Submitted:
29 January 2024
Posted:
30 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
- LBP is used to extract more discriminative features to classify retinal diseases using a multi-scale discriminative feature extraction approach.
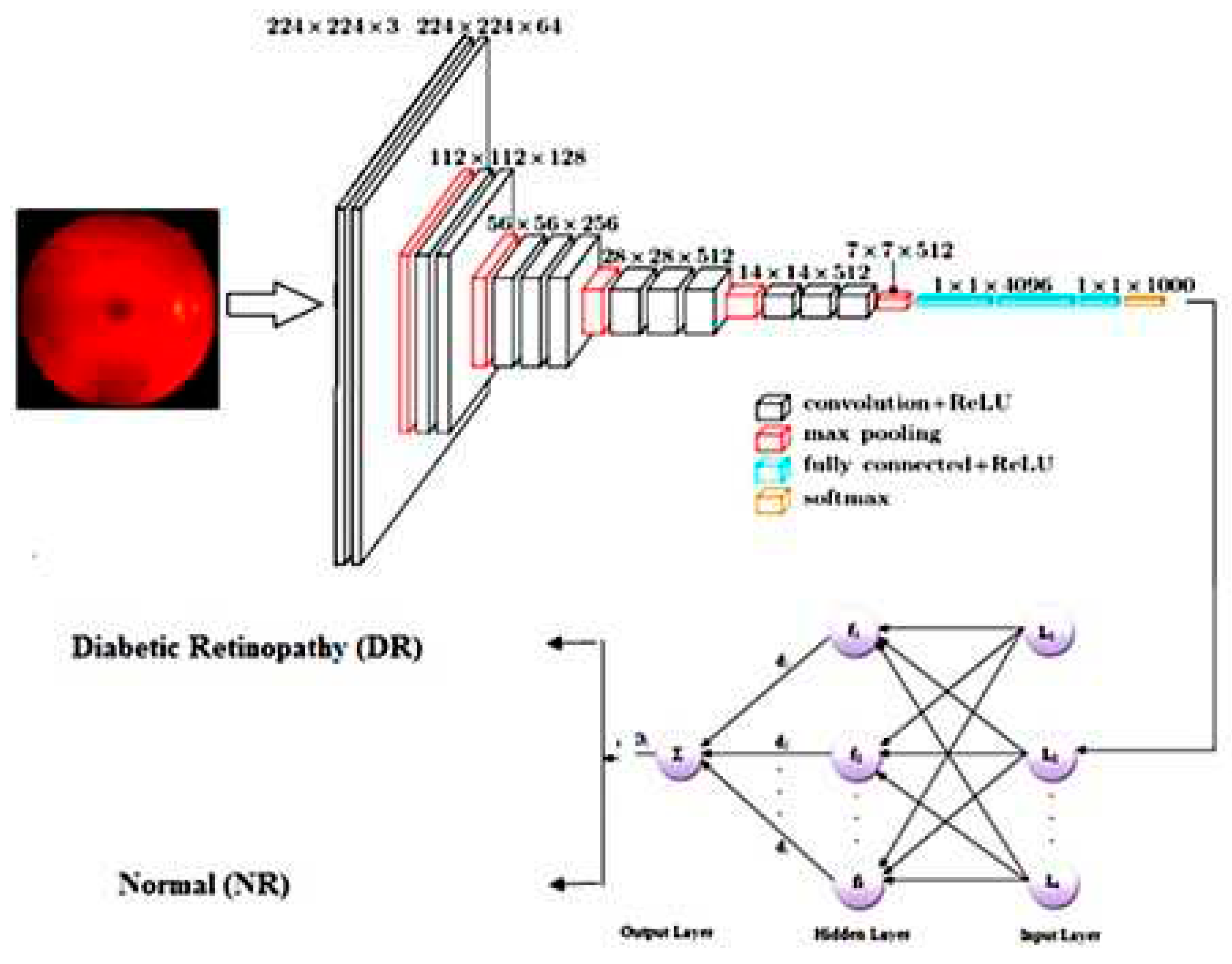
- Deep learning techniques were used to develop a new classifier using CNNs and RBFs
- In comparison with the state-of-the-art methods reported in the literature, we achieved better results in retinal disease classification.
2. Related Works
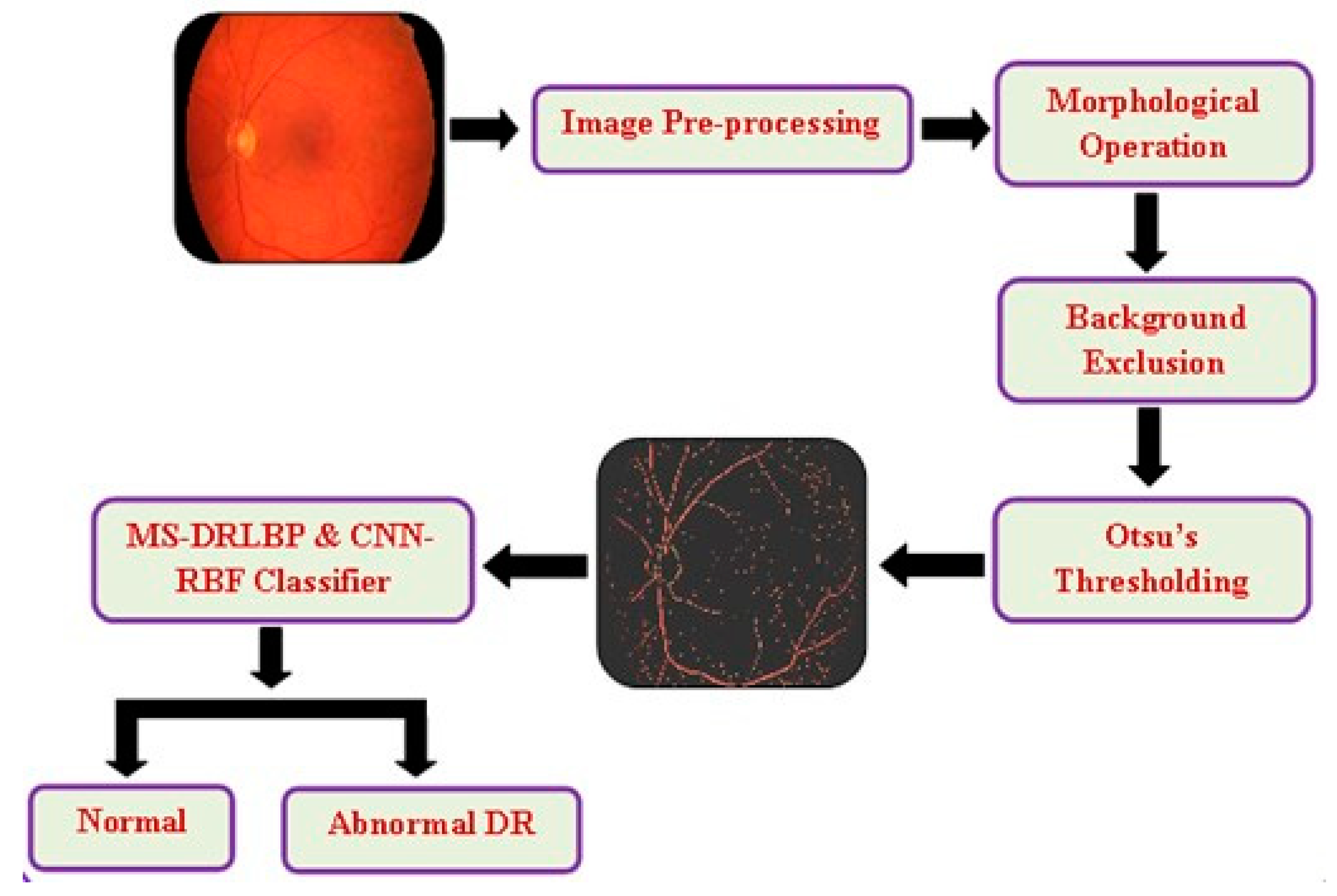
3. Materials and Methods
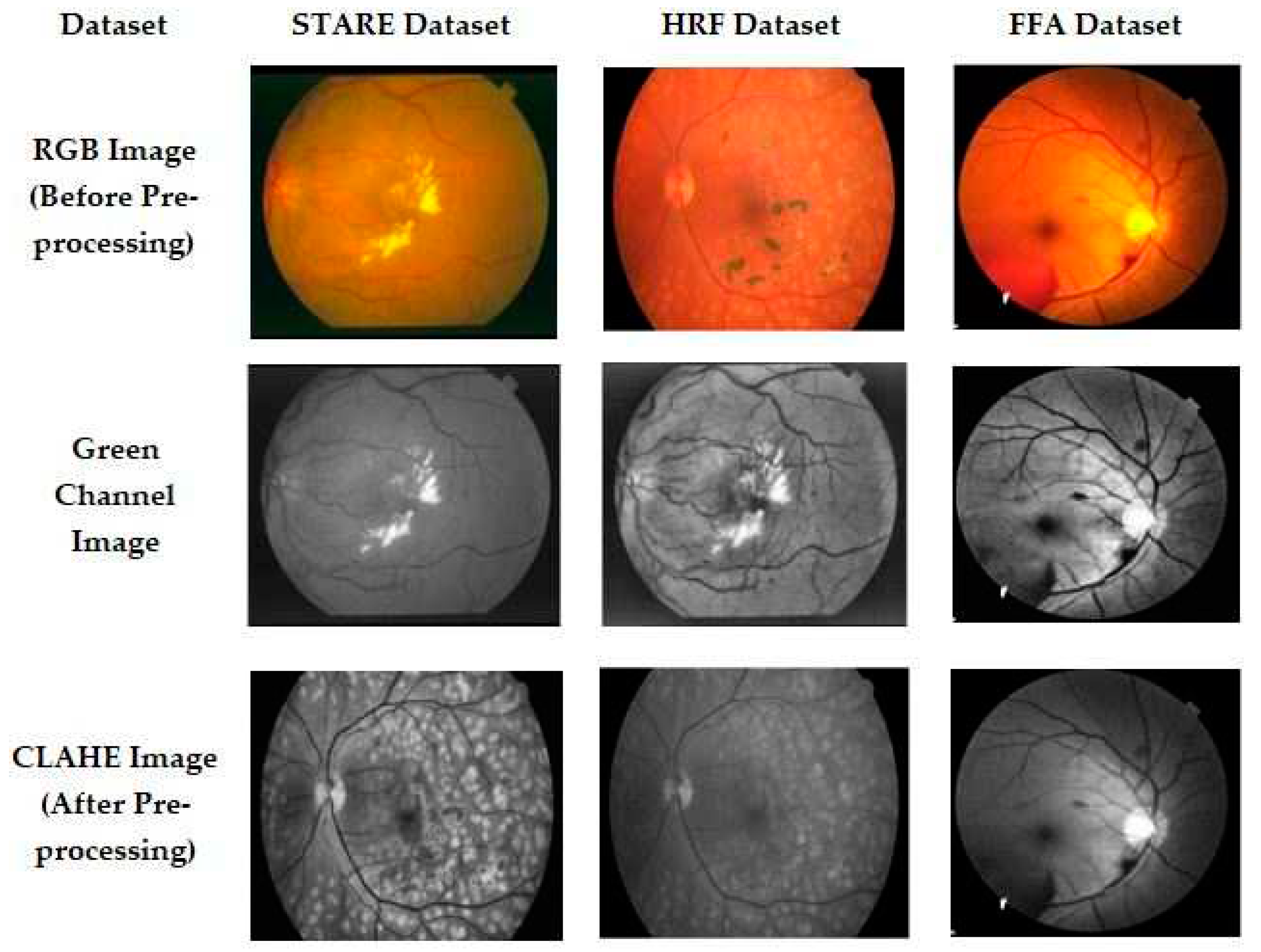
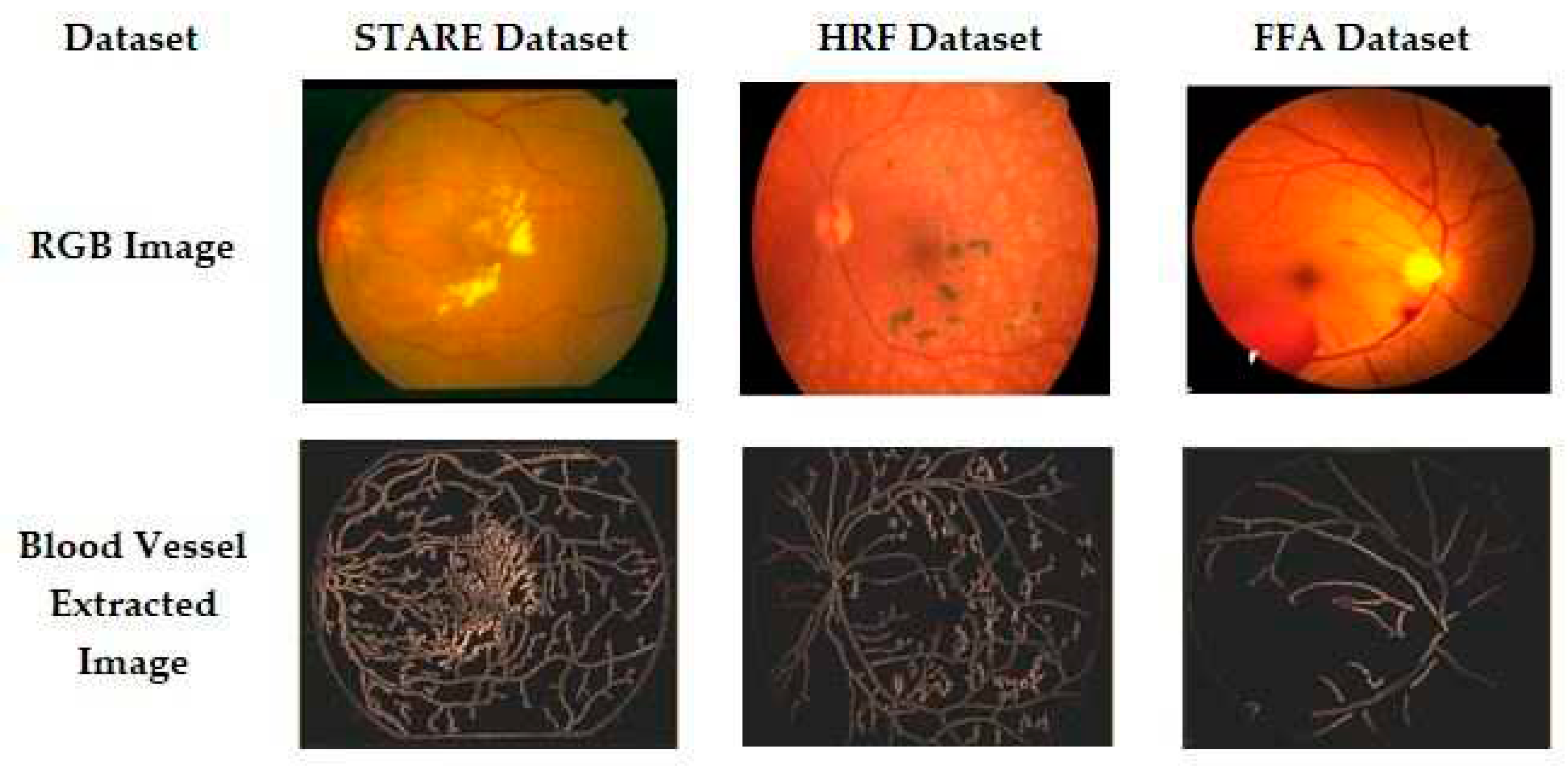
3.1. Preprocessing
3.1.1. Morphological Operations
3.1.2. Background Exclusion
3.1.3. Otsu’s Thresholding
3.2. Network Model and Training
4. Results and Discussion
- True positive (TP): DR image correctly identified as DR image.
- False positive (FP): Normal image (NR) incorrectly identified as DR image.
- True negative (TN): Normal image (NR) correctly identified as Normal image (NR).
- False negative (FN): DR image incorrectly identified as Normal image (NR).
4.1. Limitation and Future Works
- The proposed methodology has been developed and tested using only three open-source or publicly available datasets. Consequently, the results may not be more robust if they are tested with unknown datasets. A larger number of images from a variety of datasets needs to be tested to validate and generalize the proposed methodology.
- The present work only considered binary classification due to the limited number of images in each class of retinal diseases (DME, DR, CNV, AMD). The proposed methodology, however, must be trained with many multi-class data for a better clinical interpretation.
5. Conclusions
Author Contributions
Funding
Ethical Statement
Data Availability Statement
Conflicts of Interest
References
- World Health Organization. Blindness and vision impairment. Available online: https://www.who.int/news-room/fact-sheets/detail/blindness-and-visual-impairment (accessed on 28 December 2023).
- Murugappan, M.; Prakash, N.; Jeya, R.; Mohanarathinam, A.; Hemalakshmi, G. A Novel Attention Based Few-shot Classification Framework for Diabetic Retinopathy Detection and Grading. Measurement 2022, 111485. [Google Scholar] [CrossRef]
- Mutawa, A.M.; Alnajdi, S.; Sruthi, S. Transfer Learning for Diabetic Retinopathy Detection: A Study of Dataset Combination and Model Performance. Applied Sciences 2023, 13. [Google Scholar] [CrossRef]
- Qin, Q.; Chen, Y. A review of retinal vessel segmentation for fundus image analysis. Engineering Applications of Artificial Intelligence 2024, 128, 107454. [Google Scholar] [CrossRef]
- Kar, S.S.; Maity, S.P. Automatic Detection of Retinal Lesions for Screening of Diabetic Retinopathy. IEEE Transactions on Biomedical Engineering 2018, 65, 608–618. [Google Scholar] [CrossRef] [PubMed]
- Bai, C.; Huang, L.; Pan, X.; Zheng, J.; Chen, S. Optimization of deep convolutional neural network for large scale image retrieval. Neurocomputing 2018, 303, 60–67. [Google Scholar] [CrossRef]
- Khan, M.B.; Ahmad, M.; Yaakob, S.B.; Shahrior, R.; Rashid, M.A.; Higa, H. Automated Diagnosis of Diabetic Retinopathy Using Deep Learning: On the Search of Segmented Retinal Blood Vessel Images for Better Performance. Bioengineering 2023, 10, 413. [Google Scholar] [CrossRef] [PubMed]
- Marín, D.; Aquino, A.; Gegundez-Arias, M.E.; Bravo, J.M. A New Supervised Method for Blood Vessel Segmentation in Retinal Images by Using Gray-Level and Moment Invariants-Based Features. IEEE Transactions on Medical Imaging 2011, 30, 146–158. [Google Scholar] [CrossRef] [PubMed]
- Eladawi, N.; Elmogy, M.; Helmy, O.; Aboelfetouh, A.; Riad, A.; Sandhu, H.; Schaal, S.; El-Baz, A. Automatic blood vessels segmentation based on different retinal maps from OCTA scans. Comput Biol Med 2017, 89, 150–161. [Google Scholar] [CrossRef] [PubMed]
- Hassan, G.; El-Bendary, N.; Hassanien, A.E.; Fahmy, A.; Abullah M, S.; Snasel, V. Retinal Blood Vessel Segmentation Approach Based on Mathematical Morphology. Procedia Computer Science 2015, 65, 612–622. [Google Scholar] [CrossRef]
- Fraz, M.M.; Remagnino, P.; Hoppe, A.; Uyyanonvara, B.; Rudnicka, A.R.; Owen, C.G.; Barman, S.A. Blood vessel segmentation methodologies in retinal images--a survey. Comput Methods Programs Biomed 2012, 108, 407–433. [Google Scholar] [CrossRef]
- Roychowdhury, S.; Koozekanani, D.D.; Parhi, K.K. Blood Vessel Segmentation of Fundus Images by Major Vessel Extraction and Subimage Classification. IEEE J Biomed Health Inform 2015, 19, 1118–1128. [Google Scholar] [CrossRef] [PubMed]
- Barkana, B.D.; Saricicek, I.; Yildirim, B. Performance analysis of descriptive statistical features in retinal vessel segmentation via fuzzy logic, ANN, SVM, and classifier fusion. Knowledge-Based Systems 2017, 118, 165–176. [Google Scholar] [CrossRef]
- Marin, D.; Aquino, A.; Gegundez, M.; Bravo, J.M. A New Supervised Method for Blood Vessel Segmentation in Retinal Images by Using Gray-Level and Moment Invariants-Based Features. IEEE Transactions on Medical Imaging 2011, 30, 146–158. [Google Scholar] [CrossRef] [PubMed]
- Kavitha, M.; Palani, S. Blood Vessel, Optical Disk and Damage Area-Based Features for Diabetic Detection from Retinal Images. Arabian Journal for Science and Engineering 2014, 39, 7059–7071. [Google Scholar] [CrossRef]
- Liskowski, P.; Krawiec, K. Segmenting Retinal Blood Vessels With Deep Neural Networks. IEEE Trans Med Imaging 2016, 35, 2369–2380. [Google Scholar] [CrossRef] [PubMed]
- Vasanthi, S.; Banu, R.W. Automatic segmentation and classification of hard exudates to detect Macular Edema in fundus images. Journal of Theoretical and Applied Information Technology 2014, 66, 684–690. [Google Scholar]
- Morales, S.; Engan, K.; Naranjo, V.; Colomer, A. Retinal Disease Screening Through Local Binary Patterns. IEEE J Biomed Health Inform 2017, 21, 184–192. [Google Scholar] [CrossRef] [PubMed]
- Morales, S.; Naranjo, V.; Angulo, J.; Fuertes, J.; Alcañiz Raya, M. Segmentation and analysis of retinal vascular tree from fundus image processing. In Proceedings of the International Conference on Bio-inspired Systems and Signal Processing (BIOSIGNALS-2012) 2012; 2012; pp. 321–324. [Google Scholar] [CrossRef]
- Sangeethaa, S.N.; Uma Maheswari, P. An Intelligent Model for Blood Vessel Segmentation in Diagnosing DR Using CNN. J Med Syst 2018, 42, 175. [Google Scholar] [CrossRef]
- Alkhaleefah, M.; Wu, C.-C. A Hybrid CNN and RBF-Based SVM Approach for Breast Cancer Classification in Mammograms. 2018 IEEE International Conference on Systems, Man, and Cybernetics (SMC), 2018; 894–899. [Google Scholar] [CrossRef]
- Balasubramanian, K.; Ananthamoorthy, N. Robust retinal blood vessel segmentation using convolutional neural network and support vector machine. Journal of Ambient Intelligence and Humanized Computing 2021, 12. [Google Scholar] [CrossRef]
- Dai, L.; Wu, L.; Li, H.; Cai, C.; Wu, Q.; Kong, H.; Liu, R.; Wang, X.; Hou, X.; Liu, Y.; et al. A deep learning system for detecting diabetic retinopathy across the disease spectrum. Nat Commun 2021, 12, 3242. [Google Scholar] [CrossRef]
- Nadeem, M.W.; Goh, H.G.; Hussain, M.; Liew, S.-Y.; Andonovic, I.; Khan, M.A. Deep Learning for Diabetic Retinopathy Analysis: A Review, Research Challenges, and Future Directions. Sensors (Basel, Switzerland) 2022, 22. [Google Scholar] [CrossRef] [PubMed]
- Kanagaraj, R.; Karuna, Y. Retinal vessel segmentation to diagnose diabetic retinopathy using fundus images: A survey. International Journal of Imaging Systems and Technology 2023, 34, n/a-n/a. [Google Scholar] [CrossRef]
- Prabha, S.; Sasikumar, S.; Leela Manikanta, C. Diabetic Retinopathy Detection Using Automated Segmentation Techniques. Journal of Physics: Conference Series 2022, 2325. [Google Scholar] [CrossRef]
- Sivapriya, G.; Manjula Devi, R.; Keerthika, P.; Praveen, V. Automated diagnostic classification of diabetic retinopathy with microvascular structure of fundus images using deep learning method. Biomedical Signal Processing and Control 2024, 88, 105616. [Google Scholar] [CrossRef]
- Kaur, J.; Kaur, P. Automated Computer-Aided Diagnosis of Diabetic Retinopathy Based on Segmentation and Classification using K-nearest neighbor algorithm in retinal images. The Computer Journal 2022, 66, 2011–2032. [Google Scholar] [CrossRef]
- Li, Z.; Jia, M.; Yang, X.; Xu, M. Blood Vessel Segmentation of Retinal Image Based on Dense-U-Net Network. Micromachines 2021, 12, 1478. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Adarsh, A.; Kumar, B.; Singh, A.K. An automated early diabetic retinopathy detection through improved blood vessel and optic disc segmentation. Optics & Laser Technology 2020, 121, 105815. [Google Scholar] [CrossRef]
- Kamran, S.A.; Hossain, K.F.; Tavakkoli, A.; Zuckerbrod, S.L.; Sanders, K.M.; Baker, S.A. RV-GAN: Segmenting Retinal Vascular Structure in Fundus Photographs Using a Novel Multi-scale Generative Adversarial Network. In Proceedings of the Medical Image Computing and Computer Assisted Intervention – MICCAI 2021, Cham; 2021; pp. 34–44. [Google Scholar]
- Ma, Z.; Li, X. An improved supervised and attention mechanism-based U-Net algorithm for retinal vessel segmentation. Computers in Biology and Medicine 2024, 168, 107770. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.; Yang, K.-F.; Zhao, S.-X.; Wang, J.; Liu, L.; Li, Y.-J. Deep matched filtering for retinal vessel segmentation. Knowledge-Based Systems 2024, 283, 111185. [Google Scholar] [CrossRef]
- He, X.; Wang, T.; Yang, W. Research on Retinal Vessel Segmentation Algorithm Based on a Modified U-Shaped Network. Applied Sciences 2024, 14, 465. [Google Scholar] [CrossRef]
- Soomro, T.A.; Khan, T.M.; Khan, M.A.U.; Gao, J.; Paul, M.; Zheng, L. Impact of ICA-Based Image Enhancement Technique on Retinal Blood Vessels Segmentation. IEEE Access 2018, 6, 3524–3538. [Google Scholar] [CrossRef]
- STructured Analysis of the Retina. Available online: https://cecas.clemson.edu/~ahoover/stare/ (accessed on 2 June 2023).
- High-Resolution Fundus (HRF) Image Database. Available online: https://www5.cs.fau.de/research/data/fundus-images/ (accessed on 3 July 2023).
- Fundus Photography for Health Technicians Manual. Available online: https://wwwn.cdc.gov/nchs/data/nhanes3/manuals/fundus.pdf (accessed on 10 July 2023).
| Dataset | Normal Image | DR image |
|---|---|---|
| STARE | 38 | 52 |
| HRF | 15 | 15 |
| FFA | 30 | 40 |
| Total | 83 | 107 |
| Dataset | STARE | HRF | FFA | ALL | ||||
|---|---|---|---|---|---|---|---|---|
| Normal | DR | Normal | DR | Normal | DR | Normal | DR | |
| Training | 22 | 31 | 9 | 9 | 18 | 24 | 49 | 64 |
| Testing | 16 | 21 | 6 | 6 | 12 | 16 | 34 | 43 |
| Total | 38 | 52 | 15 | 15 | 30 | 40 | 83 | 107 |
| Techniques | Precision (%) | Recall (%) | F-Score (%) | Sensitivity (%) | Specificity (%) | Accuracy (%) |
|---|---|---|---|---|---|---|
| Proposed CNN-RBF | 100.00 | 96.49 | 98.21 | 96.49 | 100.00 | 97.22 |
| CNN | 98.11 | 91.23 | 94.55 | 91.23 | 93.33 | 91.67 |
| RBF | 94.64 | 92.98 | 93.81 | 92.98 | 80.00 | 90.28 |
| ANFIS | 98.04 | 87.72 | 92.59 | 87.72 | 93.33 | 88.89 |
| NN | 96.08 | 85.96 | 90.74 | 85.96 | 86.67 | 86.11 |
| NB | 93.75 | 78.95 | 85.71 | 78.95 | 80.00 | 79.17 |
| SVM | 97.87 | 80.70 | 88.46 | 80.70 | 93.33 | 83.33 |
| DR | AMD | CNV | NR | |
|---|---|---|---|---|
| DR | 20 | - | - | 1 |
| AMD | - | 18 | 2 | 1 |
| CNV | 3 | - | 17 | 1 |
| NR | 2 | 3 | 1 | 15 |
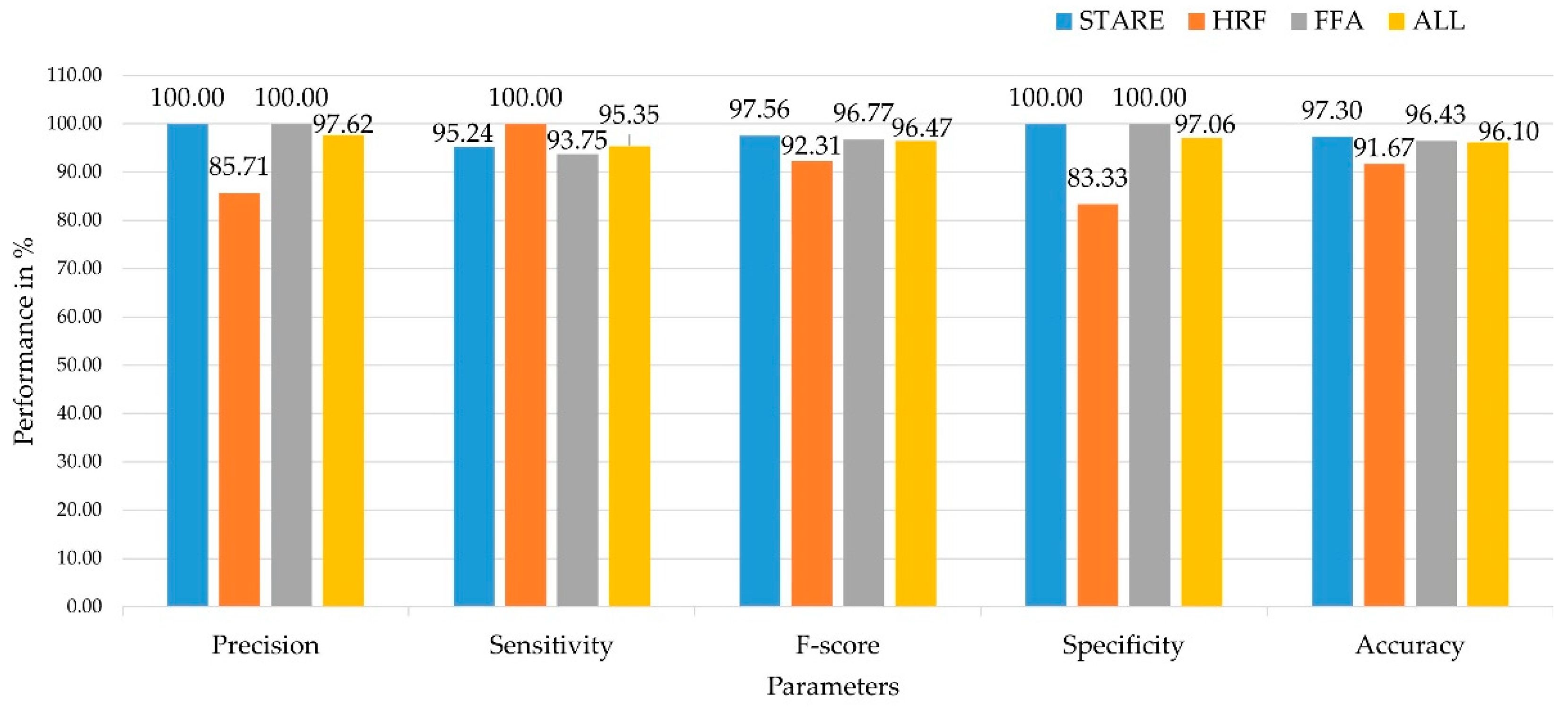
| Techniques | TP | TN | FP | FN | Precision (%) | Sensitivity (%) | Specificity (%) | Accuracy (%) | Dataset |
|---|---|---|---|---|---|---|---|---|---|
| CNN-RBF | 20 | 16 | 0 | 1 | 100.00 | 95.24 | 100.00 | 97.30 | STARE |
| CNN | 19 | 15 | 1 | 2 | 95.00 | 90.48 | 93.75 | 91.89 | |
| RBF | 18 | 15 | 1 | 3 | 94.74 | 85.71 | 93.75 | 89.19 | |
| ANFIS | 18 | 14 | 2 | 3 | 90.00 | 85.71 | 87.50 | 86.49 | |
| NN | 17 | 14 | 2 | 4 | 89.47 | 80.95 | 87.50 | 83.78 | |
| NB | 16 | 13 | 3 | 5 | 84.21 | 76.19 | 81.25 | 78.38 | |
| SVM | 18 | 13 | 3 | 3 | 85.71 | 85.71 | 81.25 | 83.78 | |
| CNN-RBF | 6 | 5 | 1 | 0 | 85.71 | 100.00 | 83.33 | 91.67 | HRF |
| CNN | 5 | 5 | 1 | 1 | 83.33 | 83.33 | 83.33 | 83.33 | |
| RBF | 4 | 5 | 1 | 2 | 80.00 | 66.67 | 83.33 | 75.00 | |
| ANFIS | 5 | 3 | 3 | 1 | 62.50 | 83.33 | 50.00 | 66.67 | |
| NN | 3 | 4 | 2 | 3 | 60.00 | 50.00 | 66.67 | 58.33 | |
| NB | 3 | 3 | 3 | 3 | 50.00 | 50.00 | 50.00 | 50.00 | |
| SVM | 4 | 5 | 1 | 2 | 80.00 | 66.67 | 83.33 | 75.00 | |
| CNN-RBF | 15 | 12 | 0 | 1 | 100.00 | 93.75 | 100.00 | 96.43 | |
| CNN | 14 | 12 | 0 | 2 | 100.00 | 87.50 | 100.00 | 92.86 | |
| RBF | 14 | 11 | 1 | 2 | 93.33 | 87.50 | 91.67 | 89.29 | |
| ANFIS | 13 | 11 | 1 | 3 | 92.86 | 81.25 | 91.67 | 85.71 | FFA |
| NN | 12 | 10 | 2 | 4 | 85.71 | 75.00 | 83.33 | 78.57 | |
| NB | 13 | 10 | 2 | 3 | 86.67 | 81.25 | 83.33 | 82.14 | |
| SVM | 15 | 12 | 0 | 1 | 100.00 | 93.75 | 100.00 | 96.43 | |
| CNN-RBF | 41 | 33 | 1 | 2 | 97.62 | 95.35 | 97.06 | 96.10 | ALL (STARE, HRF, FFA) |
| CNN | 38 | 32 | 2 | 5 | 95.00 | 88.37 | 94.12 | 90.91 | |
| RBF | 36 | 31 | 3 | 7 | 92.31 | 83.72 | 91.18 | 87.01 | |
| ANFIS | 36 | 28 | 6 | 7 | 85.71 | 83.72 | 82.35 | 83.12 | |
| NN | 32 | 28 | 6 | 11 | 84.21 | 74.42 | 82.35 | 77.92 | |
| NB | 32 | 26 | 8 | 11 | 80.00 | 74.42 | 76.47 | 75.32 | |
| SVM | 37 | 30 | 4 | 6 | 90.24 | 86.05 | 88.24 | 87.01 |
| Reference | Method | Dataset | F-score (%) | Sensitivity (%) | Specificity (%) | Accuracy (%) |
|---|---|---|---|---|---|---|
| Barkana [13] | Fuzzy+ANN+SVM | STARE | - | 70.14 | 98.46 | 95.53 |
| Soomro [35] | ICA | STARE | - | 78.60 | 98.20 | 96.70 |
| Kumar [30] | RBFNN | DIARETDB1 | - | 87.00 | 93.00 | - |
| Kamran [31] | RV-GAN | STARE | 83.23 | 83.56 | 98.64 | 97.54 |
| Li [29] | Dense-U-Net | DRIVE | - | 79.31 | 98.96 | 96.98 |
| Jaspreet [28] | KNN | DIARETDB1 | - | 92.60 | 87.56 | 95.00 |
| Sivapriya [27] | ResEAD2Net | STARE | - | 90.24 | 99.01 | 98.07 |
| Zhendi [32] | U-Net | STARE | 82.98 | 78.11 | 98.80 | 96.60 |
| Yubo [33] | WS-DMF | STARE | - | 84.48 | 98.54 | 96.13 |
| HRF | - | 83.78 | 99.75 | 95.71 | ||
| Xialan [34] | MU-Net | STARE | - | 82.64 | 98.21 | 96.93 |
| Our proposed model | CNN-RBF | STARE | 97.56 | 95.24 | 100.00 | 97.30 |
| HRF | 92.31 | 100.00 | 83.33 | 91.37 | ||
| FFA | 96.77 | 93.75 | 100.00 | 96.43 | ||
| ALL | 96.47 | 95.35 | 97.06 | 96.10 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




