Introduction:
The number of patients with an allergic response to their environment is increasing and is now more than 40% in numerous populations in the United States and Europe.[
1] Allergic rhinitis refers to an allergic response that occurs in the nose.[
2] The number of patients suffering from allergic rhinitis, commonly known as hay fever, in the United States is approximately 30%.[
3] Allergic rhinitis contributes to lost or unproductive time at work and school, sleep problems, and reduced participation in outdoor activities. The ability to control asthma and allergic rhinitis in individuals with asthma has been linked to the control of allergic rhinitis.[
4]
According to published studies, most individuals with asthma develop rhinitis.[
5] In addition, the presence of allergic rhinitis (seasonal or perennial) significantly increases the likelihood of asthma; up to 40% of people with allergic rhinitis have or will have asthma.[
6] It is also essential to define how breathing is affected in patients with obstructive sleep apnea (OSA).[
7] Breathing starts at the nose, and while OSA is characterized by the collapse of the muscles of the oropharyngeal airways, nasal obstruction and OSA are usually co-existing conditions that worsen each individual condition.[
8,
9,
10] Therapies to improve airflow through the nose in compromised patients significantly reduce daytime and nighttime symptoms.[
11] The nose is responsible for almost 50% of the resistance when transporting air from the nose to the lungs and plays an essential role in humidification, heating, and air filtration.[
12] The tissues inside the nose, known as the nasal mucosa, are dynamic organs regulated by the autonomic nervous system.[
13] Periodic nasal congestion and decongestion are termed the "nasal cycle."[
14]. In patients with a permanent one-sided nasal obstruction, the nasal cycle can make getting air inside the body difficult.[
15]
Nasal sprays are used by a large percentage of the population, especially with recent news reports demonstrating the inhibitory action of xylitol nasal sprays and other ingredients on the attachment (then endocytosis) of Sars-CoV-2 virus to nasal cells.[
16,
17,
18] Xylitol has also been reported to inhibit the oncogenesis of oral cells, making xylitol nasal sprays even more desirable for this population.[
19] However, no study has analyzed the potential shift in the nasal microbiome after using xylitol nasal spray.
Nasal and lung microbiomes may be affected by the gut microbiome.[
20] Formula-fed infants are at an increased risk of infections.[
21] Owing to the crosstalk between the mucosal systems of the gastrointestinal and respiratory tracts, adding synbiotics (prebiotics and probiotics) to infant formulas may prevent infections even at distant sites.[
22,
23] In a study on infants who were born full-term and weaned from breast milk, they were randomized to either prebiotic formula (fructo- and galactooligosaccharides) or the same prebiotic formula along with
Lactobacillus paracasei ssp. paracasei F19 (synbiotics) from 1 to 6 months of age. [
22] With the synbiotic effects on gut microbiota development,
L. fermentum PCC and
L. reuteri RC-14 were more resistant to gastric conditions, and their survival rate was further improved in the presence of 5 out of 10 tested pectins. Additionally, two pectins positively affected the viability of the less resistant
L. rhamnosus LGG and
L. paracasei F-19.[
24]
Since the advent of high-throughput sequencing, PCR-amplified 16S sequences have typically been clustered based on their similarity to generate operational taxonomic units (OTUs), and representative OTU sequences have been compared with reference databases to infer the likely taxonomy. However, convenient and powerful usage of 16S rRNA has necessitated certain assumptions, such as the now historical assumption that sequences of > 95% identity represent the same genus. In contrast, sequences with > 97% identity represent the same species.[
25,
26]
Objective:
The principal objective of this study was to discover the normal complete nasal microbiome after using an over-the-counter nasal spray, which, as of this date, had yet to be completed. In addition, more information is needed on the microbiome shifts that occur in response to xylitol nasal spray. Before the widespread use of nasal sprays, what may be considered normal should be determined. Shifts in microbial diversity and richness should be regarded as likely.
Materials and Methods:
This study performed a nasal swab test before and after using the most popular OTC nasal spray in 10 participants aged 18–48 years. All participants had non-significant medical histories with no history of allergies or medication use that could affect the nasal microbiome. The participants consisted of five men and five women who were randomly assigned a number using randomizing software. The participants signed an informed consent form (submitted to the IRB), and the principal investigator kept all personal protected information in a hard copy safely locked in a secure cabinet.
An e-mail newsletter recruited participants from all patients of the principal investigator, and recruiting posters were placed in the reception area of the PI office. No inducements or compensation were provided to the participants. The study protocol, informed consent, medical history, and sampling are discussed separately. The samples were preserved and shipped by FedEx to Northwestern University’s NUSeq Center for Whole Genome Deep Sequencing. After 21 days of OTC nasal spray use twice daily, the participants returned for another sampling of their nasal microbiome, which FedEx then shipped to NUSeq for deep sequencing. The participants were instructed to use the spray before dismissal during the first visit. The spray was obtained as samples normally distributed to clinics at no charge and was already in the inventory of the principal investigator. These samples were often administered to pediatric patients diagnosed with sleep apnea.
Sampling consisted of placing a nasal swab atraumatically into the anterior nasal cavity, similar to self-performed sampling for COVID-19 tests. No injury was reported, and only slight discomfort, if any, was reported. The swab was placed approximately one inch into the nares, rotated for 15 seconds, and then placed into the other nares for another rotation of 15 seconds. The swabs were placed in a preservative and sealed safely. Biosafety envelopes were used for shipping according to previous protocols. After the second sampling, patients were excluded from the study. All personal information was destroyed after two years.
Xlear Sinus Care is an OTC xylitol solution. Xylitol is a natural sugar (pentose aldose) that inhibits biofilm formation.[
27] Pure xylitol is a white crystalline substance found in many fruits and vegetables.[
28] According to the manufacturer, the Xlear solution cleanses, hydrates, dries, and irritates tissues. Xlear products use a patented xylitol solution that inhibits bacteria and pulls moisture into the nasal cavity. According to recently published research, xylitol also inhibits the viral invasion of cells.[
17]
Ingredients:
Purified water
Xylitol
Saline
Grapefruit seed extract
Study Data
Microbiome Analysis. The quality of the reads in the FASTQ format was evaluated using FastQC. Reads were trimmed to remove Illumina adapters from the 3’ ends using Cutadapt.[
29] Paired-end trimmed reads were aligned using Kraken2 with the standard database using default parameters, except for 12 threads.[
30] Alignments that could not be classified into known taxa were excluded. Taxon abundance estimates were performed using Bracken, except for 12 threads, using the default parameters.[
31] Normalization and differential expression were calculated using DESeq2.[
32]
Results:
The analysis determined 2,558 taxa with greater abundances of the following top taxa: Xanthobacter autotrophicus, Streptomyces sp. ICC1, Streptomyces armeniacus, Streptococcus thermophilus, Streptococcus sanguinis, Streptococcus pyogenes, Streptococcus oralis, Staphylococcus epidermidis, Staphylococcus aureus, Salmonella enterica, Rothia aeria, Ralstonia pickettii, Pseudomonas aeruginosa, Peptoniphilus harei, Lawsonella clevelandensis, Klebsiella pneumoniae, Finegoldia magna, Escherichia coli, Enterococcus faecium, Dolosigranulum pigrum, Delftia lacustris, Cutibacterium granulosum, Cutibacterium acnes, Corynebacterium tuberculostearicum, Corynebacterium striatum, Corynebacterium segmentosum, Corynebacterium propinquum, Corynebacterium macginleyi, Corynebacterium kefirresidentii, Corynebacterium glutamicum, Corynebacterium diphtheriae, Burkholderia dolosa, Brevibacillus brevis, Bartonella krasnovii, Anaerococcus prevotii, Aeromonas caviae, and [Haemophilus] ducreyi
The routine use of xylitol nasal spray changed the relative abundance of ten taxa (
Table 1). This result is not surprising because recently published research has demonstrated that even a few species that are being changed have a cascading effect throughout the microbiome.[
33] Xylitol should reduce the number of pathogenic bacteria, which would also affect the pathogens' synergistic co-pathogens[
34,
35,
36] Reports have previously been published that the oral microbiome has a “downstream effect” on the gut microbiome.[
37,
38] It would not be surprising if the nasal gateway microbiome did not have the same effect, with a reduction in pathogens of the anterior nasal passageway strain/species shifting the respiratory microbiome.
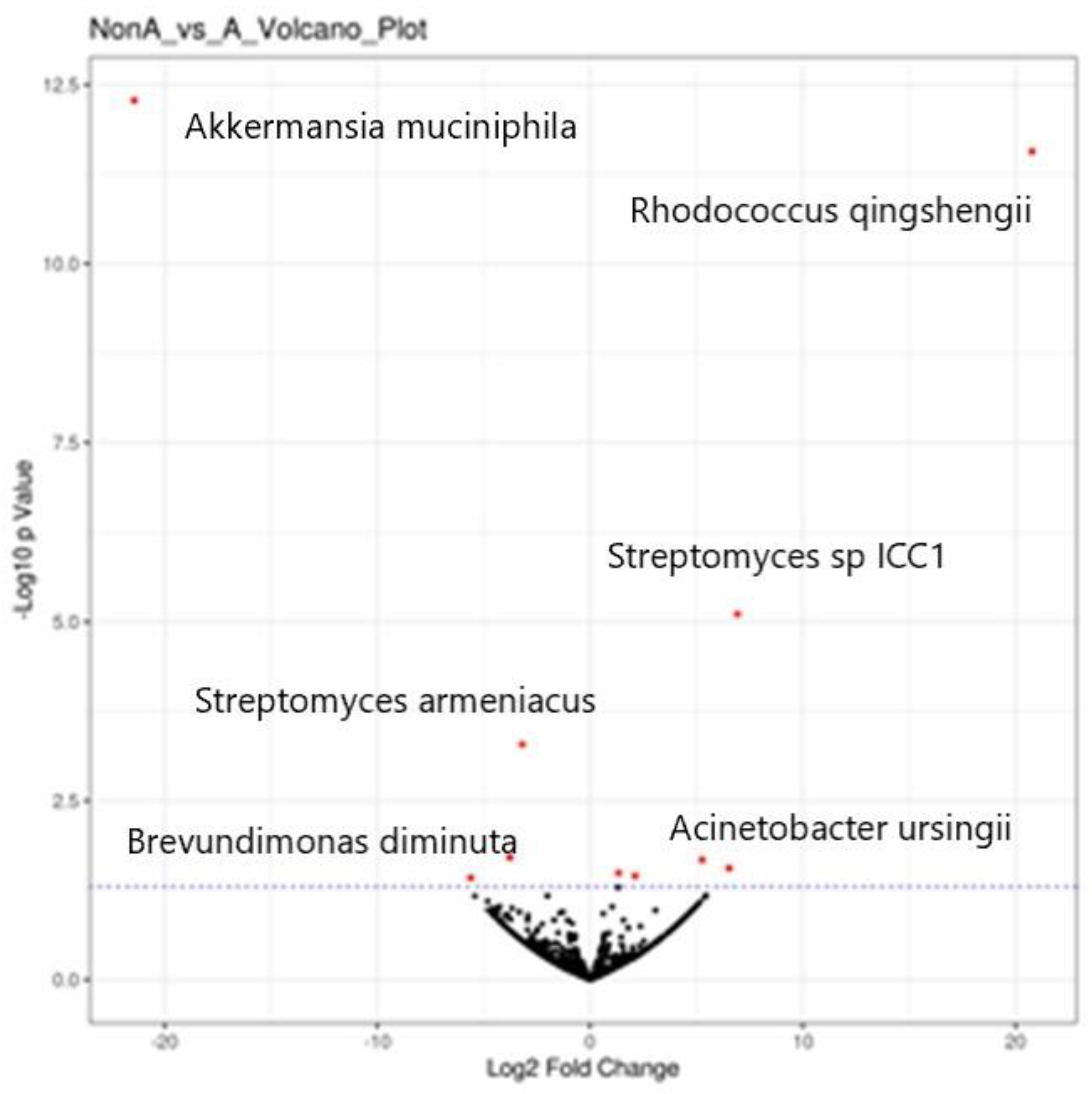
Four species decreased, with six increasing in abundance (
Figure 1. Volcano plot of A versus non-A, after xylitol and before xylitol).
Rhodococcus qingshengii (soil organism with anti-fungal properties) significantly increased, but
Akkermansia muciniphilia substantially decreased. Akkermansia muciniphilia is considered a probiotic in the gut and the oral cavity.
Brevundimonas diminuta was also decreased and is ubiquitous in humans but has been associated with infections in immunocompromised and cancer patients.
Acinetobacter ursingii increased in predominance and has been associated with immunocompromised patient infections (see
Table 1). It is essential to note that almost all microbes may cause an illness when they are in the wrong place at the wrong time. We lack sufficient knowledge of the nasal gateway microbiome to properly judge which bacteria are acceptable commensals and which are pathobionts. For instance,
Streptomyces armeniacus is a spore-forming soil organism, a probiotic producing Streptopyrrole, and the abundance is increased by xylitol therapy. The presence of so many soil-based microorganisms is to be expected as they may become airborne via dust particles. One can only imagine the bacterial load a runner takes in nasally as they sprint on a dirt path. The Streptopyrrole probiotic bacteria
Streptomyces armeniacus may be an important link in nasal health.
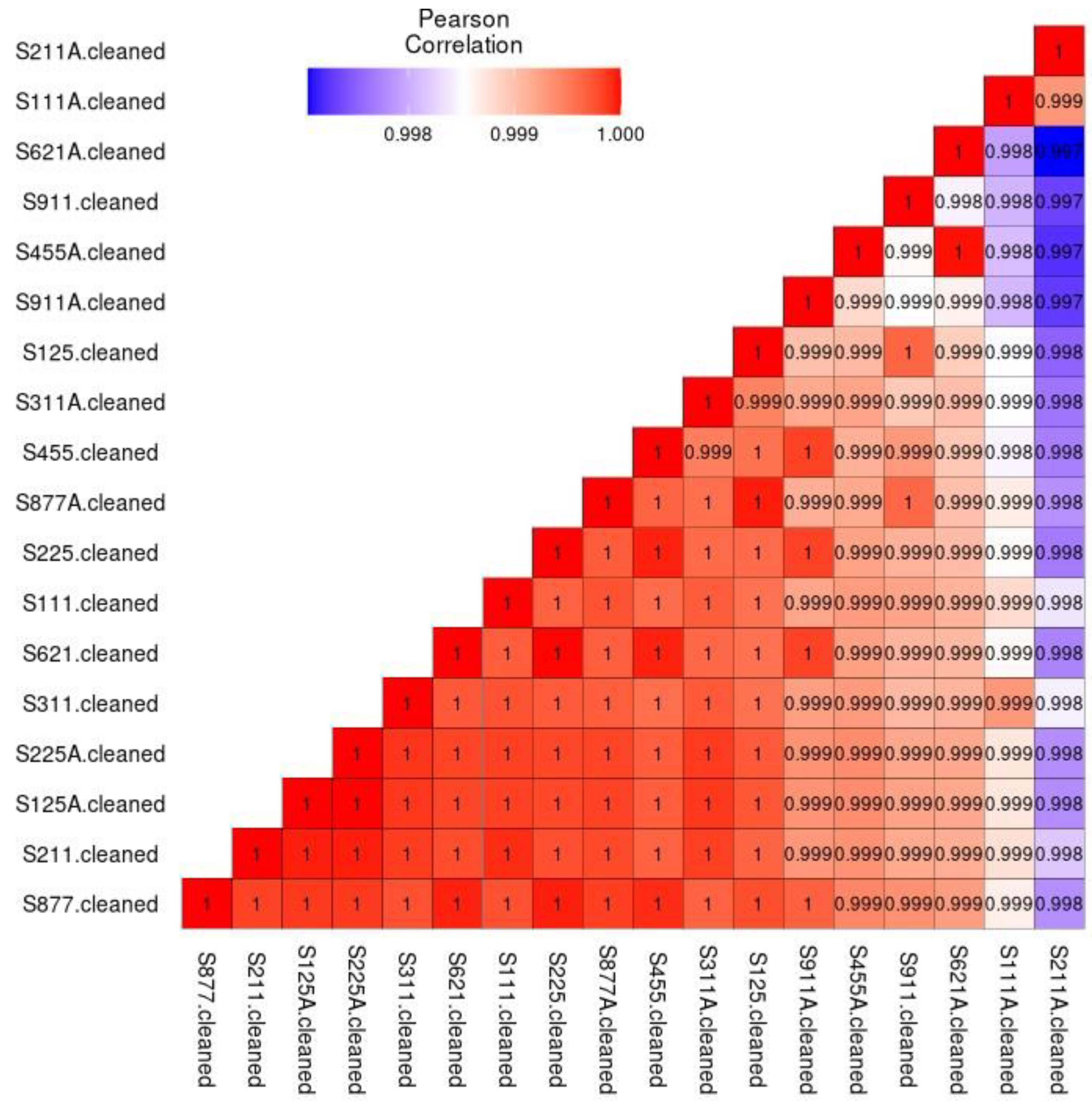
Pearson Correlation demonstrated statistical differences between the A and Non-A groups. A group refers to the subjects using a xylitol spray for 21 days, Non-A was the group before using the spray. A and non-A nomenclature was used to blind the laboratory personnel to the nature of the research (see
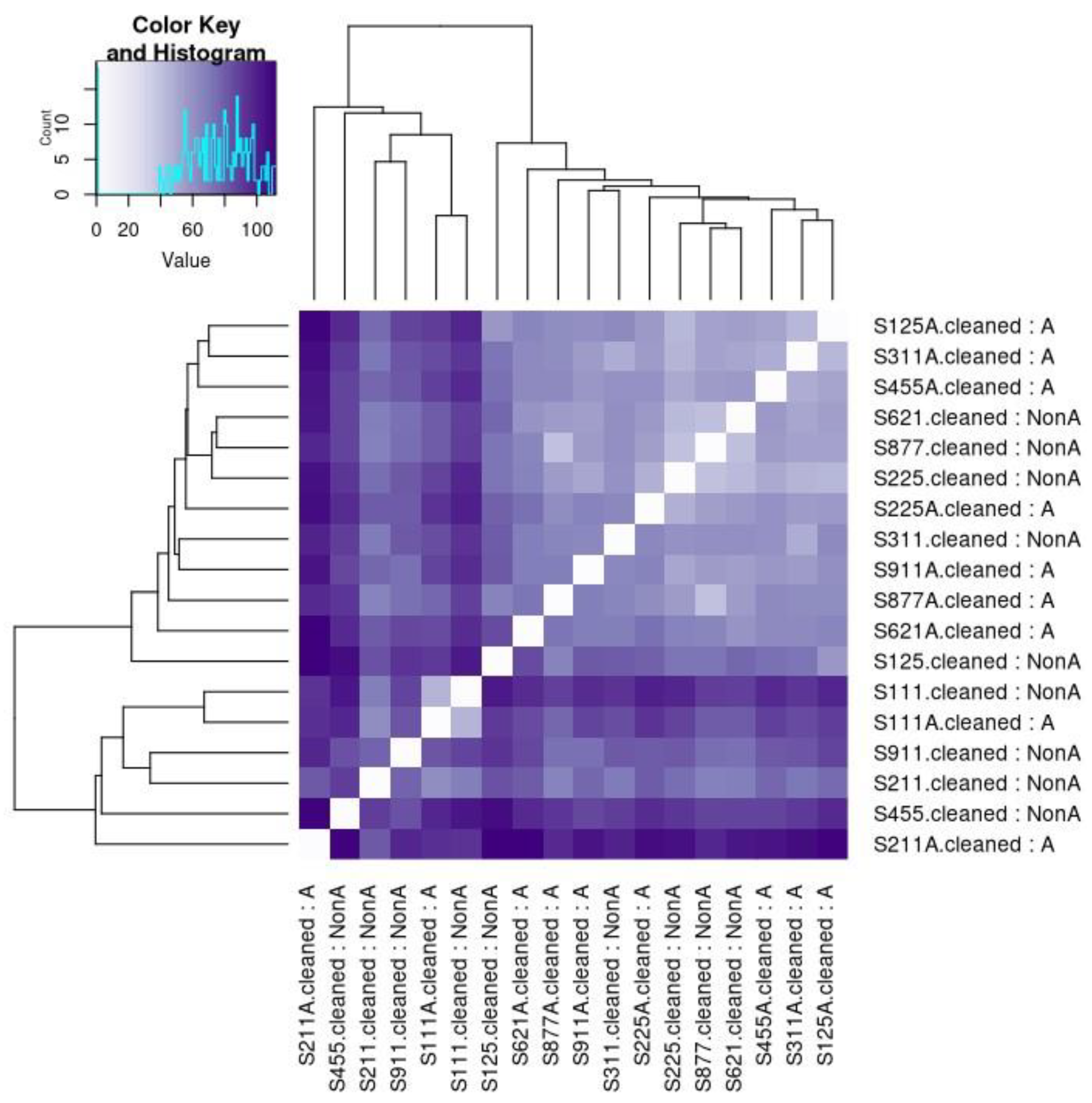
Figure 2). Similar to a correlation matrix, a heatmap demonstrated the differences between the A and non-A groups (see
Figure 3).
Discussion:
The microbial analysis included all bacteria, archaea, viruses, molds, and yeasts obtained via deep sequencing for species analysis. Artificial intelligence analyses by the Northwestern University Sequencing Center revealed shifts in species and strains due to the use of the OTC nasal spray, resulting in changes in diversity or richness. Deep sequencing and artificial intelligence analysis may be used to determine changes in Richness and Shannon diversity. Genus, species, and strain shifts determined the precise response of the nasal microbiome to the most common OTC nasal spray. Moreover, the complete nasal microbiome by deep sequencing was yet to be discovered. In addition, more information was needed on the microbiome shifts that occur in response to xylitol nasal spray. Before the widespread use of COVID-19 precautions, what may be considered the normal nasal microbiome needed to be determined. The nasal microbiome of healthy adults before and after xylitol exposure had yet to be analyzed using whole-genome deep sequencing methods.
The large number of taxa found may not represent the total number of taxa that are likely to be present in the oral cavity. The study ended at the number reported mainly owing to the economic restraints of the study, and the use of artificial intelligence with machine language had demands on technology resources. In addition, there are often constraints owing to the availability of DNA libraries for comparison. Routine xylitol nasal spray use significantly affects the prevalence of at least ten taxa.
Several probiotic bacteria have been identified in the nasal cavity; oral or gut probiotics are often abundant. The nasal microbiome also contains several skin commensals, such as
Staphylococcus epidermidis and
Cutibacterium acnes. This indicates that the nasal microbiome combines the features of both the oral and skin microbiomes. Additionally, bacteria are more specific to the nasal cavity. Previous studies have shown that the most abundant bacteria are
Staphylococcus aureus, Staphylococcus epidermidis, and
Propionibacterium acnes.[
39,
40]
Of evolutionary importance,
Bartonella kraznovii is associated with sub-Saharan black African rats.
41 During the great “throttling,”
Homo sapiens moved into caves along the seacoast because of significant climate changes (Marine Isotope Stage 6).[
42] These surviving H. sapiens' diet included tubers, often roasted in caves.[
43] The roasted tubers were consumed as a survivor's food, but any leftovers would have been available for the cave rodents. The association of this nasal bacterium with rodents that share
Homo sapiens caves should not be surprising and is an interesting result of this study.
Burkholderia dolosa is a species of bacteria. It is a member of the
Burkholderia cepacia complex. This strain is highly drug-resistant and primarily found in immunocompromised patients.[
44]
B. dolosa chronic infection in cystic fibrosis is associated with an accelerated loss of lung function and decreased survival.[
45] However, this was observed in several participants. Another rare pathogen found in patients with cystic fibrosis,
Achromobacter xylosoxidans, was also found in a few participants.[
46] However, as in all diseases, it is not just the presence of pathogens but the absence of compensatory commensals and probiotic microorganisms that influence the expression of pathology.[
47] Xylitol solutions may inhibit pathogens without negatively reducing the levels of probiotic bacteria, as has often been reported in the literature.[
48] Xylitol nebulizers have already been proven effective in patients with cystic fibrosus.[
49]
Nasal microbiome changes and sleep-disturbed breathing may be correlated, which warrants significant research.[
50] This research project utilized young adults, not children, but examined the possible correlation between the nasal microbiome and nasal obstruction due to chronic inflammation, which leads to nasal airway obstruction.[
51] The connection between the airway and microbiome has been well established. Although many researchers believe that the airway affects the microbiome, they do not consider that it affects respiration, albeit indirectly. For instance, the oral microbiome produces nitrites from nitrates, which are eventually processed by stomach acid into nitric oxide.[
52] Salivary nitric oxide inhibits decay, reduces periodontal disease, lowers CRP, and potentially affects airway resistance.[
53] Systemic serum levels are associated with normosystolic blood pressure during pregnancy.[
54] Nitrate-reducing oral bacterial levels are also linked to a normal pregnancy, and oral dysbiosis due to
Porphyromonas gingivalis causes pre-eclampsia.[
55,
56,
57]
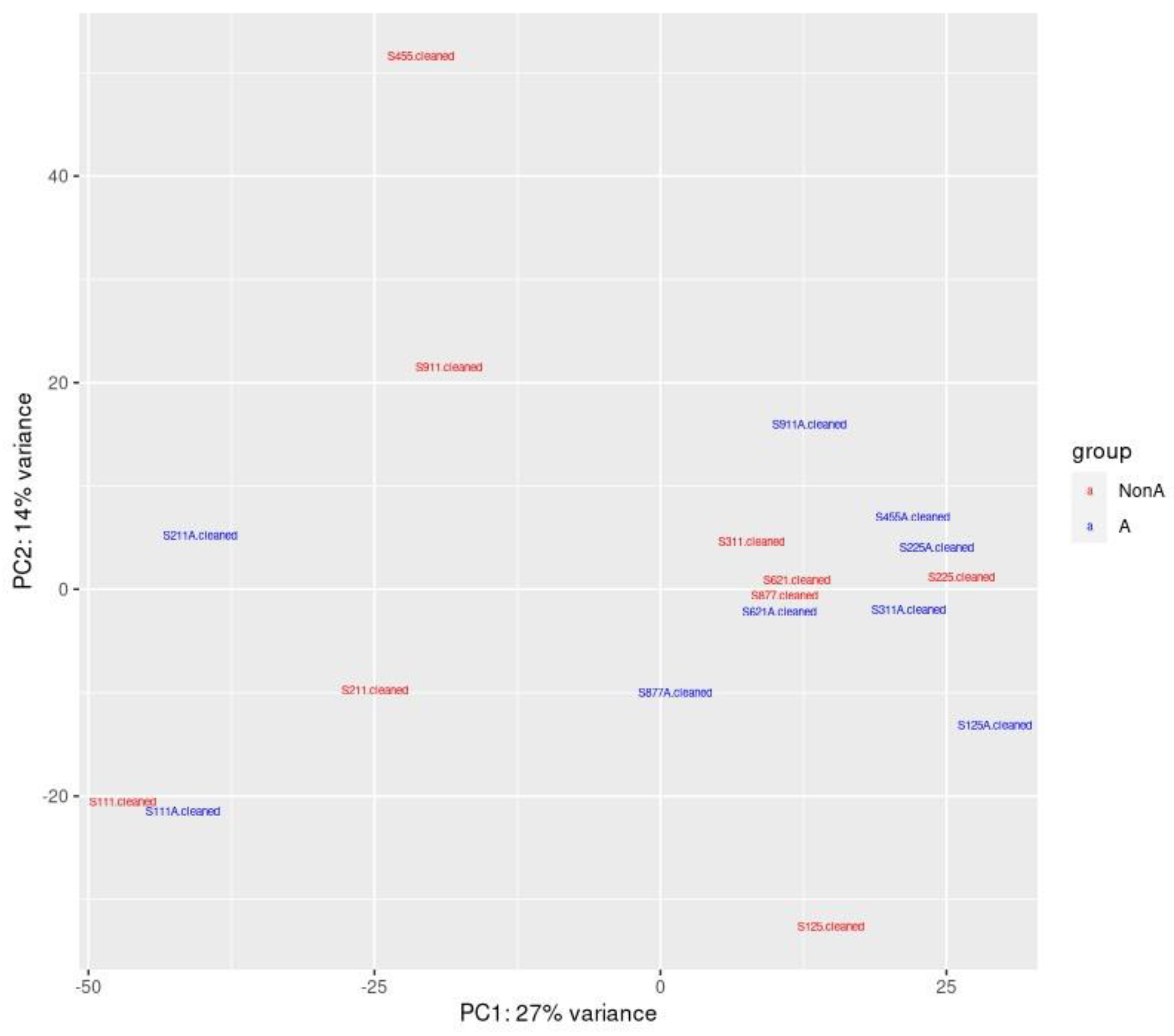
Ideally, future studies should use larger population samples to examine the association between diseases and microbiome shifts. The first step was to analyze the healthy microbiome before and after a 21-day course of xylitol (see
Figure 4). All patients were asymptomatic; therefore, no clinical correlations could be established. Animal studies have demonstrated an association between the gut and feline upper respiratory tract disease, specifically in felines. FURTD, which is often caused by infectious etiologies, is a multifactorial syndrome that affects feline populations. Using the eco-phylogenetic method, 136 and 89 microbial features were identified within the gut and nasal microbiomes, which were significantly associated with active FURTD clinical signs.[
58] Nasal and gut microbial community members are associated with a chronic clinical course.[
59] Studies have shown that endogenous microbiome dysbiosis can affect mucosal health and disease severity. Some bacterial species exhibit protective properties, whereas others are pathogenic.[
60] Antimicrobial agents can create a similar disruption, affect the nasal microbiome balance, and provoke allergic responses.[
61] Probiotics offer a promising avenue for developing systemic and topical therapies to strategically manipulate the biological host load, thereby augmenting immune homeostasis.[
62] Combining probiotics indigenous to oral and nasal cavities with prebiotics, such as xylitol, may inhibit pathogens and restore nasal health. This combination therapy may benefit the developing fetus and infant from maternal usage and prevent pathogenic microbiome development.[
63,
64,
65] An example of a potential nasal probiotic therapy is the discovery that
Lactobacilli taxa can be present on the facial skin, in the nasal cavity and vagina. Hypothetically, multiple microniches provide commensal protection to multiple microniches.[
66] Further discovery that
Lactobacilli casei (AMBR2) is present in healthy individuals but not in those with chronic rhinosinusitis led to the new designation of “keystone” probiotic commensal.[
67]
Conclusions:
The nasal microbiome is rich and diverse, containing taxa often associated with oral and skin microbiomes. Microbial interventions, even with OTC nasal rinses, may have significant effects.
Author Contributions
Principal Investigator- Cannon ML, Conceptualization, Investigation, Writing original Draft. Laboratory- Schipma M, Metabolomics- Data Curating, Methodology Ferrar- Editing Tesch- Editing
Institutional Review Board Statement
Pearl IRB, Indianapolis IN USA "Metabolomics services were performed by the Metabolomics Core Facility at Robert H. Lurie Comprehensive Cancer Center of Northwestern University."
Conflicts of Interest
Cannon- none, Schipma- none, Ferrer- Tesch- none
References
- Salo, P.M.; Arbes, S.J., Jr.; Jaramillo, R.; Calatroni, A.; Weir, C.H.; Sever, M.L.; Hoppin, J.A.; Rose, K.M.; Liu, A.H.; Gergen, P.J.; et al. Prevalence of allergic sensitization in the United States: Results from the National Health and Nutrition Examination Survey (NHANES) 2005–2006. J. Allergy Clin. Immunol. 2014, 134, 350–359. [Google Scholar] [CrossRef] [PubMed]
- Wheatley, L.M.; Togias, A. Clinical practice. Allergic rhinitis. N. Engl. J. Med. 2015, 372, 456–463. [Google Scholar] [CrossRef] [PubMed]
- Settipane, R.A. Demographics and epidemiology of allergic and nonallergic rhinitis. Allergy Asthma Proc. 2001, 22, 185–189. [Google Scholar] [PubMed]
- Bousquet, J.; Khaltaev, N.; Cruz, A.A.; Denburg, J.; Fokkens, W.J.; Togias, A.; Zuberbier, T.; Baena-Cagnani, C.E.; Canonica, G.W.; van Weel, C. Allergic Rhinitis and its Impact on Asthma (ARIA) 2008 update (in collaboration with the World Health Organization, GA(2)LEN and AllerGen). Allergy 2008, 63 (Suppl 86), 8–160. [Google Scholar] [CrossRef] [PubMed]
- Cruz, A.A.; Popov, T.; Pawankar, R.; Annesi-Maesano, I.; Fokkens, W.; Kemp, J.; Ohta, K.; Price, D.; Bousquet, J.; ARIA Initiative Scientific Committee. Common characteristics of upper and lower airways in rhinitis and asthma: ARIA update, in collaboration with GA(2)LEN. Allergy 2007, 62 (Suppl 84), 1–41. [Google Scholar] [CrossRef] [PubMed]
- Shaaban, R.; Zureik, M.; Soussan, D.; Neukirch, C.; Heinrich, J.; Sunyer, J.; Wjst, M.; Cerveri, I.; Pin, I.; Bousquet, J.; et al. Rhinitis and onset of asthma: A longitudinal population-based study. Lancet 2008, 372, 1049–1057. [Google Scholar] [CrossRef] [PubMed]
- Chirakalwasan, N.; Ruxrungtham, K. The linkage of allergic rhinitis and obstructive sleep apnea. Asian Pac. J. Allergy Immunol. 2014, 32, 276–286. [Google Scholar] [PubMed]
- Zheng, M.; Wang, X.; Ge, S.; Gu, Y.; Ding, X.; Zhang, Y.; Ye, J.; Zhang, L. Allergic and non-allergic rhinitis are common in obstructive sleep apnea but not associated with disease severity. J. Clin. Sleep Med. 2017, 13, 959–966. [Google Scholar] [CrossRef] [PubMed]
- Zheng, M.; Wang, X.; Zhang, L. Association between allergic and nonallergic rhinitis and obstructive sleep apnea. Curr. Opin. Allergy Clin. Immunol. 2018, 18, 16–25. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Wu, S.; Zhang, L.; Yang, Y.; Cao, S.; Li, Q. Association of allergic rhinitis with obstructive sleep apnea: A meta-analysis. Medicine 2018, 97, e13783. [Google Scholar] [CrossRef] [PubMed]
- Lavigne, F.; Petrof, B.J.; Johnson, J.R.; Lavigne, P.; Binothman, N.; Kassissia, G.O.; Al Samri, M.; Giordano, C.; Dubé, N.; Hercz, D.; et al. Effect of topical corticosteroids on allergic airway inflammation and disease severity in obstructive sleep apnoea. Clin. Exp. Allergy 2013, 43, 1124–1133. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K.; Jiang, J. What is normal nasal airflow? A computational study of 22 healthy adults. Int. Forum Allergy Rhinol. 2014, 4, 435–446. [Google Scholar] [CrossRef] [PubMed]
- Seppänen, T.M.; Alho, O.P.; Seppänen, T. Concomitant dynamic changes in autonomic nervous system function and nasal airflow resistance during allergen provocation. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc.. Annual International Conference of the IEEE Engineering in Medicine and Biology Society. Annual International Conference 2015, 2015, 3339–3342. [Google Scholar] [CrossRef]
- Hanif, J.; Jawad, S.S.; Eccles, R. The nasal cycle in health and disease. Clin. Otolaryngol. Allied Sci. 2000, 25, 461–467. [Google Scholar] [CrossRef] [PubMed]
- Letzel, J.; Darbinjan, A.; Hummel, T. The nasal cycle before and after nasal septoplasty. Eur. Arch. Otorhinolaryngol. 2022, 279, 4961–4968. [Google Scholar] [CrossRef] [PubMed]
- Soler, E.; de Mendoza, A.; Cuello, V.I.; Silva-Vetri, M.G.; Núñez, Z.H.; Ortega, R.G.; Rizvi, S.A.; Sanchez-Gonzalez, M.; Ferrer, G. Intranasal xylitol for the treatment of COVID-19 in the outpatient setting: A pilot study. Cureus 2022, 14, e27182. [Google Scholar] [CrossRef] [PubMed]
- Cannon, M.L.; Westover, J.B.; Bleher, R.; Sanchez-Gonzalez, M.A.; Ferrer, G. In vitro Analysis of the antiviral Potential of nasal spray constituents against SARS-CoV-2. bioRxiv 2020, 12.02.408575. [Google Scholar] [CrossRef]
- Winchester, S.; John, S.; Jabbar, K.; John, I. Clinical efficacy of nitric oxide nasal spray (NONS) for the treatment of mild COVID-19 infection. J. Infect. 2021, 83, 237–279. [Google Scholar] [CrossRef]
- Tomonobu, N.; Komalasari, N.L.G.Y.; Sumardika, I.W.; Jiang, F.; Chen, Y.; Yamamoto, K.I.; Kinoshita, R.; Murata, H.; Inoue, Y.; Sakaguchi, M. Xylitol acts as an anticancer monosaccharide to induce selective cancer death via regulation of the glutathione level. Chem. Biol. Interact. 2020, 324, 109085. [Google Scholar] [CrossRef] [PubMed]
- Shi, C.Y.; Yu, C.H.; Yu, W.Y.; Ying, H.Z. Gut-lung microbiota in chronic pulmonary diseases: Evolution, pathogenesis, and therapeutics. Can. J. Infect. Dis. Med. Microbiol. 2021, 2021, 9278441. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Dee, D.; Li, C.M.; Hoffman, H.J.; Grummer-Strawn, L.M. Breastfeeding and risk of infections at 6 years. Pediatrics 2014, 134 (Suppl 1), S13–S20. [Google Scholar] [CrossRef] [PubMed]
- Rashidi, K.; Darand, M.; Garousi, N.; Dehghani, A.; Alizadeh, S. Effect of infant formula supplemented with prebiotics and probiotics on incidence of respiratory tract infections: A systematic review and meta-analysis of randomized clinical trials. Complement. Ther. Med. 2021, 63, 102795. [Google Scholar] [CrossRef] [PubMed]
- Chan, C.K.Y.; Tao, J.; Chan, O.S.; Li, H.B.; Pang, H. Preventing respiratory tract infections by synbiotic interventions: A systematic review and meta-analysis of randomized controlled trials. Adv. Nutr. 2020, 11, 979–988. [Google Scholar] [CrossRef] [PubMed]
- Larsen, N.; Cahú, T.B.; Isay Saad, S.M.; Blennow, A.; Jespersen, L. The effect of pectins on survival of probiotic Lactobacillus spp. in gastrointestinal juices is related to their structure and physical properties. Food Microbiol. 2018, 74, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Konstantinidis, K.T.; Ramette, A.; Tiedje, J.M. The bacterial species definition in the genomic era. Phil. Trans. R. Soc. B 2006, 361, 1929–1940. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.S.; Spakowicz, D.J.; Hong, B.Y.; Petersen, L.M.; Demkowicz, P.; Chen, L.; Leopold, S.R.; Hanson, B.M.; Agresta, H.O.; Gerstein, M.; et al. Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis. Nat. Commun. 2019, 10, 5029. [Google Scholar] [CrossRef] [PubMed]
- Silva, C.F.F.S.D.; Silva, F.E.R.D.; Pauna, H.F.; Hurtado, J.G.G.M.; Dos Santos, M.C.J. Symptom assessment after nasal irrigation with xylitol in the postoperative period of endonasal endoscopic surgery. Braz. J. Orl. 2022, 88, 243–250. [Google Scholar] [CrossRef] [PubMed]
- Salli, K.; Lehtinen, M.J.; Tiihonen, K.; Ouwehand, A.C. Xylitol’s health benefits beyond Dental Health: A comprehensive review. Nutrients 2019, 11, 1813. [Google Scholar] [CrossRef]
- Martin, M. ISSN 2226-6089. http://journal.embnet.org/index.php/embnetjournal/article/view/200/458. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal, [S.l.] 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Wood, D.E.; Lu, J.; Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 2019, 20, 257. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Breitwieser, F.P.; Thielen, P.; Salzberg, S.L. Bracken: Estimating species abundance in metagenomics data. PeerJ Comput. Sci. 2017, 3, e104. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Sjödin, K.S.; Sjödin, A.; Ruszczyński, M.; Kristensen, M.B.; Hernell, O.; Szajewska, H.; West, C.E. Targeting the gut-lung axis by synbiotic feeding to infants in a randomized controlled trial. BMC Biol. 2023, 21, 38. [Google Scholar] [CrossRef] [PubMed]
- Azarpazhooh, A.; Lawrence, H.P.; Shah, P.S. Xylitol for preventing acute otitis media in children up to 12 years of age. Cochrane Database Syst. Rev. 2016, 2016, CD007095. [Google Scholar] [CrossRef] [PubMed]
- Uhari, M.; Tapiainen, T.; Kontiokari, T. Xylitol in preventing acute otitis media. Vaccine 2000, 19 (Suppl 1), S144–S147. [Google Scholar] [CrossRef] [PubMed]
- Hernández, P.; Sánchez, M.C.; Llama-Palacios, A.; Ciudad, M.J.; Collado, L. Strategies to combat caries by maintaining the integrity of biofilm and homeostasis during the rapid phase of supragingival plaque formation. Antibiotics (Basel, Switzerland) 2022, 11, 880. [Google Scholar] [CrossRef] [PubMed]
- Gasmi Benahmed, A.; Gasmi, A.; Doşa, A.; Chirumbolo, S.; Mujawdiya, P.K.; Aaseth, J.; Dadar, M.; Bjørklund, G. Association between the gut and oral microbiome with obesity. Anaerobe 2021, 70, 102248. [Google Scholar] [CrossRef]
- Olsen, I.; Yamazaki, K. Can oral bacteria affect the microbiome of the gut? J. Oral Microbiol. 2019, 11, 1586422. [Google Scholar] [CrossRef] [PubMed]
- Bassis, C.M.; Tang, A.L.; Young, V.B.; Pynnonen, M.A. The nasal cavity microbiota of healthy adults. Microbiome 2014, 2, 27. [Google Scholar] [CrossRef] [PubMed]
- Rawls, M.; Ellis, A.K. The microbiome of the nose. Ann. Allergy Asthma Immunol. 2019, 122, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, R.; Shalit, T.; Markus, B.; Yuan, C.; Nachum-Biala, Y.; Elad, D.; Harrus, S. Bartonella kosoyi sp. nov. and Bartonella krasnovii sp. nov., two novel species closely related to the zoonotic Bartonella elizabethae, isolated from black rats and wild desert rodent-fleas. Int. J. Syst. Evol. Microbiol. 2020, 70, 1656–1665. [Google Scholar] [CrossRef] [PubMed]
- Marean, C.W. When the sea saved humanity. Sci. Am. 2010, 303, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Marlowe, F.W.; Berbesque, J.C. Tubers as fallback foods and their impact on Hadza hunter-gatherers. Am. J. Phys. Anthropol. 2009, 140, 751–758. [Google Scholar] [CrossRef] [PubMed]
- Jones, A.M.; Dodd, M.E.; Webb, A.K. Burkholderia cepacia: Current clinical issues, environmental controversies and ethical dilemmas. Eur. Respir. J. 2001, 17, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Kalish, L.A.; Waltz, D.A.; Dovey, M.; Potter-Bynoe, G.; McAdam, A.J.; LiPuma, J.J.; Gerard, C.; Goldmann, D. Impact of Burkholderia dolosa on lung function and survival in cystic fibrosis. Am. J. Respir. Crit. Care Med. 2006, 173, 421–425. [Google Scholar] [CrossRef] [PubMed]
- Isler, B.; Kidd, T.J.; Stewart, A.G.; Harris, P.; Paterson, D.L. Achromobacter infections and treatment options. Antimicrob. Agents Chemother. 2020, 64, e01025–20. [Google Scholar] [CrossRef] [PubMed]
- Lyons, K.E.; Ryan, C.A.; Dempsey, E.M.; Ross, R.P.; Stanton, C. Breast milk, a source of beneficial microbes and associated benefits for infant health. Nutrients 2020, 12, 1039. [Google Scholar] [CrossRef] [PubMed]
- Sakallioğlu, Ö.; Güvenç, I.A.; Cingi, C. Xylitol and its usage in ENT practice. J. Laryngol. Otol. 2014, 128, 580–585. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Hornick, D.; Fedler, J.; Launspach, J.L.; Teresi, M.E.; Santacroce, T.R.; Cavanaugh, J.E.; Horan, R.; Nelson, G.; Starner, T.D.; et al. Randomized controlled study of aerosolized hypertonic xylitol versus hypertonic saline in hospitalized patients with pulmonary exacerbation of cystic fibrosis. J. Cyst. Fibros. 2020, 19, 108–113. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Goldberg, A.N.; Chang, J.L. The nose and nasal breathing in sleep apnea. Otolaryngol. Clin. North Am. 2020, 53, 385–395. [Google Scholar] [CrossRef] [PubMed]
- Nosetti, L.; Piacentini, G.; Macchi, A.; De Bernardi, F.; Simoncini, D.; Nicoloso, M.; Agosti, M.; Zaffanello, M. Nasal cytology in children with primary snoring and obstructive sleep apnoea syndrome. Int. J. PediatrInt. J. Pediatr. Otorhinolaryngol. Orl. 2019, 122, 133–137. [Google Scholar] [CrossRef] [PubMed]
- Rosier, B.T.; Takahashi, N.; Zaura, E.; Krom, B.P.; Martínez-Espinosa, R.M.; van Breda, S.G.J.; Marsh, P.D.; Mira, A. The importance of nitrate reduction for Oral Health. J. Dent. Res. 2022, 101, 887–897. [Google Scholar] [CrossRef]
- de Farias, J.O.; de Freitas Lima, S.M.; Rezende, T.M.B. Physiopathology of nitric oxide in the oral environment and its biotechnological potential for new oral treatments: A literature review. Clin. Oral Investig. 2020, 24, 4197–4212. [Google Scholar] [CrossRef] [PubMed]
- Altemani, F.; Barrett, H.L.; Callaway, L.K.; McIntyre, H.D.; Dekker Nitert, M. Reduced abundance of nitrate-reducing bacteria in the oral microbiota of women with future preeclampsia. Nutrients 2022, 14, 1139. [Google Scholar] [CrossRef] [PubMed]
- Owusu Darkwa, E.; Djagbletey, R.; Sottie, D.; Owoo, C.; Vanderpuye, N.M.; Essuman, R.; Aryee, G. Serum nitric oxide levels in healthy pregnant women: A case- control study in a tertiary facility in Ghana. Matern. Health Neonatol. Perinatol. 2018, 4, 3. [Google Scholar] [CrossRef] [PubMed]
- Vanterpool, S.F.; Been, J.V.; Houben, M.L.; Nikkels, P.G.; De Krijger, R.R.; Zimmermann, L.J.; Kramer, B.W.; Progulske-Fox, A.; Reyes, L. Porphyromonas gingivalis within Placental Villous Mesenchyme and Umbilical Cord stroma Is Associated with Adverse Pregnancy Outcome. PLOS ONE 2016, 11, e0146157. [Google Scholar] [CrossRef] [PubMed]
- León, R.; Silva, N.; Ovalle, A.; Chaparro, A.; Ahumada, A.; Gajardo, M.; Martinez, M.; Gamonal, J. Detection of Porphyromonas gingivalis in the amniotic fluid in pregnant women with a diagnosis of threatened premature labor. J. Periodontol. 2007, 78, 1249–1255. [Google Scholar] [CrossRef] [PubMed]
- Arnold, H.K.; Hanselmann, R.; Duke, S.M.; Sharpton, T.J.; Beechler, B.R. Chronic clinical signs of upper respiratory tract disease associate with gut and respiratory microbiomes in a cohort of domestic felines. PLOS ONE 2022, 17, e0268730. [Google Scholar] [CrossRef] [PubMed]
- Miraglia Del Giudice, M.; Parisi, G.F.; Indolfi, C.; Manti, S.; Leonardi, S.; Decimo, F.; Ciprandi, G. Nasal microbiome in chronic rhinosinusitis. Minerva Pediatr. 2022, 74, 586–592. [Google Scholar] [CrossRef] [PubMed]
- Cho, D.Y.; Hunter, R.C.; Ramakrishnan, V.R. The microbiome and chronic rhinosinusitis. Immunol. Allergy Clin. North Am. 2020, 40, 251–263. [Google Scholar] [CrossRef] [PubMed]
- Tramper-Stranders, G.; Ambrożej, D.; Arcolaci, A.; Atanaskovic-Markovic, M.; Boccabella, C.; Bonini, M.; Karavelia, A.; Mingomataj, E.; O’ Mahony, L.; Sokolowska, M.; et al. Dangerous liaisons: Bacteria, antimicrobial therapies, and allergic diseases. Allergy 2021, 76, 3276–3291. [Google Scholar] [CrossRef] [PubMed]
- Luo, C.; Peng, S.; Li, M.; Ao, X.; Liu, Z. The efficacy and safety of probiotics for allergic rhinitis: A systematic review and meta-analysis. Front. Immunol. 2022, 13, 848279. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.; Jeckel, H.; Simon-Soro, A.; Xiang, Z.; Liu, Y.; Cavalcanti, I.M.; Xiao, J.; Tin, N.N.; Hara, A.; Drescher, K.; et al. Interkingdom assemblages in human saliva display group-level surface mobility and disease-promoting emergent functions. Proc. Natl. Acad. Sci. U. S. A. 2022, 119, e2209699119. [Google Scholar] [CrossRef] [PubMed]
- Alamoudi, N.M.; Hanno, A.G.; Sabbagh, H.J.; Masoud, M.I.; Almushayt, A.S.; El Derwi, D.A. Impact of maternal xylitol consumption on mutans streptococci, plaque and caries levels in children. J. Clin. Pediatr. Dent. 2012, 37, 163–166. [Google Scholar] [CrossRef] [PubMed]
- Talattof, Z.; Azad, A.; Zahed, M.; Shahradnia, N. Antifungal activity of xylitol against Candida albicans: An in vitro study. J. Contemp. Dent. Pract. 2018, 19, 125–129. [Google Scholar] [CrossRef] [PubMed]
- Lebeer, S.; Oerlemans, E.F.M.; Claes, I.; Wuyts, S.; Henkens, T.; Spacova, I.; van den Broek, M.F.L.; Tuyaerts, I.; Wittouck, S.; De Boeck, I.; Allonsius, C.N.; Kiekens, F.; Lambert, J. Topical cream with live lactobacilli modulates the skin microbiome and reduce acne symptoms. bioRxiv 2018, 463307. [Google Scholar] [CrossRef]
- De Boeck, I.; van den Broek, M.F.L.; Allonsius, C.N.; Spacova, I.; Wittouck, S.; Martens, K.; Wuyts, S.; Cauwenberghs, E.; Jokicevic, K.; Vandenheuvel, D.; et al. Lactobacilli have a niche in the human nose. Cell Rep. 2020, 31, 107674. [Google Scholar] [CrossRef] [PubMed]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).