Submitted:
27 March 2024
Posted:
28 March 2024
You are already at the latest version
Abstract
Keywords:
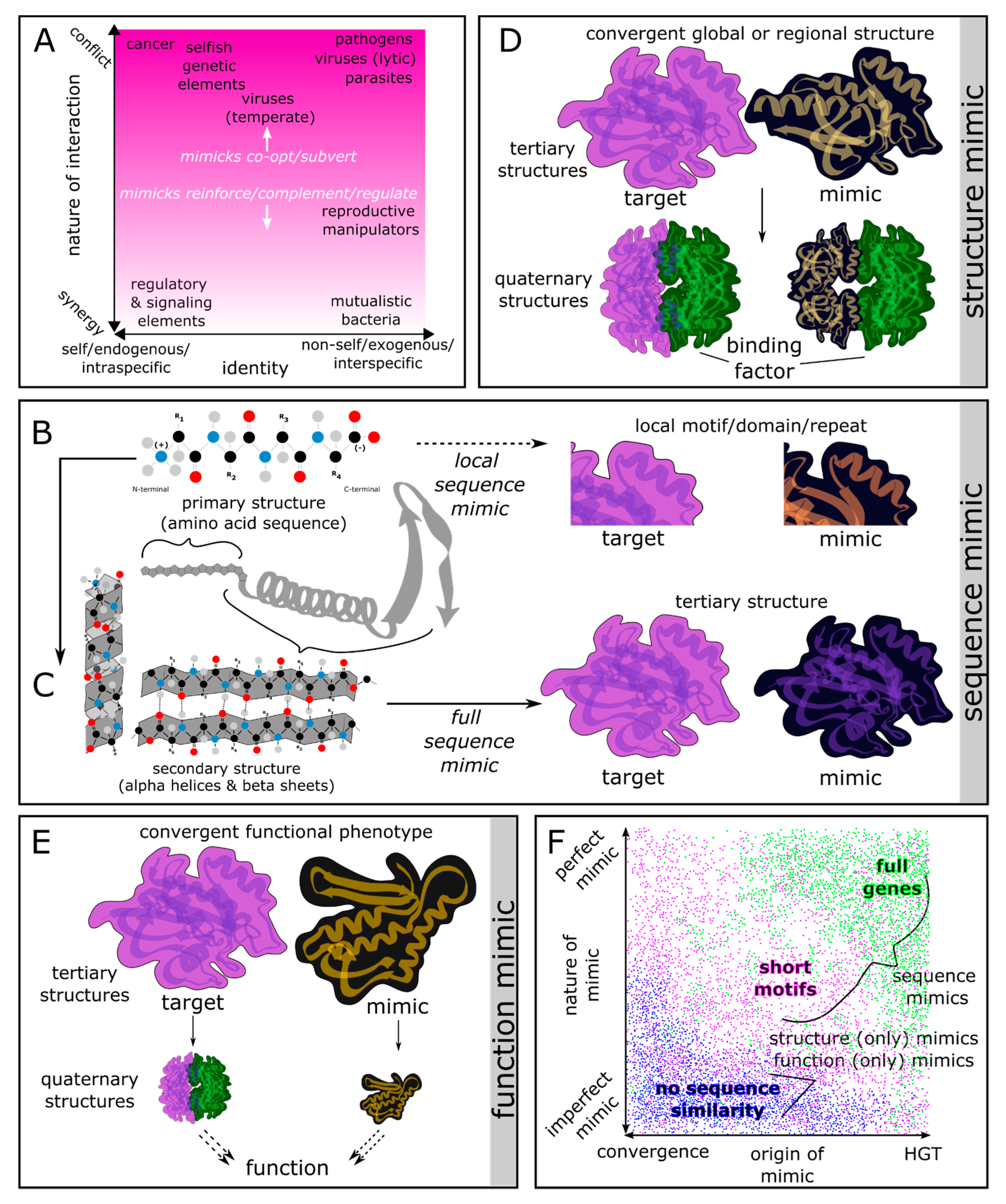
1. Introduction to Genetic Conflict and Molecular Mimicry
2. Distribution of Molecular Mimicry across Life: Expectations from Molecular Evolutionary Theory
3. Empirical examples of molecular mimicry in nature
3.1. Sequence-Based Mimics
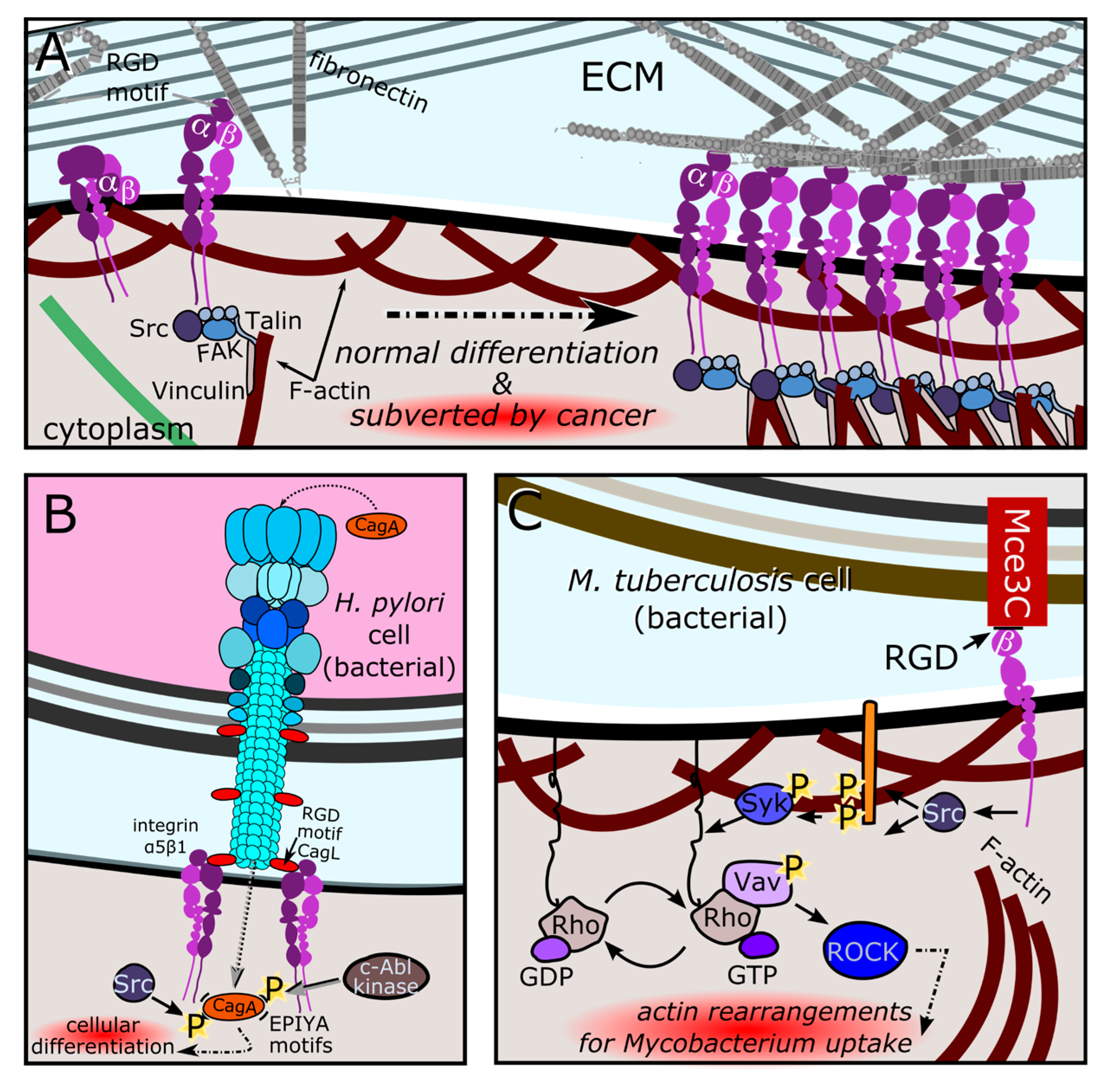
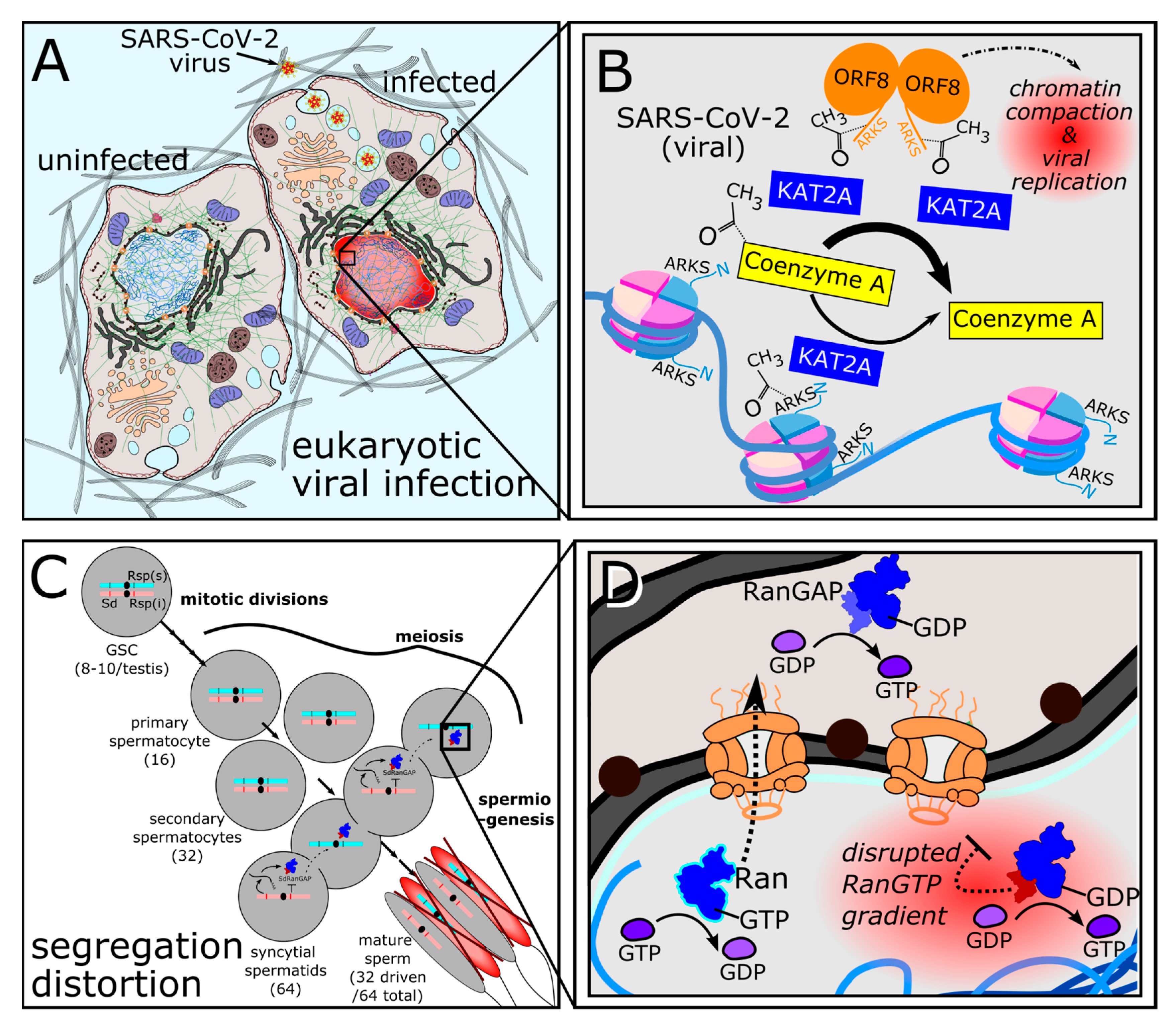
3.1.1. Recurrently Evolved SLiM Ligand Mimics Hijack Eukaryotic Cell Biology
3.1.2. De novo Full-Length Sequence Mimics Arise from Gene Duplications
3.1.3. Horizontally Transmitted Full-Gene Mimics Are Repurposed for Conflict across Life
3.2. Structural Mimics
3.2.1. Repurposed Ancient Domains for Molecular Mimicry
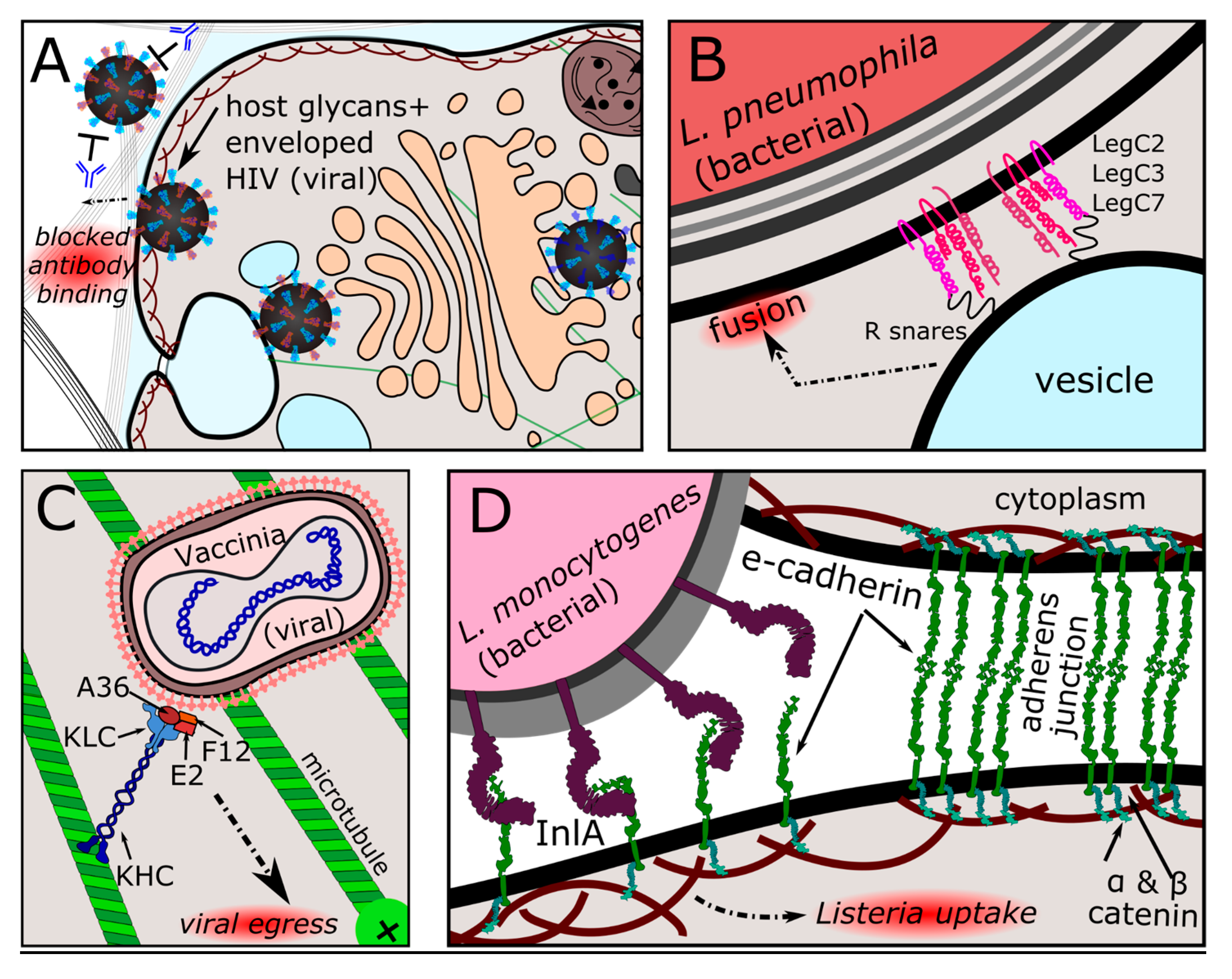
3.2.2. Convergent Structural Mimics among Membrane-Bound and Secreted Effector Proteins
3.3. Functional Mimics
3.3.1. Convergent Charge Distributions for DNA Mimicry
3.3.2. Horizontally Transferred Extracellular Matrix Functional Mimics
3.3.3. Bacteria Convergently Evolved a Functional Mimic System for Modifying Ubiquitination
4. Detecting molecular mimicry in genetic conflict computationally
4.1. Detecting Sequence-Level Mimics
4.2. Detecting Structure-Level Mimics
4.3. Detecting Function-Level Mimics
5. Conclusion
Supplementary Materials
Author Contributions
Acknowledgments
References
- Werren, J.H. Selfish genetic elements, genetic conflict, and evolutionary innovation. Proc. Natl. Acad. Sci. 2011, 108, 10863–10870. [Google Scholar] [CrossRef]
- Massey, S.E.; Mishra, B. Origin of biomolecular games: deception and molecular evolution. J. R. Soc. Interface 2018, 15, 20180429. [Google Scholar] [CrossRef] [PubMed]
- McLaughlin, R.N.; Malik, H.S. Genetic conflicts: the usual suspects and beyond. J. Exp. Biol. 2017, 220, 6–17. [Google Scholar] [CrossRef]
- Ågren, J.A.; Clark, A.G. Selfish genetic elements. PLOS Genet. 2018, 14, e1007700. [Google Scholar] [CrossRef] [PubMed]
- Elde, N.C.; Malik, H.S. The evolutionary conundrum of pathogen mimicry. Nat. Rev. Microbiol. 2009, 7, 787–797. [Google Scholar] [CrossRef] [PubMed]
- Qin, S.; Liu, Y.; Tempel, W.; Eram, M.S.; Bian, C.; Liu, K.; Senisterra, G.; Crombet, L.; Vedadi, M.; Min, J. Structural basis for histone mimicry and hijacking of host proteins by influenza virus protein NS1. Nat. Commun. 2014, 5, 3952. [Google Scholar] [CrossRef]
- Aravind, L.; Iyer, L.M.; Burroughs, A.M. Discovering Biological Conflict Systems Through Genome Analysis: Evolutionary Principles and Biochemical Novelty. Annu. Rev. Biomed. Data Sci. 2022, 5, 367–391. [Google Scholar] [CrossRef] [PubMed]
- de Groot, N.S.; Burgas, M.T. Bacteria use structural imperfect mimicry to hijack the host interactome. PLOS Comput. Biol. 2020, 16, e1008395. [Google Scholar] [CrossRef]
- Holen, H.; Johnstone, R.A. The Evolution of Mimicry under Constraints. Am. Nat. 2004, 164, 598–613. [Google Scholar] [CrossRef]
- Alcami, A. Viral mimicry of cytokines, chemokines and their receptors. Nat. Rev. Immunol. 2003, 3, 36–50. [Google Scholar] [CrossRef]
- Stebbins, C.E.; Galán, J.E. Structural mimicry in bacterial virulence. Nature 2001, 412, 701–705. [Google Scholar] [CrossRef]
- Burstein, D.; Zusman, T.; Degtyar, E.; Viner, R.; Segal, G.; Pupko, T. Genome-Scale Identification of Legionella pneumophila Effectors Using a Machine Learning Approach. PLOS Pathog. 2009, 5, e1000508. [Google Scholar] [CrossRef]
- Xu, S.; Zhang, C.; Miao, Y.; Gao, J.; Xu, D. Effector prediction in host-pathogen interaction based on a Markov model of a ubiquitous EPIYA motif. BMC Genom. 2010, 11, 1–15. [Google Scholar] [CrossRef]
- Doxey, A.C.; McConkey, B.J. Prediction of molecular mimicry candidates in human pathogenic bacteria. Virulence 2013, 4, 453–466. [Google Scholar] [CrossRef]
- Drayman, N.; Glick, Y.; Ben-Nun-Shaul, O.; Zer, H.; Zlotnick, A.; Gerber, D.; Schueler-Furman, O.; Oppenheim, A. Pathogens Use Structural Mimicry of Native Host Ligands as a Mechanism for Host Receptor Engagement. Cell Host Microbe 2013, 14, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Bryant, P.; Pozzati, G.; Elofsson, A. Improved prediction of protein-protein interactions using AlphaFold2. Nat. Commun. 2022, 13, 1–11. [Google Scholar] [CrossRef]
- Muthye, V.; Wasmuth, J.D. Proteome-wide comparison of tertiary protein structures reveals molecular mimicry in Plasmodium-human interactions. Front. Parasitol. 2023, 2, 1162697. [Google Scholar] [CrossRef]
- Mondino, S.; Schmidt, S.; Buchrieser, C. Molecular Mimicry: a Paradigm of Host-Microbe Coevolution Illustrated by Legionella. mBio 2020, 11. [Google Scholar] [CrossRef]
- Davey, N.E.; Travé, G.; Gibson, T.J. How viruses hijack cell regulation. Trends Biochem. Sci. 2011, 36, 159–169. [Google Scholar] [CrossRef]
- Frank, A.C. Molecular host mimicry and manipulation in bacterial symbionts. FEMS Microbiol. Lett. 2019, 366. [Google Scholar] [CrossRef]
- Ma, J.; Ratan, A.; Raney, B.J.; Suh, B.B.; Miller, W.; Haussler, D. The infinite sites model of genome evolution. Proc. Natl. Acad. Sci. USA 2008, 105, 14254–14261. [Google Scholar] [CrossRef] [PubMed]
- Harpak, A.; Bhaskar, A.; Pritchard, J.K. Mutation Rate Variation is a Primary Determinant of the Distribution of Allele Frequencies in Humans. PLOS Genet. 2016, 12, e1006489. [Google Scholar] [CrossRef] [PubMed]
- Stern, D.L. The genetic causes of convergent evolution. Nat. Rev. Genet. 2013, 14, 751–764. [Google Scholar] [CrossRef]
- Markov, P.V.; Ghafari, M.; Beer, M.; Lythgoe, K.; Simmonds, P.; Stilianakis, N.I.; Katzourakis, A. The evolution of SARS-CoV-2. Nat. Rev. Microbiol. 2023, 21, 361–379. [Google Scholar] [CrossRef]
- Giulieri, S.G.; Guérillot, R.; Duchene, S.; Hachani, A.; Daniel, D.; Seemann, T.; Davis, J.S.; Tong, S.Y.; Young, B.C.; Wilson, D.J.; et al. Niche-specific genome degradation and convergent evolution shaping Staphylococcus aureus adaptation during severe infections. eLife 2022, 11. [Google Scholar] [CrossRef]
- Tompa, P.; Davey, N.E.; Gibson, T.J.; Babu, M.M. A Million Peptide Motifs for the Molecular Biologist. Mol. Cell 2014, 55, 161–169. [Google Scholar] [CrossRef]
- Kajala, K.; Gouran, M.; Shaar-Moshe, L.; Mason, G.A.; Rodriguez-Medina, J.; Kawa, D.; Pauluzzi, G.; Reynoso, M.; Canto-Pastor, A.; Manzano, C.; et al. Innovation, conservation, and repurposing of gene function in root cell type development. Cell 2021, 184, 3333–3348.e19. [Google Scholar] [CrossRef] [PubMed]
- Nitta, K.R.; Jolma, A.; Yin, Y.; Morgunova, E.; Kivioja, T.; Akhtar, J.; Hens, K.; Toivonen, J.; Deplancke, B.; Furlong, E.E.M.; et al. Conservation of transcription factor binding specificities across 600 million years of bilateria evolution. eLife 2015, 4, e04837. [Google Scholar] [CrossRef]
- Li, S.; Dohlman, H.G. Evolutionary conservation of sequence motifs at sites of protein modification. J. Biol. Chem. 2023, 299, 104617. [Google Scholar] [CrossRef]
- E Martyn, J.; Gomez-Valero, L.; Buchrieser, C. The evolution and role of eukaryotic-like domains in environmental intracellular bacteria: the battle with a eukaryotic cell. FEMS Microbiol. Rev. 2022, 46. [Google Scholar] [CrossRef]
- Kumar, M.; Michael, S.; Alvarado-Valverde, J.; Mészáros, B.; Sámano-Sánchez, H.; Zeke, A.; Dobson, L.; Lazar, T.; Örd, M.; Nagpal, A.; et al. The Eukaryotic Linear Motif resource: 2022 release. Nucleic Acids Res. 2021, 50, D497–D508. [Google Scholar] [CrossRef] [PubMed]
- Sámano-Sánchez, H.; Gibson, T.J. Mimicry of Short Linear Motifs by Bacterial Pathogens: A Drugging Opportunity. Trends Biochem. Sci. 2020, 45, 526–544. [Google Scholar] [CrossRef] [PubMed]
- Courret, C.; Chang, C.-H.; Wei, K.H.-C.; Montchamp-Moreau, C.; Larracuente, A.M. Meiotic drive mechanisms: lessons fromDrosophila. Proc. R. Soc. B: Biol. Sci. 2019, 286, 20191430. [Google Scholar] [CrossRef]
- Levasseur, A.; Pontarotti, P. The role of duplications in the evolution of genomes highlights the need for evolutionary-based approaches in comparative genomics. Biol. Direct 2011, 6, 11. [Google Scholar] [CrossRef]
- A Vogan, A.; Ament-Velásquez, S.L.; Granger-Farbos, A.; Svedberg, J.; Bastiaans, E.; Debets, A.J.; Coustou, V.; Yvanne, H.; Clavé, C.; Saupe, S.J.; et al. Combinations of Spok genes create multiple meiotic drivers in Podospora. eLife 2019, 8. [Google Scholar] [CrossRef]
- Botham, C.M.; Wandler, A.M.; Guillemin, K. A Transgenic Drosophila Model Demonstrates That the Helicobacter pylori CagA Protein Functions as a Eukaryotic Gab Adaptor. PLOS Pathog. 2008, 4, e1000064. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, J.; Li, B.; Wang, J.; Liu, C.H. Mycobacterium tuberculosisMce3C promotes mycobacteria entry into macrophages through activation of β2 integrin-mediated signalling pathway. Cell. Microbiol. 2017, 20, e12800. [Google Scholar] [CrossRef]
- Kee, J.; Thudium, S.; Renner, D.M.; Glastad, K.; Palozola, K.; Zhang, Z.; Li, Y.; Lan, Y.; Cesare, J.; Poleshko, A.; et al. SARS-CoV-2 disrupts host epigenetic regulation via histone mimicry. Nature 2022, 610, 381–388. [Google Scholar] [CrossRef]
- Davey, N.E.; Cyert, M.S.; Moses, A.M. Short linear motifs – ex nihilo evolution of protein regulation. Cell Commun. Signal. 2015, 13, 1–15. [Google Scholar] [CrossRef]
- Hamidi, H.; Ivaska, J. Every step of the way: integrins in cancer progression and metastasis. Nat. Rev. Cancer 2018, 18, 533–548. [Google Scholar] [CrossRef]
- Conradi, J.; Tegtmeyer, N.; Woźna, M.; Wissbrock, M.; Michalek, C.; Gagell, C.; Cover, T.L.; Frank, R.; Sewald, N.; Backert, S. An RGD Helper Sequence in CagL of Helicobacter pylori Assists in Interactions with Integrins and Injection of CagA. Front. Cell. Infect. Microbiol. 2012, 2, 70. [Google Scholar] [CrossRef]
- Hayashi, T.; Morohashi, H.; Hatakeyama, M. Bacterial EPIYA effectors - Where do they come from? What are they? Where are they going? Cell. Microbiol. 2012, 15, 377–385. [Google Scholar] [CrossRef]
- Norris, E.G.; Pan, X.S.; Hocking, D.C. Receptor-binding domain of SARS-CoV-2 is a functional αv-integrin agonist. J. Biol. Chem. 2023, 299, 102922. [Google Scholar] [CrossRef] [PubMed]
- Simons, P.; Rinaldi, D.A.; Bondu, V.; Kell, A.M.; Bradfute, S.; Lidke, D.S.; Buranda, T. Integrin activation is an essential component of SARS-CoV-2 infection. Sci. Rep. 2021, 11, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, U.; Ho, J.S.; Prinjha, R.K.; Tarakhovsky, A. The "Histone Mimicry" by Pathogens. Cold Spring Harb. Symp. Quant. Biol. 2013, 78, 81–90. [Google Scholar] [CrossRef]
- Marazzi, I.; Ho, J.S.Y.; Kim, J.; Manicassamy, B.; Dewell, S.; Albrecht, R.A.; Seibert, C.W.; Schaefer, U.; Jeffrey, K.L.; Prinjha, R.K.; et al. Suppression of the antiviral response by an influenza histone mimic. Nature 2012, 483, 428–433. [Google Scholar] [CrossRef]
- Shi, J.; Wang, Y.; Zeng, L.; Wu, Y.; Deng, J.; Zhang, Q.; Lin, Y.; Li, J.; Kang, T.; Tao, M.; et al. Disrupting the Interaction of BRD4 with Diacetylated Twist Suppresses Tumorigenesis in Basal-like Breast Cancer. Cancer Cell 2014, 25, 210–225. [Google Scholar] [CrossRef] [PubMed]
- Gingell, L.F.; McLean, J.R. A Protamine Knockdown Mimics the Function ofSdinDrosophila melanogaster. G3 Genes|Genomes|Genetics 2020, 10, 2111–2115. [Google Scholar] [CrossRef]
- Herbette, M.; Wei, X.; Chang, C.-H.; Larracuente, A.M.; Loppin, B.; Dubruille, R. Distinct spermiogenic phenotypes underlie sperm elimination in the Segregation Distorter meiotic drive system. PLOS Genet. 2021, 17, e1009662. [Google Scholar] [CrossRef]
- Malone, C.D.; Lehmann, R.; Teixeira, F.K. The cellular basis of hybrid dysgenesis and Stellate regulation in Drosophila. Curr. Opin. Genet. Dev. 2015, 34, 88–94. [Google Scholar] [CrossRef]
- Burga, A.; Ben-David, E.; Kruglyak, L. Toxin-Antidote Elements Across the Tree of Life. Annu. Rev. Genet. 2020, 54, 387–415. [Google Scholar] [CrossRef]
- Silpe, J.E.; Bassler, B.L. A Host-Produced Quorum-Sensing Autoinducer Controls a Phage Lysis-Lysogeny Decision. Cell 2018, 176, 268–280.e13. [Google Scholar] [CrossRef]
- Jurėnas, D.; Fraikin, N.; Goormaghtigh, F.; Van Melderen, L. Biology and evolution of bacterial toxin–antitoxin systems. Nat. Rev. Microbiol. 2022, 20, 335–350. [Google Scholar] [CrossRef]
- Blower, T.R.; Evans, T.J.; Przybilski, R.; Fineran, P.C.; Salmond, G.P.C. Viral Evasion of a Bacterial Suicide System by RNA–Based Molecular Mimicry Enables Infectious Altruism. PLOS Genet. 2012, 8, e1003023. [Google Scholar] [CrossRef] [PubMed]
- Núñez, M.A.B.; Lange, J.J.; Zanders, S.E. A suppressor of a wtf poison-antidote meiotic driver acts via mimicry of the driver’s antidote. PLOS Genet. 2018, 14, e1007836. [Google Scholar] [CrossRef]
- English, J.; Patrick, S.; Stewart, L.D. The potential role of molecular mimicry by the anaerobic microbiota in the aetiology of autoimmune disease. Anaerobe 2023, 80, 102721. [Google Scholar] [CrossRef] [PubMed]
- de Groot, N.S.; Burgas, M.T. Bacteria use structural imperfect mimicry to hijack the host interactome. PLOS Comput. Biol. 2020, 16, e1008395. [Google Scholar] [CrossRef] [PubMed]
- Beaudoin, C.A.; Jamasb, A.R.; Alsulami, A.F.; Copoiu, L.; van Tonder, A.J.; Hala, S.; Bannerman, B.P.; Thomas, S.E.; Vedithi, S.C.; Torres, P.H.; et al. Predicted structural mimicry of spike receptor-binding motifs from highly pathogenic human coronaviruses. Comput. Struct. Biotechnol. J. 2021, 19, 3938–3953. [Google Scholar] [CrossRef] [PubMed]
- Lasso, G.; Honig, B.; Shapira, S.D. A Sweep of Earth’s Virome Reveals Host-Guided Viral Protein Structural Mimicry and Points to Determinants of Human Disease. Cell Syst. 2020, 12, 82–91.e3. [Google Scholar] [CrossRef]
- Böhmer, F.; Szedlacsek, S.; Tabernero, L.; Östman, A.; Hertog, J.D. Protein tyrosine phosphatase structure–function relationships in regulation and pathogenesis. FEBS J. 2012, 280, 413–431. [Google Scholar] [CrossRef] [PubMed]
- Salomon, D.; Orth, K. What Pathogens Have Taught Us About Posttranslational Modifications. Cell Host Microbe 2013, 14, 269–279. [Google Scholar] [CrossRef]
- Beaudoin, C.A.; Jamasb, A.R.; Alsulami, A.F.; Copoiu, L.; van Tonder, A.J.; Hala, S.; Bannerman, B.P.; Thomas, S.E.; Vedithi, S.C.; Torres, P.H.; et al. Predicted structural mimicry of spike receptor-binding motifs from highly pathogenic human coronaviruses. Comput. Struct. Biotechnol. J. 2021, 19, 3938–3953. [Google Scholar] [CrossRef] [PubMed]
- Custódio, C.A.; Mano, J.F. Cell Surface Engineering to Control Cellular Interactions. Chemnanomat 2016, 2, 376–384. [Google Scholar] [CrossRef] [PubMed]
- Varki, A.; Kannagi, R.; Toole, B.; Stanley, P. Glycosylation Changes in Cancer. In Essent. Glycobiol., 3rd ed.; Varki, A., Cummings, R.D., Esko, J.D., Stanley, P., Hart, G.W., Aebi, M., Darvill, A.G., Kinoshita, T., Packer, N.H., Prestegard, J.H., Schnaar, R.L., Seeberger, P.H., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2015; Available online: http://www.ncbi.nlm.nih.gov/books/NBK453023/ (accessed on 31 October 2023).
- de Jong, H.; Wösten, M.M.S.M.; Wennekes, T. Sweet impersonators: Molecular mimicry of host glycans by bacteria. Glycobiology 2021, 32, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Miller, N.L.; Clark, T.; Raman, R.; Sasisekharan, R. Glycans in Virus-Host Interactions: A Structural Perspective. Front. Mol. Biosci. 2021, 8. [Google Scholar] [CrossRef]
- Amara, A.; Mercer, J. Viral apoptotic mimicry. Nat. Rev. Microbiol. 2015, 13, 461–469. [Google Scholar] [CrossRef]
- Shi, X.; Halder, P.; Yavuz, H.; Jahn, R.; Shuman, H.A. Direct targeting of membrane fusion by SNARE mimicry: Convergent evolution ofLegionellaeffectors. Proc. Natl. Acad. Sci. 2016, 113, 8807–8812. [Google Scholar] [CrossRef] [PubMed]
- Seo, D.; Gammon, D.B. Manipulation of Host Microtubule Networks by Viral Microtubule-Associated Proteins. Viruses 2022, 14, 979. [Google Scholar] [CrossRef] [PubMed]
- Lamason, R.L.; Welch, M.D. Actin-based motility and cell-to-cell spread of bacterial pathogens. Curr. Opin. Microbiol. 2016, 35, 48–57. [Google Scholar] [CrossRef]
- Yesudhas, D.; Batool, M.; Anwar, M.A.; Panneerselvam, S.; Choi, S. Proteins Recognizing DNA: Structural Uniqueness and Versatility of DNA-Binding Domains in Stem Cell Transcription Factors. Genes 2017, 8, 192. [Google Scholar] [CrossRef]
- Wang, H.-C.; Ho, C.-H.; Hsu, K.-C.; Yang, J.-M.; Wang, A.H.-J. DNA Mimic Proteins: Functions, Structures, and Bioinformatic Analysis. Biochemistry 2014, 53, 2865–2874. [Google Scholar] [CrossRef] [PubMed]
- Hartsock, A.; Nelson, W.J. Adherens and tight junctions: Structure, function and connections to the actin cytoskeleton. Biochim. et Biophys. Acta (BBA) - Biomembr. 2008, 1778, 660–669. [Google Scholar] [CrossRef]
- Shibata-Seki, T.; Nagaoka, M.; Goto, M.; Kobatake, E.; Akaike, T. Direct visualization of the extracellular binding structure of E-cadherins in liquid. Sci. Rep. 2020, 10, 1–10. [Google Scholar] [CrossRef]
- Bonazzi, M.; Lecuit, M.; Cossart, P. Listeria monocytogenesinternalin and E-cadherin: from structure to pathogenesis. Cell. Microbiol. 2009, 11, 693–702. [Google Scholar] [CrossRef]
- Ragon, M.; Wirth, T.; Hollandt, F.; Lavenir, R.; Lecuit, M.; Le Monnier, A.; Brisse, S. A New Perspective on Listeria monocytogenes Evolution. PLOS Pathog. 2008, 4, e1000146. [Google Scholar] [CrossRef] [PubMed]
- Grau-Bové, X.; Sebé-Pedrós, A.; Ruiz-Trillo, I. The Eukaryotic Ancestor Had a Complex Ubiquitin Signaling System of Archaeal Origin. Mol. Biol. Evol. 2014, 32, 726–739. [Google Scholar] [CrossRef] [PubMed]
- Franklin, T.G.; Pruneda, J.N. Bacteria make surgical strikes on host ubiquitin signaling. PLOS Pathog. 2021, 17, e1009341. [Google Scholar] [CrossRef]
- Hebert, F.O.; Phelps, L.; Samonte, I.; Panchal, M.; Grambauer, S.; Barber, I.; Kalbe, M.; Landry, C.R.; Aubin-Horth, N. Identification of candidate mimicry proteins involved in parasite-driven phenotypic changes. Parasites Vectors 2015, 8, 225. [Google Scholar] [CrossRef] [PubMed]
- Ludin, P.; Nilsson, D.; Mäser, P. Genome-Wide Identification of Molecular Mimicry Candidates in Parasites. PLOS ONE 2011, 6, e17546. [Google Scholar] [CrossRef]
- Hraber, P.; O’maille, P.E.; Silberfarb, A.; Davis-Anderson, K.; Generous, N.; McMahon, B.H.; Fair, J.M. Resources to Discover and Use Short Linear Motifs in Viral Proteins. Trends Biotechnol. 2019, 38, 113–127. [Google Scholar] [CrossRef]
- Lu, J.; Wu, T.; Zhang, B.; Liu, S.; Song, W.; Qiao, J.; Ruan, H. Types of nuclear localization signals and mechanisms of protein import into the nucleus. Cell Commun. Signal. 2021, 19, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Bernhofer, M.; Goldberg, T.; Wolf, S.; Ahmed, M.; Zaugg, J.; Boden, M.; Rost, B. NLSdb—major update for database of nuclear localization signals and nuclear export signals. Nucleic Acids Res. 2017, 46, D503–D508. [Google Scholar] [CrossRef] [PubMed]
- Goubert, C.; Craig, R.J.; Bilat, A.F.; Peona, V.; Vogan, A.A.; Protasio, A.V. A beginner’s guide to manual curation of transposable elements. Mob. DNA 2022, 13, 1–19. [Google Scholar] [CrossRef]
- Varadi, M.; Anyango, S.; Deshpande, M.; Nair, S.; Natassia, C.; Yordanova, G.; Yuan, D.; Stroe, O.; Wood, G.; Laydon, A.; et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 2022, 50, D439–D444. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- van Kempen, M.; Kim, S.S.; Tumescheit, C.; Mirdita, M.; Lee, J.; Gilchrist, C.L.M.; Söding, J.; Steinegger, M. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. 2023, 42, 243–246. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.J.; Choy, W.-Y.; Karttunen, M. AlphaFold2: A Role for Disordered Protein/Region Prediction? Int. J. Mol. Sci. 2022, 23, 4591. [Google Scholar] [CrossRef]
- Gozashti, L.; Roy, S.W.; Thornlow, B.; Kramer, A.; Ares, M.; Corbett-Detig, R. Transposable elements drive intron gain in diverse eukaryotes. Proc. Natl. Acad. Sci. 2022, 119. [Google Scholar] [CrossRef]
- Corbett-Detig, R.; Medina, P.; Frérot, H.; Blassiau, C.; Castric, V. Bulk pollen sequencing reveals rapid evolution of segregation distortion in the male germline of Arabidopsis hybrids. Evol. Lett. 2019, 3, 93–103. [Google Scholar] [CrossRef]
- Berube, B.; Ernst, E.; Cahn, J.; Roche, B.; de Santis Alves, C.; Lynn, J.; Scheben, A.; Siepel, A.; Ross-Ibarra, J.; Kermicle, J.; et al. Teosinte Pollen Drive guides maize domestication and evolution by RNAi. BioRxiv 2023. [Google Scholar] [CrossRef]
- Aagaard, J.E.; Springer, S.A.; Soelberg, S.D.; Swanson, W.J. Duplicate Abalone Egg Coat Proteins Bind Sperm Lysin Similarly, but Evolve Oppositely, Consistent with Molecular Mimicry at Fertilization. PLOS Genet. 2013, 9, e1003287. [Google Scholar] [CrossRef]
- Chen, S.; Qin, R.; Mahal, L.K. Sweet systems: technologies for glycomic analysis and their integration into systems biology. Crit. Rev. Biochem. Mol. Biol. 2021, 56, 301–320. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



