Submitted:
28 May 2024
Posted:
28 May 2024
You are already at the latest version
Abstract
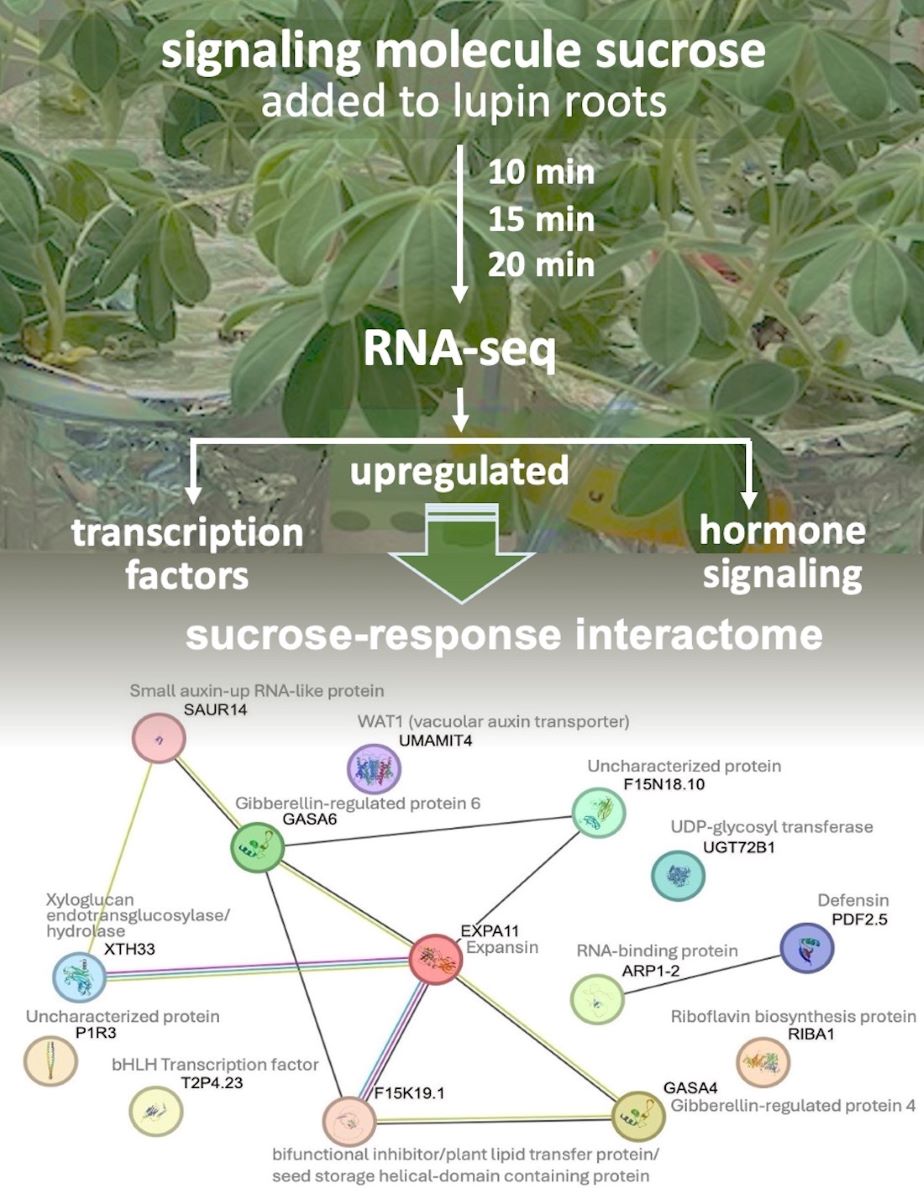
Keywords:
1. Introduction
2. Results
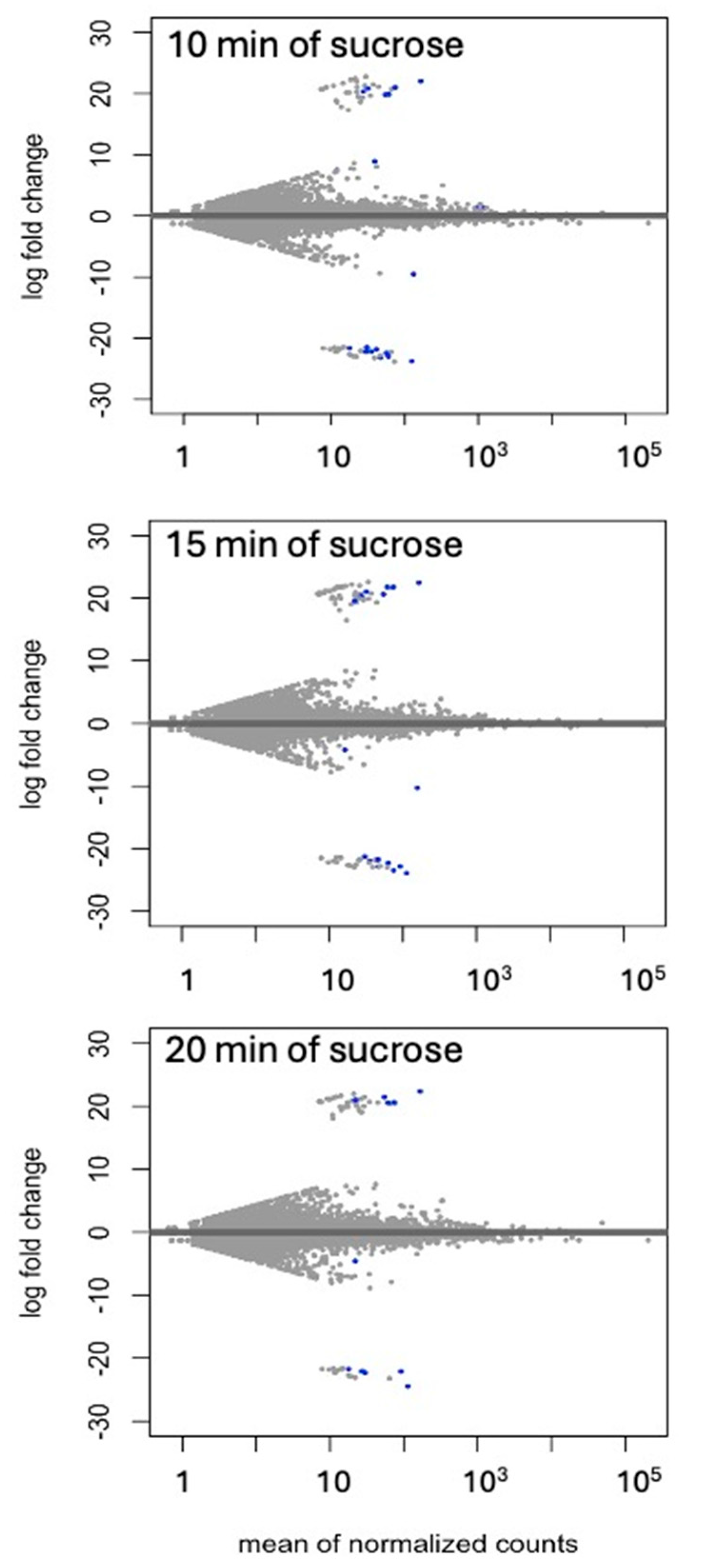
2.1. Nanopore cDNA Sequencing to Assess Short-Term Responses to Sucrose Resulted in 35.5 Million Reads
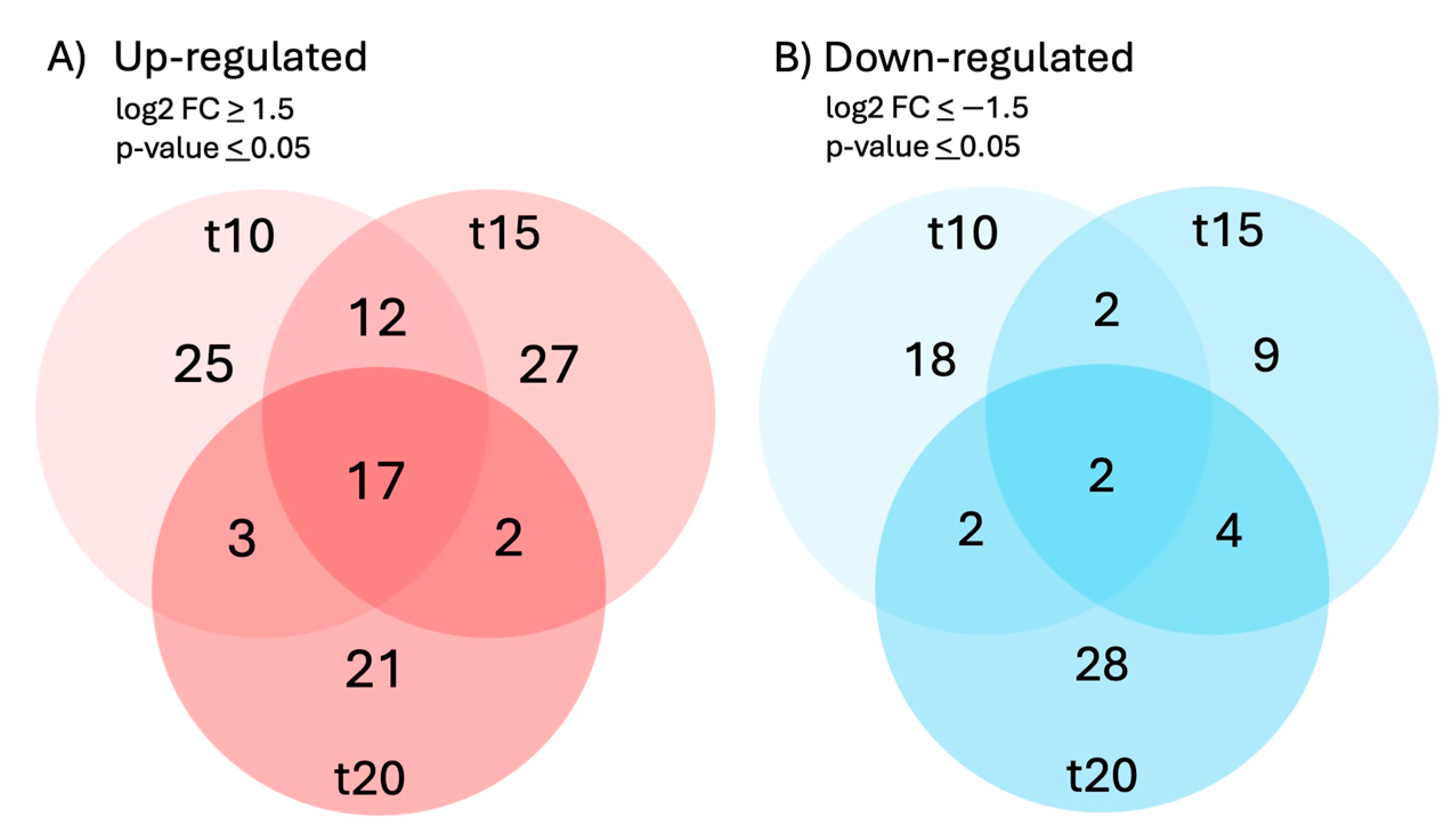
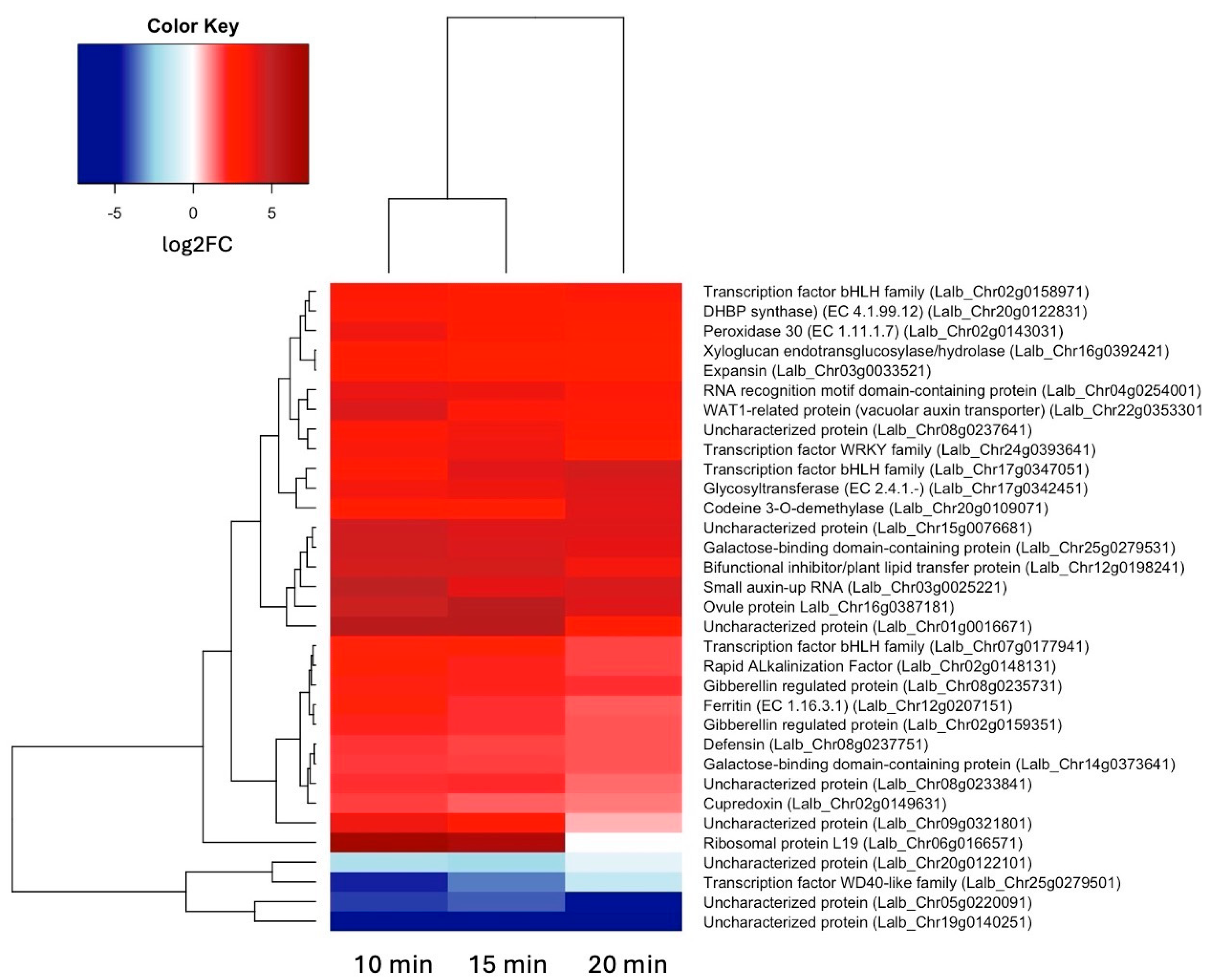
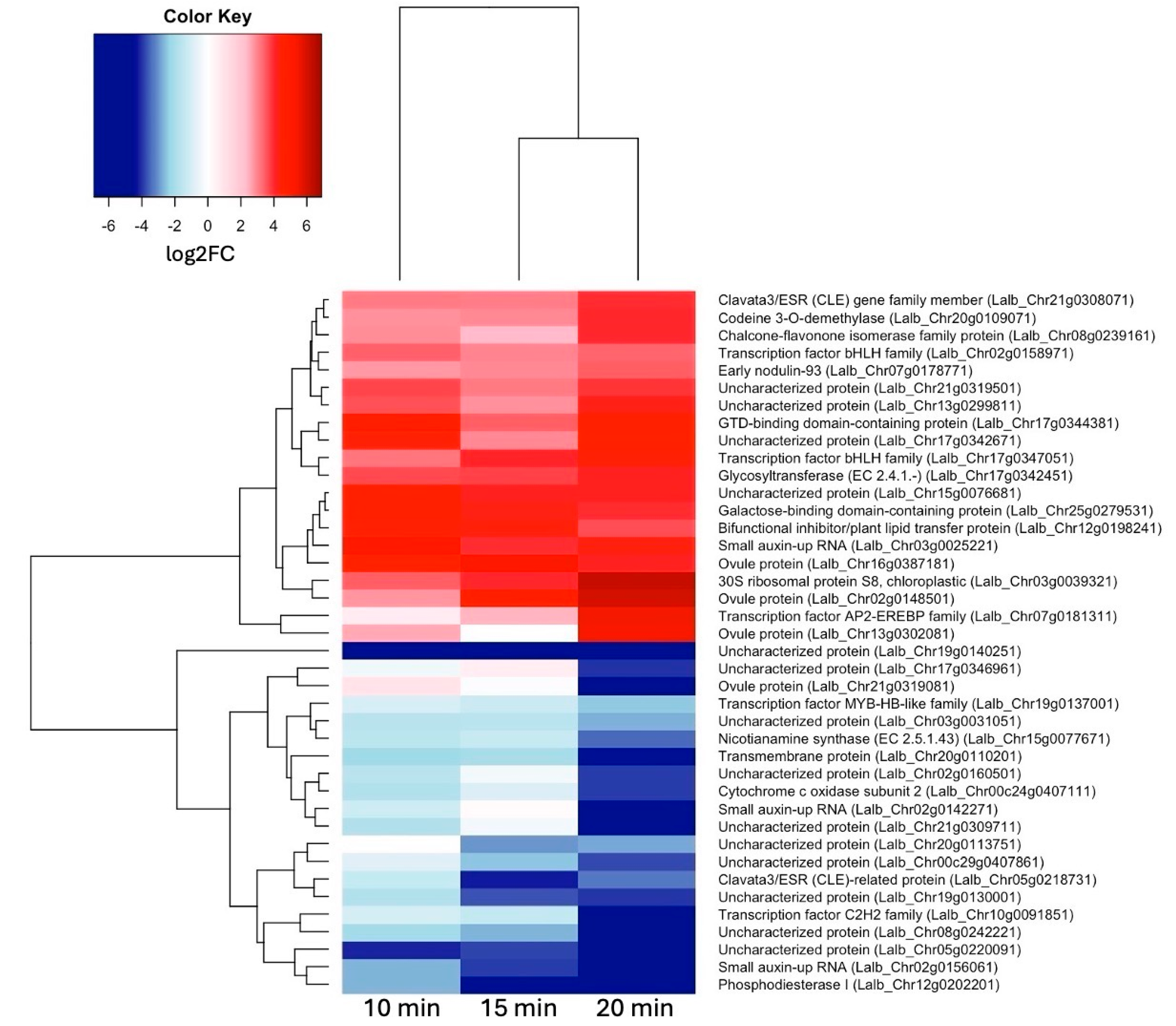
2.2. A Set of 17 Genes Was Up-Regulated at All Three Time Points of Sucrose Exposure
2.3. Auxin- and Gibberellin-Responsive Genes and Two bHLH Transcription Factors Are among the Earliest Up-Regulated Genes
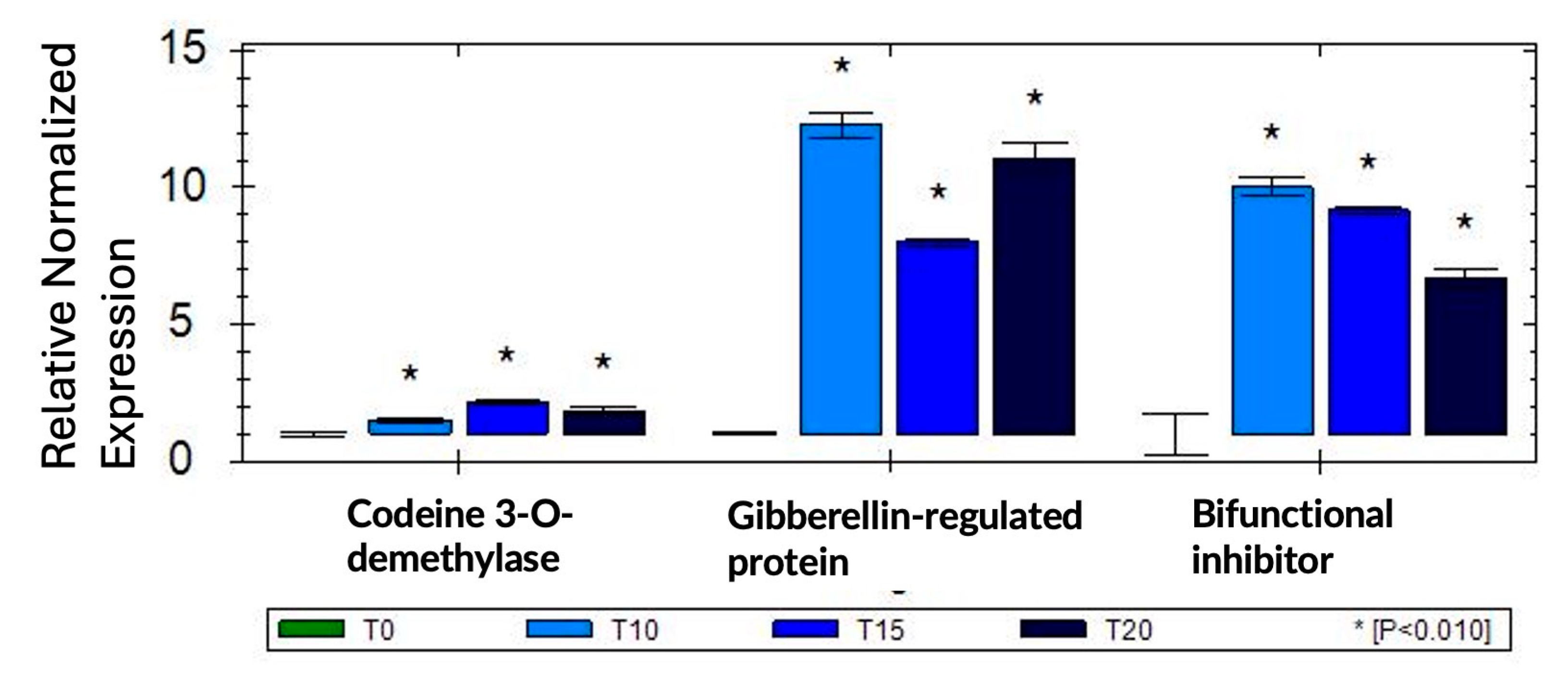
2.4. qRT-PCR Confirms Differential Expression of Selected Genes
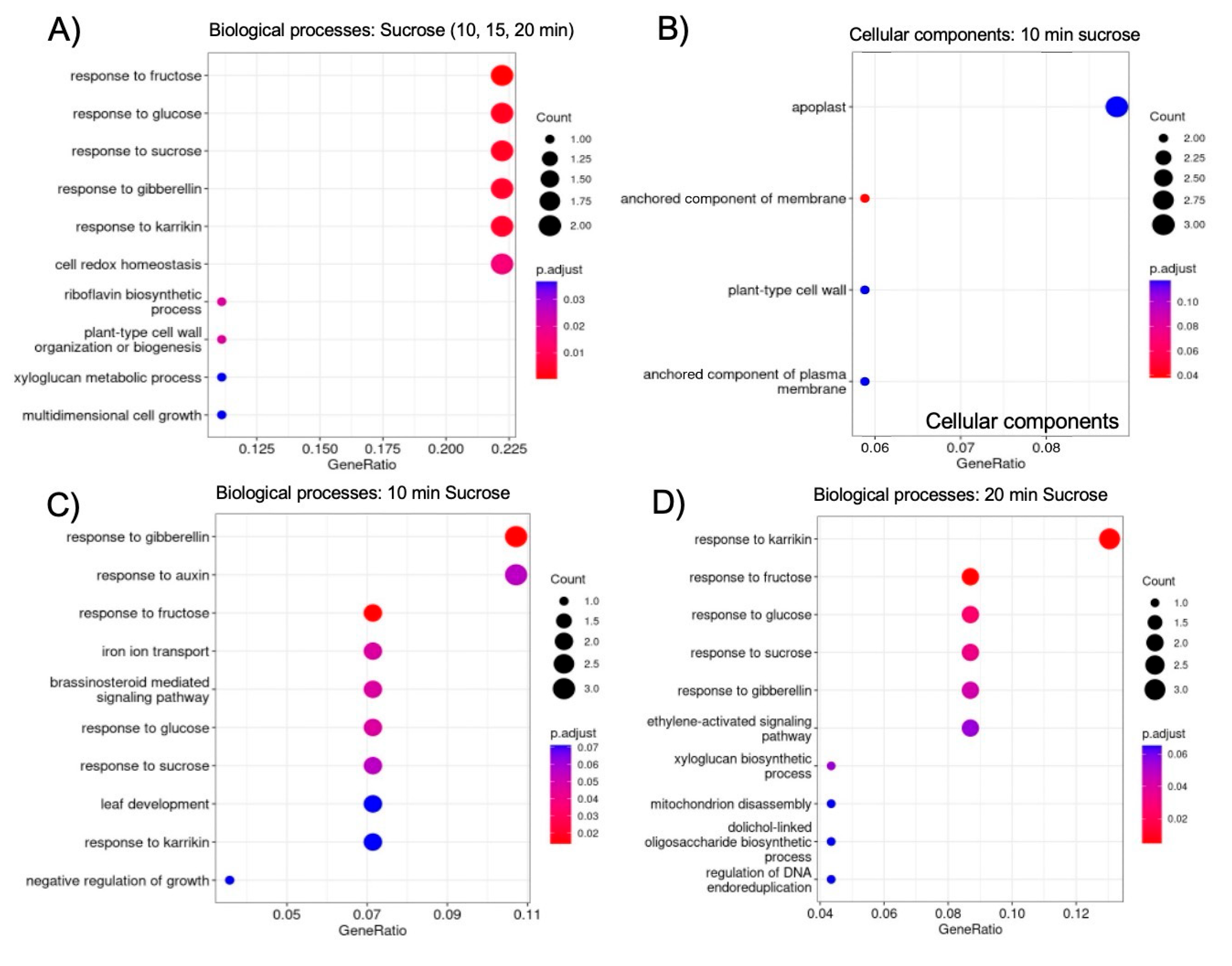
2.5. GO Analysis Reveals Enrichment of Sugar- and Hormone-Responsive Genes
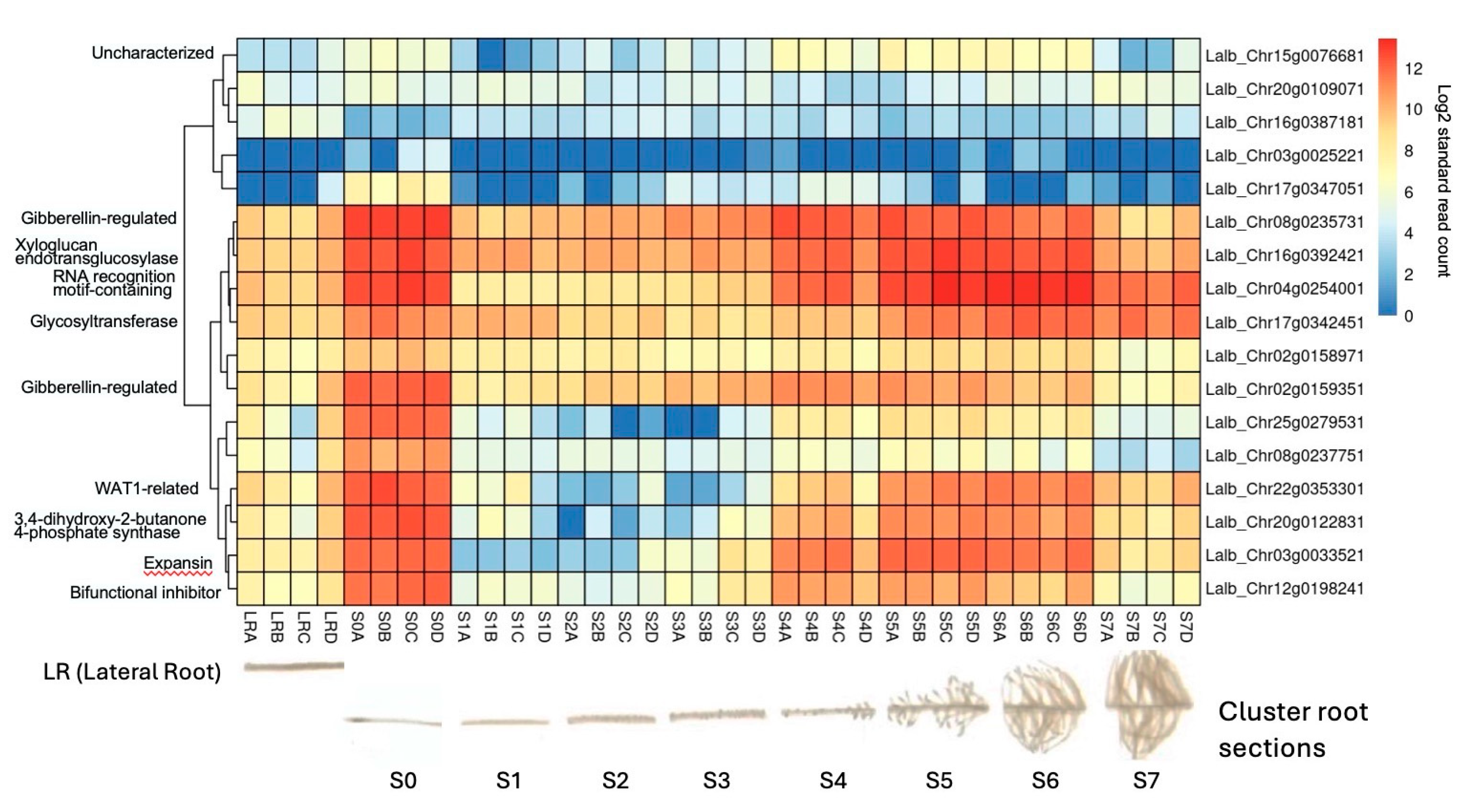
2.6. Several Sucrose-Induced Genes Are Also Expressed in Cluster Roots
3. Discussion
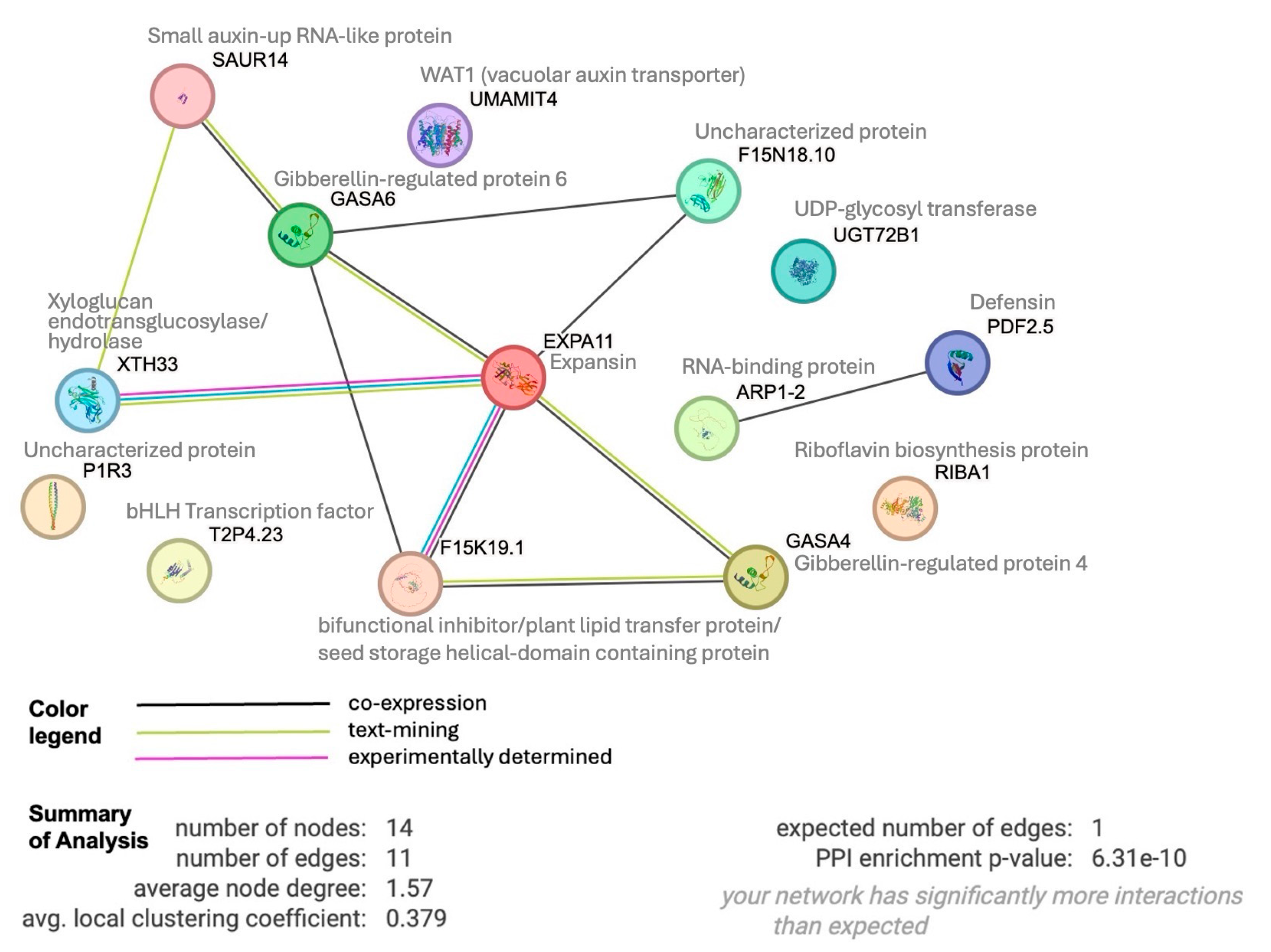
3.1. A Network of Sucrose-Responsive Genes Is Involved in Cell Growth and Differentiation
3.2. Timing and Coordination of Sucrose Responses
4. Materials and Methods
4.1. Plant Growth and Treatments
4.2. RNA Isolation and Quality Check
4.3. cDNA Library Preparation and RNA-Sequencing
4.4. RNA-seq Data Analysis
4.5. Bioinformatic Analysis
4.6. Validation by qRT-PCR
Author Contributions
Funding
Data Availability Statement
References
- Vance, C.P.; Uhde-Stone, C.; Allan, D.L. Phosphorus acquisition and use: critical adaptations by plants for securing a nonrenewable resource. New Phytologist 2003, 157, 423–447. [Google Scholar] [CrossRef]
- Howarth, R.W. Phosphorus in all its formsThe Devil's Element: Phosphorus and a World Out of Balance Dan Egan Norton, 2023. 256 pp. Science 2023, 379, 1096. [Google Scholar] [CrossRef]
- Ahmad, N.; Usman, M.; Ahmad, H.R.; Sabir, M.; Farooqi, Z.U.R.; Shehzad, M.T. Environmental implications of phosphate-based fertilizer industrial waste and its management practices. Environ Monit Assess 2023, 195, 1326. [Google Scholar] [CrossRef]
- Uhde-Stone, C. White lupin: a model system for understanding plant adaptation to low phosphorus availability. In Legume Nitrogen Fixation in Soils with Low Phosphorus Availability; Springer: 2017; pp. 243-280.
- Zanin, L.; Venuti, S.; Marroni, F.; Franco, A.; Morgante, M.; Pinton, R.; Tomasi, N. Physiological and RNA sequencing data of white lupin plants grown under Fe and P deficiency. Data Brief 2019, 25, 104069. [Google Scholar] [CrossRef]
- Neumann, G.; Massonneau, A.; Langlade, N.; Dinkelaker, B.; Hengeler, C.; Römheld, V.; Martinoia, E. Physiological aspects of cluster root function and development in phosphorus-deficient white lupin (Lupinus albus L.). Annals of botany 2000, 85, 909–919. [Google Scholar] [CrossRef]
- Massonneau, A.; Langlade, N.; Léon, S.; Smutny, J.; Vogt, E.; Neumann, G.; Martinoia, E. Metabolic changes associated with cluster root development in white lupin (Lupinus albus L.): relationship between organic acid excretion, sucrose metabolism and energy status. Planta 2001, 213, 534–542. [Google Scholar] [CrossRef]
- Sas, L.; Rengel, Z.; Tang, C. Excess cation uptake, and extrusion of protons and organic acid anions by Lupinus albus under phosphorus deficiency. Plant Sci 2001, 160, 1191–1198. [Google Scholar] [CrossRef]
- Wang, B.; Tang, X.; Cheng, L.; Zhang, A.; Zhang, W.; Zhang, F.; Liu, J.; Cao, Y.; Allan, D.; Vance, C. Nitric oxide is involved in phosphorus deficiency-induced cluster-root development and citrate exudation in white lupin. New Phytologist 2010, 187, 1112–1123. [Google Scholar] [CrossRef]
- Johnson, J.; Allan, D.; Vance, C. Phosphorus Stress-Induced Proteoid Roots Show Altered Metabolism in Lupinus albus. Plant physiology 1994, 104, 657–665. [Google Scholar] [CrossRef]
- Zhou, Y.; Olt, P.; Neuhauser, B.; Moradtalab, N.; Bautista, W.; Uhde-Stone, C.; Neumann, G.; Ludewig, U. Loss of LaMATE impairs isoflavonoid release from cluster roots of phosphorus-deficient white lupin. Physiol Plant 2021, 173, 1207–1220. [Google Scholar] [CrossRef]
- Uhde-Stone, C.; Zinn, K.E.; Ramirez-Yanez, M.; Li, A.; Vance, C.P.; Allan, D.L. Nylon filter arrays reveal differential gene expression in proteoid roots of white lupin in response to phosphorus deficiency. Plant physiology 2003, 131, 1064–1079. [Google Scholar] [CrossRef]
- O'Rourke, J.A.; Yang, S.S.; Miller, S.S.; Bucciarelli, B.; Liu, J.; Rydeen, A.; Bozsoki, Z.; Uhde-Stone, C.; Tu, Z.J.; Allan, D.; et al. An RNA-Seq transcriptome analysis of orthophosphate-deficient white lupin reveals novel insights into phosphorus acclimation in plants. Plant physiology 2013, 161, 705–724. [Google Scholar] [CrossRef]
- Secco, D.; Shou, H.; Whelan, J.; Berkowitz, O. RNA-seq analysis identifies an intricate regulatory network controlling cluster root development in white lupin. BMC Genomics 2014, 15, 230. [Google Scholar] [CrossRef]
- Wang, Z.; Straub, D.; Yang, H.; Kania, A.; Shen, J.; Ludewig, U.; Neumann, G. The regulatory network of cluster-root function and development in phosphate-deficient white lupin (Lupinus albus) identified by transcriptome sequencing. Physiologia plantarum 2014, 151, 323–338. [Google Scholar] [CrossRef]
- Shu, L.; Shen, J.; Rengel, Z.; Tang, C.; Zhang, F. Cluster root formation by Lupinus albus is modified by stratified application of phosphorus in a split-root system. Journal of plant nutrition 2007, 30, 271–288. [Google Scholar] [CrossRef]
- Hammond, J.; White, P. Sucrose transport in the phloem: integrating root responses to phosphorus starvation. J Exp Bot 2008, 59, 93–109. [Google Scholar] [CrossRef]
- Hammond, J.P.; White, P.J. Sugar signaling in root responses to low phosphorus availability. Plant physiology 2011, 156, 1033–1040. [Google Scholar] [CrossRef]
- Lin, X.Y.; Ye, Y.Q.; Fan, S.K.; Jin, C.W.; Zheng, S.J. Increased Sucrose Accumulation Regulates Iron-Deficiency Responses by Promoting Auxin Signaling in Arabidopsis Plants. Plant physiology 2016, 170, 907–920. [Google Scholar] [CrossRef]
- Pontis, H.G. On the scent of the riddle of sucrose. Trends Biochem Sci 1978, 3, 137–139. [Google Scholar] [CrossRef]
- Zhou, K.; Yamagishi, M.; Osaki, M.; Masuda, K. Sugar signalling mediates cluster root formation and phosphorus starvation-induced gene expression in white lupin. J Exp Bot 2008, 59, 2749–2756. [Google Scholar] [CrossRef]
- Wang, Z.; Shen, J.; Ludewig, U.; Neumann, G. A re-assessment of sucrose signaling involved in cluster-root formation and function in phosphate-deficient white lupin (Lupinus albus). Physiologia plantarum 2015, 154, 407–419. [Google Scholar] [CrossRef]
- Lei, M.; Liu, D. Sucrose regulates plant responses to deficiencies in multiple nutrients. Plant Signal Behav 2011, 6, 1247–1249. [Google Scholar] [CrossRef]
- Ruffel, S. Nutrient-related Long-Distance Signals: common players and possible crosstalk. Plant Cell Physiol 2018. [CrossRef]
- Thibaud, M.C.; Gineste, S.; Nussaume, L.; Robaglia, C. Sucrose increases pathogenesis-related PR-2 gene expression in Arabidopsis thaliana through an SA-dependent but NPR1-independent signaling pathway. Plant Physiol Biochem 2004, 42, 81–88. [Google Scholar] [CrossRef]
- Morkunas, I.; Marczak, L.; Stachowiak, J.; Stobiecki, M. Sucrose-induced lupine defense against Fusarium oxysporum. Sucrose-stimulated accumulation of isoflavonoids as a defense response of lupine to Fusarium oxysporum. Plant Physiol Biochem 2005, 43, 363–373. [Google Scholar] [CrossRef]
- Tun, W.; Yoon, J.; Vo, K.T.X.; Cho, L.H.; Hoang, T.V.; Peng, X.; Kim, E.J.; Win, K.; Lee, S.W.; Jung, K.H.; et al. Sucrose preferentially promotes expression of OsWRKY7 and OsPR10a to enhance defense response to blast fungus in rice. Front Plant Sci 2023, 14, 1117023. [Google Scholar] [CrossRef]
- Nidumolu, L.C.M.; Lorilla, K.M.; Chakravarty, I.; Uhde-Stone, C. Soybean Root Transcriptomics: Insights into Sucrose Signaling at the Crossroads of Nutrient Deficiency and Biotic Stress Responses. Plants (Basel) 2023, 12. [Google Scholar] [CrossRef]
- Yoon, J.; Cho, L.H.; Tun, W.; Jeon, J.S.; An, G. Sucrose signaling in higher plants. Plant Sci 2021, 302, 110703. [Google Scholar] [CrossRef]
- Hufnagel, B.; Marques, A.; Soriano, A.; Marques, L.; Divol, F.; Doumas, P.; Sallet, E.; Mancinotti, D.; Carrere, S.; Marande, W.; et al. High-quality genome sequence of white lupin provides insight into soil exploration and seed quality. Nature communications 2020, 11, 492. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 2014, 15, 550. [Google Scholar] [CrossRef]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 2002, 3, RESEARCH0034. [Google Scholar] [CrossRef]
- Johnson, J.; Vance, C.; Allan, D. Phosphorus deficiency in Lupinus albus. Altered lateral root development and enhanced expression of phosphoenolpyruvate carboxylase. Plant physiology 1996, 112, 31–41. [Google Scholar]
- Wang, Z.; Rahman, A.B.; Wang, G.; Ludewig, U.; Shen, J.; Neumann, G. Hormonal interactions during cluster-root development in phosphate-deficient white lupin (Lupinus albus L.). J Plant Physiol 2015, 177, 74–82. [Google Scholar] [CrossRef]
- Liu, J.; Samac, D.; Bucciarelli, B.; Allan, D.; Vance, C. Signaling of phosphorus deficiency-induced gene expression in white lupin requires sugar and phloem transport. the Plant journal : for cell and molecular biology 2005, 41, 257–268. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Kirsch, R.; Koutrouli, M.; Nastou, K.; Mehryary, F.; Hachilif, R.; Gable, A.L.; Fang, T.; Doncheva, N.T.; Pyysalo, S.; et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic acids research 2022. [CrossRef] [PubMed]
- Stortenbeker, N.; Bemer, M. The SAUR gene family: the plant's toolbox for adaptation of growth and development. J Exp Bot 2019, 70, 17–27. [Google Scholar] [CrossRef]
- Bacete, L.; Melida, H.; Miedes, E.; Molina, A. Plant cell wall-mediated immunity: cell wall changes trigger disease resistance responses. The Plant journal : for cell and molecular biology 2018, 93, 614–636. [Google Scholar] [CrossRef] [PubMed]
- Kircher, S.; Schopfer, P. Photosynthetic sucrose drives the lateral root clock in Arabidopsis seedlings. Current biology : CB 2023, 33, 2201-2212 e2203. [CrossRef]
- Liu, F.; Zhang, X.; Lu, C.; Zeng, X.; Li, Y.; Fu, D.; Wu, G. Non-specific lipid transfer proteins in plants: presenting new advances and an integrated functional analysis. J Exp Bot 2015, 66, 5663–5681. [Google Scholar] [CrossRef] [PubMed]
- Kader, J.C. Lipid-Transfer Proteins in Plants. Annu Rev Plant Physiol Plant Mol Biol 1996, 47, 627–654. [Google Scholar] [CrossRef] [PubMed]
- Thumuluri, V.; Almagro Armenteros, J.J.; Johansen, A.R.; Nielsen, H.; Winther, O. DeepLoc 2. Nucleic acids research 2022, 50, W228–W234. [Google Scholar] [CrossRef]
- Zhou, X.; Xiang, Y.; Li, C.; Yu, G. Modulatory Role of Reactive Oxygen Species in Root Development in Model Plant of Arabidopsis thaliana. Front Plant Sci 2020, 11, 485932. [Google Scholar] [CrossRef] [PubMed]
- Kucukoglu, M.; Nilsson, O. CLE peptide signaling in plants–the power of moving around. Physiologia plantarum 2015, 155, 74–87. [Google Scholar] [CrossRef]
- Okamoto, S.; Kawasaki, A.; Makino, Y.; Ishida, T.; Sawa, S. Long-distance translocation of CLAVATA3/ESR-related 2 peptide and its positive effect on roots sucrose status. Plant physiology 2022, 189, 2357–2367. [Google Scholar] [CrossRef] [PubMed]
- Nakagami, S.; Aoyama, T.; Sato, Y.; Kajiwara, T.; Ishida, T.; Sawa, S. CLE3 and its homologs share overlapping functions in the modulation of lateral root formation through CLV1 and BAM1 in Arabidopsis thaliana. The Plant journal : for cell and molecular biology 2023, 113, 1176-1191. [CrossRef]
- Olt, P.; Ding, W.; Schulze, W.X.; Ludewig, U. The LaCLE35 peptide modifies rootlet density and length in cluster roots of white lupin. Plant Cell Environ 2024, 47, 1416–1431. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Shi, P.T.; Zhou, M.; Liu, H.Z.; Xu, X.J.; Liu, W.T.; Chen, K.M. Rapid alkalinization factor: function, regulation, and potential applications in agriculture. Stress Biol 2023, 3, 16. [Google Scholar] [CrossRef] [PubMed]
- Hoagland, D.R.; Arnon, D.I. The water-culture method for growing plants without soil. Circular. California agricultural experiment station 1950, 347. [Google Scholar]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3--new capabilities and interfaces. Nucleic acids research 2012, 40, e115. [Google Scholar] [CrossRef]
|
cDNA library - biological replicate |
Number of sequence reads |
Number of mapped sequences |
Quality mean |
Length range (after trim) |
Length mean (after trim) |
Number of mapped genes |
| t0 - rep 1 | 3,578,071 | 1,946,056 | 11 | 62-8,182 | 700 | 22,960 |
| t0 - rep 2 | 3,820,082 | 1,903,893 | 11 | 61-7,175 | 557 | 21,454 |
| t0 - rep 3 | 1,771,992 | 1,104,448 | 11 | 62-9,278 | 748 | 21,427 |
| t10 - rep 1 | 3,118,626 | 2,063,375 | 11 | 61-8,182 | 755 | 22,639 |
| t10 - rep 2 | 3,682,223 | 2,417,422 | 11 | 61-6,392 | 774 | 23,322 |
| t10 - rep 3 | 2,230,808 | 1,296,886 | 11 | 61-11,327 | 699 | 22,243 |
| t15 - rep 1 | 2,177,882 | 1,043,045 | 11 | 61-6,158 | 556 | 18,894 |
| t15 - rep 2 | 2,424,918 | 1,609,515 | 12 | 62-7,296 | 765 | 22,278 |
| t15 - rep 3 | 4,093,433 | 2,059,972 | 11 | 62-11,327 | 620 | 24,655 |
| t20 - rep 1 | 3,996,371 | 2,656,449 | 11 | 62- 8,486 | 849 | 24,302 |
| t20 - rep 2 | 1,914,530 | 1,328,517 | 11 | 61-6,542 | 843 | 21,933 |
| t20 - rep 3 | 2736983 | 1,450,390 | 11 | 62-12,545 | 578 | 22,544 |
| Gene ID, description |
Relevant GO terms (where available) | 10 min log2FC (pvalue) |
15 min log2FC (pvalue) |
20 min log2FC (pvalue) |
|---|---|---|---|---|
| Lalb_Chr03g0025221, Small auxin-up RNA (auxin-induced protein) |
GO:0009733, response to auxin |
5.1 (0.001) |
4.0 (0.011) |
4.3 (0.006) |
| Lalb_Chr16g0387181, Ovule protein |
GO:0016020 , membrane |
4.6 (0.015) |
5.3 (0.005) |
4.2 (0.030) |
| Lalb_Chr25g0279531, Galactose-binding domain-containing protein |
4.5 (0.003) |
4.3 (0.005) |
4.0 (0.009) |
|
| Lalb_Chr15g0076681, Uncharacterized protein |
4.5 (0.006) |
4.2 (0.012) |
4.2 (0.011) |
|
| Lalb_Chr12g0198241, Bifunctional inhibitor/… domain -containing protein |
4.4 (0.000) |
4.4 (0.000) |
3.7 (0.003) |
|
| Lalb_Chr22g0353301, WAT1-related (vacuolar auxin transport) |
GO:0022857, transmembrane transporter |
4.3 (0.004) |
3.3 (0.029) |
3.2 (0.034) |
| Lalb_Chr04g0254001, RNA recognition motif domain-containing |
GO:0006397 (mRNA processing) | 3.9 (0.004) |
3.8 (0.007) |
3.3 (0.017) |
| Lalb_Chr17g0342451, Glycosyltransferase |
GO:0008194 (UDP-glycosyltransferase activity) |
3.7 (0.014) |
3.8 (0.012) |
4.2 (0.005) |
| Lalb_Chr02g0158971, Transcription factor bHLH UPBEAT1 |
GO:0006355, regulation of DNA-templated transcription | 3.4 (0.000) |
3.0 (0.002) |
3.3 (0.000) |
| Lalb_Chr20g0122831, 3,4-dihydroxy-2-butanone 4-phosphate synthase |
GO:0009231 (riboflavin biosynthetic process) | 3.2 (0.026) |
3.0 (0.040) |
2.9 (0.043) |
| Lalb_Chr17g0347051, Transcription factor bHLH family 61 |
GO:0006355, regulation of DNA-templated transcription | 3.2 (0.037) |
4.1 (0.007) |
4.4 (0.003) |
| Lalb_Chr03g0033521, Expansin |
GO:0009664 (plant-type cell wall organization) | 3.0 (0.011) |
2.6 (0.030) |
2.5 (0.038) |
| Lalb_Chr16g0392421, Xyloglucan endotransglucosylase/hydrolase |
GO:0071555 (cell wall organization) |
3.0 (0.001) |
2.6 (0.005) |
2.5 (0.006) |
| Lalb_Chr20g0109071, Codeine 3-O-demethylase |
GO:0008168, methyltransferase activity | 2.9 (0.029) |
2.9 (0.026) |
4.0 (0.002) |
| Lalb_Chr08g0235731, Gibberellin regulated protein |
GO:0009744 (response to sucrose) GO:0009739 (response to gibberellin) |
2.2 (0.010) |
2.1 (0.014) |
2.0 (0.020) |
| Lalb_Chr02g0159351, Gibberellin regulated protein |
2.1 (0.004) |
2.0 (0.008) |
1.6 (0.029) |
|
| Lalb_Chr08g0237751, Defensin |
GO:0006952, defense response |
1.9 (0.014) |
1.7 (0.024) |
1.6 (0.033) |
| Lalb_Chr19g0140251 Uncharacterized membrane protein |
-6.6 (0.006) |
-6.4 (0,007) |
-6.6 (0.005) |
|
| Lalb_Chr05g0220091 Uncharacterized |
-4.2 (0.015) |
-3.8 (0.024) |
-4.7 (0.008) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




