Submitted:
31 May 2024
Posted:
03 June 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
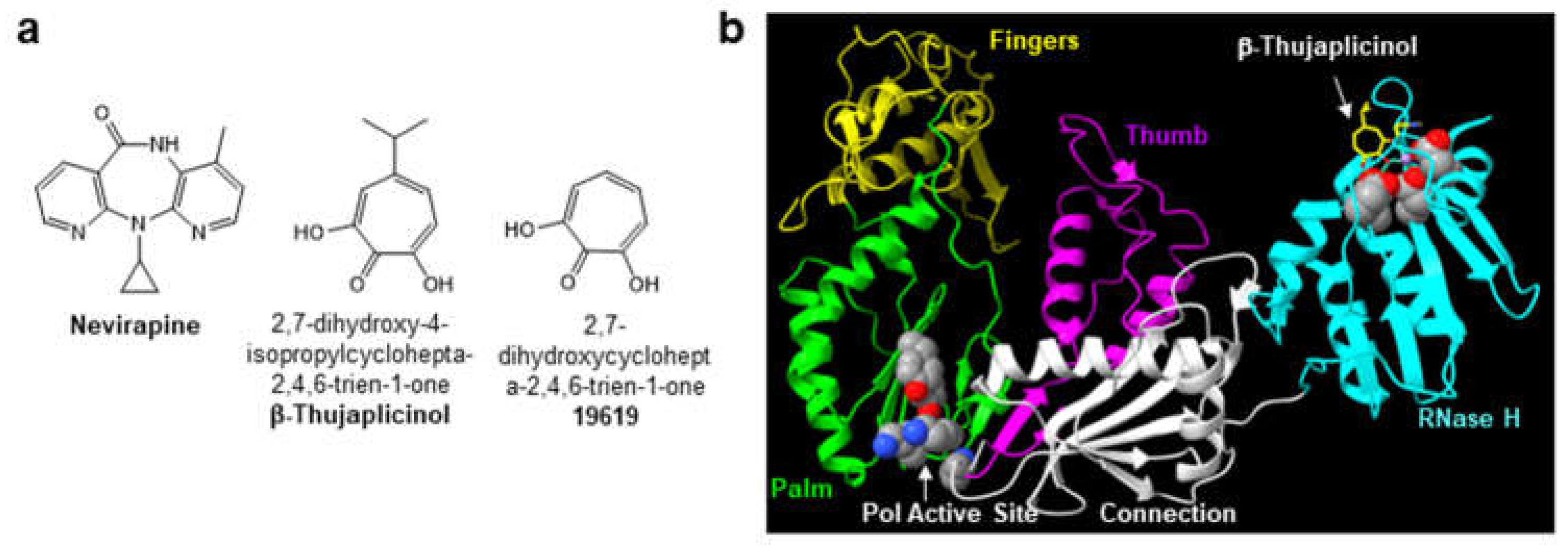
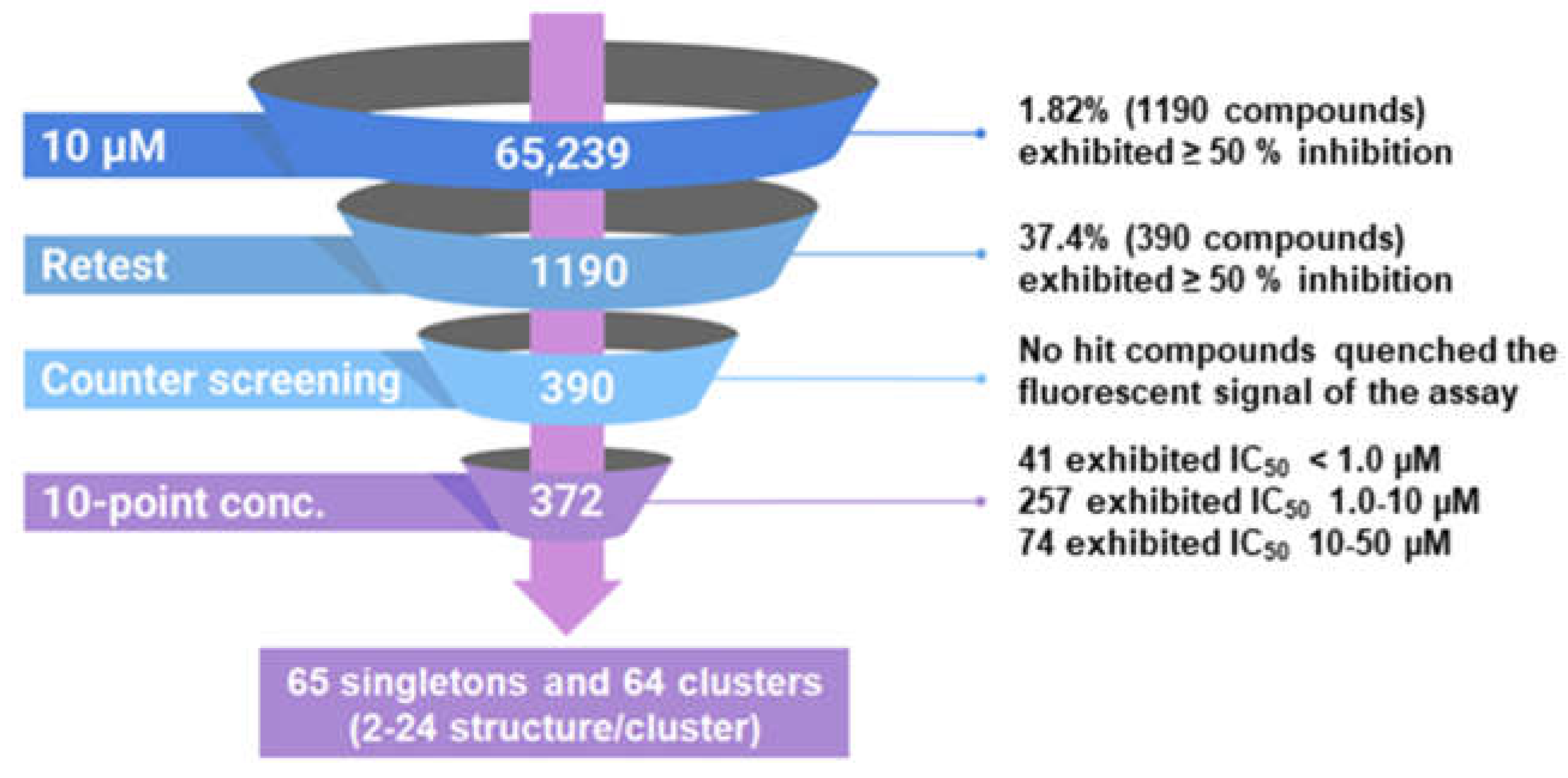
2.1. High Throughput Screen (HTS) to Identify HIV-1 RT RNase H Inhibitors
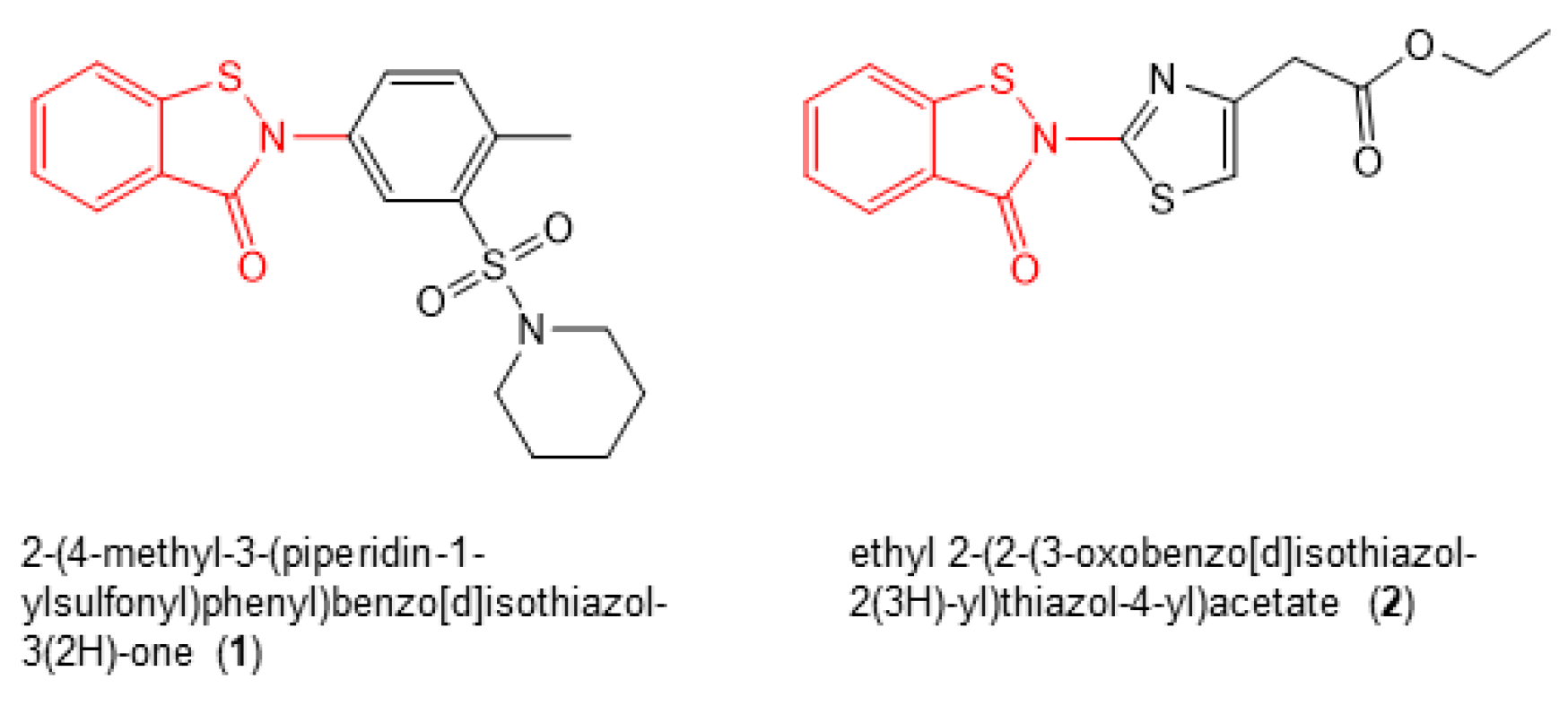
2.2. Identification of RT RNase H Inhibitors with Antiviral Activity
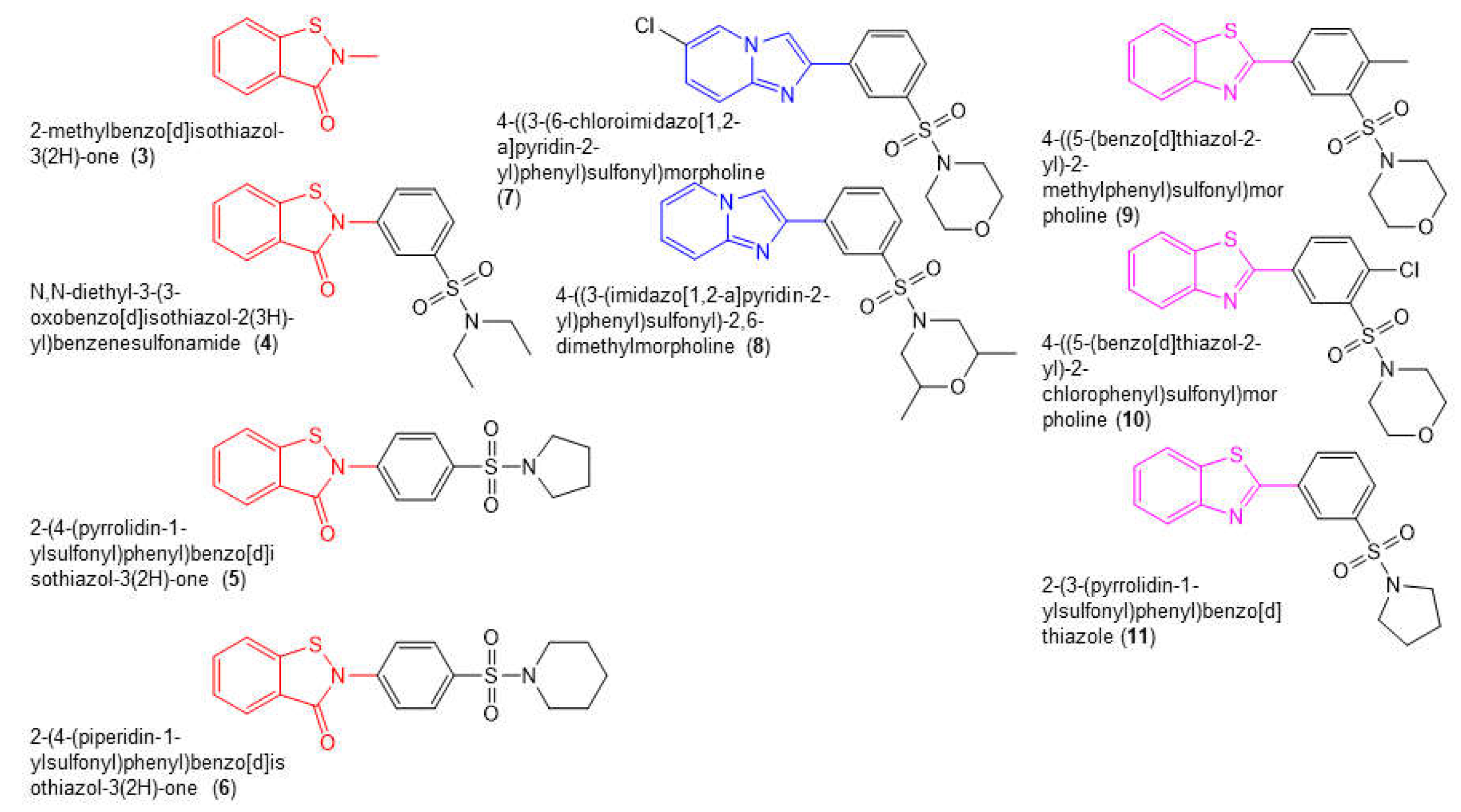
2.3. Structure Activity Relationships (SAR)
2.4. Inhibitory Activity of 1, 2, 3 and 4 against Wild-Type (WT) HIV-1 RT and HIV-1 RT Containing NNRTI Resistance Mutations
3. Discussion
4. Materials and Methods
- Materials
- Compound Library
- Primary HTS Assay.
- T/P substrates
- FRET-based RNase H cleavage assays
- Fluorescence-based DDDP Assay
- HIV-1 drug susceptibility assays.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Xavier Ruiz, F.; Arnold, E. Evolving understanding of HIV-1 reverse transcriptase structure, function, inhibition, and resistance. Curr Opin Struct Biol. 2020, 61, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Sluis-Cremer, N. Retroviral reverse transcriptase: Structure, function and inhibition. Enzymes 2021, 50, 179–194. [Google Scholar] [CrossRef]
- Singh, A.K.; Das, K. Insights into HIV-1 Reverse Transcriptase (RT) Inhibition and Drug Resistance from Thirty Years of Structural Studies. Viruses 2022, 14, 1027. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, C.; Pannecouque, C.; De Clercq, E.; Chen, F. Development of non-nucleoside reverse transcriptase inhibitors (NNRTIs): our past twenty years. Acta Pharm Sin B. 2020, 10, 961–978. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, Y.; Honma, M.; Kimura, Y.; Abe, H. Structure, Synthesis and Inhibition Mechanism of Nucleoside Analogues as HIV-1 Reverse Transcriptase Inhibitors (NRTIs). ChemMedChem. 2021, 16, 743–766. [Google Scholar] [CrossRef] [PubMed]
- Ilina, T.V.; Brosenitsch, T.; Sluis-Cremer, N.; Ishima, R. Retroviral RNase H: Structure, mechanism, and inhibition. Enzymes. 2021, 50, 227–247. [Google Scholar] [CrossRef] [PubMed]
- Beilhartz, G.L.; Wendeler, M.; Baichoo, N.; Rausch, J.; Le Grice, S.; Götte, M. HIV-1 reverse transcriptase can simultaneously engage its DNA/RNA substrate at both DNA polymerase and RNase H active sites: implications for RNase H inhibition. J Mol Biol. 2009, 388, 462–474. [Google Scholar] [CrossRef] [PubMed]
- Herman BD, Sluis-Cremer N. Transient kinetic analyses of the ribonuclease H cleavage activity of HIV-1 reverse transcriptase in complex with efavirenz and/or a beta-thujaplicinol analogue. Biochem J. 2013, 455, 179–184. [CrossRef]
- Hang, J.Q.; Li, Y.; Yang, Y.; Cammack, N.; Mirzadegan, T.; Klumpp, K. Substrate-dependent inhibition or stimulation of HIV RNase H activity by non-nucleoside reverse transcriptase inhibitors (NNRTIs). Biochem Biophys Res Commun. 2007, 352, 341–350. [Google Scholar] [CrossRef]
- Shaw-Reid, C.A.; Feuston, B.; Munshi, V.; Getty, K.; Krueger, J.; Hazuda, D.J.; Parniak, M.A.; Miller, M.D.; Lewis, D. Dissecting the effects of DNA polymerase and ribonuclease H inhibitor combinations on HIV-1 reverse-transcriptase activities. Biochemistry. 2005, 44, 1595–1606. [Google Scholar] [CrossRef]
- Radzio, J.; Sluis-Cremer, N. Efavirenz accelerates HIV-1 reverse transcriptase ribonuclease H cleavage, leading to diminished zidovudine excision. Mol Pharmacol. 2008, 73, 601–606. [Google Scholar] [CrossRef] [PubMed]
- Parniak, M.A.; Min, K.L.; Budihas, S.R.; Le Grice, S.F.; Beutler, .JA. A fluorescence-based high-throughput screening assay for inhibitors of human immunodeficiency virus-1 reverse transcriptase-associated ribonuclease H activity. Anal Biochem. 2003, 322, 33–39. [Google Scholar] [CrossRef] [PubMed]
- Barnard, J.P.; Huber, K.D.; Sluis-Cremer, N. Nonnucleoside Reverse Transcriptase Inhibitor Hypersusceptibility and Resistance by Mutation of Residue 181 in HIV-1 Reverse Transcriptase. Antimicrob Agents Chemother. 2019, 63, e00676–19. [Google Scholar] [CrossRef] [PubMed]
- Budihas, S.R.; Gorshkova, I.; Gaidamakov, S.; Wamiru, A.; Bona, M.K.; Parniak, M.A.; Crouch, R.J.; McMahon, J.B.; Beutler, J.A.; Le Grice, S.F. Selective inhibition of HIV-1 reverse transcriptase-associated ribonuclease H activity by hydroxylated tropolones. Nucleic Acids Res. 2005, 33, 1249–1256. [Google Scholar] [CrossRef] [PubMed]
- Himmel, D.M.; Maegley, K.A.; Pauly, T.A.; Bauman, J.D.; Das, K.; Dharia, C.; Clark, A.D. Jr.; Ryan, K.; Hickey, M.J.; Love, R.A.; Hughes, S.H.; Bergqvist, S.; Arnold, E. Structure of HIV-1 reverse transcriptase with the inhibitor beta-Thujaplicinol bound at the RNase H active site. Structure. 2009, 17, 1625–1635. [Google Scholar] [CrossRef] [PubMed]
- Di Grandi, M.; Olson, M.; Prashad, A.S.; Bebernitz, G.; Luckay, A.; Mullen, S.; Hu, Y.; Krishnamurthy, G.; Pitts, K.; O'Connell, J. Small molecule inhibitors of HIV RT Ribonuclease H. Bioorg Med Chem Lett. 2010, 20, 398–402. [Google Scholar] [CrossRef] [PubMed]
- Kirschberg, T.A.; Balakrishnan, M.; Huang, W.; Hluhanich, R.; Kutty, N.; Liclican, A.C.; McColl, D.J.; Squires, N.H.; Lansdon, E.B. Triazole derivatives as non-nucleoside inhibitors of HIV-1 reverse transcriptase-structure-activity relationships and crystallographic analysis. Bioorg. Med. Chem. Lett. 2008, 18, 1131–1134. [Google Scholar] [CrossRef] [PubMed]
- Himmel, D.M.; Sarafianos, S.G.; Dharmasena, S.; Hossain, M.M.; McCoy-Simandle, K.; Ilina, T.; Clark, A.D., Jr.; Knight, J.L.; Julias, J.G.; Clark, P.K.; Krogh-Jespersen, K.; Levy, R.M.; Hughes, S.H.; Parniak, M.A.; Arnold, E. HIV-1 reverse transcriptase structure with RNase H inhibitor dihydroxy benzoyl naphthyl hydrazone bound at a novel site. ACS Chem. Biol. 2006, 1, 702–712. [Google Scholar] [CrossRef]
- Wendeler, M.; Lee, H.-F.; Bermingham, A.; Miller, J.T.; Chertov, O.; Bona, M.K.; Baichoo, N.S.; Ehtesham, M.; Beutler, J.A.; O'Keefe, B.R.; Gotte, M.; Kvaratskhelia, M.; Le Grice, S.F.J. Vinylogous ureas as a novel class of reverse transcriptase-associated ribonuclease H activity. ACS Chem. Biol. 2008, 3, 635–644. [Google Scholar] [CrossRef]
- Chung, S.; Miller, J.T.; Johnson, B.C.; Hughes, S.H.; Le Grice, S.F. Mutagenesis of human immunodeficiency virus reverse transcriptase p51 subunit defines residues contributing to vinylogous urea inhibition of ribonuclease H activity. J. Biol. Chem. 2012, 287, 4066–4075. [Google Scholar] [CrossRef]
- Archer, R.H.; Dykes, C.; Gerondelis, P.; Lloyd, A.; Fay, P.; Reichman, R.C.; Bambara, R.A.; Demeter, L.M. Mutants of human immunodeficiency virus type 1 (HIV-1) reverse transcriptase resistant to nonnucleoside reverse transcriptase inhibitors demonstrate altered rates of RNase H cleavage that correlate with HIV-1 replication fitness in cell culture. J. Virol. 2000, 74, 8390–8401. [Google Scholar] [CrossRef] [PubMed]
- Archer, R.H.; Wisniewski, M.; Bambara, R.A.; Demeter, L.M. The Y181C mutant of HIV-1 reverse transcriptase resistant to nonnucleoside reverse transcriptase inhibitors alters the size distribution of RNase H cleavages. Biochemistry 2001, 40, 4087–4095. [Google Scholar] [CrossRef] [PubMed]
- Le Grice, S.F.; Grüninger-Leitch, F. Rapid purification of homodimer and heterodimer HIV-1 reverse transcriptase by metal chelate affinity chromatography. Eur J Biochem. 1990, 187, 307–314. [Google Scholar] [CrossRef] [PubMed]
- Mansouri, M.; Rumrill, S.; Dawson, S.; Johnson, A.; Pinson, J.A.; Gunzburg, M.J.; Latham, C.F.; Barlow, N.; Mbogo, G.W.; Ellenberg, P.; Headey, S.J.; Sluis-Cremer, N.; Tyssen, D.; Bauman, J.D.; Ruiz, F.X.; Arnold, E.; Chalmers, D.K.; Tachedjian, G. Targeting HIV-1 Reverse Transcriptase Using a Fragment-Based Approach. Molecules. 2023, 28, 3103. [Google Scholar] [CrossRef]
- Giacobbi, N.S.; Sluis-Cremer, N. In Vitro Cross-Resistance Profiles of Rilpivirine, Dapivirine, and MIV-150, Nonnucleoside Reverse Transcriptase Inhibitor Microbicides in Clinical Development for the Prevention of HIV-1 Infection. Antimicrob Agents Chemother. 2017, 61, e00277–17. [Google Scholar] [CrossRef]
| RNase H Activity | Polymerase Activity | Antiviral Activity | Cytotoxicity | |
| IC50 (µM) | IC50 (µM) | EC50 (µM) | CC50 (µM) | |
| RNase H | DNA Polymerase | HIV-1 | ||
| 1 | 0.16 ± 0.03 | 5.97 ± 3.10 | 1.68 ± 0.94 | ≥ 100 |
| 2 | 0.13 ± 0.04 | 2.64 ± 1.85 | 2.68 ± 0.54 | ≥ 100 |
| 3 | 2.5±0.2 | > 81 | 28.45 ± 12.26 | ≥ 100 |
| 4 | 0.26±0.08 | 1.1±0.3 | 3.27 ± 0.49 | ≥ 100 |
| 5 | 15.41±9.2 | 25±25 | ||
| 6 | 39.21±20.99 | 12.2±3.7 | ||
| 7 | > 81 | > 81 | ||
| 8 | > 81 | > 81 | ||
| 9 | > 81 | > 81 | ||
| 10 | > 81 | > 81 | ||
| 11 | > 81 | 62±38 |
| Inhibition of RT RNase H Activity | ||||||||||
| RT | 19619 | 1 | 2 | 3 | 4 | |||||
| IC50 (µM) | Fold-R1 | IC50 (µM) | Fold-R | IC50 (µM) | Fold-R | IC50 (µM) | Fold-R | IC50 (µM) | Fold-R | |
| WT | 8.7±3.0 | - | 0.16±0.03 | - | 0.13±0.04 | - | 2.5±0.2 | 0.26±0.08 | ||
| Y181C | 12.1±1.6 | 1.4 | 0.47±0.05 | 2.9 | 0.55±0.02 | 4.2 | 2.1±0.3 | 0.8 | 0.27±0.03 | 1.0 |
| K103N | 9.4±0.9 | 1.1 | 0.34±0.04 | 2.1 | 0.24±0.12 | 1.8 | 2.4±0.2 | 1.0 | 0.83±0.23 | 3.2 |
| G190A | 7.1±1.9 | 0.8 | 0.21±0.02 | 1.3 | 0.23±0.02 | 1.8 | 2.4±0.4 | 1.0 | 0.31±0.04 | 1.2 |
| K101E | 8.1±1.7 | 0.9 | 0.25±0.06 | 1.6 | 0.28±0.02 | 2.2 | 2.2±0.3 | 0.9 | 0.09±0.03 | 0.3 |
| Y188C | 5.5±2.4 | 0.6 | 0.32±0.01 | 2.0 | 0.59±0.04 | 4.5 | 5.7±1.2 | 2.3 | 1.10±0.32 | 4.2 |
| P236L | 8.9±2.5 | 1.0 | 0.22±0.01 | 1.4 | 0.21±0.01 | 1.6 | 2.2±0.2 | 0.9 | 0.31±0.04 | 1.2 |
| Inhibition of RT DNA polymerase activity | ||||||||||
| NVP | 1 | 2 | 3 | 4 | ||||||
| WT | 4.96±3.19 | - | 5.97±3.10 | - | 2.64±1.85 | - | - | 1.1±0.3 | - | |
| Y181C | > 81 | > 16 | 0.61±0.06 | 0.1 | 0.71±0.10 | 0.3 | - | 0.5±0.2 | 0.45 | |
| K103N | > 81 | > 16 | 3.66±1.84 | 0.6 | 3.28±1.53 | 1.2 | - | 2.1±0.7 | 1.9 | |
| G190A | > 81 | > 16 | 1.60±0.88 | 0.2 | 1.93±0.49 | 0.7 | - | 1.7±1.0 | 1.5 | |
| K101E | > 81 | > 16 | 7.33±7.05 | 1.2 | 4.88±1.56 | 1.8 | - | 1.8±0.7 | 1.6 | |
| Y188C | > 81 | > 16 | 0.14±0.01 | 0.02 | 0.53±0.08 | 0.2 | - | 0.3±0.1 | 0.3 | |
| P236L | > 81 | > 16 | 2.22±0.18 | 0.4 | 5.13±1.68 | 1.9 | - | 1.6±0.9 | 1.5 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




