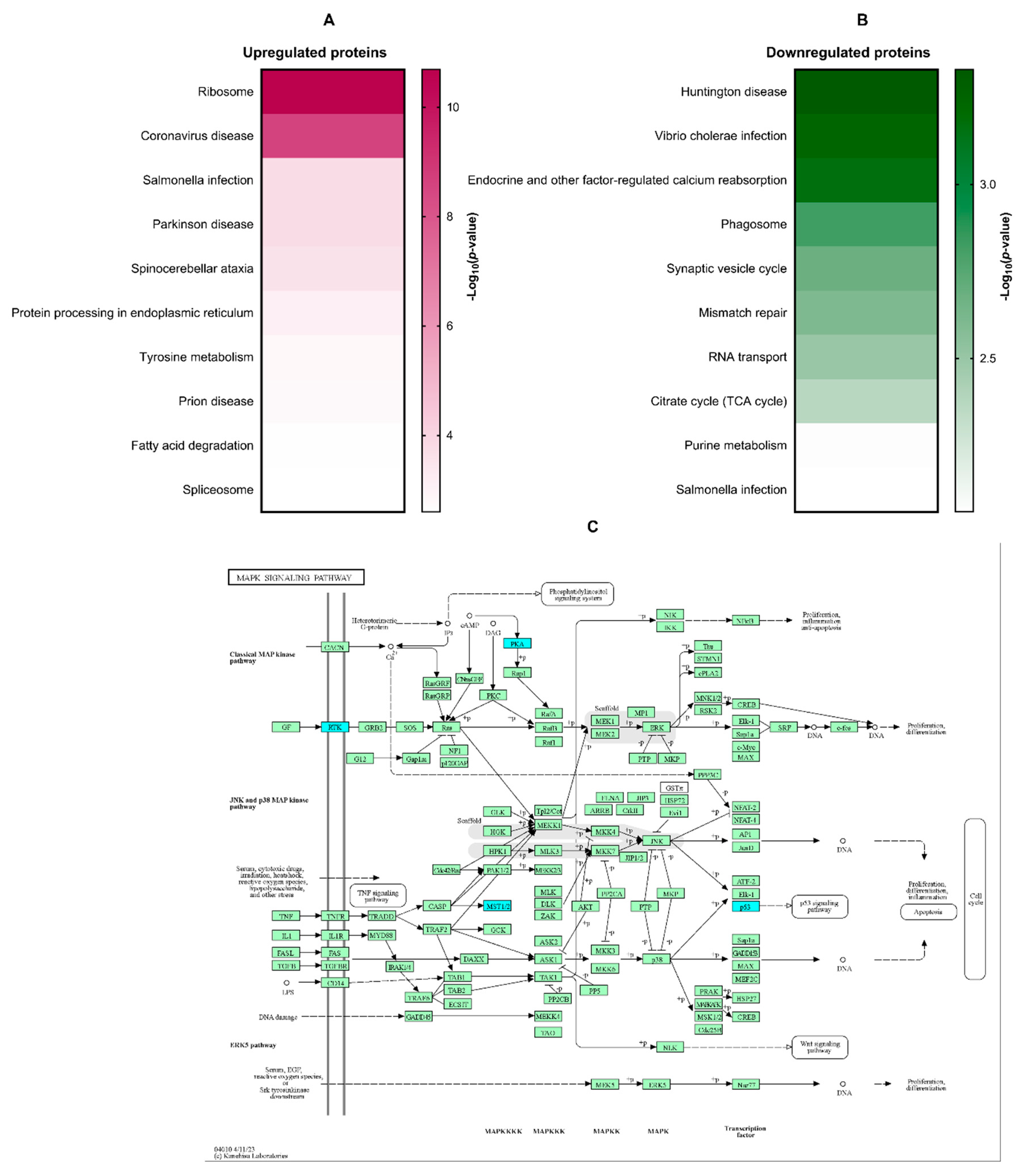
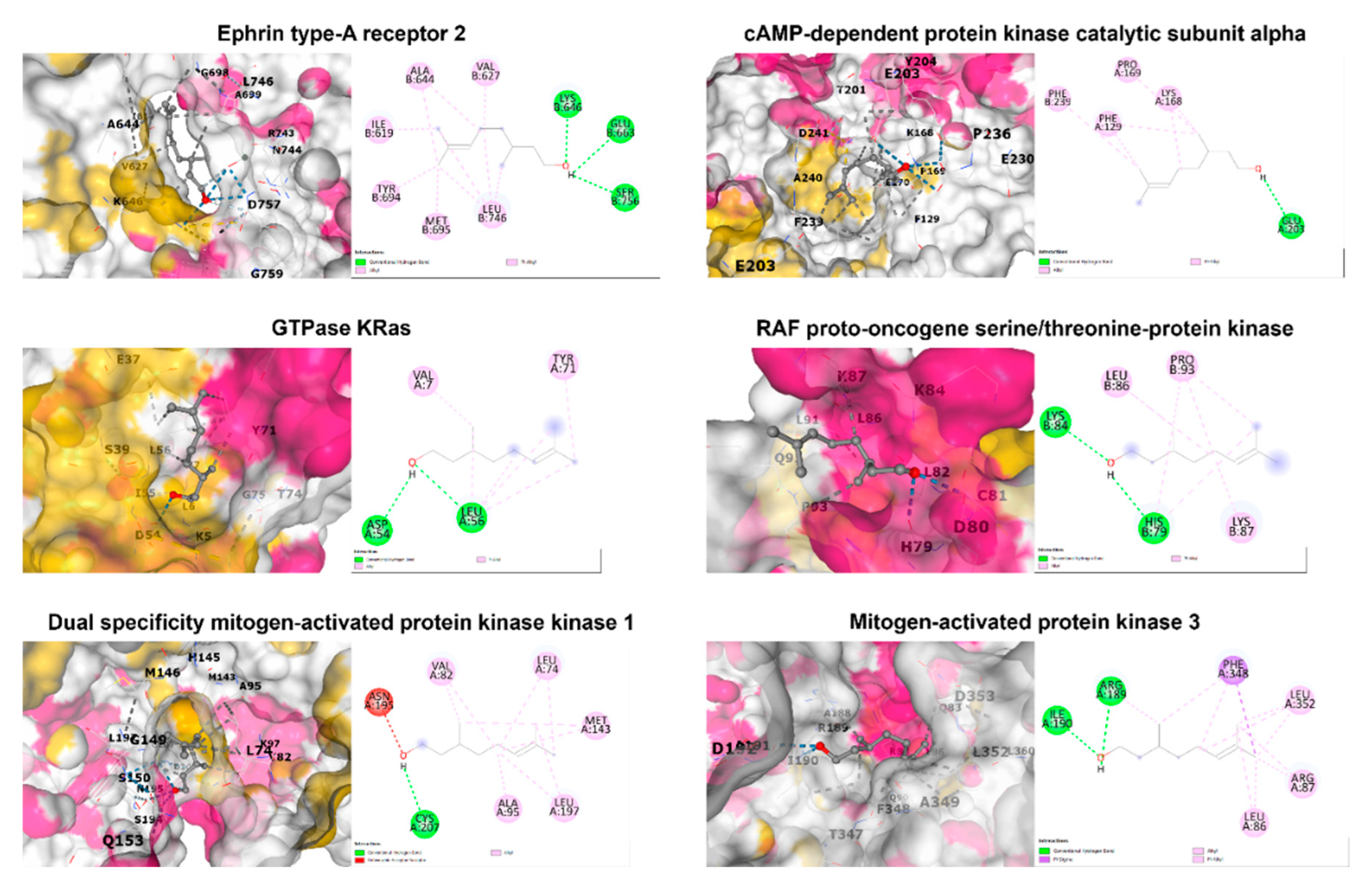
3.4. In silico molecular docking analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Devarbhavi, H.; Asrani, S.K.; Arab, J.P.; Nartey, Y.A.; Pose, E.; Kamath, P.S. Global burden of liver disease: 2023 update. J Hepatol 2023, 79, 516–537. [Google Scholar] [CrossRef] [PubMed]
- Luangmonkong, T.; Parichatikanond, W.; Olinga, P. Targeting collagen homeostasis for the treatment of liver fibrosis: Opportunities and challenges. Biochem Pharmacol 2023, 215, 115740. [Google Scholar] [CrossRef] [PubMed]
- Acharya, P.; Chouhan, K.; Weiskirchen, S.; Weiskirchen, R. Cellular Mechanisms of Liver Fibrosis. Front Pharmacol 2021, 12, 671640. [Google Scholar] [CrossRef] [PubMed]
- Tsuchida, T.; Friedman, S.L. Mechanisms of hepatic stellate cell activation. Nat Rev Gastroenterol Hepatol 2017, 14, 397–411. [Google Scholar] [CrossRef] [PubMed]
- Dhar, D.; Baglieri, J.; Kisseleva, T.; Brenner, D.A. Mechanisms of liver fibrosis and its role in liver cancer. Exp Biol Med 2020, 245, 96–108. [Google Scholar] [CrossRef]
- Zhang, C.Y.; Liu, S.; Yang, M. Treatment of liver fibrosis: Past, current, and future. World J Hepatol 2023, 15, 755–774. [Google Scholar] [CrossRef] [PubMed]
- Agouillal, F.; Taher, Z.; Moghrani, H.; Nasrallah, N.; El Enshasy, H. A Review of Genetic Taxonomy, Biomolecules Chemistry and Bioactivities of Citrus hystrix DC. Biosci Biotechnol Res Asia 2017, 14, 285–305. [Google Scholar] [CrossRef]
- Kidarn, S.; Saenjum, C.; Hongwiset, D.; Phrutivorapongkul, A. Furanocoumarins from Kaffir lime and their inhibitory effects on inflammatory mediator production. Cogent Chem 2018, 4, 1529259. [Google Scholar] [CrossRef]
- Buakaew, W.; Pankla Sranujit, R.; Noysang, C.; Thongsri, Y.; Potup, P.; Nuengchamnong, N.; Suphrom, N.; Usuwanthim, K. Phytochemical Constituents of Citrus hystrix DC. Leaves Attenuate Inflammation via NF-κB Signaling and NLRP3 Inflammasome Activity in Macrophages. Biomolecules 2021, 11, 105. [Google Scholar] [CrossRef]
- Ratanachamnong, P.; Chunchaowarit, Y.; Namchaiw, P.; Niwaspragrit, C.; Rattanacheeworn, P.; Jaisin, Y. HPLC analysis and in vitro antioxidant mediated through cell migration effect of C. hystrix water extract on human keratinocytes and fibroblasts. Heliyon 2023, 9, 13068. [Google Scholar] [CrossRef]
- Abirami, A.; Nagarani, G.; Siddhuraju, P. In vitro antioxidant, anti-diabetic, cholinesterase and tyrosinase inhibitory potential of fresh juice from Citrus hystrix and C. maxima fruits. Food Sci Hum Wellness 2014, 3, 16–25. [Google Scholar] [CrossRef]
- Srifuengfung, S.; Bunyapraphatsara, N.; Satitpatipan, V.; Tribuddharat, C.; Junyaprasert, V.B.; Tungrugsasut, W.; Srisukh, V. Antibacterial oral sprays from kaffir lime (Citrus hystrix DC.) fruit peel oil and leaf oil and their activities against respiratory tract pathogens. J Tradit Complement Med 2020, 10, 594–598. [Google Scholar] [CrossRef] [PubMed]
- Sreepian, P.M.; Rattanasinganchan, P.; Sreepian, A. Antibacterial efficacy of Citrus hystrix (makrut lime) essential oil against clinical multidrug-resistant methicillin-resistant and methicillin-susceptible Staphylococcus aureus isolates. Saudi Pharm J 2023, 31, 1094–1103. [Google Scholar] [CrossRef] [PubMed]
- Tunjung, W.A.S.; Cinatl, J.; Michaelis, M.; Smales, C.M. Anti-Cancer Effect of Kaffir Lime (Citrus Hystrix DC) Leaf Extract in Cervical Cancer and Neuroblastoma Cell Lines. Procedia Chem 2015, 14, 465–468. [Google Scholar] [CrossRef]
- Ho, Y.; Suphrom, N.; Daowtak, K.; Potup, P.; Thongsri, Y.; Usuwanthim, K. Anticancer Effect of Citrus hystrix DC. Leaf Extract and Its Bioactive Constituents Citronellol and, Citronellal on the Triple Negative Breast Cancer MDA-MB-231 Cell Line. Pharmaceuticals 2020, 13, 476. [Google Scholar] [CrossRef] [PubMed]
- Abolmaesoomi, M.; Mat Junit, S.; Mohd Ali, J.; Chik, Z.B.; Abdul Aziz, A. Effects of polyphenolic-rich extracts from Citrus hystrix on proliferation and oxidative stress in breast and colorectal cancer. Turk Biyokim Derg 2023, 48, 110–118. [Google Scholar] [CrossRef]
- Abirami, A.; Nagarani, G.; Siddhuraju, P. Hepatoprotective effect of leaf extracts from Citrus hystrix and C. maxima against paracetamol induced liver injury in rats. Food Sci Hum Wellness 2015, 4, 35–41. [Google Scholar] [CrossRef]
- Buakaew, W.; Pankla Sranujit, R.; Noysang, C.; Krobthong, S.; Yingchutrakul, Y.; Thongsri, Y.; Potup, P.; Daowtak, K.; Usuwanthim, K. Proteomic Analysis Reveals Proteins Involved in the Mode of Action of β-Citronellol Identified From Citrus hystrix DC. Leaf Against Candida albicans. Front Microbiol 2022, 13, 894637. [Google Scholar] [CrossRef] [PubMed]
- Rao, X.; Huang, X.; Zhou, Z.; Lin, X. An improvement of the 2 -ΔΔCT method for quantitative real-time polymerase chain reaction data analysis. Biostat Bioinforma Biomath 2013, 3, 71–85. [Google Scholar]
- Robert, S.; Gicquel, T.; Bodin, A.; Lagente, V.; Boichot, E. Characterization of the MMP/TIMP Imbalance and Collagen Production Induced by IL-1β or TNF-α Release from Human Hepatic Stellate Cells. PLoS One 2016, 11, 153118. [Google Scholar] [CrossRef]
- Krobthong, S.; Yingchutrakul, Y.; Samutrtai, P.; Hitakarun, A.; Siripattanapipong, S.; Leelayoova, S.; Mungthin, M.; Choowongkomon, K. Utilizing Quantitative Proteomics to Identify Species-Specific Protein Therapeutic Targets for the Treatment of Leishmaniasis. ACS Omega 2022, 7, 12580–12588. [Google Scholar] [CrossRef] [PubMed]
- Griss, J.; Viteri, G.; Sidiropoulos, K.; Nguyen, V.; Fabregat, A.; Hermjakob, H. ReactomeGSA - Efficient Multi-Omics Comparative Pathway Analysis. Mol Cell Proteomics 2020, 19, 2115–2125. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Berman, H.; Henrick, K.; Nakamura, H. Announcing the worldwide Protein Data Bank. Nat Struct Mol Biol 2003, 10, 980–980. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B.A.; Thiessen, P.A.; Yu, B.; et al. PubChem 2023 update. Nucleic Acids Res 2023, 51, 1373–1380. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Yang, X.; Gan, J.; Chen, S.; Xiao, Z.-X.; Cao, Y. CB-Dock2: improved protein–ligand blind docking by integrating cavity detection, docking and homologous template fitting. Nucleic Acids Res 2022, 50, 159–164. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A visualization system for exploratory research and analysis. J Comput Chem 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Dewidar, B.; Meyer, C.; Dooley, S.; Meindl, B.; Nadja. TGF-β in Hepatic Stellate Cell Activation and Liver Fibrogenesis—Updated 2019. Cells 2019, 8, 1419. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Zhang, Y.; Sun, B. The Molecular Mechanisms of Liver Fibrosis and Its Potential Therapy in Application. Int J Mol Sci 2022, 23, 12572. [Google Scholar] [CrossRef]
- Westenberger, G.; Sellers, J.; Fernando, S.; Junkins, S.; Han, S.M.; Min, K.; Lawan, A. Function of Mitogen-Activated Protein Kinases in Hepatic Inflammation. J Cell Signal 2021, 2, 172–180. [Google Scholar]
- Darling, T.K.; Lamb, T.J. Emerging Roles for Eph Receptors and Ephrin Ligands in Immunity. Front Immunol 2019, 10, 1473. [Google Scholar] [CrossRef] [PubMed]
- Mekala, S.; Dugam, P.; Das, A. Ephrin–Eph receptor tyrosine kinases for potential therapeutics against hepatic pathologies. J Cell Commun Signal 2023, 17, 549–561. [Google Scholar] [CrossRef] [PubMed]
- Creeden, J.F.; Kipp, Z.A.; Xu, M.; Flight, R.M.; Moseley, H.N.B.; Martinez, G.J.; Lee, W.H.; Alganem, K.; Imami, A.S.; McMullen, M.R.; et al. Hepatic kinome atlas: An in-depth identification of kinase pathways in liver fibrosis of humans and rodents. Hepatology 2022, 76, 1376–1388. [Google Scholar] [CrossRef] [PubMed]
| Gene | Description | Forward primer (3’ → 5’) |
Reverse primer (3’ → 5’) |
|---|---|---|---|
| ACTA2 | actin alpha 2, smooth muscle | CATCCTCATCCTCCCTTGAG | ATGAAGGATGGCTGGAACAG |
| COL1A1 | Collagen type I alpha 1 chain | CCGGCTCCTGCTCCTCTTAGCG | CGTTCTGTACGCAGGTGATTGGTGG |
| TIMP1 | TIMP metallopeptidase inhibitor 1 | CAAGATGTATAAAGGGTTCCAAGC | TCCATCCTGCAGTTTTCCAG |
| SMAD2 | SMAD family member 2 | TGCTCTGAAATTTGGGGACTGA | GACGACCATCAAGAGACCTGG |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase | ATGACATCAAGAAGGTGGTG | CATACCAGGAAATGAGCTTG |
| Uniprot ID | Gene ID | Description | Log2 fold change |
-Log10 p-value |
|---|---|---|---|---|
| Top-10 upregulated proteins | ||||
| O15355 | PPM1G | Protein phosphatase, Mg2+/Mn2+ dependent 1G | 4.57 | 4.95 |
| Q9Y4W6 | AFG3L2 | AFG3-like protein 2 | 3.11 | 1.68 |
| P04066 | FUCA1 | Tissue alpha-L-fucosidase | 3.05 | 2.29 |
| Q16537 | PPP2R5E | Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit epsilon isoform | 3.04 | 1.57 |
| Q12904 | AIMP1 | Aminoacyl tRNA synthase complex-interacting multifunctional protein 1 | 2.87 | 2.98 |
| O00139 | KIF2A | Kinesin-like protein KIF2A | 2.85 | 2.80 |
| Q16186 | ADRM1 | Proteasomal ubiquitin receptor ADRM1 | 2.80 | 2.24 |
| P24844 | MYL9 | Myosin regulatory light polypeptide 9 | 2.76 | 3.42 |
| Q0VDF9 | HSPA14 | Heat shock 70 kDa protein 14 | 2.69 | 2.32 |
| Q9GZT3 | SLIRP | SRA stem-loop-interacting RNA-binding protein, mitochondrial | 2.66 | 3.31 |
| Top-10 downregulated proteins | ||||
| Q9Y5P6 | GMPPB | Mannose-1-phosphate guanyltransferase beta | -4.91 | 3.65 |
| O75431 | MTX2 | Metaxin-2 | -2.92 | 2.20 |
| Q9P0L0 | VAPA | Vesicle-associated membrane protein-associated protein A | -2.42 | 7.19 |
| P04264 | KRT1 | Keratin, type II cytoskeletal 1 | -2.32 | 2.38 |
| O76021 | RSL1D1 | Ribosomal L1 domain-containing protein 1 | -2.24 | 1.67 |
| P13995 | MTHFD2 | Bifunctional methylenetetrahydrofolate dehydrogenase/cyclohydrolase, mitochondrial | -2.14 | 4.90 |
| P51398 | DAP3 | 28S ribosomal protein S29, mitochondrial | -1.96 | 1.56 |
| Q9P2R3 | ANKFY1 | Rabankyrin-5 | -1.89 | 1.66 |
| O00291 | HIP1 | Huntingtin-interacting protein 1 | -1.78 | 1.95 |
| PDB ID | Protein name | Binding score (Kcal/mol) | Interacting amino acid |
|---|---|---|---|
| 1MQB | Ephrin type-A receptor 2 | -5.0 | Ile619, Val627, Ala644, Lys646, Glu663, Tyr694, Met695, Leu746, Ser756 |
| 7Y1G | cAMP-dependent protein kinase catalytic subunit alpha | -5.4 | Phe129, Lys168, Pro169, Glu203, Phe239 |
| 8FMI | GTPase KRas | -4.8 | Val7, Asp54, Leu56, Tyr71 |
| 1C1Y | RAF proto-oncogene serine/threonine-protein kinase | -4.2 | His79, Lys84, Leu86, Lys87, Pro93 |
| 1S9J | Dual specificity mitogen-activated protein kinase kinase 1 | -5.3 | Leu74, Val82, Ala95, Met143, Asn195, Leu197, Cys207 |
| 2ZOQ | Mitogen-activated protein kinase 3 | -5.7 | Leu86, Arg87, Arg189, Ile190, Phe348, Leu352 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




