Submitted:
18 February 2024
Posted:
19 February 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Identification and analysis of TDC genes in cucumber
2.2. Analysis of cis-elements in the promoter regions
2.3. Plant materials and treatments
2.4. RNA extraction and qRT-PCR analysis
2.5. Subcellular localization analysis
2.6. Transient overexpression of CsTDC genes in tobacco leaves
2.7. Determination of TDC activity and melatonin content
2.8. Transient overexpression of CsTDC genes and abiotic stress treatments
2.9. Statistical analyses
3. Results
3.1. Identification and characterization of cucumber TDC genes
3.2. Identification of cis-elements in CsTDC promoters
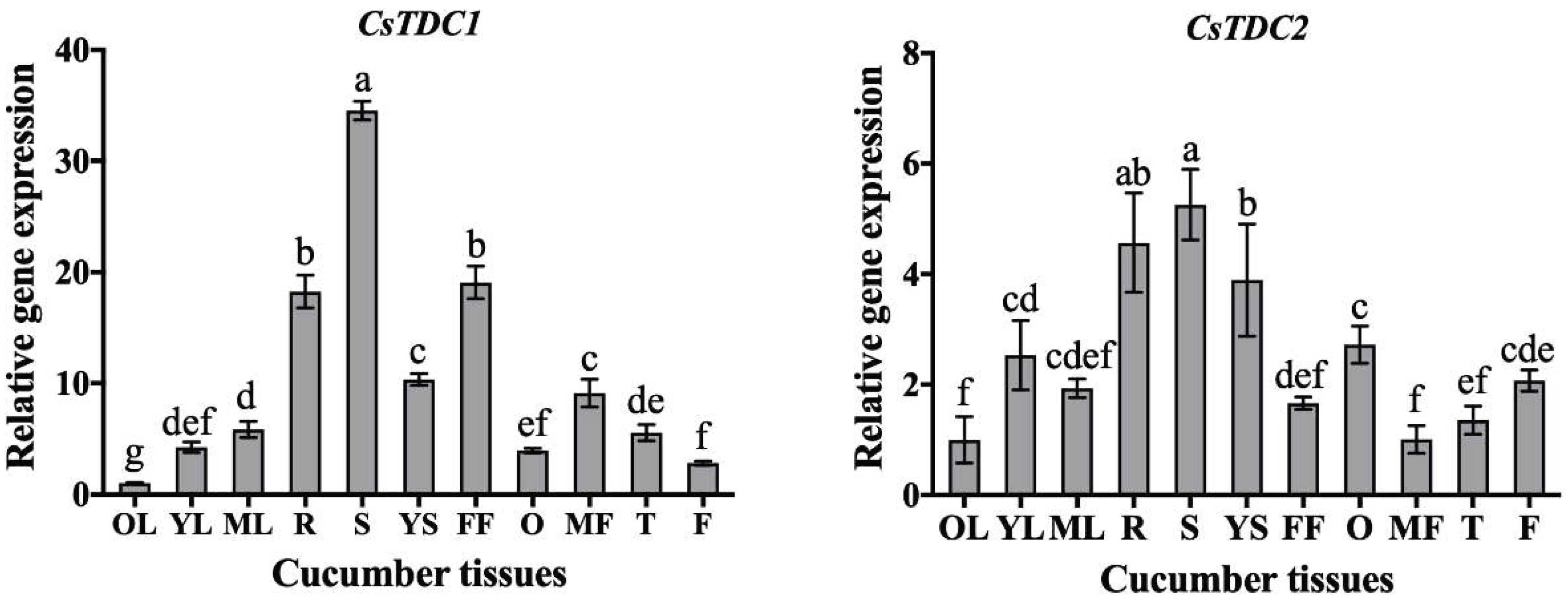
3.3. CsTDC expression profiles in different tissues
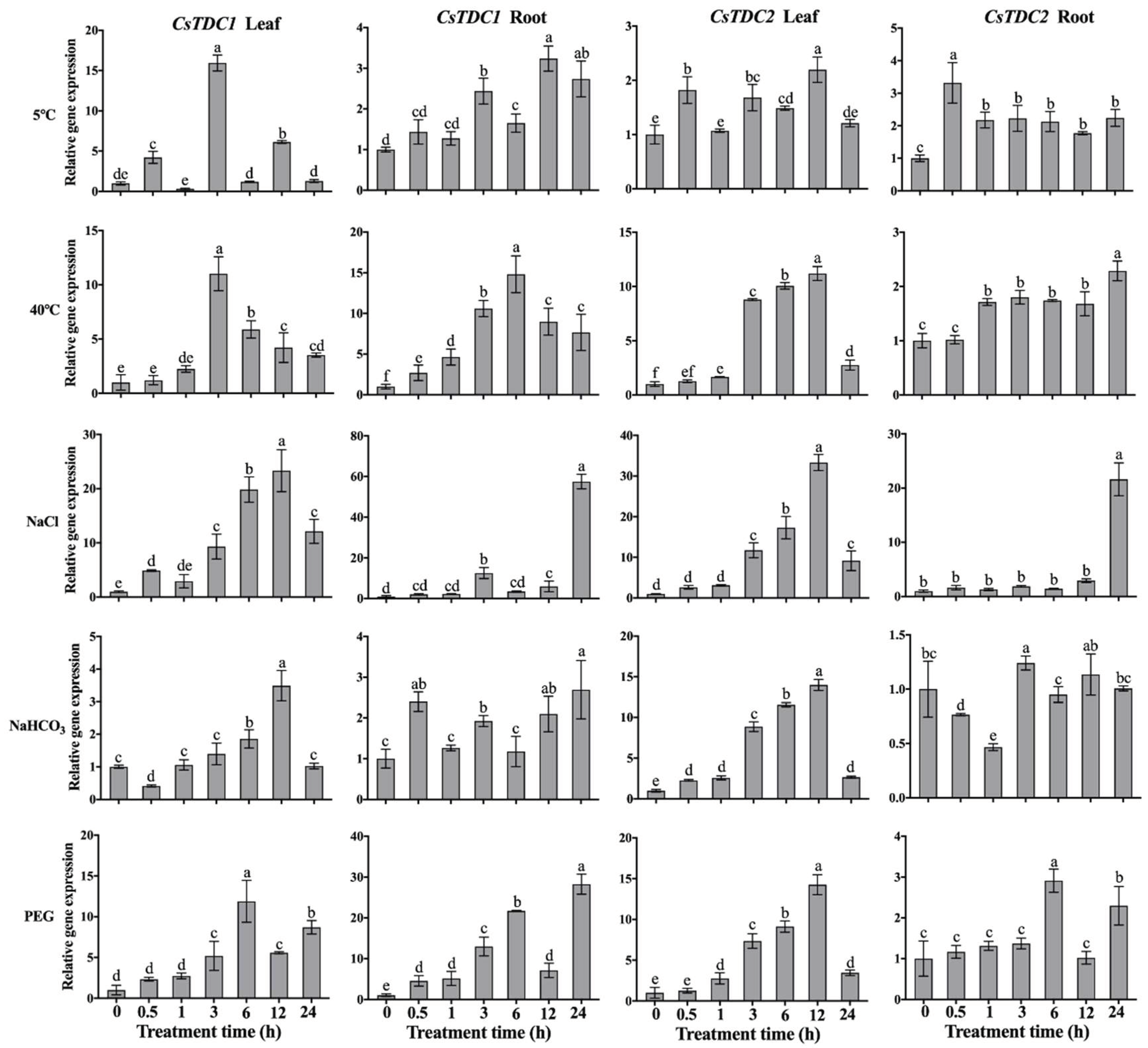
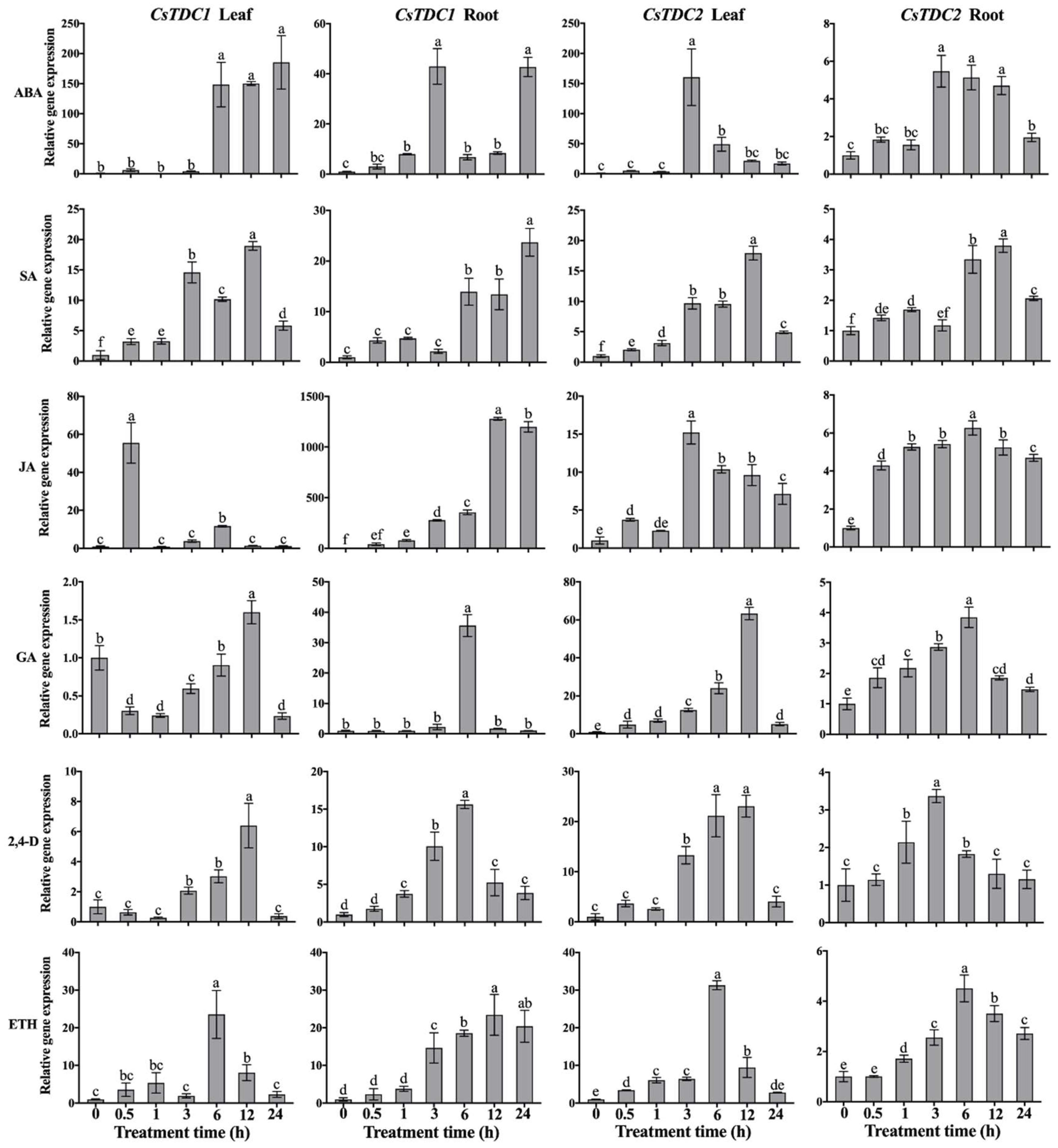
3.4. CsTDC expression profiles in response to various abiotic stresses and exogenous phytohormones
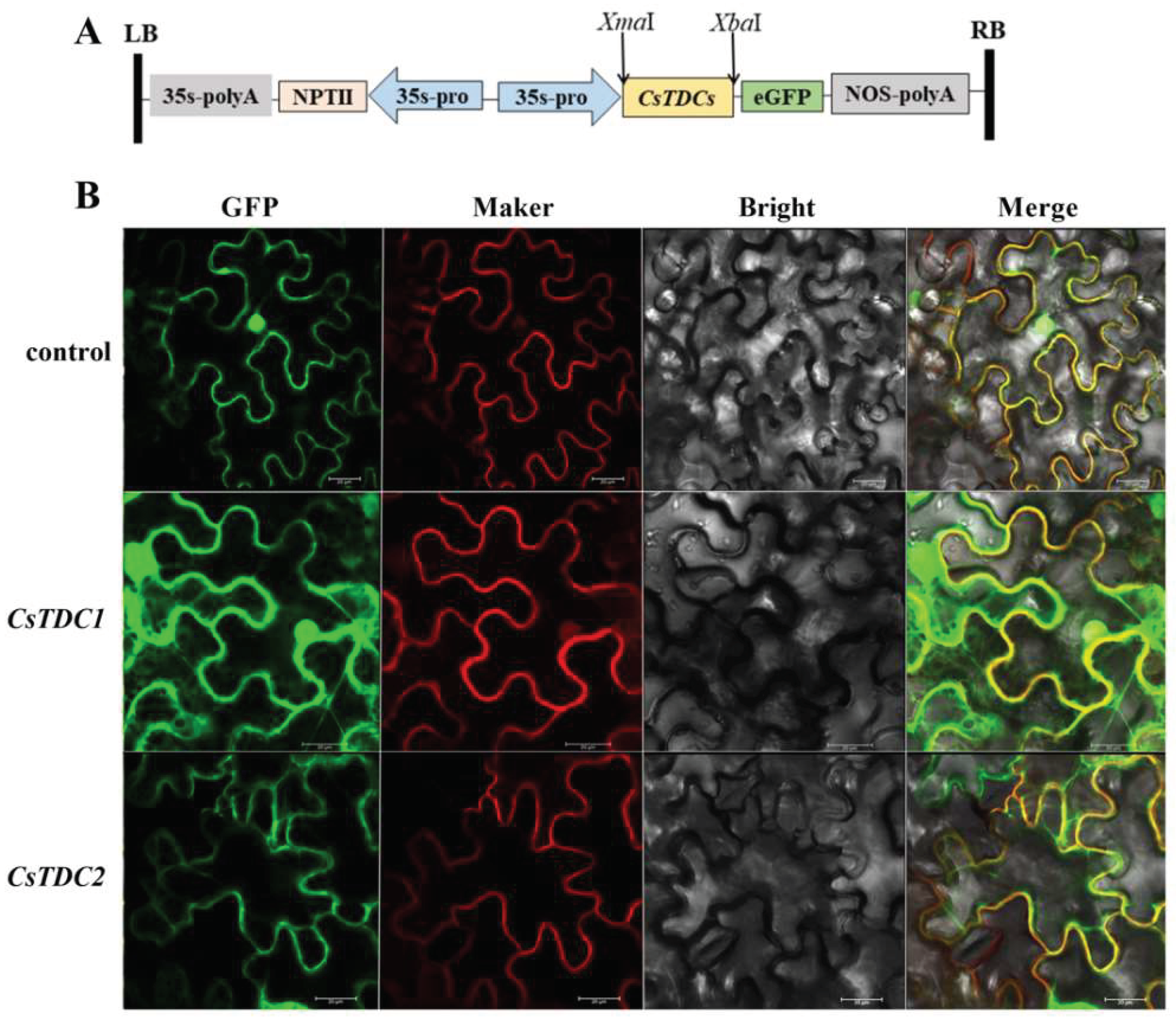
3.5. Subcellular localization of CsTDC1 and CsTDC2
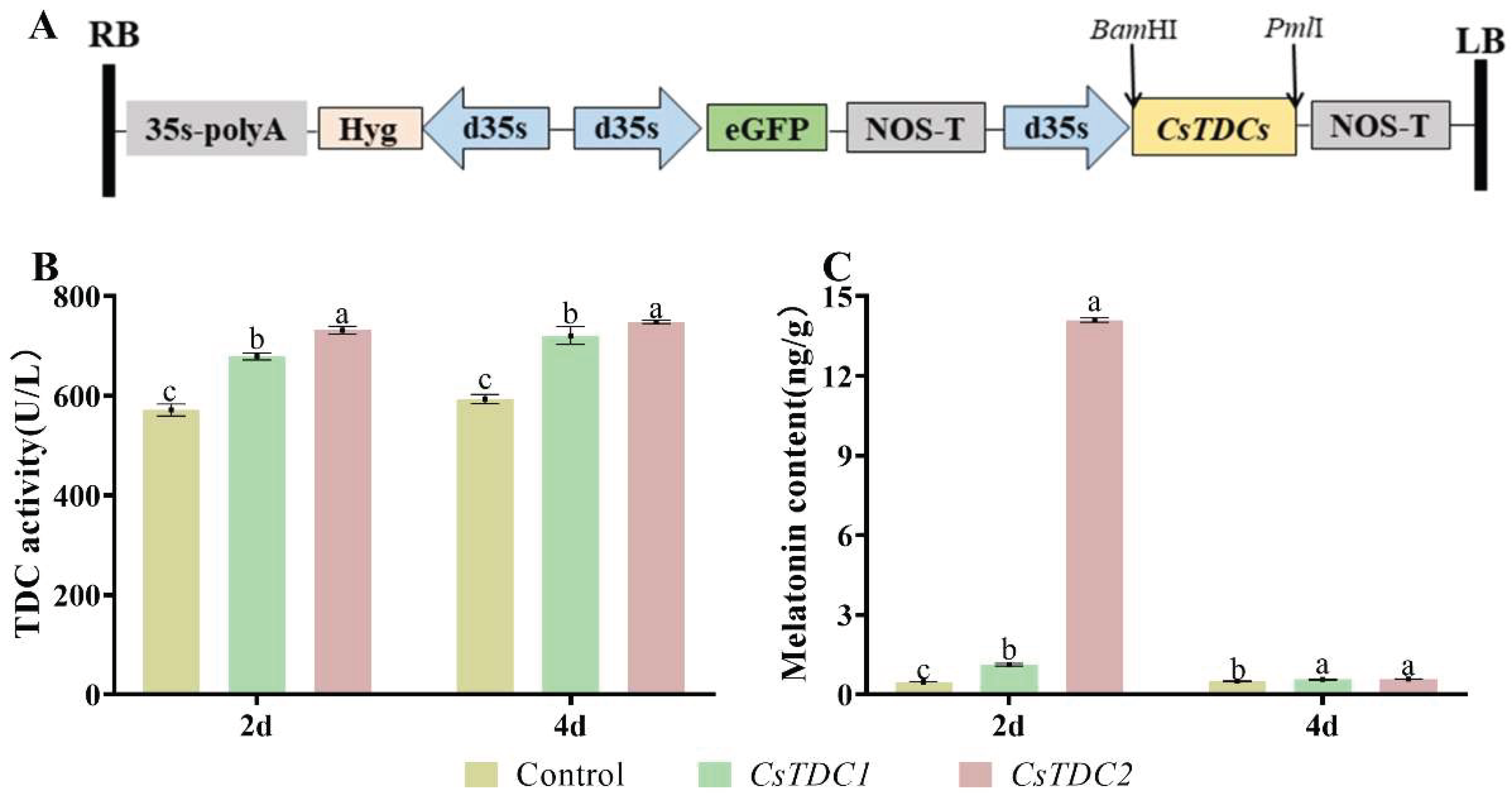
3.6. Transient overexpression of CsTDC genes in tobacco leaves promoted melatonin biosynthesis
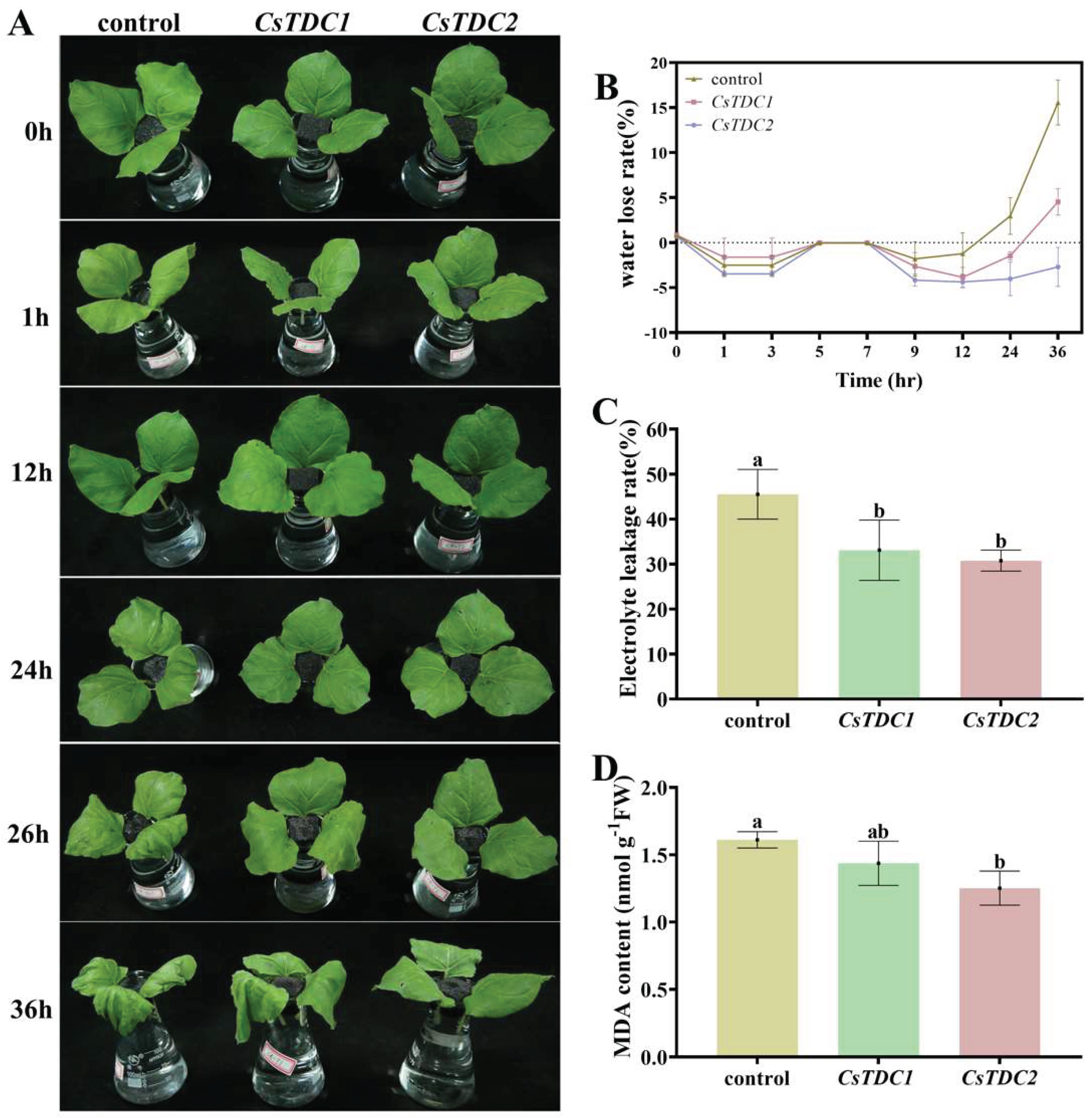
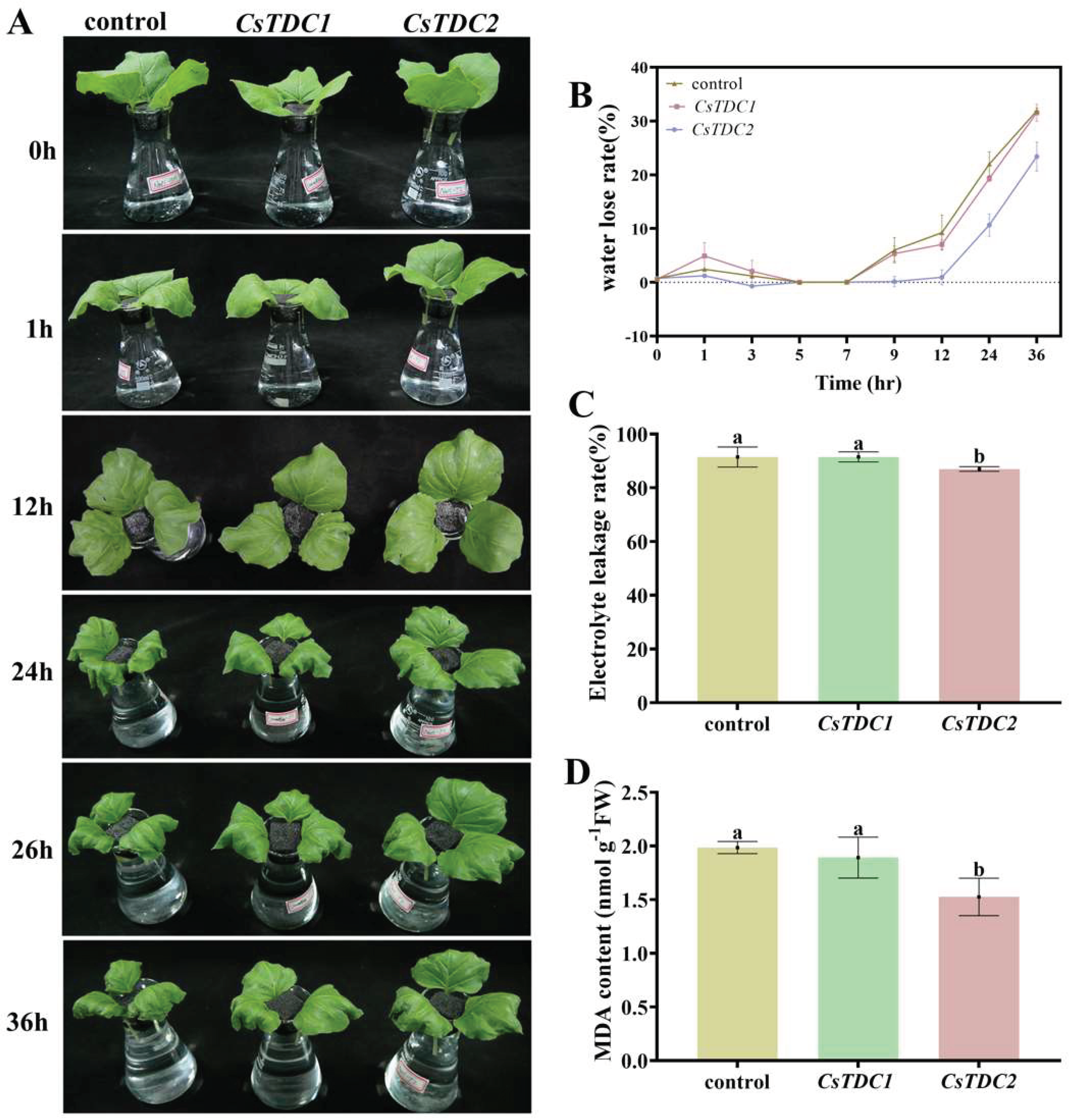
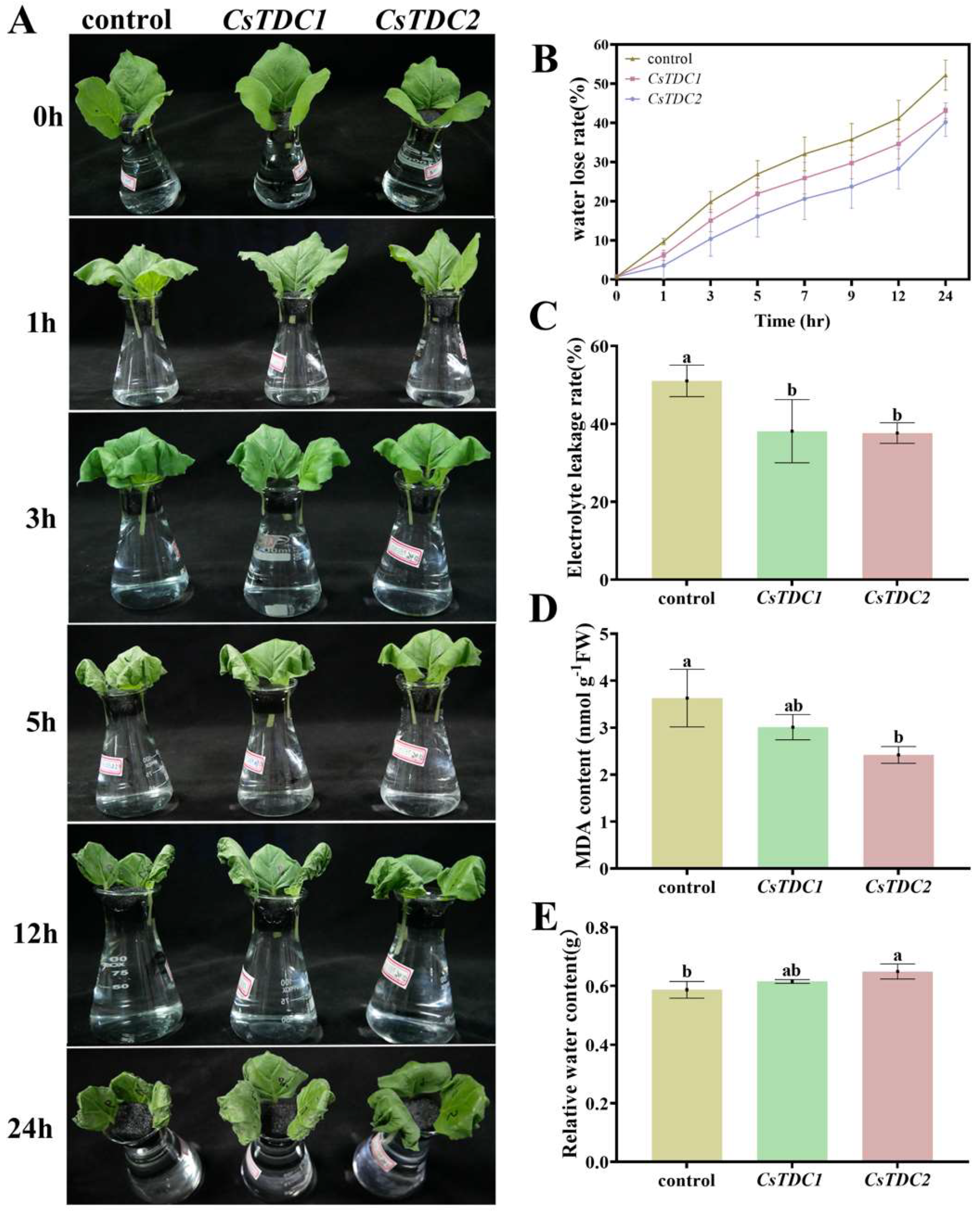
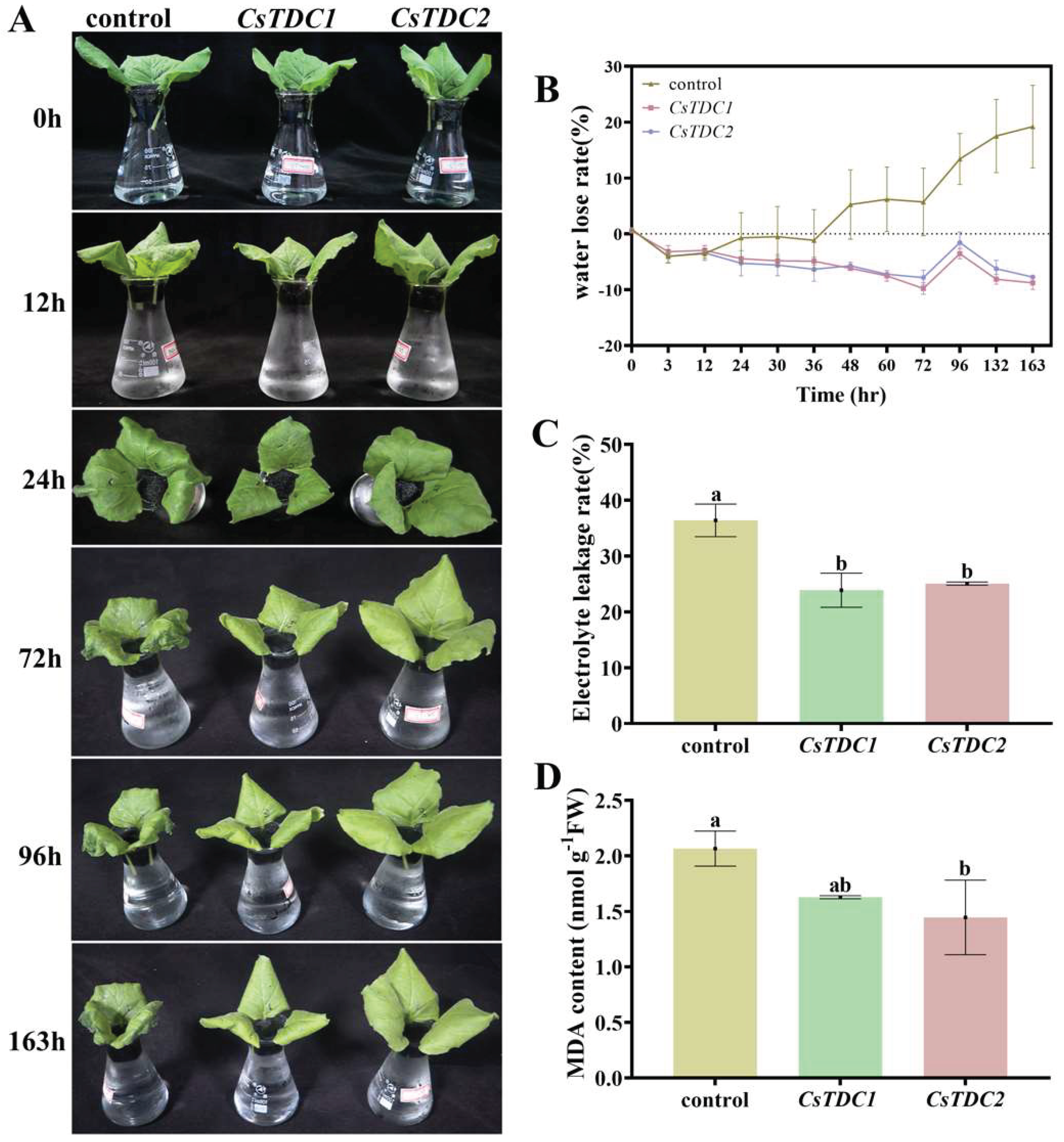
3.7. Transient overexpression of CsTDC genes enhanced abiotic stress tolerance in tobacco
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Dubbels, R.; Reiter, R.; Klenke, E.; Goebel, A.; Schnakenberg, E.; Ehlers, C.; Schiwara, H.; Schloot, W. Melatonin in edible plants identified by radioimmunoassay and by high performance liquid chromatography-mass spectrometry. J Pineal Res 1995, 18, 28-31. [CrossRef]
- Hattori, A.; Migitaka, H.; Iigo, M.; Itoh, M.; Yamamoto, K.; Ohtani-Kaneko, R.; Hara, M.; Suzuki, T.; Reiter, R.J. Identification of melatonin in plants and its effects on plasma melatonin levels and binding to melatonin receptors in vertebrates. Biochem Mol Biol Int 1995, 35, 627-634.
- Wang, K.; Xing, Q.; Ahammed, G.J.; Zhou, J. Functions and prospects of melatonin in plant growth, yield, and quality. J Exp Bot 2022, 73, 5928-5946. [CrossRef]
- Pan, Y.; Xu, X.; Li, L.; Sun, Q.; Wang, Q.; Huang, H.; Tong, Z.; Zhang, J. Melatonin-mediated development and abiotic stress tolerance in plants. Front Plant Sci 2023, 14, 1100827. [CrossRef]
- Huang, X.; Tanveer, M.; Min, Y.; Shabala, S. Melatonin as a regulator of plant ionic homeostasis: implications for abiotic stress tolerance. J Exp Bot 2022, 73, 5886-5902. [CrossRef]
- Liu, N.; Gong, B.; Jin, Z.; Wang, X.; Wei, M.; Yang, F.; Li, Y.; Shi, Q. Sodic alkaline stress mitigation by exogenous melatonin in tomato needs nitric oxide as a downstream signal. J Plant Physiol 2015, 186, 68-77. [CrossRef]
- Zhao, D.; Zhang, X.; Wang, R.; Liu, D.; Sun, J.; Tao, J. Herbaceous peony tryptophan decarboxylase confers drought and salt stresses tolerance. Environ Exp Bot 2019, 162, 345-356. [CrossRef]
- Sun, C.; Liu, L.; Wang, L.; Li, B.; Jin, C.; Lin, X. Melatonin: A master regulator of plant development and stress responses. J Integr Plant Biol 2021, 63, 126-145. [CrossRef]
- Back, K.; Tan, D.X.; Reiter, R.J. Melatonin biosynthesis in plants: multiple pathways catalyze tryptophan to melatonin in the cytoplasm or chloroplasts. J Pineal Res 2016, 61, 426-437. [CrossRef]
- Han, S.-W.; Shin, J.-S. Aromatic L-amino acid decarboxylases: mechanistic features and microbial applications. Appl Microbiol Biot 2022, 106, 4445-4458. [CrossRef]
- Facchini, P.J.; Huber-Allanach, K.L.; Tari, L.W. Plant aromatic L-amino acid decarboxylases: evolution, biochemistry, regulation, and metabolic engineering applications. Phytochem 2000, 54, 121-138. [CrossRef]
- Torrens-Spence, M.P.; Chiang, Y.-C.; Smith, T.; Vicent, M.A.; Wang, Y.; Weng, J.-K. Structural basis for divergent and convergent evolution of catalytic machineries in plant aromatic amino acid decarboxylase proteins. PNAS 2020, 117, 10806-10817. [CrossRef]
- Back, K. Melatonin metabolism, signaling and possible roles in plants. Plant J 2021, 105, 376-391. [CrossRef]
- De Luca, V.; Marineau, C.; Brisson, N. Molecular cloning and analysis of cDNA encoding a plant tryptophan decarboxylase: comparison with animal dopa decarboxylases. PNAS 1989, 86, 2582-2586. [CrossRef]
- Lee, K.; Back, K. Melatonin-deficient rice plants show a common semidwarf phenotype either dependent or independent of brassinosteroid biosynthesis. J Pineal Res 2019, 66, e12537. [CrossRef]
- You, D.; Feng, Y.; Wang, C.; Sun, C.; Wang, Y.; Zhao, D.; Kai, G. Cloning, characterization, and enzymatic identification of a new tryptophan decarboxylase from Ophiorrhiza pumila. Biotechnol Appl Bioc 2021, 68, 381-389.
- De Masi, L.; Castaldo, D.; Pignone, D.; Servillo, L.; Facchiano, A. Experimental evidence and in silico identification of tryptophan decarboxylase in Citrus genus. Molecules 2017, 22, 272. [CrossRef]
- Li, L.; Zheng, M.; Long, H.; Deng, G.; Ishihara, A.; Liu, F.; Liang, J.; Pan, Z.; Yu, M. Molecular cloning and characterization of two genes encoding tryptophan decarboxylase from Aegilops variabilis with resistance to the cereal cyst nematode (Heterodera avenae) and root-knot nematode (Meloidogyne naasi). Plant Mol Biol Rep 2016, 34, 273-282. [CrossRef]
- Qiao, C.; Chen, F.; Liu, Z.; Huang, T.; Li, W.; Zhang, G.; Luo, Y. Functional characterization of a catalytically promiscuous tryptophan decarboxylase from camptothecin-producing Camptotheca acuminata. Front Plant Sci 2022, 13, 987348. [CrossRef]
- Tsunoda, Y.; Hano, S.; Imoto, N.; Shibuya, T.; Ikeda, H.; Amagaya, K.; Kato, K.; Shirakawa, H.; Aso, H.; Kanayama, Y. Physiological roles of tryptophan decarboxylase revealed by overexpression of SlTDC1 in tomato. Sci Hortic-amsterdam 2021, 275, 109672. [CrossRef]
- Byeon, Y.; Park, S.; Lee, H.Y.; Kim, Y.S.; Back, K. Elevated production of melatonin in transgenic rice seeds expressing rice tryptophan decarboxylase. J Pineal Res 2014, 56, 275-282. [CrossRef]
- Kanjanaphachoat, P.; Wei, B.-Y.; Lo, S.-F.; Wang, I.-W.; Wang, C.-S.; Yu, S.-M.; Yen, M.-L.; Chiu, S.-H.; Lai, C.-C.; Chen, L.-J. Serotonin accumulation in transgenic rice by over-expressing tryptophan decarboxlyase results in a dark brown phenotype and stunted growth. Plant Mol Biol 2012, 78, 525-543. [CrossRef]
- Li, S.; Miao, L.; Huang, B.; Gao, L.; He, C.; Yan, Y.; Wang, J.; Yu, X.; Li, Y. Genome-wide identification and characterization of cucumber BPC transcription factors and their responses to abiotic stresses and exogenous phytohormones. Int J Mol Sci 2019, 20, 5048. [CrossRef]
- Letunic, I.; Bork, P. 20 years of the SMART protein domain annotation resource. NAR 2018, 46, D493-D496. [CrossRef]
- Bateman, A.; Coin, L.; Durbin, R.; Finn, R.D.; Hollich, V.; Griffiths-Jones, S.; Khanna, A.; Marshall, M.; Moxon, S.; Sonnhammer, E.L. The Pfam protein families database. NAR 2004, 32, D138-D141.
- Yu, C.S.; Chen, Y.C.; Lu, C.H.; Hwang, J.K. Prediction of protein subcellular localization. Proteins 2006, 64, 643-651.
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. NAR 2002, 30, 325-327. [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 2001, 25, 402-408.
- Hong, Y.; Pan, X.; Welti, R.; Wang, X. Phospholipase Dα3 is involved in the hyperosmotic response in Arabidopsis. Plant Cell 2008, 20, 803-816. [CrossRef]
- Li, S.; Sun, M.; Miao, L.; Di, Q.; Lv, L.; Yu, X.; Yan, Y.; He, C.; Wang, J.; Shi, A. Multifaceted regulatory functions of CsBPC2 in cucumber under salt stress conditions. Hortic Res 2023, 10, uhad051. [CrossRef]
- Wang, Q.; An, B.; Shi, H.; Luo, H.; He, C. High concentration of melatonin regulates leaf development by suppressing cell proliferation and endoreduplication in Arabidopsis. Int J Mol Sci 2017, 18, 991. [CrossRef]
- Wang, H.; Song, C.; Fang, S.; Wang, Z.; Song, S.; Jiao, J.; Wang, M.; Zheng, X.; Bai, T. Genome-wide identification and expression analysis of the ASMT gene family reveals their role in abiotic stress tolerance in apple. Sci Hortic-amsterdam 2022, 293, 110683. [CrossRef]
- Chang, J.; Guo, Y.; Yan, J.; Zhang, Z.; Yuan, L.; Wei, C.; Zhang, Y.; Ma, J.; Yang, J.; Zhang, X. The role of watermelon caffeic acid O-methyltransferase (ClCOMT1) in melatonin biosynthesis and abiotic stress tolerance. Hortic Res 2021, 8.
- Commisso, M.; Negri, S.; Gecchele, E.; Fazion, E.; Pontoriero, C.; Avesani, L.; Guzzo, F. Indolamine accumulation and TDC/T5H expression profiles reveal the complex and dynamic regulation of serotonin biosynthesis in tomato (Solanum lycopersicum L.). Front Plant Sci 2022, 13, 975434. [CrossRef]
- Qi, C.; Zhang, H.; Liu, Y.; Wang, X.; Dong, D.; Yuan, X.; Li, X.; Zhang, X.; Li, X.; Zhang, N. CsSNAT positively regulates salt tolerance and growth of cucumber by promoting melatonin biosynthesis. Environ Exp Bot 2020, 175, 104036. [CrossRef]
- Zhao, D.; Luan, Y.; Shi, W.; Tang, Y.; Huang, X.; Tao, J. Melatonin enhances stem strength by increasing lignin content and secondary cell wall thickness in herbaceous peony. J Exp Bot 2022, 73, 5974-5991. [CrossRef]
- Lv, Y.; Pan, J.; Wang, H.; Reiter, R.J.; Li, X.; Mou, Z.; Zhang, J.; Yao, Z.; Zhao, D.; Yu, D. Melatonin inhibits seed germination by crosstalk with abscisic acid, gibberellin, and auxin in Arabidopsis. J Pineal Res 2021, 70, e12736. [CrossRef]
- Zhang, N.; Zhao, B.; Zhang, H.J.; Weeda, S.; Yang, C.; Yang, Z.C.; Ren, S.; Guo, Y.D. Melatonin promotes water-stress tolerance, lateral root formation, and seed germination in cucumber (Cucumis sativus L.). J Pineal Res 2013, 54, 15-23.
- Yang, W.-Y.; Zheng, Y.; Bahn, S.C.; Pan, X.-Q.; Li, M.-Y.; Vu, H.S.; Roth, M.R.; Scheu, B.; Welti, R.; Hong, Y.-Y. The patatin-containing phospholipase A pPLAIIα modulates oxylipin formation and water loss in Arabidopsis thaliana. Mol plant 2012, 5, 452-460. [CrossRef]
- Yang, N.; Sun, K.; Wang, X.; Wang, K.; Kong, X.; Gao, J.; Wen, D. Melatonin participates in selenium-enhanced cold tolerance of cucumber seedlings. Front Plant Sci 2021, 12, 786043. [CrossRef]
- Feng, Y.; Fu, X.; Han, L.; Xu, C.; Liu, C.; Bi, H.; Ai, X. Nitric oxide functions as a downstream signal for melatonin-induced cold tolerance in cucumber seedlings. Front Plant Sci 2021, 12, 686545. [CrossRef]
- Wang, D.; Chen, Q.; Chen, W.; Guo, Q.; Xia, Y.; Wang, S.; Jing, D.; Liang, G. Physiological and transcription analyses reveal the regulatory mechanism of melatonin in inducing drought resistance in loquat (Eriobotrya japonica Lindl.) seedlings. Environ Exp Bot 2021, 181, 104291. [CrossRef]
- Xu, L.; Xiang, G.; Sun, Q.; Ni, Y.; Jin, Z.; Gao, S.; Yao, Y. Melatonin enhances salt tolerance by promoting MYB108A-mediated ethylene biosynthesis in grapevines. Hortic Res 2019, 6. [CrossRef]
- Yan, Y.; Jing, X.; Tang, H.; Li, X.; Gong, B.; Shi, Q. Using transcriptome to discover a novel melatonin-induced sodic alkaline stress resistant pathway in Solanum lycopersicum L. Plant Cell Physiol 2019, 60, 2051-2064. [CrossRef]
- Fan, H.; Wang, S.; Wang, H.; Sun, M.; Wu, S.; Bao, W. Melatonin ameliorates the toxicity induced by deoxynivalenol in murine ovary granulosa cells by antioxidative and anti-inflammatory effects. Antioxidants 2021, 10, 1045. [CrossRef]
| Gene ID | Length (aa) | Molecular Weight (KD) | Chromosome | Location | pI | Strand Direction | Subcellular Location |
|---|---|---|---|---|---|---|---|
| CsaV3_1G036910 | 499 | 55.7 | 1 | 22824145-22827250 | 6.34 | – | cytoplasm, plasmamembrane |
| CsaV3_3G028450 | 486 | 54.6 | 3 | 24862649-24886753 | 5.79 | – | cytoplasm, plasmamembrane |
| Cis-element | Function | CsTDC1 CsTDC2 |
|---|---|---|
| Stress-related | ||
| ARE | cis-element essential for the anaerobic induction | 2 5 |
| WRE3 | wound response elements | 1 1 |
| LTR | cis-element involved in low-temperature responsiveness | 0 1 |
| STRE | stress response elements | 4 2 |
| MYB | stress response elements | 1 3 |
| MYC | drought and cold responsive elements | 1 3 |
| Hormone-related | ||
| ERE | ethylene-responsive element | 1 5 |
| ABRE | cis-element involved in the abscisic acid responsiveness | 1 1 |
| CGTCA-motif | cis-element involved in the MeJA-responsiveness | 0 2 |
| TCA-element | cis-element involved in salicylic acid responsiveness | 1 0 |
| TGA-element | auxin-responsive element | 0 1 |
| TGACG-motif | cis-element involved in the MeJA-responsiveness | 0 2 |
| Development-related | ||
| AACA_motif | involved in endosperm-specific negative expression | 1 0 |
| circadian | cis-acting regulatory element involved in circadian control | 1 0 |
| as-1 | cis-element involved in the root-specific expression | 0 2 |
| CAT-box | cis-acting regulatory element related to meristem expression | 0 2 |
| Light-related | ||
| AE-box | part of a module for light response | 0 1 |
| Box 4 | a conserved DNA module involved in light responsiveness | 4 7 |
| GATA-motif | part of a light responsive element | 1 0 |
| GT1-motif | light responsive element | 0 3 |
| TCCC-motif | part of a light responsive element | 1 0 |
| TCT-motif | part of a light responsive element | 1 0 |
| G-Box | cis-element involved in light responsiveness | 1 2 |
| G-box | cis-element involved in light responsiveness | 0 1 |
| GA-motif | part of a light responsive element | 0 1 |
| MRE | MYB binding site involved in light responsiveness | 4 1 |
| AAAC-motif | light responsive element | 1 0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




