Submitted:
07 May 2024
Posted:
07 May 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
- The proposal of a novel weakly-supervised and self-guided fine-tuning method of SAM for colonoscopy polyp segmentation. This method reduces the dependence on precise annotations by fully utilizing SAM’s zero-shot capability to use weak annotations for guiding the generation of segmentation masks. These masks are then treated as pseudo-labels, which are then utilized for self-guided fine-tuning, avoiding the need to introduce additional segmentation models.
- The introduction of a series of pseudo-label enhancement strategies to generate high-quality pseudo-labels. These enhancement strategies, including multi-augmentation fusion, pixel-level weighting, and mask post-processing techniques, enable the acquisition of more accurate pseudo-labels.
- Experimental results on five medical image datasets demonstrate that WSPoly-SAM outperforms current fully-supervised mainstream polyp segmentation networks on the Kvasir-SEG, ColonDB, CVC-300, and ETIS datasets. Specifically, on the ColonDB dataset, our method demonstrated an improvement of 9.4% in mDice score and 9.6% in mIoU score compared to state-of-the-art networks, representing a significant breakthrough in the field of colonoscopy polyp segmentation. Furthermore, by using different amounts of training data in weakly-supervised and fully-supervised experiments, it is found that weakly-supervised fine-tuning can save 70% to 73% of annotation time costs compared to fully-supervised fine-tuning.
2. Related works
2.1. Polyp Segmentation
2.2. Segment Anything Model(SAM) Related
3. Methods
3.1. Background
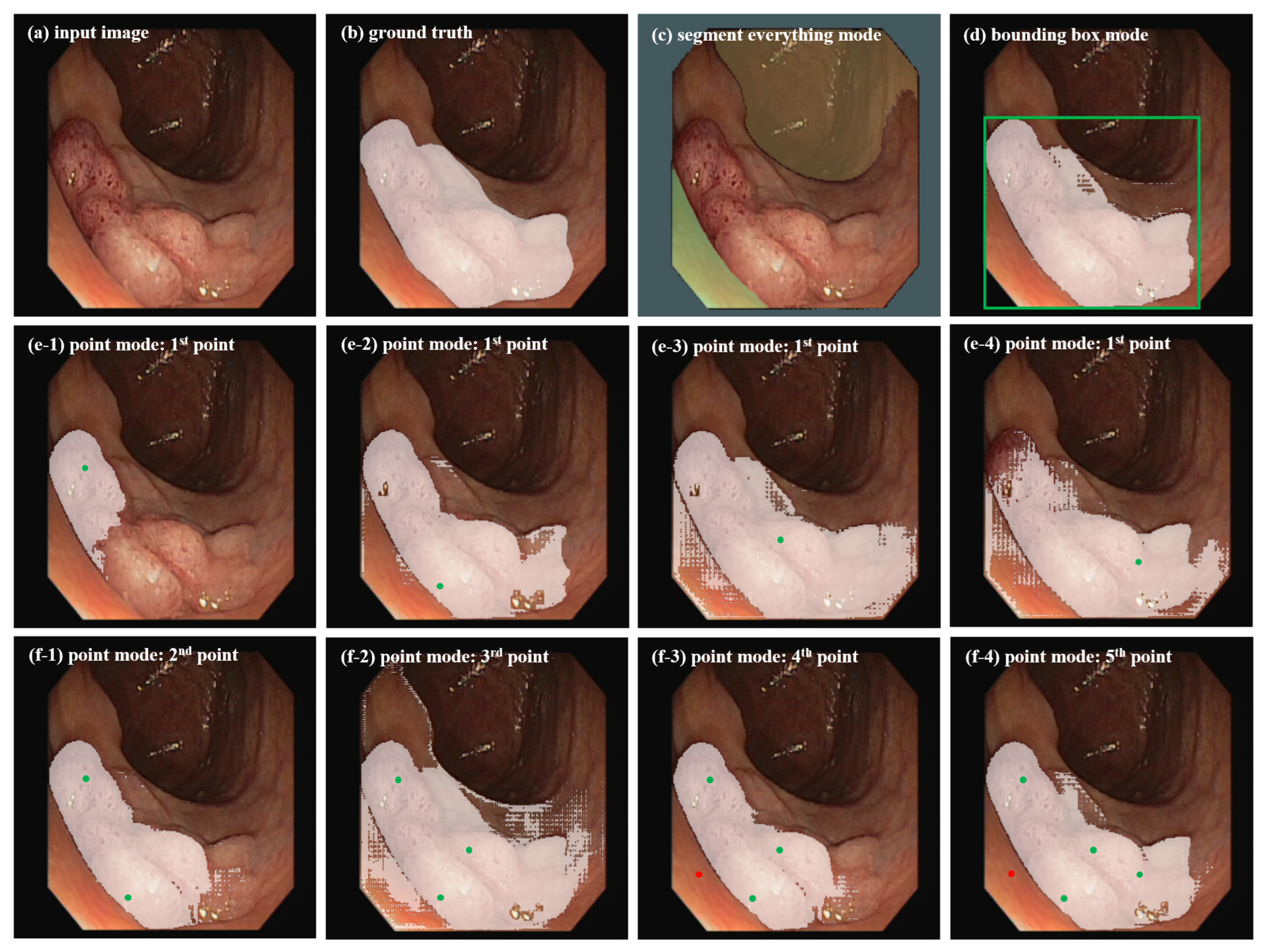
3.2. Prompt Selection
3.3. Self-Guided Pseudo-Label Generation
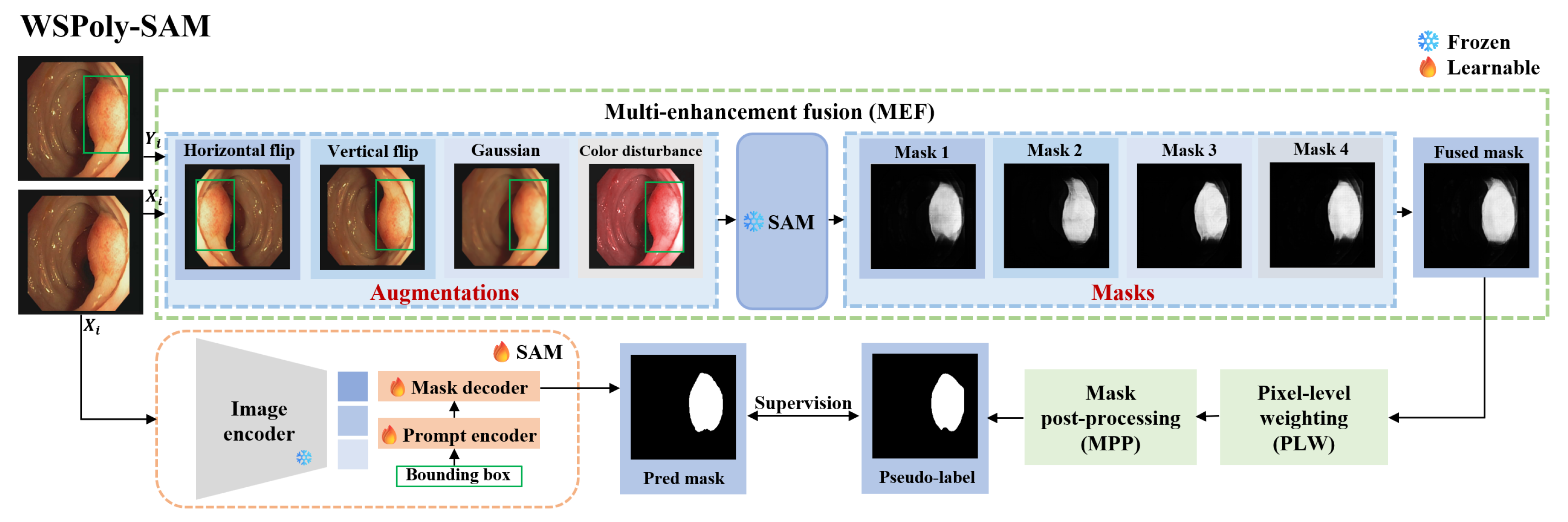
3.4. Weakly-Supervised Fine-Tuning
4. Experiments
4.1. Datasets
- Kvasir-SEG [47]: This dataset gathered by the Vestre Viken Health Trust in Norway, comprises 1000 colonoscopy video sequences showcasing polyp images, each accompanied by its respective annotations. The image resolution varies from 332×487 to 1920×1072 pixels. Annotations are meticulously labeled by medical professionals and validated by seasoned gastroenterologists.
- ClinicDB [48]: This dataset comprises 612 images drawn from 29 colonoscopy video sequences, with a resolution of 288×384 pixels. It is created in partnership with the Hospital Clinic of Barcelona in Spain.
- ColonDB [49]: This dataset includes 380 polyp images along with their corresponding annotations, with each image having a resolution of 500×570 pixels. These images are extracted from 15 distinct videos, with frames meticulously chosen by experts to showcase diverse perspectives. The annotations are manually crafted with precision.
- CVC-300 [50]: The dataset contains 60 polyp images, each with a resolution of 500×574 pixels.
- ETIS [51]: This dataset is released by the MIC-CAI Polyp Detection Subchallenge in 2017, encompassing 196 polyp images extracted from colonoscopy videos, each paired with its respective label. The images boast a resolution of 966×1225 pixels.
4.2. Implementation Details
4.3. Results
4.3.1. Quantitative Results
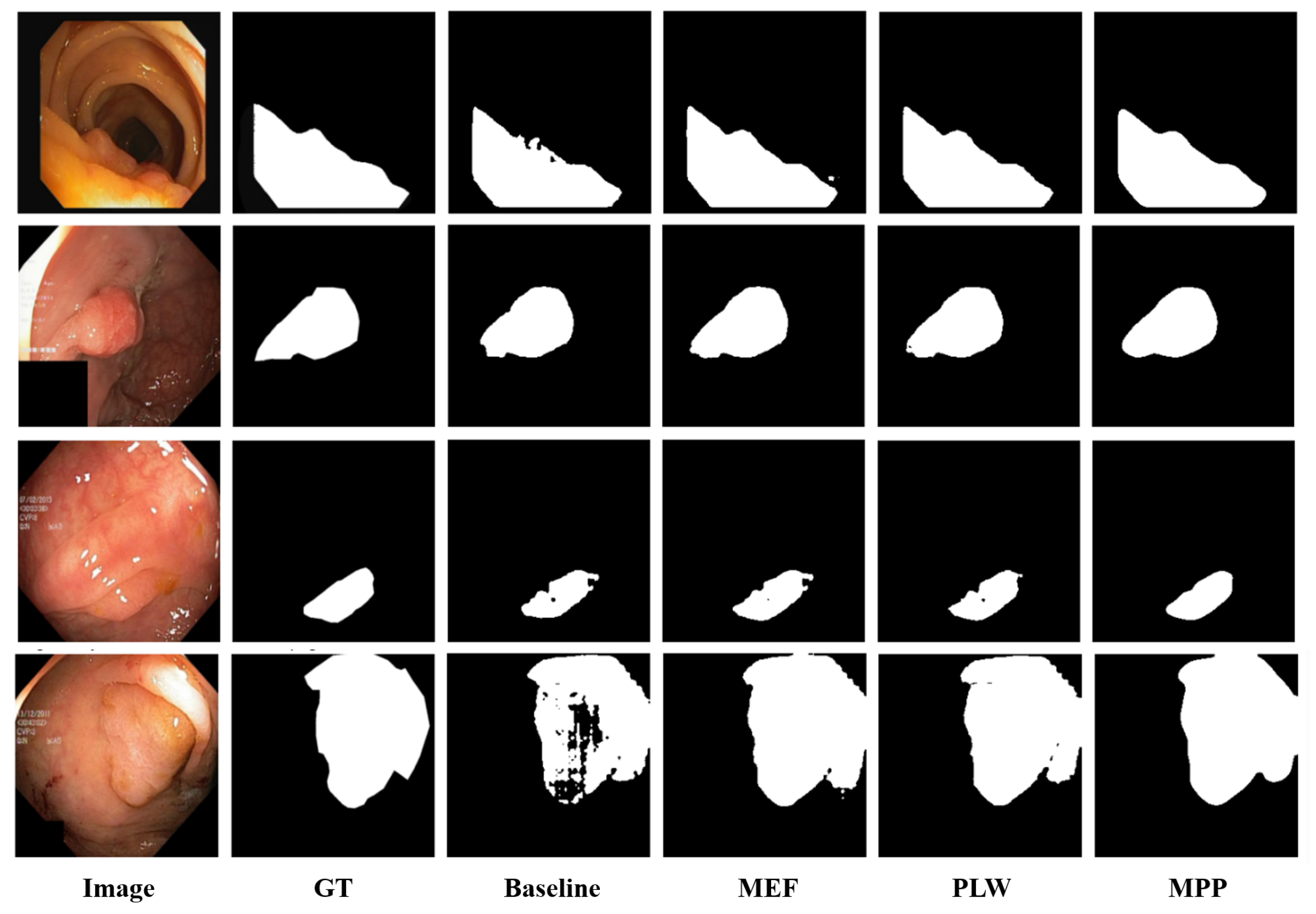
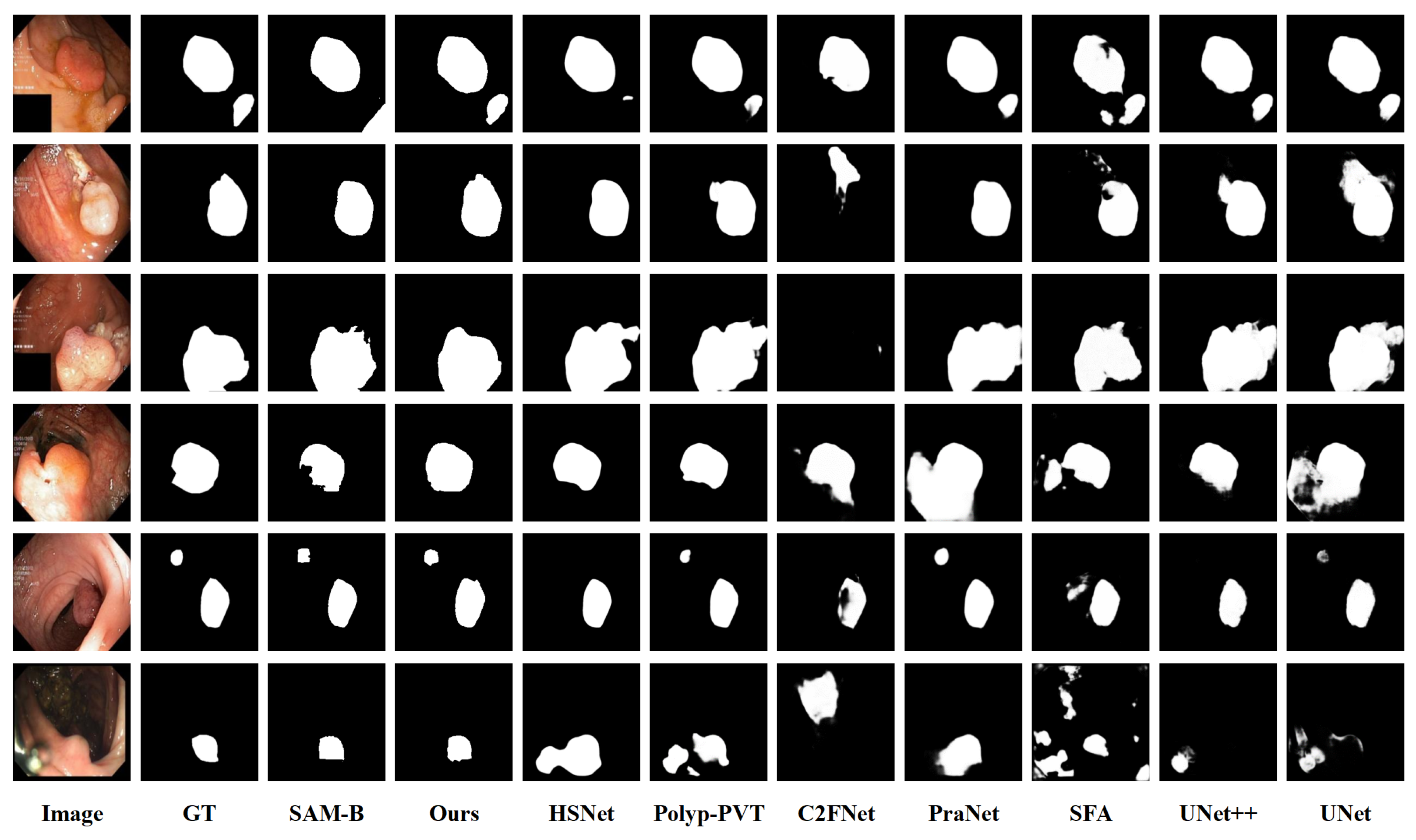
4.3.2. Qualitative Results
4.3.3. Ablation Study
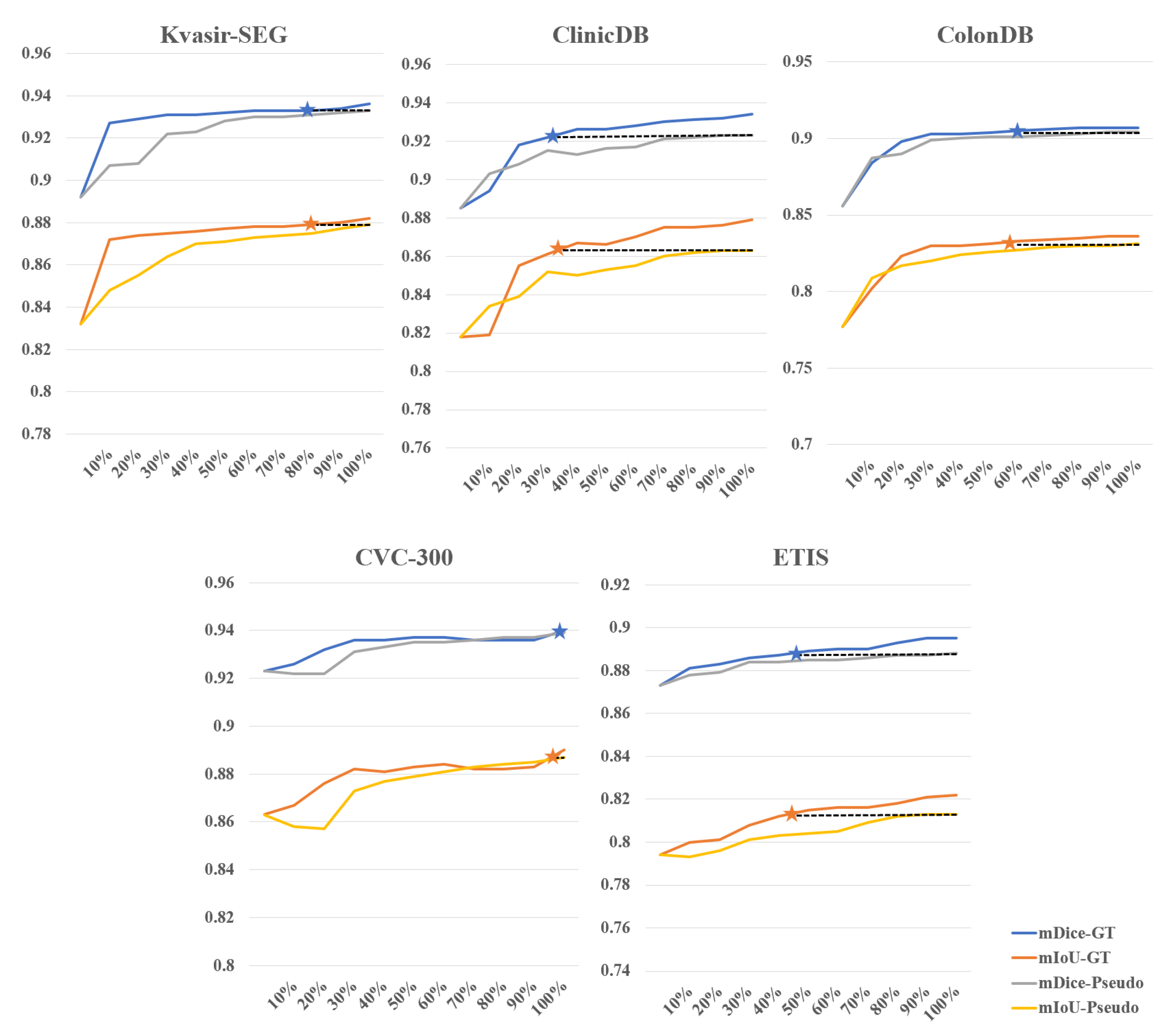
4.3.4. Further Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Isensee, F.; Jaeger, P.F.; Kohl, S.A.; Petersen, J.; Maier-Hein, K.H. nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation. Nature Methods 2021, 18, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, D.; He, B.; Ghorbani, A.; Yuan, N.; Ebinger, J.; Langlotz, C.P.; Heidenreich, P.A.; Harrington, R.A.; Liang, D.H.; Ashley, E.A.; others. Video-based AI for beat-to-beat assessment of cardiac function. Nature 2020, 580, 252–256. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Shi, J.; Qi, X.; Wang, X.; Jia, J. Pyramid scene parsing network. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2017, pp. 2881–2890.
- Chen, L.C.; Papandreou, G.; Kokkinos, I.; Murphy, K.; Yuille, A.L. Deeplab: Semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected crfs. IEEE Transactions on Pattern Analysis and Machine Intelligence 2017, 40, 834–848. [Google Scholar] [CrossRef] [PubMed]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; Van Der Laak, J.A.; Van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Medical Image Analysis 2017, 42, 60–88. [Google Scholar] [CrossRef] [PubMed]
- Asgari Taghanaki, S.; Abhishek, K.; Cohen, J.P.; Cohen-Adad, J.; Hamarneh, G. Deep semantic segmentation of natural and medical images: A review. Artificial Intelligence Review 2021, 54, 137–178. [Google Scholar] [CrossRef]
- Zhang, R.; Lai, P.; Wan, X.; Fan, D.J.; Gao, F.; Wu, X.J.; Li, G. Lesion-aware dynamic kernel for polyp segmentation. International Conference on Medical Image Computing and Computer-Assisted Intervention. Springer, 2022, pp. 99–109.
- Zhou, T.; Zhou, Y.; Gong, C.; Yang, J.; Zhang, Y. Feature aggregation and propagation network for camouflaged object detection. IEEE Transactions on Image Processing 2022, 31, 7036–7047. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; He, Y.; Li, F.; Han, L.; You, C.; Wang, B. Segment anything in medical images. Nature Communications 2024, 15, 654. [Google Scholar] [CrossRef] [PubMed]
- Ding, N.; Qin, Y.; Yang, G.; Wei, F.; Yang, Z.; Su, Y.; Hu, S.; Chen, Y.; Chan, C.M.; Chen, W.; others. Parameter-efficient fine-tuning of large-scale pre-trained language models. Nature Machine Intelligence 2023, 5, 220–235. [Google Scholar] [CrossRef]
- Kirillov, A.; Mintun, E.; Ravi, N.; Mao, H.; Rolland, C.; Gustafson, L.; Xiao, T.; Whitehead, S.; Berg, A.C.; Lo, W.Y. ; others. Segment anything. Proceedings of the IEEE/CVF International Conference on Computer Vision, 2023, pp. 4015–4026.
- Hu, M.; Li, Y.; Yang, X. Skinsam: Empowering skin cancer segmentation with segment anything model. arXiv 2023, arXiv:2304.13973. [Google Scholar]
- Wu, J.; Fu, R.; Fang, H.; Liu, Y.; Wang, Z.; Xu, Y.; Jin, Y.; Arbel, T. Medical sam adapter: Adapting segment anything model for medical image segmentation. arXiv 2023, arXiv:2304.12620. [Google Scholar]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2015, pp. 3431–3440.
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference, Munich, Germany, 5-9, October 2015, Proceedings, Part III 18. Springer, 2015, pp. 234–241.
- Zhou, Z.; Siddiquee, M.M.R.; Tajbakhsh, N.; Liang, J. Unet++: Redesigning skip connections to exploit multiscale features in image segmentation. IEEE Transactions on Medical Imaging 2019, 39, 1856–1867. [Google Scholar] [CrossRef]
- Yang, X.; Li, X.; Ye, Y.; Lau, R.Y.; Zhang, X.; Huang, X. Road detection and centerline extraction via deep recurrent convolutional neural network U-Net. IEEE Transactions on Geoscience and Remote Sensing 2019, 57, 7209–7220. [Google Scholar] [CrossRef]
- Qin, X.; Zhang, Z.; Huang, C.; Dehghan, M.; Zaiane, O.R.; Jagersand, M. U2-Net: Going deeper with nested U-structure for salient object detection. Pattern Recognition 2020, 106, 107404. [Google Scholar] [CrossRef]
- Sun, X.; Zhang, P.; Wang, D.; Cao, Y.; Liu, B. Colorectal polyp segmentation by U-Net with dilation convolution. 2019 18th IEEE International Conference on Machine Learning and Applications (ICMLA). IEEE, 2019, pp. 851–858.
- Chen, L.C.; Zhu, Y.; Papandreou, G.; Schroff, F.; Adam, H. Encoder-decoder with atrous separable convolution for semantic image segmentation. Proceedings of the European Conference on Computer Vision (ECCV), 2018, pp. 801–818.
- Liu, S.; Huang, D. ; others. Receptive field block net for accurate and fast object detection. Proceedings of the European Conference on Computer Vision (ECCV), 2018, pp. 385–400.
- Fang, Y.; Chen, C.; Yuan, Y.; Tong, K.y. Selective feature aggregation network with area-boundary constraints for polyp segmentation. Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, 13–17, October 2019, Proceedings, Part I 22. Springer, 2019, pp. 302–310.
- Zhang, R.; Li, G.; Li, Z.; Cui, S.; Qian, D.; Yu, Y. Adaptive context selection for polyp segmentation. Medical Image Computing and Computer Assisted Intervention–MICCAI 2020: 23rd International Conference, Lima, Peru, 4–8, October 2020, Proceedings, Part VI 23. Springer, 2020, pp. 253–262.
- Nguyen, T.C.; Nguyen, T.P.; Diep, G.H.; Tran-Dinh, A.H.; Nguyen, T.V.; Tran, M.T. CCBANet: Cascading context and balancing attention for polyp segmentation. Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, 27 September–1 October 2021, Proceedings, Part I 24. Springer, 2021, pp. 633–643.
- Deng, R.; Cui, C.; Liu, Q.; Yao, T.; Remedios, L.W.; Bao, S.; Landman, B.A.; Wheless, L.E.; Coburn, L.A.; Wilson, K.T. ; others. Segment anything model (sam) for digital pathology: Assess zero-shot segmentation on whole slide imaging. arXiv 2023, arXiv:2304.04155. [Google Scholar]
- Hu, C.; Li, X. When sam meets medical images: An investigation of segment anything model (sam) on multi-phase liver tumor segmentation. arXiv 2023, arXiv:2304.08506. [Google Scholar]
- He, S.; Bao, R.; Li, J.; Grant, P.E.; Ou, Y. Accuracy of segment-anything model (sam) in medical image segmentation tasks. arXiv 2023, arXiv:2304.09324. [Google Scholar]
- Roy, S.; Wald, T.; Koehler, G.; Rokuss, M.R.; Disch, N.; Holzschuh, J.; Zimmerer, D.; Maier-Hein, K.H. Sam. md: Zero-shot medical image segmentation capabilities of the segment anything model. arXiv 2023, arXiv:2304.05396. [Google Scholar]
- Zhou, T.; Zhang, Y.; Zhou, Y.; Wu, Y.; Gong, C. Can sam segment polyps? arXiv 2023, arXiv:2304.07583. [Google Scholar]
- Mohapatra, S.; Gosai, A.; Schlaug, G. Brain extraction comparing segment anything model (sam) and fsl brain extraction tool. arXiv 2023, arXiv:2304.04738. [Google Scholar]
- Mazurowski, M.A.; Dong, H.; Gu, H.; Yang, J.; Konz, N.; Zhang, Y. Segment anything model for medical image analysis: An experimental study. Medical Image Analysis 2023, 89, 102918. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Bai, X. Learning to “segment anything” in thermal infrared images through knowledge distillation with a large scale dataset satir. arXiv 2023, arXiv:2304.07969. [Google Scholar]
- Tang, L.; Xiao, H.; Li, B. Can sam segment anything? when sam meets camouflaged object detection. arXiv 2023, arXiv:2304.04709. [Google Scholar]
- Ji, G.P.; Fan, D.P.; Xu, P.; Cheng, M.M.; Zhou, B.; Van Gool, L. SAM Struggles in Concealed Scenes–Empirical Study on” Segment Anything“. arXiv 2023, arXiv:2304.06022. [Google Scholar]
- Ji, W.; Li, J.; Bi, Q.; Li, W.; Cheng, L. Segment anything is not always perfect: An investigation of sam on different real-world applications. arXiv 2023, arXiv:2304.05750. [Google Scholar]
- Ma, J.; He, Y.; Li, F.; Han, L.; You, C.; Wang, B. Segment anything in medical images. Nature Communications 2024, 15, 654. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Ye, J.; Deng, Z.; Chen, J.; Li, T.; Wang, H.; Su, Y.; Huang, Z.; Chen, J.; Jiang, L. ; others. Sam-med2d. arXiv 2023, arXiv:2308.16184. [Google Scholar]
- Jiang, P.T.; Yang, Y. Segment anything is a good pseudo-label generator for weakly supervised semantic segmentation. arXiv 2023, arXiv:2305.01275. [Google Scholar]
- He, C.; Li, K.; Zhang, Y.; Xu, G.; Tang, L.; Zhang, Y.; Guo, Z.; Li, X. Weakly-supervised concealed object segmentation with sam-based pseudo labeling and multi-scale feature grouping. Advances in Neural Information Processing Systems 2024, 36. [Google Scholar]
- Reedha, R.; Dericquebourg, E.; Canals, R.; Hafiane, A. Transformer neural network for weed and crop classification of high resolution UAV images. Remote Sensing 2022, 14, 592. [Google Scholar] [CrossRef]
- Brown, T.; Mann, B.; Ryder, N.; Subbiah, M.; Kaplan, J.D.; Dhariwal, P.; Neelakantan, A.; Shyam, P.; Sastry, G.; Askell, A.; others. Language models are few-shot learners. Advances in Neural Information Processing Systems 2020, 33, 1877–1901. [Google Scholar]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S. ; others. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv 2020, arXiv:2010.11929. [Google Scholar]
- He, K.; Chen, X.; Xie, S.; Li, Y.; Dollár, P.; Girshick, R. Masked autoencoders are scalable vision learners. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2022, pp. 16000–16009.
- Zhang, W.; Fu, C.; Zheng, Y.; Zhang, F.; Zhao, Y.; Sham, C.W. HSNet: A hybrid semantic network for polyp segmentation. Computers in Biology and Medicine 2022, 150, 106173. [Google Scholar] [CrossRef] [PubMed]
- Isensee, F.; Jaeger, P.F.; Kohl, S.A.; Petersen, J.; Maier-Hein, K.H. nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation. Nature Methods 2021, 18, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Chen, J.; Ng, M.; Huang, R.; Li, Y.; Li, C.; Yang, X.; Martel, A.L. Loss odyssey in medical image segmentation. Medical Image Analysis 2021, 71, 102035. [Google Scholar] [CrossRef]
- Jha, D.; Smedsrud, P.H.; Riegler, M.A.; Halvorsen, P.; De Lange, T.; Johansen, D.; Johansen, H.D. Kvasir-seg: A segmented polyp dataset. MultiMedia Modeling: 26th International Conference, MMM 2020, Daejeon, South Korea, 5–8, January 2020, Proceedings, Part II 26. Springer, 2020, pp. 451–462.
- Bernal, J.; Sánchez, F.J.; Fernández-Esparrach, G.; Gil, D.; Rodríguez, C.; Vilariño, F. WM-DOVA maps for accurate polyp highlighting in colonoscopy: Validation vs. saliency maps from physicians. Computerized Medical Imaging and Graphics 2015, 43, 99–111. [Google Scholar] [CrossRef]
- Tajbakhsh, N.; Gurudu, S.R.; Liang, J. Automated polyp detection in colonoscopy videos using shape and context information. IEEE Transactions on Medical Imaging 2015, 35, 630–644. [Google Scholar] [CrossRef] [PubMed]
- Vázquez, D.; Bernal, J.; Sánchez, F.J.; Fernández-Esparrach, G.; López, A.M.; Romero, A.; Drozdzal, M.; Courville, A. A benchmark for endoluminal scene segmentation of colonoscopy images. Journal of Healthcare Engineering 2017, 2017, 4037190. [Google Scholar] [CrossRef] [PubMed]
- Silva, J.; Histace, A.; Romain, O.; Dray, X.; Granado, B. Toward embedded detection of polyps in wce images for early diagnosis of colorectal cancer. International Journal of Computer Assisted Radiology and Surgery 2014, 9, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Dong, B.; Wang, W.; Fan, D.P.; Li, J.; Fu, H.; Shao, L. Polyp-pvt: Polyp segmentation with pyramid vision transformers. arXiv 2021, arXiv:2108.06932. [Google Scholar]
- Zhang, W.; Fu, C.; Zheng, Y.; Zhang, F.; Zhao, Y.; Sham, C.W. HSNet: A hybrid semantic network for polyp segmentation. Computers in Biology and Medicine 2022, 150, 106173. [Google Scholar] [CrossRef]
- Fan, D.P.; Ji, G.P.; Zhou, T.; Chen, G.; Fu, H.; Shen, J.; Shao, L. Pranet: Parallel reverse attention network for polyp segmentation. International Conference on Medical Image Computing and Computer-Assisted Intervention. Springer, 2020, pp. 263–273.
- Sun, Y.; Chen, G.; Zhou, T.; Zhang, Y.; Liu, N. Context-aware cross-level fusion network for camouflaged object detection. arXiv 2021, arXiv:2105.12555. [Google Scholar]

| Models | Kvasir-SEG | ClinicDB | ColonDB | CVC-300 | ETIS | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| mDice | mIoU | mDice | mIoU | mDice | mIoU | mDice | mIoU | mDice | mIoU | |
| U-Net | 0.818 | 0.746 | 0.823 | 0.755 | 0.504 | 0.436 | 0.710 | 0.627 | 0.398 | 0.335 |
| U-Net++ | 0.821 | 0.744 | 0.794 | 0.729 | 0.482 | 0.408 | 0.707 | 0.624 | 0.401 | 0.344 |
| SFA | 0.723 | 0.611 | 0.700 | 0.607 | 0.456 | 0.337 | 0.467 | 0.329 | 0.297 | 0.271 |
| PraNet | 0.898 | 0.840 | 0.899 | 0.849 | 0.712 | 0.640 | 0.871 | 0.797 | 0.628 | 0.567 |
| C2FNet | 0.886 | 0.831 | 0.919 | 0.872 | 0.724 | 0.650 | 0.874 | 0.801 | 0.699 | 0.624 |
| Polyp-PVT | 0.917 | 0.864 | 0.948 | 0.905 | 0.808 | 0.727 | 0.900 | 0.833 | 0.787 | 0.706 |
| HSNet | 0.926 | 0.877 | 0.937 | 0.887 | 0.810 | 0.735 | 0.903 | 0.839 | 0.808 | 0.734 |
| SAM-B | 0.892 | 0.832 | 0.884 | 0.812 | 0.856 | 0.777 | 0.922 | 0.862 | 0.873 | 0.794 |
| WSPoly-SAM-B | 0.933 | 0.879 | 0.920 | 0.858 | 0.904 | 0.831 | 0.933 | 0.878 | 0.888 | 0.813 |
| 0.041 | 0.047 | 0.036 | 0.046 | 0.048 | 0.054 | 0.011 | 0.016 | 0.015 | 0.019 | |
| Baseline | MEF | PLW | MPP | Kvasir-SEG | ClinicDB | ColonDB | CVC-300 | ETIS | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| mDice | mIoU | mDice | mIoU | mDice | mIoU | mDice | mIoU | mDice | mIoU | ||||
| - | - | - | - | 0.892 | 0.832 | 0.884 | 0.812 | 0.856 | 0.777 | 0.922 | 0.862 | 0.873 | 0.794 |
| √ | 0.928 | 0.873 | 0.914 | 0.850 | 0.895 | 0.820 | 0.931 | 0.875 | 0.876 | 0.796 | |||
| √ | √ | 0.928 | 0.875 | 0.914 | 0.851 | 0.897 | 0.822 | 0.931 | 0.875 | 0.880 | 0.797 | ||
| √ | √ | √ | 0.932 | 0.878 | 0.916 | 0.851 | 0.903 | 0.830 | 0.933 | 0.878 | 0.885 | 0.805 | |
| √ | √ | √ | √ | 0.933 | 0.879 | 0.920 | 0.858 | 0.904 | 0.831 | 0.933 | 0.878 | 0.888 | 0.813 |
| Model-ViT | Mask | Kvasir-SEG | ClinicDB | ColonDB | CVC-300 | ETIS | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| mDice | mIoU | mDice | mIoU | mDice | mIoU | mDice | mIoU | mDice | mIoU | ||
| SAM-B | - | 0.892 | 0.832 | 0.884 | 0.812 | 0.856 | 0.777 | 0.922 | 0.862 | 0.873 | 0.794 |
| SAM-L | - | 0.921 | 0.865 | 0.884 | 0.817 | 0.897 | 0.823 | 0.926 | 0.868 | 0.855 | 0.782 |
| SAM-H | - | 0.923 | 0.868 | 0.890 | 0.826 | 0.895 | 0.822 | 0.914 | 0.852 | 0.866 | 0.732 |
| SAM-B | Pseudo | 0.933 | 0.879 | 0.920 | 0.858 | 0.904 | 0.831 | 0.933 | 0.878 | 0.888 | 0.813 |
| SAM-L | Pseudo | 0.932 | 0.884 | 0.925 | 0.867 | 0.906 | 0.837 | 0.934 | 0.879 | 0.881 | 0.812 |
| SAM-B | GT | 0.936 | 0.882 | 0.935 | 0.882 | 0.907 | 0.836 | 0.934 | 0.879 | 0.895 | 0.822 |
| SAM-L | GT | 0.934 | 0.886 | 0.926 | 0.876 | 0.908 | 0.842 | 0.934 | 0.880 | 0.890 | 0.815 |
| Methods | Kvasir-SEG | ClinicDB | ColonDB | CVC-300 | ETIS | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| mDice | mIoU | mDice | mIoU | mDice | mIoU | mDice | mIoU | mDice | mIoU | |
| SAM | 0.892 | 0.832 | 0.884 | 0.812 | 0.856 | 0.777 | 0.922 | 0.862 | 0.873 | 0.794 |
| Pseudo-label Generation | 0.899 | 0.844 | 0.912 | 0.844 | 0.890 | 0.814 | 0.932 | 0.876 | 0.884 | 0.810 |
| WSPoly-SAM | 0.933 | 0.879 | 0.920 | 0.858 | 0.904 | 0.831 | 0.933 | 0.878 | 0.888 | 0.813 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




