Submitted:
13 May 2024
Posted:
14 May 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
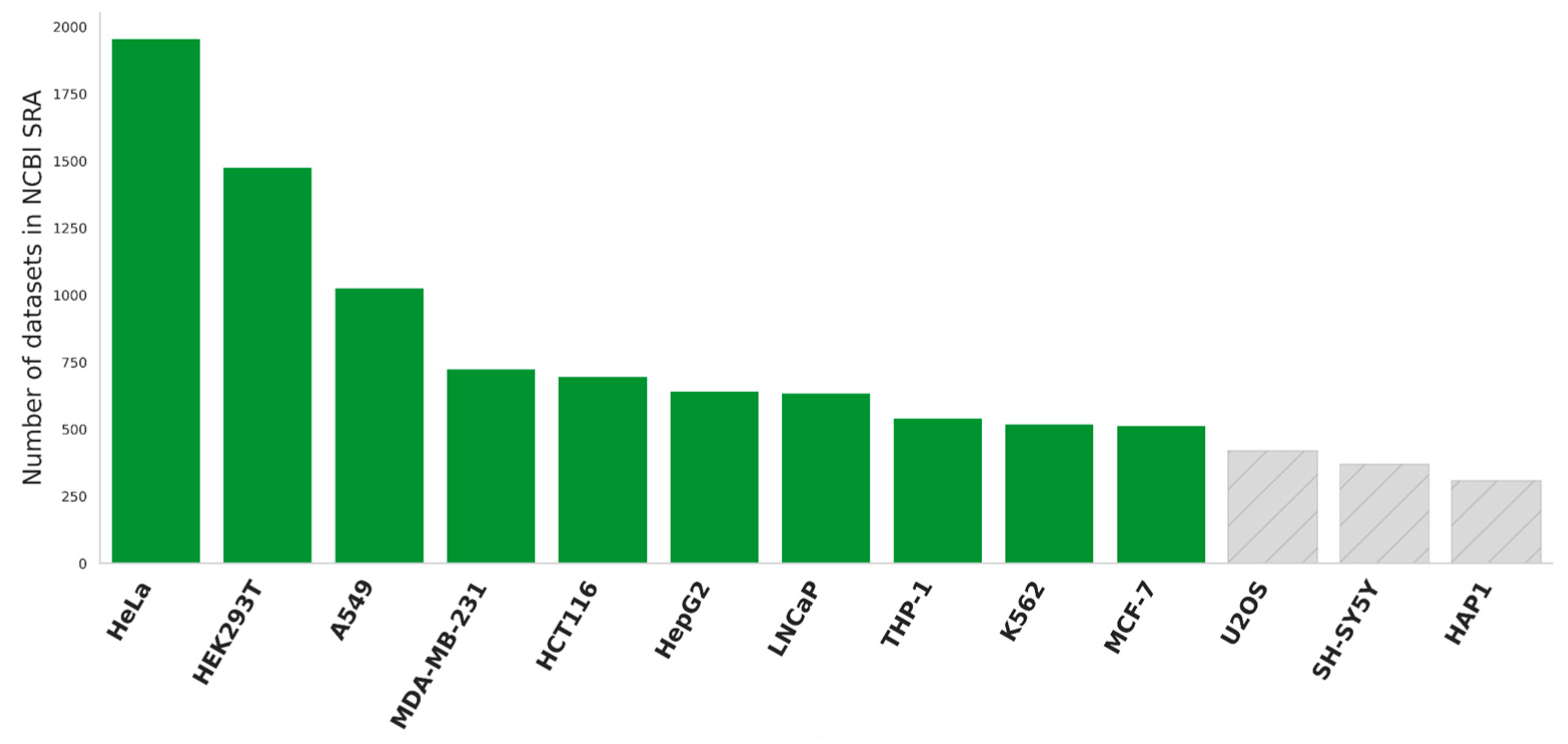
2.1. Data Description
2.2. Algorithm Description
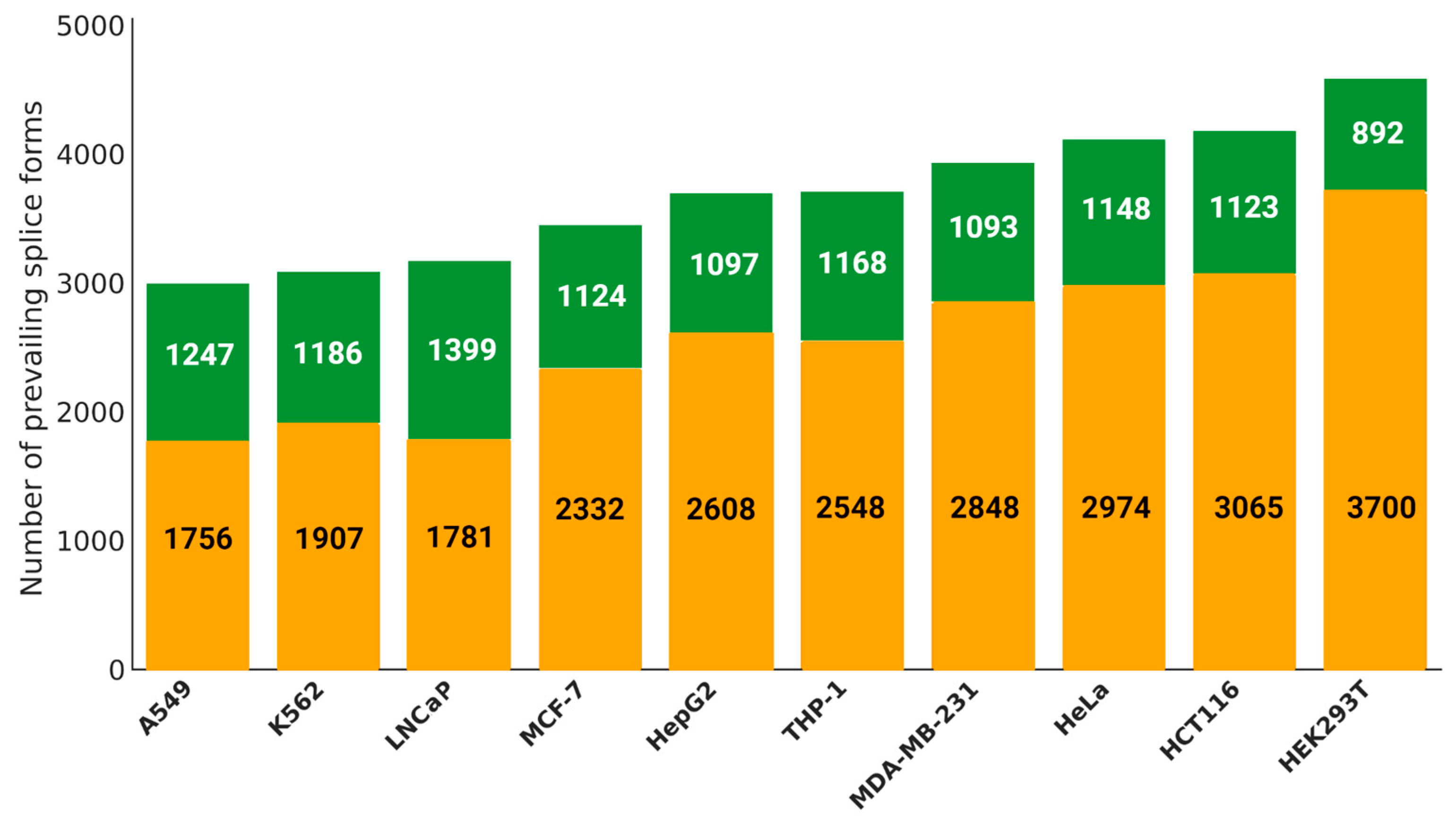
2.3. Splicing Diversity within Cell Lines
2.4. Splicing Diversity across Cell Lines
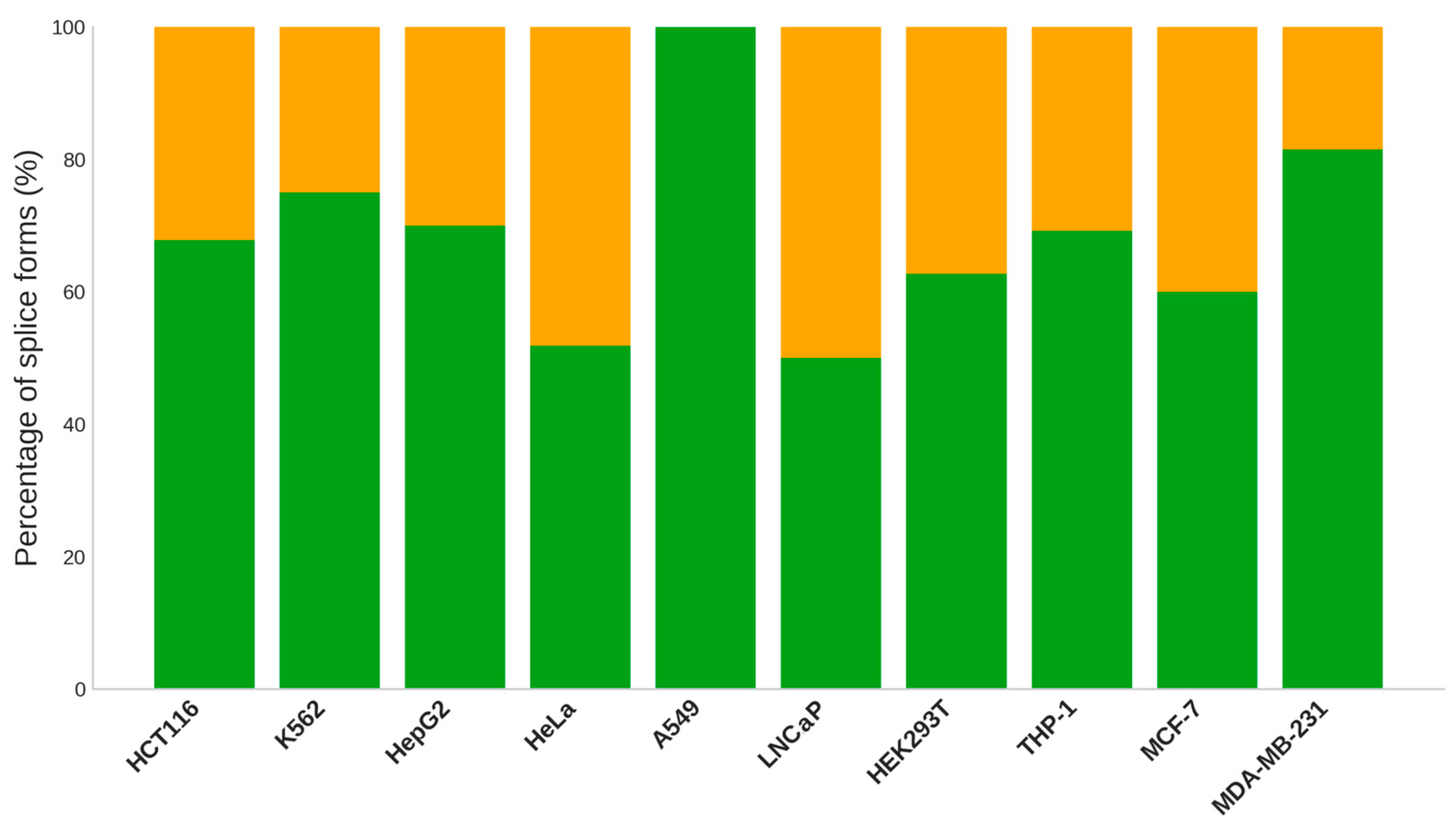
2.5. Comparative Analysis of Splice Variant Prevalence among Cell Lines
2.6. Selection of the Canonical Variant
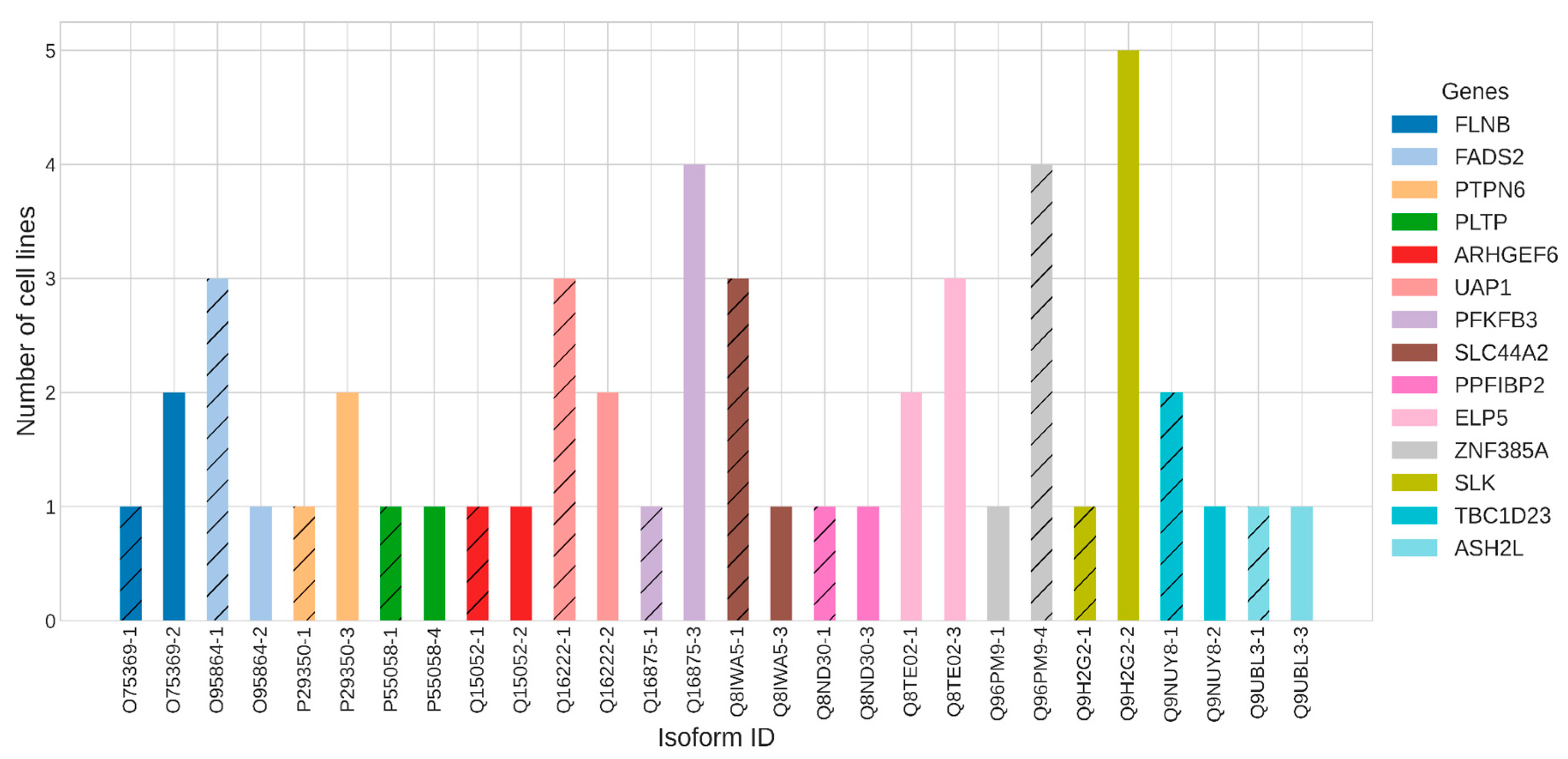
2.7. Relevance to Cancer Development
3. Discussion
4. Materials and Methods
4.1. Data
4.2. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ye, J.; Blelloch, R. Regulation of Pluripotency by RNA Binding Proteins. Cell Stem Cell 2014, 15, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Sebestyén, E.; Zawisza, M.; Eyras, E. Detection of Recurrent Alternative Splicing Switches in Tumor Samples Reveals Novel Signatures of Cancer. Nucleic Acids Res. 2015, 43, 1345–1356. [Google Scholar] [CrossRef] [PubMed]
- Reyes, A.; Huber, W. Alternative Start and Termination Sites of Transcription Drive Most Transcript Isoform Differences across Human Tissues. Nucleic Acids Res. 2018, 46, 582–592. [Google Scholar] [CrossRef] [PubMed]
- Hounkpe, B.W.; Chenou, F.; de Lima, F.; De Paula, E.V. HRT Atlas v1.0 Database: Redefining Human and Mouse Housekeeping Genes and Candidate Reference Transcripts by Mining Massive RNA-Seq Datasets. Nucleic Acids Res. 2021, 49, D947–D955. [Google Scholar] [CrossRef] [PubMed]
- Dolgalev, G.; Poverennaya, E. Quantitative Analysis of Isoform Switching in Cancer. Int. J. Mol. Sci. 2023, 24, 10065. [Google Scholar] [CrossRef] [PubMed]
- Vitting-Seerup, K.; Sandelin, A. The Landscape of Isoform Switches in Human Cancers. Mol. Cancer Res. MCR 2017, 15, 1206–1220. [Google Scholar] [CrossRef] [PubMed]
- Paronetto, M.P.; Passacantilli, I.; Sette, C. Alternative Splicing and Cell Survival: From Tissue Homeostasis to Disease. Cell Death Differ. 2016, 23, 1919–1929. [Google Scholar] [CrossRef] [PubMed]
- Arzumanian, V.A.; Kiseleva, O.I.; Poverennaya, E.V. The Curious Case of the HepG2 Cell Line: 40 Years of Expertise. Int. J. Mol. Sci. 2021, 22, 13135. [Google Scholar] [CrossRef]
- Bao, J.; Vitting-Seerup, K.; Waage, J.; Tang, C.; Ge, Y.; Porse, B.T.; Yan, W. UPF2-Dependent Nonsense-Mediated mRNA Decay Pathway Is Essential for Spermatogenesis by Selectively Eliminating Longer 3’UTR Transcripts. PLoS Genet. 2016, 12, e1005863. [Google Scholar] [CrossRef]
- Behdenna, A.; Colange, M.; Haziza, J.; Gema, A.; Appé, G.; Azencott, C.-A.; Nordor, A. pyComBat, a Python Tool for Batch Effects Correction in High-Throughput Molecular Data Using Empirical Bayes Methods. BMC Bioinform. 2023, 24, 459. [Google Scholar] [CrossRef]
- Yang, Y.; Lin, J.; Guo, S.; Xue, X.; Wang, Y.; Qiu, S.; Cui, J.; Ma, L.; Zhang, X.; Wang, J. RRM2 Protects against Ferroptosis and Is a Tumor Biomarker for Liver Cancer. Cancer Cell Int. 2020, 20, 587. [Google Scholar] [CrossRef]
- Liu, Q.; Guo, L.; Qi, H.; Lou, M.; Wang, R.; Hai, B.; Xu, K.; Zhu, L.; Ding, Y.; Li, C.; et al. A MYBL2 Complex for RRM2 Transactivation and the Synthetic Effect of MYBL2 Knockdown with WEE1 Inhibition against Colorectal Cancer. Cell Death Dis. 2021, 12, 1–11. [Google Scholar] [CrossRef]
- Liu, Y.-T.; Xu, L.; Bennett, L.; Hooks, J.C.; Liu, J.; Zhou, Q.; Liem, P.; Zheng, Y.; Skapek, S.X. Identification of De Novo Enhancers Activated by TGFβ to Drive Expression of CDKN2A and B in HeLa Cells. Mol. Cancer Res. MCR 2019, 17, 1854–1866. [Google Scholar] [CrossRef]
- Landry, J.J.M.; Pyl, P.T.; Rausch, T.; Zichner, T.; Tekkedil, M.M.; Stütz, A.M.; Jauch, A.; Aiyar, R.S.; Pau, G.; Delhomme, N.; et al. The Genomic and Transcriptomic Landscape of a HeLa Cell Line. G3 Bethesda Md 2013, 3, 1213–1224. [Google Scholar] [CrossRef]
- Gong, C.; Lin, Q.; Fang, X.; Jiang, W.; Li, J.; Li, J.; Shi, Y.; Luo, Q.; Liu, Z.; Cen, Y. CTNND1 to Mediate Bone Metastasis of Triple-Negative Breast Cancer via Regulating CXCR4. J. Clin. Oncol. 2021, 39, e13045–e13045. [Google Scholar] [CrossRef]
- Lachmann, A.; Torre, D.; Keenan, A.B.; Jagodnik, K.M.; Lee, H.J.; Wang, L.; Silverstein, M.C.; Ma’ayan, A. Massive Mining of Publicly Available RNA-Seq Data from Human and Mouse. Nat. Commun. 2018, 9, 1366. [Google Scholar] [CrossRef]
- Yuan, C.; Hu, H.; Kuang, M.; Chen, Z.; Tao, X.; Fang, S.; Sun, Y.; Zhang, Y.; Chen, H. Super Enhancer Associated RAI14 Is a New Potential Biomarker in Lung Adenocarcinoma. Oncotarget 2017, 8, 105251–105261. [Google Scholar] [CrossRef] [PubMed]
- Davidson, B.; Stavnes, H.T.; Risberg, B.; Nesland, J.M.; Wohlschlaeger, J.; Yang, Y.; Shih, I.-M.; Wang, T.-L. Gene Expression Signatures Differentiate Adenocarcinoma of Lung and Breast Origin in Effusions. Hum. Pathol. 2012, 43, 684–694. [Google Scholar] [CrossRef]
- Grayeski, P.J.; Weidmann, C.A.; Kumar, J.; Lackey, L.; Mustoe, A.M.; Busan, S.; Laederach, A.; Weeks, K.M. Global 5’-UTR RNA Structure Regulates Translation of a SERPINA1 mRNA. Nucleic Acids Res. 2022, 50, 9689–9704. [Google Scholar] [CrossRef]
- Sullivan, K.D.; Galbraith, M.D.; Andrysik, Z.; Espinosa, J.M. Mechanisms of Transcriptional Regulation by P53. Cell Death Differ. 2018, 25, 133–143. [Google Scholar] [CrossRef]
- Khan, I.; Steeg, P.S. Endocytosis: A Pivotal Pathway for Regulating Metastasis. Br. J. Cancer 2021, 124, 66–75. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.; Kim, Y.-J. RhoBTB3 Regulates Proliferation and Invasion of Breast Cancer Cells via Col1a1. Mol. Cells 2022, 45, 631–639. [Google Scholar] [CrossRef] [PubMed]
- Bertrand-Chapel, A.; Caligaris, C.; Fenouil, T.; Savary, C.; Aires, S.; Martel, S.; Huchedé, P.; Chassot, C.; Chauvet, V.; Cardot-Ruffino, V.; et al. SMAD2/3 Mediate Oncogenic Effects of TGF-β in the Absence of SMAD4. Commun. Biol. 2022, 5, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Girolami, I.; Veronese, N.; Smith, L.; Caruso, M.G.; Reddavide, R.; Leandro, G.; Demurtas, J.; Nottegar, A. The Activation Status of the TGF-β Transducer Smad2 Is Associated with a Reduced Survival in Gastrointestinal Cancers: A Systematic Review and Meta-Analysis. Int. J. Mol. Sci. 2019, 20, 3831. [Google Scholar] [CrossRef] [PubMed]
- Lv, D.; Li, Y.; Zhang, W.; Alvarez, A.A.; Song, L.; Tang, J.; Gao, W.-Q.; Hu, B.; Cheng, S.-Y.; Feng, H. TRIM24 Is an Oncogenic Transcriptional Co-Activator of STAT3 in Glioblastoma. Nat. Commun. 2017, 8, 1454. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.K.; Allton, K.; Duncan, A.D.; Barton, M.C. TRIM24 Is a P53-Induced E3-Ubiquitin Ligase That Undergoes ATM-Mediated Phosphorylation and Autodegradation during DNA Damage. Mol. Cell. Biol. 2014, 34, 2695–2709. [Google Scholar] [CrossRef]
- Khan, T.; Kryza, T.; Lyons, N.J.; He, Y.; Hooper, J.D. The CDCP1 Signaling Hub: A Target for Cancer Detection and Therapeutic Intervention. Cancer Res. 2021, 81, 2259–2269. [Google Scholar] [CrossRef] [PubMed]
- Ding, Z.-Y.; Li, R.; Zhang, Q.-J.; Wang, Y.; Jiang, Y.; Meng, Q.-Y.; Xi, Q.-L.; Wu, G.-H. Prognostic Role of Cyclin D2/D3 in Multiple Human Malignant Neoplasms: A Systematic Review and Meta-Analysis. Cancer Med. 2019, 8, 2717–2729. [Google Scholar] [CrossRef] [PubMed]
- Blanca, A.; Requena, M.J.; Alvarez, J.; Cheng, L.; Montironi, R.; Raspollini, M.R.; Reymundo, C.; Lopez-Beltran, A. FGFR3 and Cyclin D3 as Urine Biomarkers of Bladder Cancer Recurrence. Biomark. Med. 2016, 10, 243–253. [Google Scholar] [CrossRef]
- Arzumanian, V.; Pyatnitskiy, M.; Poverennaya, E. Comparative Transcriptomic Analysis of Three Common Liver Cell Lines. Int. J. Mol. Sci. 2023, 24, 8791. [Google Scholar] [CrossRef]
- Snezhkina, A.V.; Krasnov, G.S.; Zaretsky, A.R.; Zhavoronkov, A.; Nyushko, K.M.; Moskalev, A.A.; Karpova, I.Y.; Afremova, A.I.; Lipatova, A.V.; Kochetkov, D.V.; et al. Differential Expression of Alternatively Spliced Transcripts Related to Energy Metabolism in Colorectal Cancer. BMC Genomics 2016, 17, 1011. [Google Scholar] [CrossRef]
- Trivedi, U.H.; Cezard, T.; Bridgett, S.; Montazam, A.; Nichols, J.; Blaxter, M.; Gharbi, K. Quality Control of Next-Generation Sequencing Data without a Reference. Front. Genet. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Patro, R.; Duggal, G.; Love, M.I.; Irizarry, R.A.; Kingsford, C. Salmon Provides Fast and Bias-Aware Quantification of Transcript Expression. Nat. Methods 2017, 14, 417–419. [Google Scholar] [CrossRef]
- Smedley, D.; Haider, S.; Ballester, B.; Holland, R.; London, D.; Thorisson, G.; Kasprzyk, A. BioMart--Biological Queries Made Easy. BMC Genomics 2009, 10, 22. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



