Submitted:
20 February 2024
Posted:
20 February 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
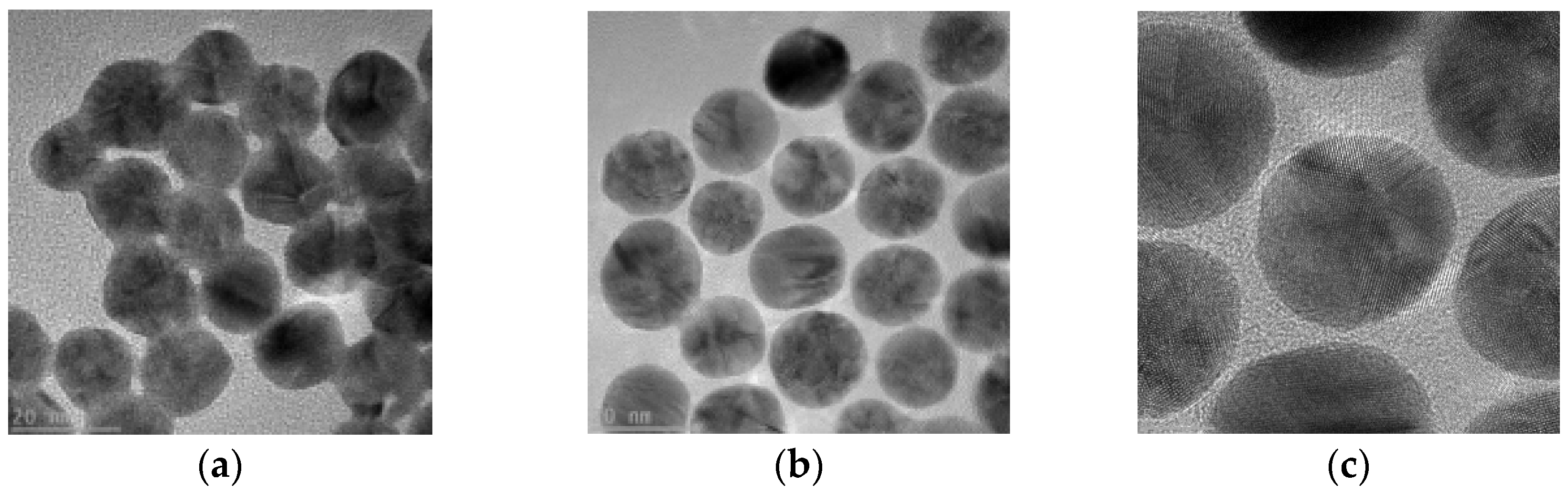
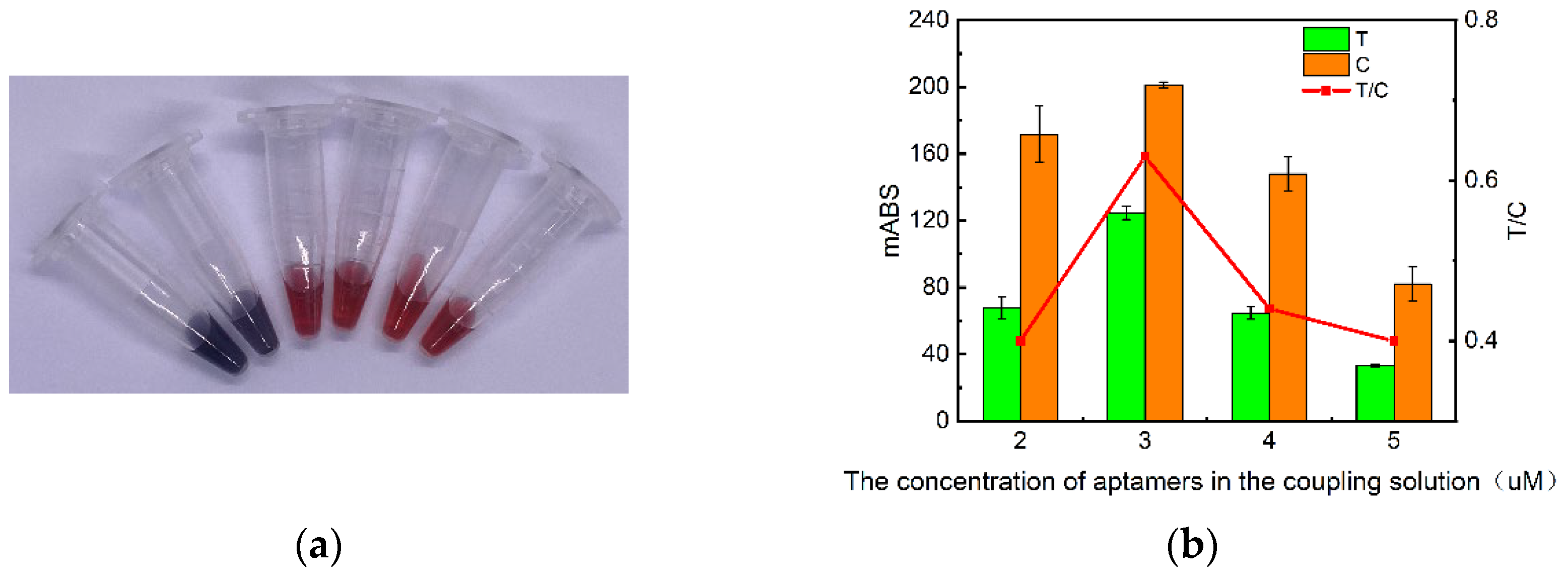
2.1. Characterization of AuNPs and determination of coupling conditions
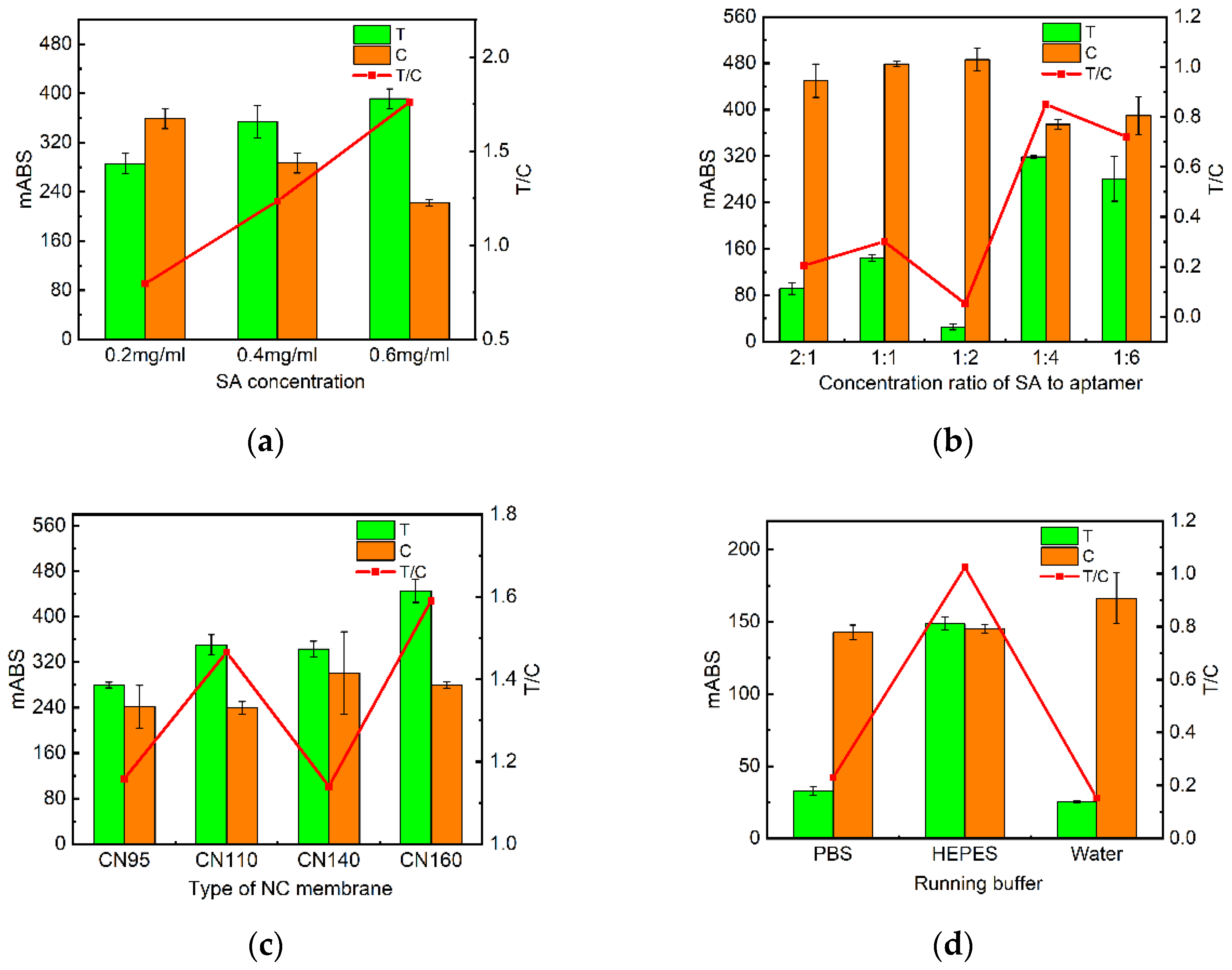
2.2. Optimization of test strip conditions
2.2.1. Optimization of SA concentrations and molar ratios of SA to Biotin-DNAT
2.2.2. Optimization of NC membranes
2.2.3. Optimization of running buffer
2.3. An aptamer engineering approach to selecting T-line complementary sequence
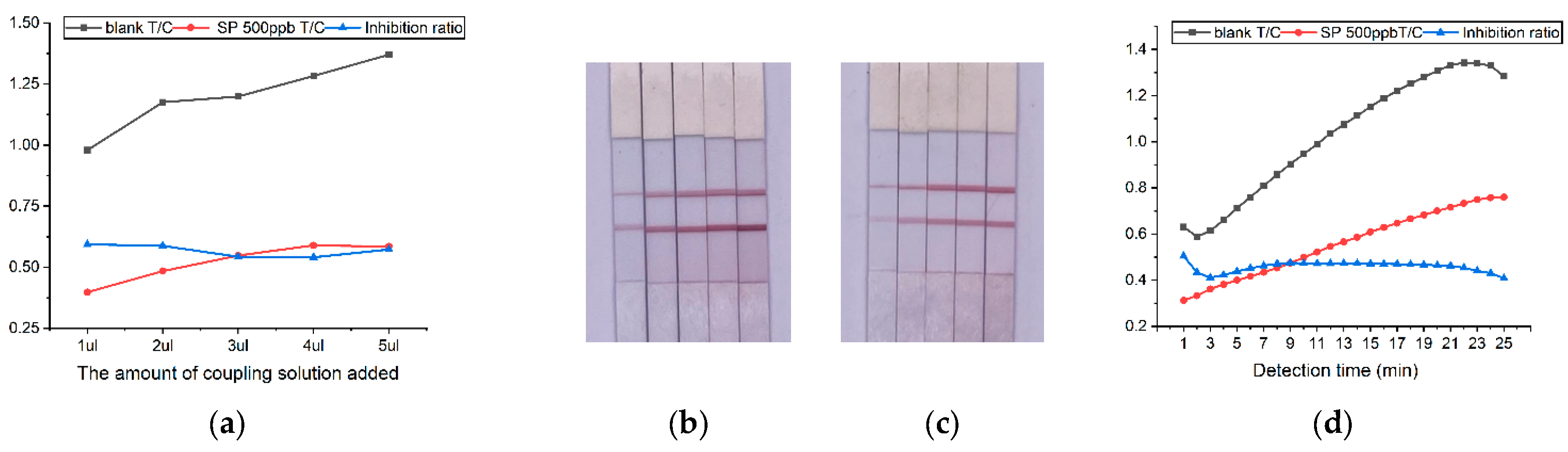
2.4. Optimization of detection conditions
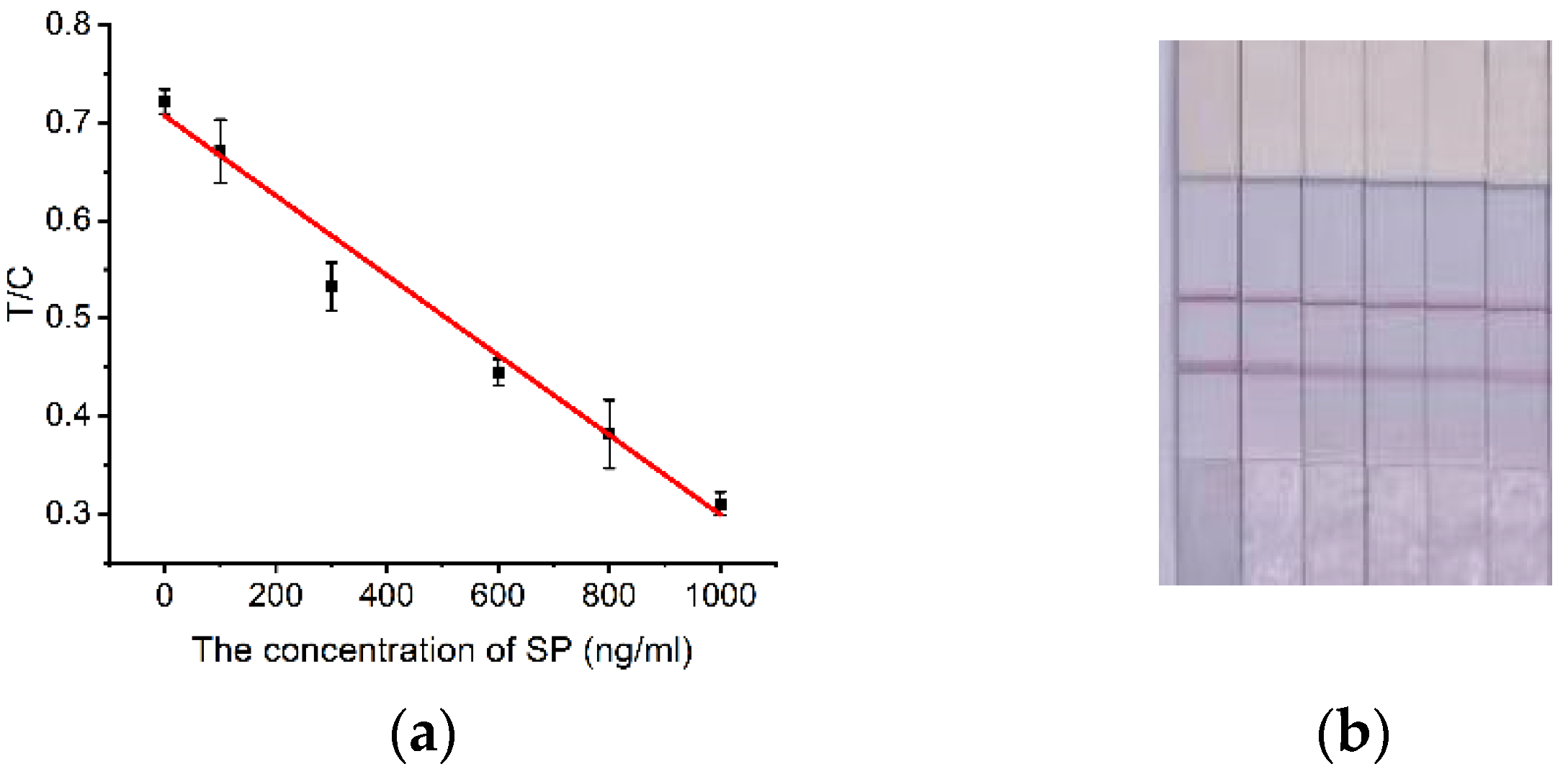
2.5. Quantitative Detection of SP by Test Strips
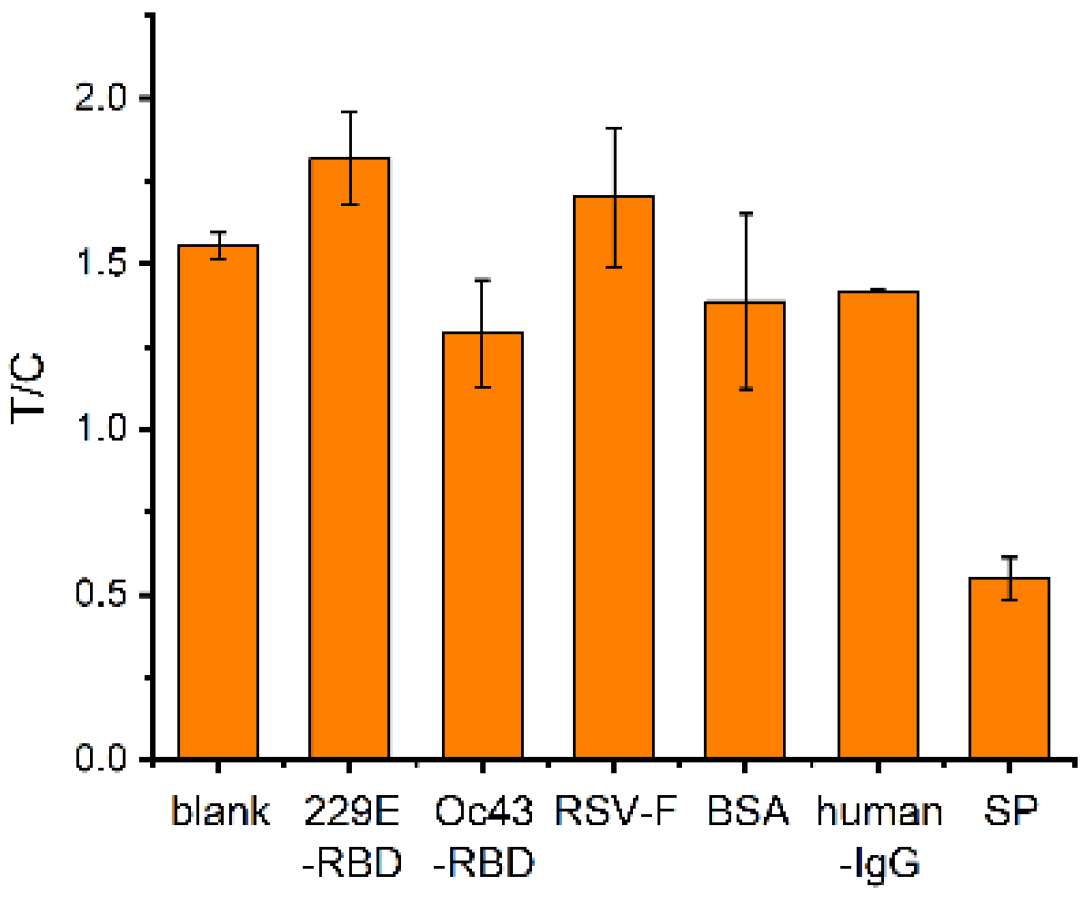
2.6. Specificity Test
2.7. Recovery assay
3. Discussion
4. Materials and Methods
4.1. Reagents and Materials
4.2. Preparation of gold nanoparticles
4.3. Conjugation of Aptamer and Nanogolds
4.4. Selection of T-line complementary sequence
4.5. Pre-treatment of test strips
4.6. Assembly of test strips
4.7. Sample Test and Evaluation Methods
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y. M.; Wang, W.; Song, Z. G.; Hu, Y.; Tao, Z. W.; Tian, J. H.; Pei, Y. Y.; Yuan, M. L.; Zhang, Y. L.; Dai, F. H.; Liu, Y.; Wang, Q. M.; Zheng, J. J.; Xu, L.; Holmes, E. C.; Zhang, Y. Z., A new coronavirus associated with human respiratory disease in China (vol 579, pg 265, 2020). Nature 2020, 580 (7803), E7-E7. [CrossRef]
- Samanta, P.; Ghosh, A. R., Environmental perspectives of COVID-19 outbreaks: A review. World J Gastroentero 2021, 27 (35). [CrossRef]
- Organization, W. H. WHO Coronavirus (COVID-19) dashboard; 2024. https://covid19.who.int/ (accessed January 2024).
- Yuen, K. S.; Ye, Z. W.; Fung, S. Y.;Chan, C. P.; Jin, D. Y., SARS-CoV-2 and COVID-19: The most important research questions. Cell Biosci 2020, 10 (1). [CrossRef]
- Han, S. L.; Liu, X. W., Can imported cold food cause COVID-19 recurrent outbreaks? A review. Environ Chem Lett 2022, 20 (1), 119-129. [CrossRef]
- Zhang, J.; Fang, X.; Mao, Y.; Qi, H. C.; Wu, J.; Liu, X. R.; You, F. S.; Zhao, W. C.; Chen, Y.; Zheng, L., Real-time, selective, and low-cost detection of trace level SARS-CoV-2 spike-protein for cold-chain food quarantine. Npj Sci Food 2021, 5 (1). [CrossRef]
- Mousavizadeh, L.; Ghasemi, S., Review Article Genotype and phenotype of COVID-19: Their roles in pathogenesis. J Microbiol Immunol 2021, 54 (2), 159-163. [CrossRef]
- Parikhani, A. B.; Bazaz, M.; Bamehr, H.; Fereshteh, S.; Amiri, S.; Salehi-Vaziri, M.; Arashkia, A.; Azadmanesh, K., The Inclusive Review on SARS-CoV-2 Biology, Epidemiology, Diagnosis, and Potential Management Options. Curr Microbiol 2021, 78 (4), 1099-1114. [CrossRef]
- Boopathi, S.; Poma, A. B.; Kolandaivel, P., Novel 2019 coronavirus structure, mechanism of action, antiviral drug promises and rule out against its treatment. J Biomol Struct Dyn 2021, 39 (9), 3409-3418. [CrossRef]
- Walls, A. C.; Park, Y. J.; Tortorici, M. A.; Wall, A.; McGuire, A. T.; Veesler, D., Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein (vol 180, 281.e1, 2020). Cell 2020, 183 (6), 1735-1735. [CrossRef]
- Ke, Z. L.; Oton, J. Q.; Qu, K.; Cortese, M.; Zila, V.; McKeane, L.; Nakane, T.; Zivanov, J.; Neufeldt, C. J.; Cerikan, B.; Lu, J. M.; Peukes, J.; Xiong, X. L.; Kräusslich, H. G.; Scheres, S. H. W.; Bartenschlager, R.; Briggs, J. A. G., Structures and distributions of SARS-CoV-2 spike proteins on intact virions. Nature 2020, 588 (7838), 498-+. [CrossRef]
- Svobodova, M.; Skouridou, V.; Jauset-Rubio, M.; Viéitez, I.; Fernández-Villar, A.; Alvargonzalez, J. J. C.; Poveda, E.; Bofill, C. B.; Sans, T.; Bashammakh, A.; Alyoubi, A. O.; O’Sullivan, C. K., Aptamer Sandwich Assay for the Detection of SARS-CoV-2 Spike Protein Antigen. Acs Omega 2021, 6 (51), 35657-35666. [CrossRef]
- Nejad, Z. H.; Fatemi, F.; Siadat, S. E. R., An outlook on coronavirus disease 2019 detection methods. J Pharm Anal 2022, 12 (2), 205-214. [CrossRef]
- Garg, A.; Ghoshal, U.; Patel, S. S.; Singh, D. V.; Arya, A. K.; Vasanth, S.; Pandey, A.; Srivastava, N., Evaluation of seven commercial RT-PCR kits for COVID-19 testing in pooled clinical specimens. J Med Virol 2021, 93 (4), 2281-2286. [CrossRef]
- Lin, Z.; Zou, Z. Y.; Pu, Z.; Wu, M. H.; Zhang, Y. Q., Application of microfluidic technologies on COVID-19 diagnosis and drug discovery. Acta Pharm Sin B 2023, 13 (7), 2877-2896. [CrossRef]
- De Felice, M.; De Falco, M.; Zappi, D.; Antonacci, A.; Scognamiglio, V., Isothermal amplification-assisted diagnostics for COVID-19. Biosens Bioelectron 2022, 205. [CrossRef]
- Ong, D. S. Y.; Fragkou, P. C.; Schweitzer, V. A.; Chemaly, R. F.; Moschopoulos, C. D.; Skevaki, C.; ESCMID; ESGR, S. G. R. V., How to interpret and use COVID-19 serology and immunology tests. Clin Microbiol Infec 2021, 27 (7), 981-986. [CrossRef]
- Organization., W. H. SARS-CoV-2 antigen-detecting rapid diagnostic tests: an implementation guide [EB/OL].
- Bahadir, E. B.; Sezgintürk, M. K., Lateral flow assays: Principles, designs and labels. Trac-Trend Anal Chem 2016, 82, 286-306. [CrossRef]
- Bijina J. Mathew1, A. K. V., Prashant Khare1, Sudheer Gupta2, Ram Kumar Nema2,; Shashwati Nema1, S. G., Shivendra K. Chaurasiya4, Debasis Biswas1,2, Anirudh K. Singh1*, Laboratory diagnosis of COVID-19: current status and challenges. Iranian Journal of Microbiology,2021, 13(1): 1. [CrossRef]
- Thongpradit, S.; Prasongtanakij, S.; Srisala, S.; Chanprasertyothin, S.; Pasomsub, E.; Ongphiphadhanakul, B., The Detection of SARS-CoV2 Antigen in Wastewater Using an Automated Chemiluminescence Enzyme Immunoassay. Int J Env Res Pub He 2022, 19 (13). [CrossRef]
- Trombetta, B. A.; Kandigian, S. E.; Kitchen, R. R.; Grauwet, K.; Webb, P. K.; Miller, G. A.; Jennings, C. G.; Jain, S.; Miller, S.; Kuo, Y. K.; Sweeney, T.; Gilboa, T.; Norman, M.; Simmons, D. P.; Ramirez, C. E.; Bedard, M.; Fink, C.; Ko, J.; Peralta, E. J. D.; Watts, G.; Gomez-Rivas, E.; Davis, V.; Barilla, R. M.; Wang, J. N.; Cunin, P.; Bates, S.; Morrison-Smith, C.; Nicholson, B.; Wong, E.; El-Mufti, L.; Kann, M.; Bolling, A.; Fortin, B.; Ventresca, H.; Zhou, W.; Pardo, S.; Kwock, M.; Hazra, A.; Cheng, L.; Ahmad, Q. R.; Toombs, J. A.; Larson, R.; Pleskow, H.; Luo, N. M.; Samaha, C.; Pandya, U. M.; De Silva, P.; Zhou, S.; Ganhadeiro, Z.; Yohannes, S.; Gay, R.; Slavik, J.; Mukerji, S. S.; Jarolim, P.; Walt, D. R.; Carlyle, B. C.; Ritterhouse, L. L.; Suliman, S., Evaluation of serological lateral flow assays for severe acute respiratory syndrome coronavirus-2 (vol 21, 580, 2021). Bmc Infect Dis 2021, 21 (1). [CrossRef]
- Chen, A. L.; Yang, S. M., Replacing antibodies with aptamers in lateral flow immunoassay. Biosens Bioelectron 2015, 71, 230-242. [CrossRef]
- Baker, M., Blame It on the Antibodies. Nature 2015, 521 (7552), 274-276. [CrossRef]
- Liu, Y. Y.; Liu, D.; Cui, S. S.; Li, C.; Yun, Z. G.; Zhang, J.; Sun, F. X., Design of a Signal-Amplified Aptamer-Based Lateral Flow Test Strip for the Rapid Detection of Ochratoxin A in Red Wine. Foods 2022, 11 (11). [CrossRef]
- Wu, S. J.; Liu, L. H.; Duan, N.; Li, Q.; Zhou, Y.; Wang, Z. P., Aptamer-Based Lateral Flow Test Strip for Rapid Detection of Zearalenone in Corn Samples. J Agr Food Chem 2018, 66 (8), 1949-1954. [CrossRef]
- Yu, Q.; Zhao, Q.; Wang, S.; Zhao, S.; Zhang, S.; Yin, Y. G.; Dong, Y. Y., Development of a lateral flow aptamer assay strip for facile identification of theranostic exosomes isolated from human lung carcinoma cells. Anal Biochem 2020, 594. [CrossRef]
- Martínez-Roque, M. A.; Franco-Urquijo, P. A.; García-Velásquez, V. M.; Choukeife, M.; Mayer, G.; Molina-Ramírez, S. R.; Figueroa-Miranda, G.; Mayer, D.; Alvarez-Salas, L. M., DNA aptamer selection for SARS-CoV-2 spike glycoprotein detection. Anal Biochem 2022, 645. [CrossRef]
- Zhang, Z. J.; Li, J. X.; Gu, J.; Amini, R.; Stacey, H. D.; Ang, J. C.; White, D.; Filipe, C. D. M.; Mossman, K.; Miller, M. S.; Salena, B. J.; Yamamura, D.; Sen, P.; Soleymani, L.; Brennan, J. D.; Li, Y. F., A Universal DNA Aptamer that Recognizes Spike Proteins of Diverse SARS-CoV-2 Variants of Concern. Chem-Eur J 2022, 28 (15). [CrossRef]
- Zhang, X.; Servos, M. R.; Liu, J. W., Surface Science of DNA Adsorption onto Citrate-Capped Gold Nanoparticles. Langmuir 2012, 28 (8), 3896-3902. [CrossRef]
- Rho, S.; Kim, S. J.; Lee, S. C.; Chang, J. H.; Kang, H. G.; Choi, J. Colorimetric Detection of SsDNA in a Solution. Curr. Appl. Phys. 2009, 9, 534−537. [CrossRef]
- Hu M, Yuan C, Tian T, et al. Single-step, salt-aging-free, and thiol-free freezing construction of AuNP-based bioprobes for advancing CRISPR-based diagnostics. Journal of the American Chemical Society, 2020, 142(16): 7506-7513. [CrossRef]
- Ku, T. H.; Zhang, T. T.; Luo, H.; Yen, T. M.; Chen, P. W.; Han, Y. Y.; Lo, Y. H., Nucleic Acid Aptamers: An Emerging Tool for Biotechnology and Biomedical Sensing. Sensors-Basel 2015, 15 (7), 16281-16313. [CrossRef]
- Zhao, Z. L.; Wang, H.; Zhai, W. L.; Feng, X. Y.; Fan, X.; Chen, A. L.; Wang, M., A Lateral Flow Strip Based on a Truncated Aptamer-Complementary Strand for Detection of Type-B Aflatoxins in Nuts and Dried Figs. Toxins 2020, 12 (2). [CrossRef]
- Gao, Y.; Deng, X.; Wen, W.; Zhang, X.; Wang, S., Ultrasensitive paper based nucleic acid detection realized by three-dimensional DNA-AuNPs network amplification. Biosensors and Bioelectronics 2017, 92, 529-535. [CrossRef]
- Cai, S. D.; Yan, J. H.; Xiong, H. J.; Liu, Y. F.; Peng, D. M.; Liu, Z. B., Investigations on the interface of nucleic acid aptamers and binding targets. Analyst 2018, 143 (22), 5317-5338. [CrossRef]
- Kim, H. S.; Seo, Y. S.; Kim, K.; Han, J. W.; Park, Y.; Cho, S., Concentration Effect of Reducing Agents on Green Synthesis of Gold Nanoparticles: Size, Morphology, and Growth Mechanism. Nanoscale Res Lett 2016, 11. [CrossRef]
- The UNA Fold Web Server. http://www.unafold.org/DNA_form.php# (accessed January 2024).
- Li, X. 3dRNA/DNA Web Server. http://biophy.hust.edu.cn/new/3dRNA(accessed January 2024).
- PubChem. https://pubchem.ncbi.nlm.nih.gov/ (accessed January 2024).
- Li, X.; Qian, Z.; Chang, R.; Peng, C.; Xie, Z.; Wang, Z., Non-thiolated nucleic acid functionalized gold nanoparticle–based aptamer lateral flow assay for rapid detection of kanamycin. Microchimica Acta 2022, 189 (7). [CrossRef]

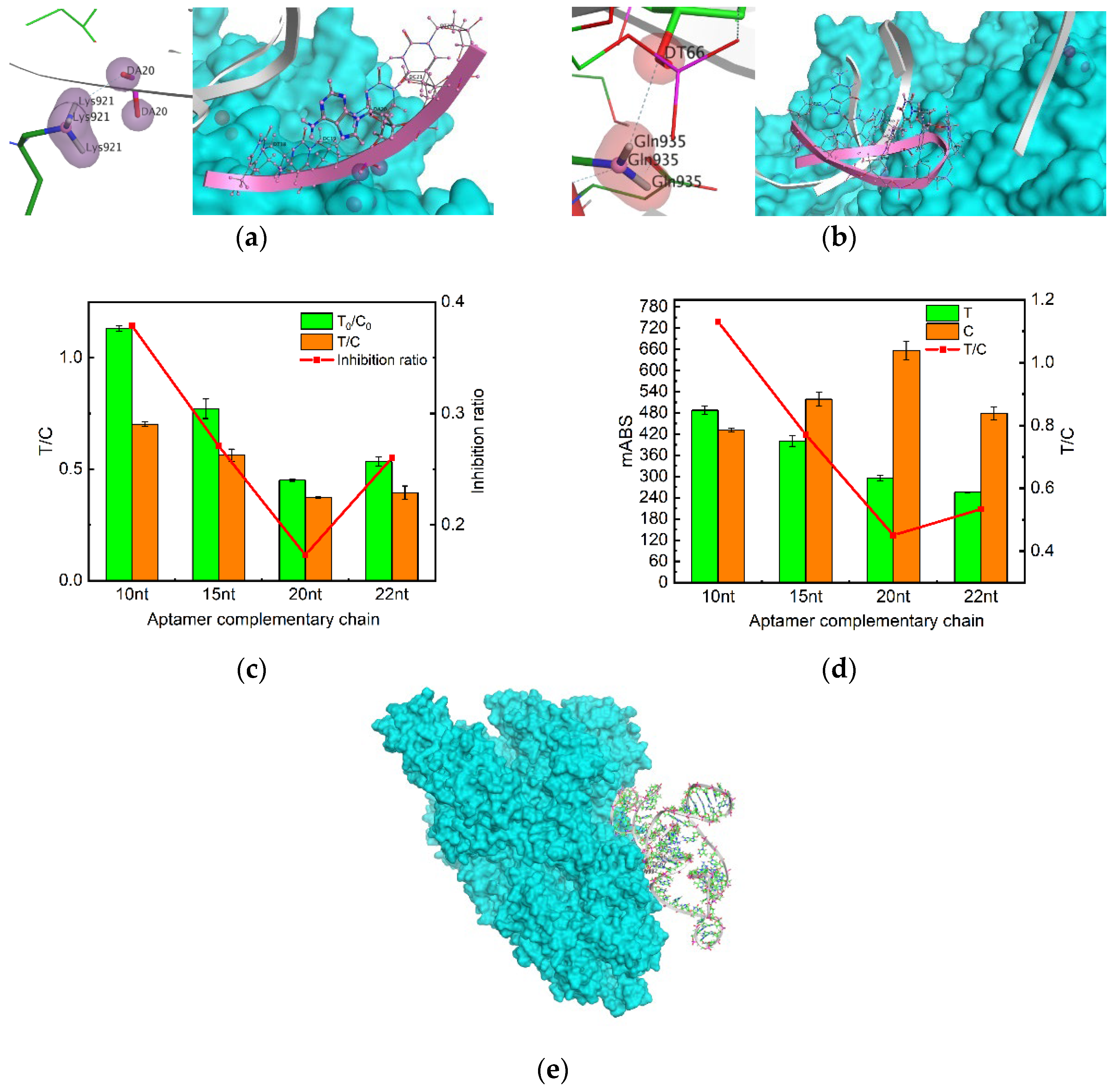

| Number | Energy | Aptamer binding site | Protein binding site | Aptamer sequence |
|---|---|---|---|---|
| 1 | -15.89 | DA20 | Lys921 | DT18-DT22 |
| 2 | -0.9 | DT66 | Gln935 | DC65-DG71 |
| 3 | -0.72 | DT80 | Lys285 | DT77-DT84 |
| 4 | -0.64 | DT82 | Lys278 | DT77-DT84 |
| Name | Sequences (5′-3′) |
|---|---|
| MSA-10 | Biotin-ACGCCAAGGA |
| MSA-15 | Biotin-ACGCCAAGGAGATGC |
| MSA-20 | Biotin-ACGCCAAGGAGATGCTTCGC |
| MSA-22 | Biotin-CGCCAGGCCCGGAGCCAAACCC |
| Control-line DNA | Biotin-AAAAAAAAAA |
| Sample | Concentration of SP (ng/mL) | Detection result | Test strip concentration (ng/mL) | Recovery rate (%) | RSD (%) |
|---|---|---|---|---|---|
| Cold chain food packaging bags—tap water rinsing (1 ml) |
0 | Negative | Undetected | Undetected | Undetected |
| 200 | Positive | 166.7 | 83.3 | 2.6 | |
| 400 | Positive | 435.2 | 108.8 | 2.3 | |
| 600 | Positive | 571.7 | 95.2 | 5.8 | |
| 800 | Positive | 752.4 | 94.1 | 6.2 | |
| 900 | Positive | 860.2 | 95.6 | 4.6 | |
| 1000 | Positive | 968.1 | 96.8 | 3.0 |
| Name | Speed (s/4cm) | Diameter (um) |
|---|---|---|
| BSK95 | 100±20 | 12-15 |
| BSK110 | 120±30 | 8-12 |
| BSK140 | 140±30 | 5-8 |
| BSK160 | 160±30 | 4-6 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




