Submitted:
07 May 2024
Posted:
10 May 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
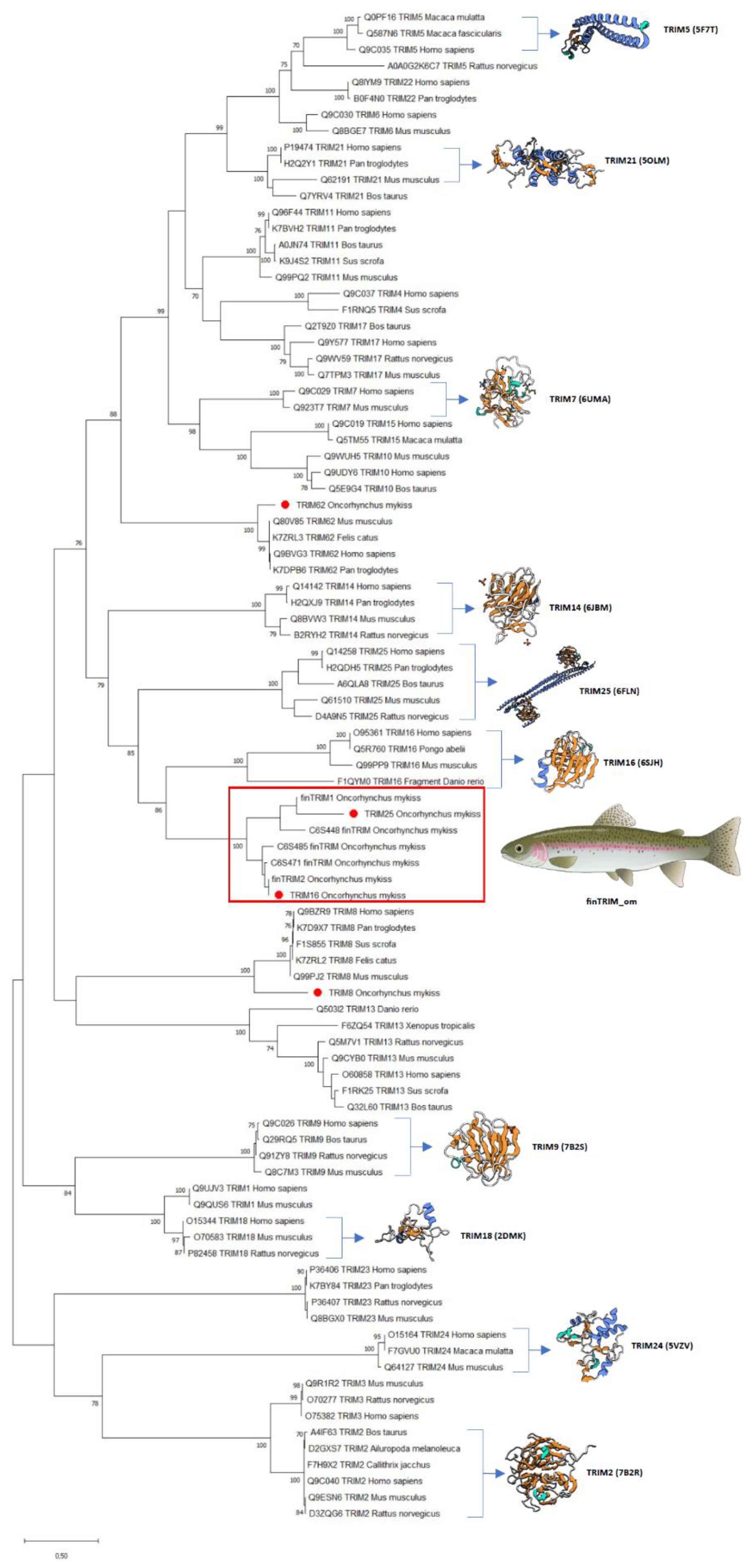
2.1. Novel TRIM-like Sequences Identified in RTgill-W1 Cells
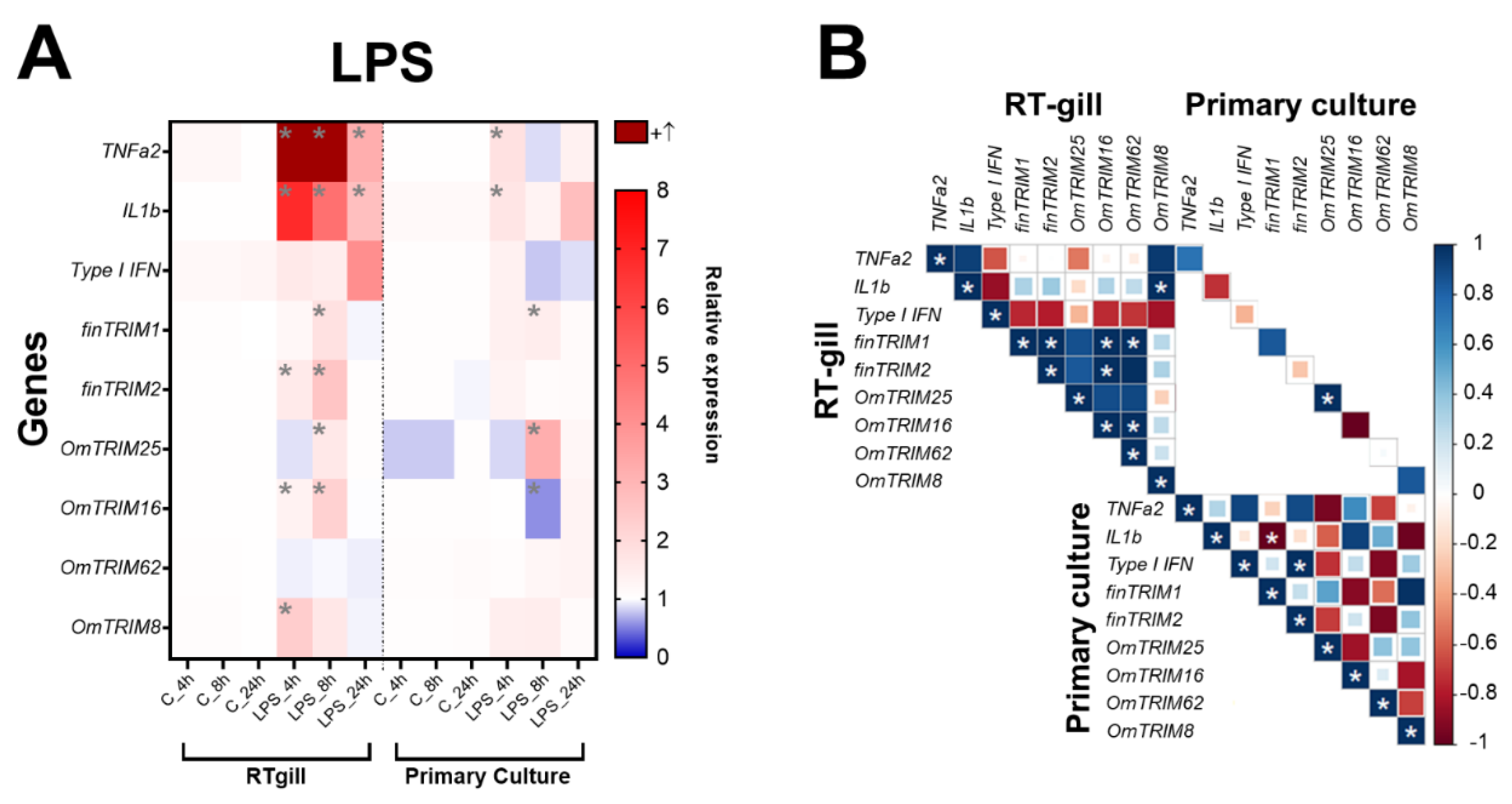
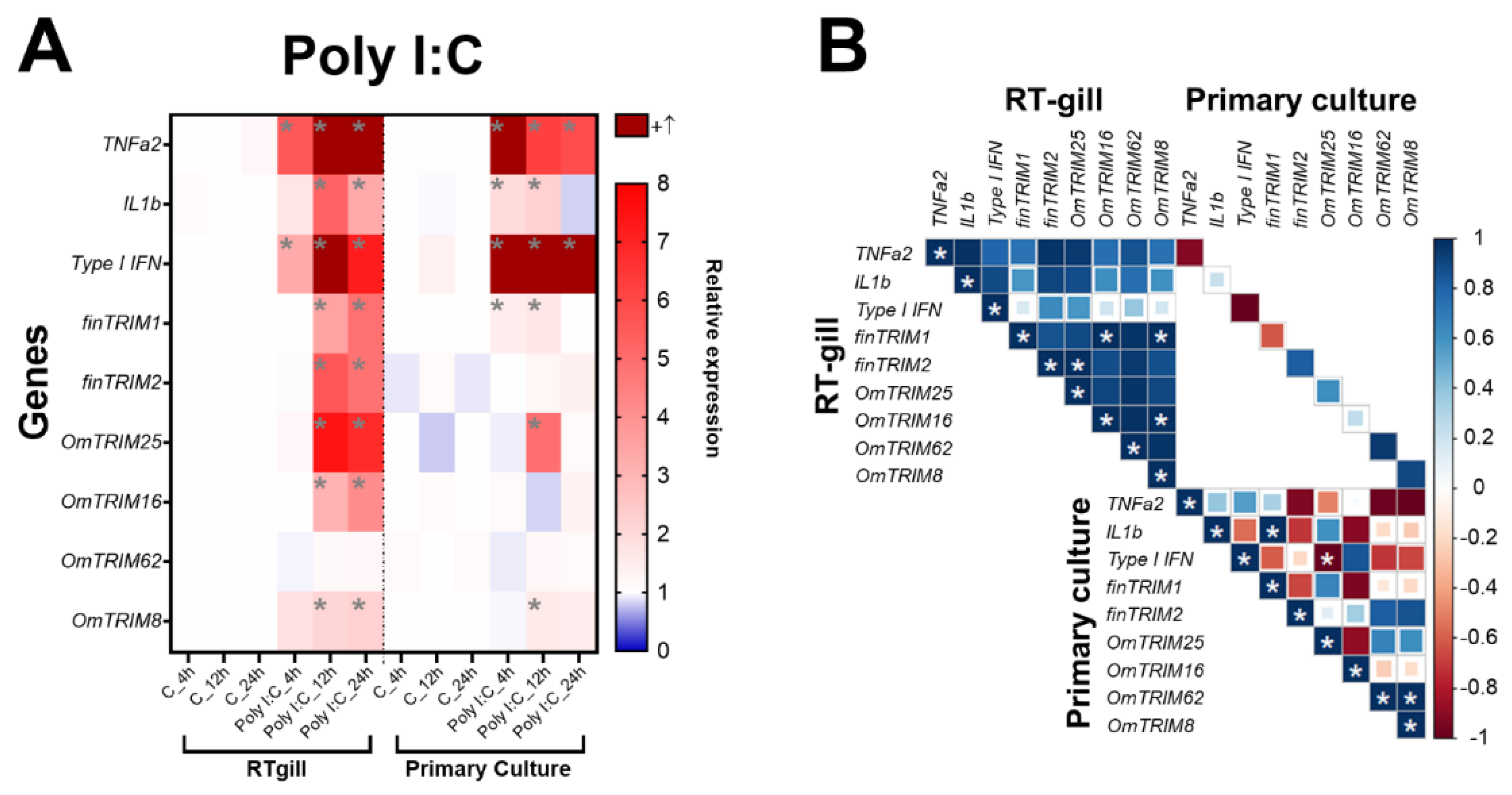
2.2. Differential Expression of Novel TRIM-like Genes upon Stimulation by MAMPs
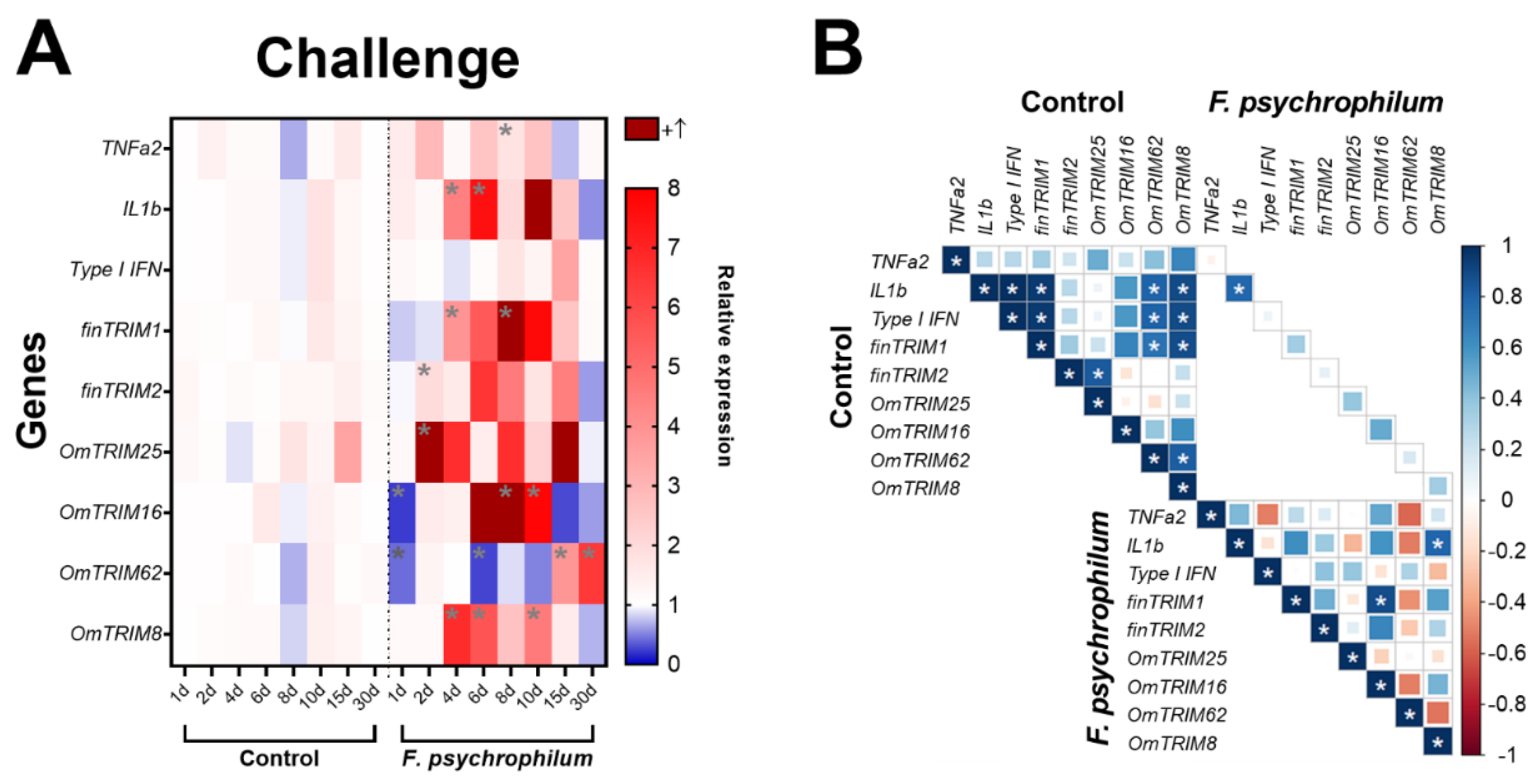
2.3. Rainbow Trout Infected with a Bacterial Pathogen Showed a Time-Depending Expression of TRIMs in Gill Tissue
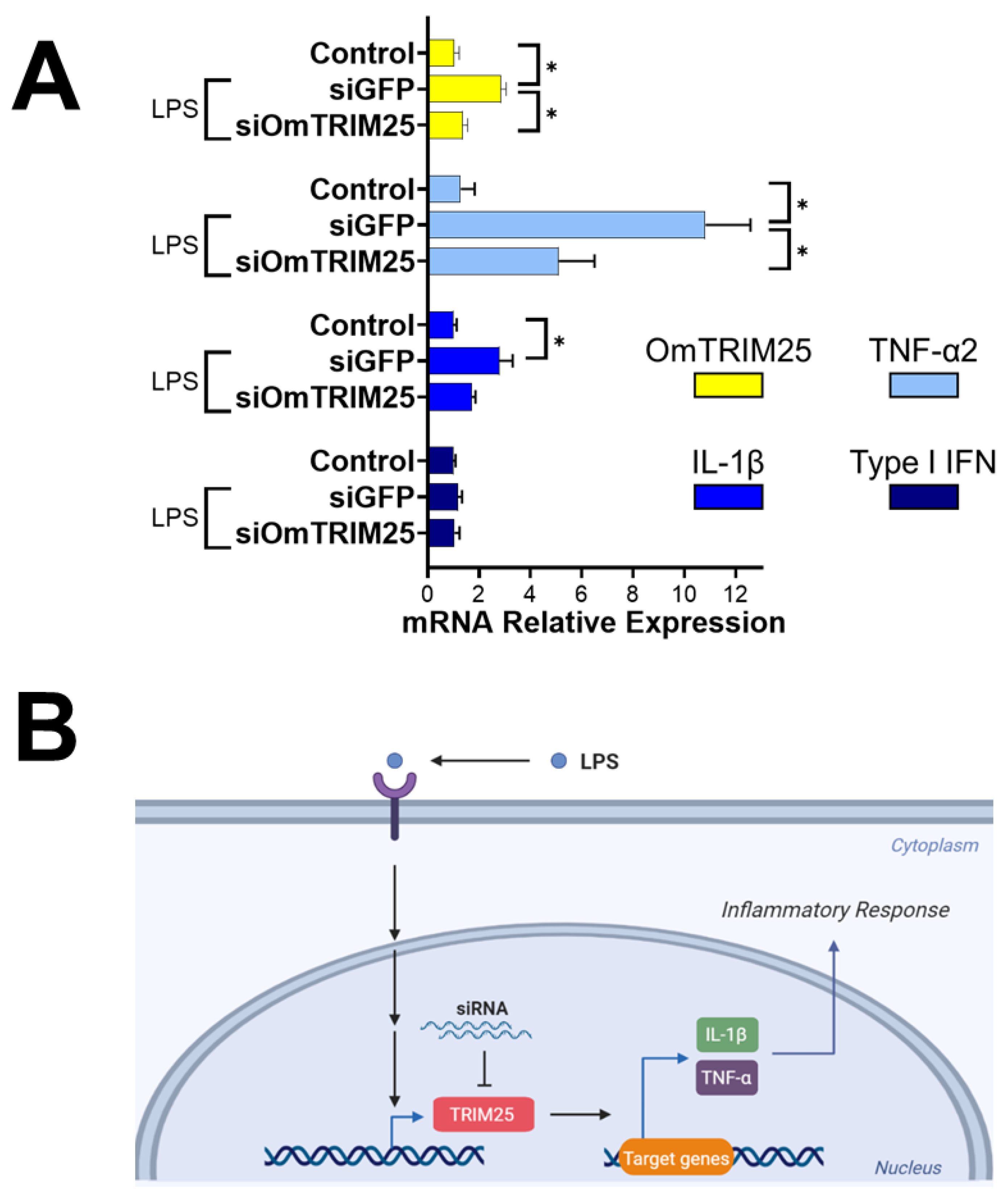
2.4. OmTRIM25 Is Required to Trigger the Expression of TNF-α2 and IL-1β in RTgill-W1 Cells during LPS Stimulation
3. Discussion
4. Materials and Methods
4.1. Chemical and Reagents
4.2. Culturing of RTgill-W1 Cells and Treatment
4.3. Primary Culturing of Rainbow Trout Gill Cells
4.4. Fish Handling and F. psychrophilum Growth Conditions
4.5. Experimental Design and Fish Sampling
4.6. Molecular Cloning and Sequencing
4.7. Identification of TRIM CDS in RTgill-W1
4.8. RNAi Gene-Silencing Assay
4.9. Total RNA Extraction, cDNA Synthesis and Reverse Transcription Quantitative Real-Time PCR (RT-qPCR)
4.10. Bioinformatics
4.11. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Malmstrøm, M.; Matschiner, M.; Tørresen, O.K.; Star, B.; Snipen, L.G.; Hansen, T.F.; Baalsrud, H.T.; Nederbragt, A.J.; Hanel, R.; Salzburger, W.; et al. Evolution of the Immune System Influences Speciation Rates in Teleost Fishes. Nat. Genet. 2016, 48, 1204–1210. [Google Scholar] [CrossRef] [PubMed]
- Yatim, K.M.; Lakkis, F.G. A Brief Journey through the Immune System. Clin. J. Am. Soc. Nephrol. CJASN 2015, 10, 1274–1281. [Google Scholar] [CrossRef]
- Plouffe, D.A.; Hanington, P.C.; Walsh, J.G.; Wilson, E.C.; Belosevic, M. Comparison of Select Innate Immune Mechanisms of Fish and Mammals. Xenotransplantation 2005, 12, 266–277. [Google Scholar] [CrossRef]
- Xu, Y.; Zhu, S.-W.; Li, Q.-W. Lamprey: A Model for Vertebrate Evolutionary Research. Zool. Res. 2016, 37, 263–269. [Google Scholar] [CrossRef]
- Zhang, X.-J.; Zhang, X.-Y.; Wang, P.; Zhang, Y.-A. Identification of Another Primordial CD80/86 Molecule in Rainbow Trout: Insights into the Origin and Evolution of CD80 and CD86 in Vertebrates. Dev. Comp. Immunol. 2018, 89, 73–82. [Google Scholar] [CrossRef]
- Hatakeyama, S. TRIM Family Proteins: Roles in Autophagy, Immunity, and Carcinogenesis. Trends Biochem. Sci. 2017, 42, 297–311. [Google Scholar] [CrossRef] [PubMed]
- Vunjak, M.; Versteeg, G.A. TRIM Proteins. Curr. Biol. CB 2019, 29, R42–R44. [Google Scholar] [CrossRef]
- Esposito, D.; Koliopoulos, M.G.; Rittinger, K. Structural Determinants of TRIM Protein Function. Biochem. Soc. Trans. 2017, 45, 183–191. [Google Scholar] [CrossRef]
- Stevens, R.V.; Esposito, D.; Rittinger, K. Characterisation of Class VI TRIM RING Domains: Linking RING Activity to C-Terminal Domain Identity. Life Sci. Alliance 2019, 2. [Google Scholar] [CrossRef]
- Ozato, K.; Shin, D.-M.; Chang, T.-H.; Morse, H.C. TRIM Family Proteins and Their Emerging Roles in Innate Immunity. Nat. Rev. Immunol. 2008, 8, 849–860. [Google Scholar] [CrossRef]
- Versteeg, G.A.; Benke, S.; García-Sastre, A.; Rajsbaum, R. InTRIMsic Immunity: Positive and Negative Regulation of Immune Signaling by Tripartite Motif Proteins. Cytokine Growth Factor Rev. 2014, 25, 563–576. [Google Scholar] [CrossRef]
- Rajsbaum, R.; García-Sastre, A.; Versteeg, G.A. TRIMmunity: The Roles of the TRIM E3-Ubiquitin Ligase Family in Innate Antiviral Immunity. J. Mol. Biol. 2014, 426, 1265–1284. [Google Scholar] [CrossRef]
- van Gent, M.; Sparrer, K.M.J.; Gack, M.U. TRIM Proteins and Their Roles in Antiviral Host Defenses. Annu. Rev. Virol. 2018, 5, 385–405. [Google Scholar] [CrossRef]
- Kimura, T.; Jain, A.; Choi, S.W.; Mandell, M.A.; Johansen, T.; Deretic, V. TRIM-Directed Selective Autophagy Regulates Immune Activation. Autophagy 2017, 13, 989–990. [Google Scholar] [CrossRef]
- Weng, L.; Mitoma, H.; Trichot, C.; Tricot, C.; Bao, M.; Liu, Y.; Zhang, Z.; Liu, Y.-J. The E3 Ubiquitin Ligase Tripartite Motif 33 Is Essential for Cytosolic RNA-Induced NLRP3 Inflammasome Activation. J. Immunol. Baltim. Md 1950 2014, 193, 3676–3682. [Google Scholar] [CrossRef]
- Du Pasquier, L. Fish “n” TRIMs. J. Biol. 2009, 8, 50. [Google Scholar] [CrossRef]
- Langevin, C.; Levraud, J.-P.; Boudinot, P. Fish Antiviral Tripartite Motif (TRIM) Proteins. Fish Shellfish Immunol. 2019, 86, 724–733. [Google Scholar] [CrossRef]
- Wang, Y.; Li, Z.; Lu, Y.; Hu, G.; Lin, L.; Zeng, L.; Zhou, Y.; Liu, X. Molecular Characterization, Tissue Distribution and Expression, and Potential Antiviral Effects of TRIM32 in the Common Carp (Cyprinus Carpio). Int. J. Mol. Sci. 2016, 17. [Google Scholar] [CrossRef]
- Wang, Y.; Kuang, M.; Lu, Y.; Lin, L.; Liu, X. Characterization and Biological Function Analysis of the TRIM47 Gene from Common Carp (Cyprinus Carpio). Gene 2017, 627, 188–193. [Google Scholar] [CrossRef] [PubMed]
- Han, R.; Wang, R.; Zhao, Q.; Han, Y.; Zong, S.; Miao, S.; Song, W.; Wang, L. Trim69 Regulates Zebrafish Brain Development by Ap-1 Pathway. Sci. Rep. 2016, 6, 24034. [Google Scholar] [CrossRef] [PubMed]
- van der Aa, L.M.; Levraud, J.-P.; Yahmi, M.; Lauret, E.; Briolat, V.; Herbomel, P.; Benmansour, A.; Boudinot, P. A Large New Subset of TRIM Genes Highly Diversified by Duplication and Positive Selection in Teleost Fish. BMC Biol. 2009, 7, 7. [Google Scholar] [CrossRef]
- Kim, J.; Kim, J.-W.; Kim, D.-G.; Nam, B.-H.; Kim, Y.-O.; Park, J.Y.; Kong, H.J. Molecular Characterization of Rhodeus Uyekii Tripartite Motif Protein 1 (TRIM1) Involved in IFN-γ/LPS-Induced NF-ΚB Signaling. Fish Shellfish Immunol. 2018, 79, 42–51. [Google Scholar] [CrossRef]
- Koppang, E.O.; Kvellestad, A.; Fischer, U. 5 - Fish Mucosal Immunity: Gill. In Mucosal Health in Aquaculture; Beck, B.H., Peatman, E., Eds.; Academic Press: San Diego, 2015; pp. 93–133. ISBN 978-0-12-417186-2. [Google Scholar]
- Huang, Y.; Yu, Y.; Yang, Y.; Yang, M.; Zhou, L.; Huang, X.; Qin, Q. Fish TRIM8 Exerts Antiviral Roles through Regulation of the Proinflammatory Factors and Interferon Signaling. Fish Shellfish Immunol. 2016, 54, 435–444. [Google Scholar] [CrossRef] [PubMed]
- Maarifi, G.; Smith, N.; Maillet, S.; Moncorgé, O.; Chamontin, C.; Edouard, J.; Sohm, F.; Blanchet, F.P.; Herbeuval, J.-P.; Lutfalla, G.; et al. TRIM8 Is Required for Virus-Induced IFN Response in Human Plasmacytoid Dendritic Cells. Sci. Adv. 2019, 5, eaax3511. [Google Scholar] [CrossRef]
- Koliopoulos, M.G.; Lethier, M.; van der Veen, A.G.; Haubrich, K.; Hennig, J.; Kowalinski, E.; Stevens, R.V.; Martin, S.R.; Reis e Sousa, C.; Cusack, S.; et al. Molecular Mechanism of Influenza A NS1-Mediated TRIM25 Recognition and Inhibition. Nat. Commun. 2018, 9, 1820. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, K.; Huang, Y.; Sun, M.; Tian, Q.; Zhang, S.; Qin, Y. TRIM25 Promotes TNF-α-Induced NF-ΚB Activation through Potentiating the K63-Linked Ubiquitination of TRAF2. J. Immunol. Baltim. Md 1950 2020, 204, 1499–1507. [Google Scholar] [CrossRef]
- Cao, Z.; Conway, K.L.; Heath, R.J.; Rush, J.S.; Leshchiner, E.S.; Ramirez-Ortiz, Z.G.; Nedelsky, N.B.; Huang, H.; Ng, A.; Gardet, A.; et al. Ubiquitin Ligase TRIM62 Regulates CARD9-Mediated Anti-Fungal Immunity and Intestinal Inflammation. Immunity 2015, 43, 715–726. [Google Scholar] [CrossRef]
- Gack, M.U.; Shin, Y.C.; Joo, C.-H.; Urano, T.; Liang, C.; Sun, L.; Takeuchi, O.; Akira, S.; Chen, Z.; Inoue, S.; et al. TRIM25 RING-Finger E3 Ubiquitin Ligase Is Essential for RIG-I-Mediated Antiviral Activity. Nature 2007, 446, 916–920. [Google Scholar] [CrossRef]
- Shen, Y.; Li, N.L.; Wang, J.; Liu, B.; Lester, S.; Li, K. TRIM56 Is an Essential Component of the TLR3 Antiviral Signaling Pathway. J. Biol. Chem. 2012, 287, 36404–36413. [Google Scholar] [CrossRef] [PubMed]
- Nematollahi, A.; Decostere, A.; Pasmans, F.; Haesebrouck, F. Flavobacterium Psychrophilum Infections in Salmonid Fish. J. Fish Dis. 2003, 26, 563–574. [Google Scholar] [CrossRef] [PubMed]
- Schulte, W.; Bernhagen, J.; Bucala, R. Cytokines in Sepsis: Potent Immunoregulators and Potential Therapeutic Targets—An Updated View. Mediators Inflamm. 2013, 2013, e165974. [Google Scholar] [CrossRef]
- Cellular and Molecular Immunology - 9th Edition. Available online: https://www.elsevier.com/books/cellular-and-molecular-immunology/abbas/978-0-323-47978-3 (accessed on 24 June 2021).
- Kawai, T.; Akira, S. Regulation of Innate Immune Signalling Pathways by the Tripartite Motif (TRIM) Family Proteins. EMBO Mol. Med. 2011, 3, 513–527. [Google Scholar] [CrossRef]
- Chikuma, S.; Suita, N.; Okazaki, I.-M.; Shibayama, S.; Honjo, T. TRIM28 Prevents Autoinflammatory T Cell Development in Vivo. Nat. Immunol. 2012, 13, 596–603. [Google Scholar] [CrossRef]
- Ahn, Y.; Hwang, J.-H.; Zheng, Z.; Bang, D.; Kim, D.-Y. Enhancement of Th1/Th17 Inflammation by TRIM21 in Behçet’s Disease. Sci. Rep. 2017, 7, 3018. [Google Scholar] [CrossRef] [PubMed]
- Venuto, S.; Merla, G. E3 Ubiquitin Ligase TRIM Proteins, Cell Cycle and Mitosis. Cells 2019, 8, E510. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.M.; Choi, S.S.; Lee, Y.H.; Khim, K.W.; Yoon, S.; Kim, B.-G.; Nam, D.; Suh, P.-G.; Myung, K.; Choi, J.H. The E3 Ubiquitin Ligase TRIM25 Regulates Adipocyte Differentiation via Proteasome-Mediated Degradation of PPARγ. Exp. Mol. Med. 2018, 50, 1–11. [Google Scholar] [CrossRef]
- Lu, M.; Zhu, X.; Yang, Z.; Zhang, W.; Sun, Z.; Ji, Q.; Chen, X.; Zhu, J.; Wang, C.; Nie, S. E3 Ubiquitin Ligase Tripartite Motif 7 Positively Regulates the TLR4-Mediated Immune Response via Its E3 Ligase Domain in Macrophages. Mol. Immunol. 2019, 109, 126–133. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.-J.; Liu, T.-T.; Ran, Y.; Li, Y.; Zhang, X.-D.; Shu, H.-B.; Wang, Y.-Y. The E3 Ubiquitin Ligase MIB1 Negatively Regulates Basal IκBα Level and Modulates NF-ΚB Activation. Cell Res. 2012, 22, 603–606. [Google Scholar] [CrossRef] [PubMed]
- Zemirli, N.; Pourcelot, M.; Dogan, N.; Vazquez, A.; Arnoult, D. The E3 Ubiquitin Ligase RNF121 Is a Positive Regulator of NF-ΚB Activation. Cell Commun. Signal. 2014, 12, 72. [Google Scholar] [CrossRef]
- Álvarez, C.A.; Ramírez-Cepeda, F.; Santana, P.; Torres, E.; Cortés, J.; Guzmán, F.; Schmitt, P.; Mercado, L. Insights into the Diversity of NOD-like Receptors: Identification and Expression Analysis of NLRC3, NLRC5 and NLRX1 in Rainbow Trout. Mol. Immunol. 2017, 87, 102–113. [Google Scholar] [CrossRef]
- 43. Enfermedades infecciosas del cultivo de salmónidos en Chile y el Mundo | ISBN 978-956-8861-01-8 - Libro.
- Urdaci, M.C.; Chakroun, C.; Faure, D.; Bernardet, J.F. Development of a Polymerase Chain Reaction Assay for Identification and Detection of the Fish Pathogen Flavobacterium Psychrophilum. Res. Microbiol. 1998, 149, 519–530. [Google Scholar] [CrossRef] [PubMed]
- Bernardet, J.-F.; Nakagawa, Y.; Holmes, B. ; Subcommittee On The Taxonomy Of Flavobacterium And Cytophaga-Like Bacteria Of The International Committee On Systematics Of Prokaryotes, null Proposed Minimal Standards for Describing New Taxa of the Family Flavobacteriaceae and Emended Description of the Family. Int. J. Syst. Evol. Microbiol. 2002, 52, 1049–1070. [Google Scholar] [CrossRef] [PubMed]
- Valdebenito, S.; Avendaño-Herrera, R. Phenotypic, Serological and Genetic Characterization of Flavobacterium Psychrophilum Strains Isolated from Salmonids in Chile. J. Fish Dis. 2009, 32, 321–333. [Google Scholar] [CrossRef] [PubMed]
- Muñoz, J.L.P.; Ocampos, D.; Poblete-Morales, M.; Oyarzún, R.; Morera, F.J.; Tapia-Cammas, D.; Avendaño-Herrera, R.; Vargas-Chacoff, L. Effect of Flavobacterium Psychrophilum on the Neuroendocrine Response of Rainbow Trout (Oncorhynchus Mykiss) in a Time Course Experiment. Comp. Biochem. Physiol. A. Mol. Integr. Physiol. 2019, 236, 110525. [Google Scholar] [CrossRef]
- Watanabe, M.; Hatakeyama, S. TRIM Proteins and Diseases. J. Biochem. (Tokyo) 2017, 161, 135–144. [Google Scholar] [CrossRef] [PubMed]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and Specific Genetic Interference by Double-Stranded RNA in Caenorhabditis Elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Kalil, S.P.; Rosa, R.D. da; Capelli-Peixoto, J.; Pohl, P.C.; Oliveira, P.L. de; Fogaça, A.C.; Daffre, S. Immune-Related Redox Metabolism of Embryonic Cells of the Tick Rhipicephalus Microplus (BME26) in Response to Infection with Anaplasma Marginale. Parasit. Vectors 2017, 10, 613. [Google Scholar] [CrossRef] [PubMed]
- Bohle, H.; Lorenzen, N.; Schyth, B.D. Species Specific Inhibition of Viral Replication Using Dicer Substrate SiRNAs (DsiRNAs) Targeting the Viral Nucleoprotein of the Fish Pathogenic Rhabdovirus Viral Hemorrhagic Septicemia Virus (VHSV). Antiviral Res. 2011, 90, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Schyth, B.D.; Lorenzen, N.; Pedersen, F.S. Antiviral Activity of Small Interfering RNAs: Specificity Testing Using Heterologous Virus Reveals Interferon-Related Effects Overlooked by Conventional Mismatch Controls. Virology 2006, 349, 134–141. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods San Diego Calif 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Drozdetskiy, A.; Cole, C.; Procter, J.; Barton, G.J. JPred4: A Protein Secondary Structure Prediction Server. Nucleic Acids Res. 2015, 43, W389–W394. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.A.; Clamp, M.; Barton, G.J. Jalview Version 2--a Multiple Sequence Alignment Editor and Analysis Workbench. Bioinforma. Oxf. Engl. 2009, 25, 1189–1191. [Google Scholar] [CrossRef] [PubMed]

| Identified TRIMs in RTgill-W1 | GenBank accession |
|---|---|
| OmTRIM25 | KY073243 |
| OmTRIM16 | KY073245 |
| OmTRIM62 | KY073247 |
| OmTRIM8 | KY073248 |
| Name | GenBank | Primers | PCR product size |
|---|---|---|---|
| finTRIM1 | AF483536 | F: ATGGCTCAACAGGGAGTTCT R: TCACTGCCTCTGTTTCTCAGTC |
606 bp |
| finTRIM2 | AM887838 | F: ATGGCTCAACAGGGAGTTCT R: TCAATGACTCTTTCTGTTCCCTT |
1227 bp |
| finTRIM3 | AM887799 | F: ATGGCTCAGCAGGGAGTTT R: CTACAGTTTAACCAGCTCAGCAGTAC |
1656 bp |
| Primer | Forward (RING) | Reverse (B30.2) |
|---|---|---|
| Set1 | TGTGGACACASTTACTGYA | AGTYCAGACCACATTCACTSA |
| Set2 | GGMTGCTGGGAYCAGGA | CAGACCACATTCACT |
| siRNA | Sense sequence | Antisense sequence |
|---|---|---|
| siGFP | rArCrArArCrArGrCrCrArCrArArCrGrUrGrUrArCrArUrCAT |
rArUrGrArUrGrUrArCrArCrGrUrUrGrUrGrGrCrUrGrUrUrGrUrArG |
| siTRIM25-1 | rGrCrArGrCrArGrArGrArGrGrArCrUrGrArGrArArArCrAGA | rUrCrUrGrUrUrUrCrUrCrArGrUrCrCrUrCrUrCrUrGrCrUrGrCrArG |
| siTRIM25-2 | rGrGrArGrGrArCrArGrUrGrArUrCrArGrArUrCrUrUrUrACT | rArGrUrArArArGrArUrCrUrGrArUrCrArCrUrGrUrCrCrUrCrCrArC |
| Gene | Forward | Reverse |
|---|---|---|
| finTRIM 1 | CTACTGAAGGAGCCGGTGG | CAGTGCAGACATCACACGC |
| finTRIM 2 | CTGGACCTGGAAATGTGACA | TGCAGGAAATTCATAGTGAGGTTT |
| OmTRIM25 | TCACCAACTGGTACCAGTTACA | AGAGCACTGGAAACTCCAGGACTT |
| OmTRIM16 | AAAGGTGACCTGTACACACCTCT | TCTCTGTTCTGCTGATGTCTTTA |
| OmTRIM62 | GATTTCCCGACCTCCAAGTACA | GCAGGTTACCATAGGCTACGAT |
| OmTRIM8 | GGAAGTGGAAGTGGCTCTCTAA | TCCATGGTACACACCAGGATCT |
| EF1α | TGGAGACTGGCACCCTGAAG | CCAACATTGTCACCAGGCATGG |
| GAPDH | CCTGCAGAAGGGAATCAAAGTCGT | TCTCATGGGGCTTCATACACTGGA |
| TNF-α2 | GTGTGGCGTTCTCTTAATAGCAGC | ATTCCGTCCTGCATCGTTGC |
| IL-1β | GTCACATTGCCAACCTCATCATCG | GTTGAGCAGGTCCTTGTCCTTGA |
| Type I IFN | GATGCTGAGTTTGAGGACAAAGTC | GTTTCATGGCAGGTGATACACAGGA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




