Submitted:
10 January 2024
Posted:
12 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Material and Methods
2.1. Samples
2.2. Immunohistochemistry and In situ Hybridisation
2.3. RNA Extraction
2.4. Gene Expression by RT-qPCR
3. Results
3.1. APIS Molecular Characteristics
3.2. Pathological and IHC Characteristics
3.3. Comparison of APIS mRNA Scores for ER, PR, HER-2, Ki-67 and IHC Based Results
4. Discussion
4.1. Clinically Relevant Discordances
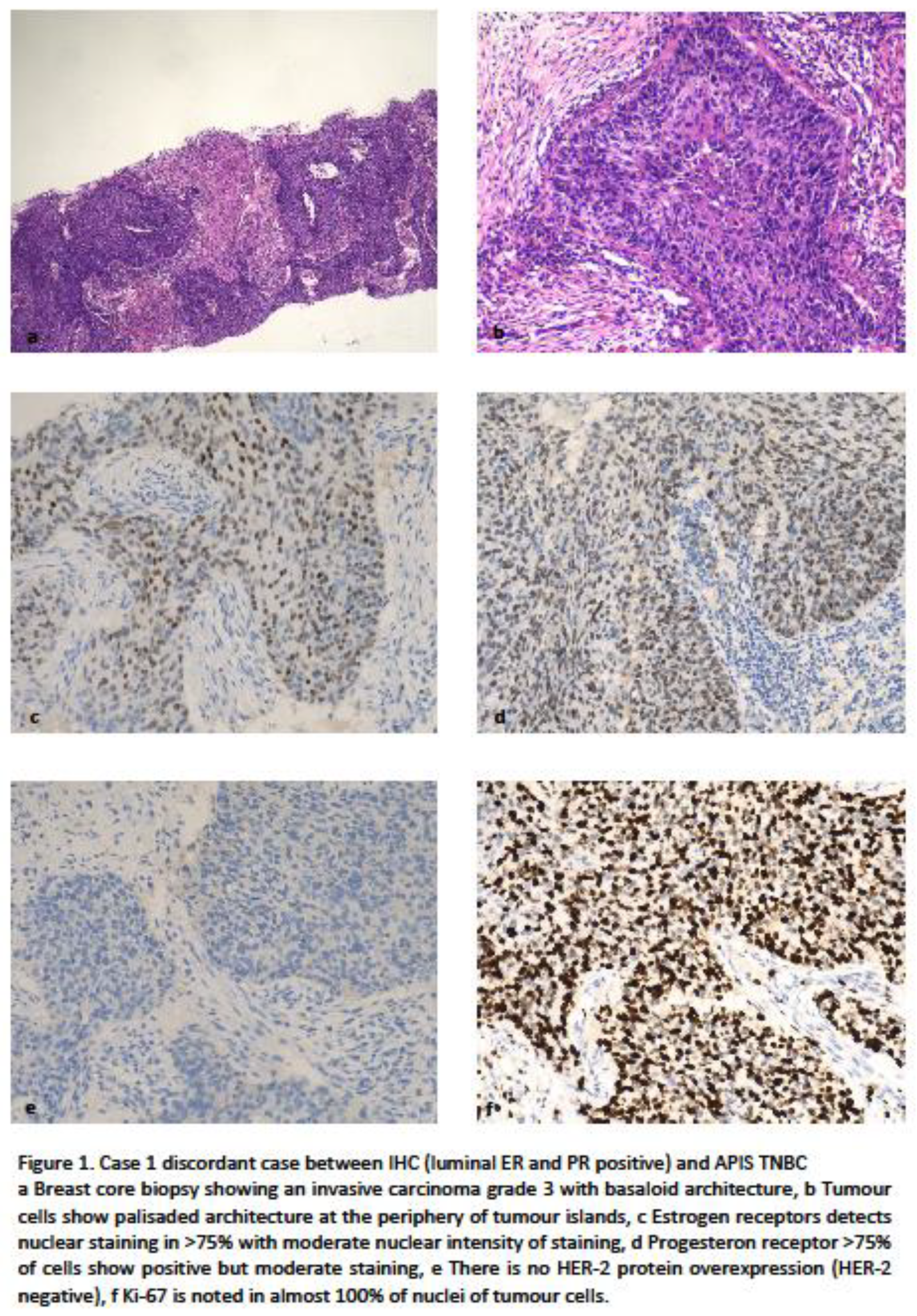
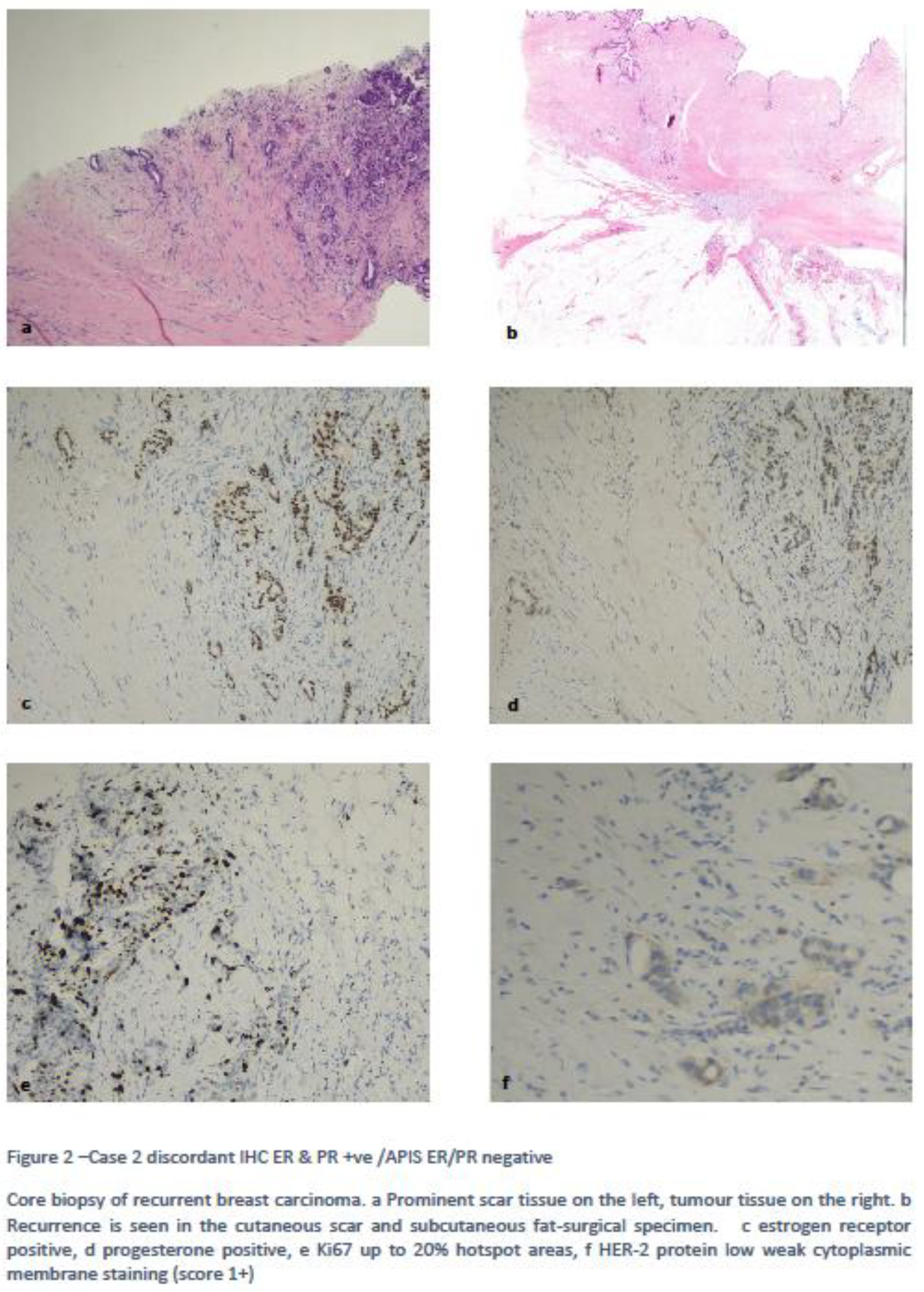
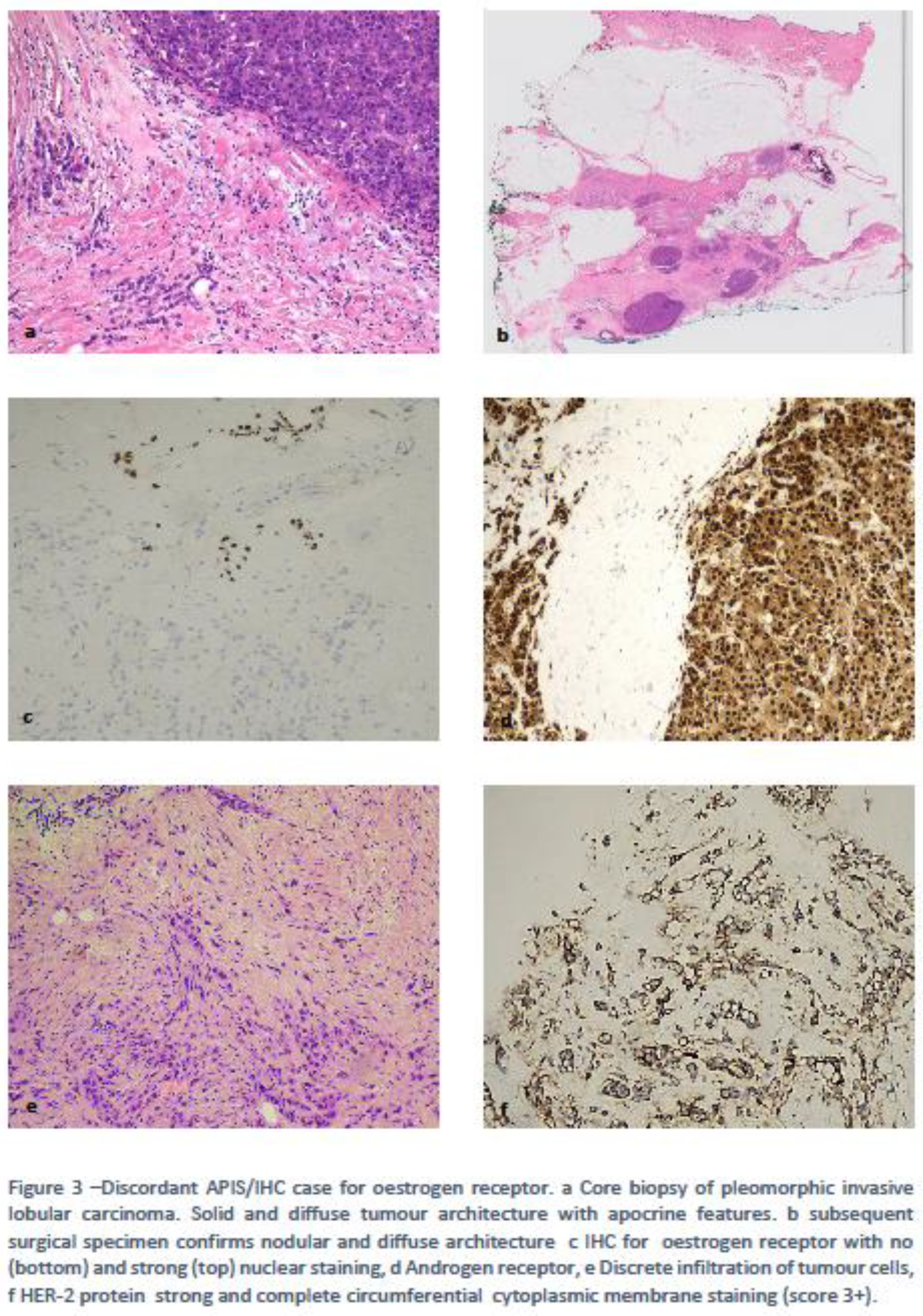
4.2. Discordances Requiring Improvements/Refinement.
Funding
Ethic approval
Patient consent
Acknowledgments
Conflicts of Interest
References
- Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, et al. Molecular portraits of human breast tumours. Nature. 2000;406:747-52.
- Prat A, Parker JS, Fan C, Perou CM. PAM50 assay and the three-gene model for identifying the major and clinically relevant molecular subtypes of breast cancer. Breast Cancer Res Treat. 2012;135:301-6.
- Nielsen TO, Parker JS, Leung S, Voduc D, Ebbert M, Vickery T, et al. A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor-positive breast cancer. Clin Cancer Res. 2010;16:5222-32.
- Di Palma S, Simpson RH, Marchiò C, et al. Salivary duct carcinomas can be classified into luminal androgen receptor-positive, HER2 and basal-like phenotypes. Histopathology. 2012;61(4):629-643. [CrossRef]
- Kim HK, Park KH, Kim Y, et al. Discordance of the PAM50 Intrinsic Subtypes Compared with Immunohistochemistry-Based Surrogate in Breast Cancer Patients: Potential Implication of Genomic Alterations of Discordance. Cancer Res Treat. 2019;51(2):737-747. [CrossRef]
- Polley MY, Leung SC, McShane LM, et al. An international Ki67 reproducibility study. J Natl Cancer Inst. 2013;105(24):1897-1906. [CrossRef]
- Zaakouk, M., Quinn, C., Provenzano, E., Boyd, C., Callagy, G., Elsheikh, S., Flint, J., Millican-Slater, R., Gunavardhan, A., Mir, Y., Makhija, P., Di Palma, S., Pritchard, S., Tanchel, B., Rakha, E., Atallah, N. M., Lee, A. H. S., Pinder, S., & Shaaban, A. M. (2023). Concordance of HER2-low scoring in breast carcinoma among expert pathologists in the United Kingdom and the republic of Ireland -on behalf of the UK national coordinating committee for breast pathology. Breast (Edinburgh, Scotland), 70, 82–91. [CrossRef]
- Erber R, Hartmann A, Fasching PA, et al. Reproducibility of mRNA-Based Testing of ESR1, PGR, ERBB2, and MKI67 Expression in Invasive Breast Cancer-A Europe-Wide External Quality Assessment. Cancers (Basel). 2021;13(18):4718. Published 2021 Sep 21. [CrossRef]
- Ahmad Fauzi MF, Wan Ahmad WSHM, Jamaluddin MF, et al. Allred Scoring of ER-IHC Stained Whole-Slide Images for Hormone Receptor Status in Breast Carcinoma. Diagnostics (Basel). 2022;12(12):3093. Published 2022 Dec 8. [CrossRef]
- Allison KH, Hammond MEH, Dowsett M, et al. Estrogen and Progesterone Receptor Testing in Breast Cancer: ASCO/CAP Guideline Update. J Clin Oncol. 2020;38(12):1346-1366. [CrossRef]
- Wolff AC, Somerfield MR, Dowsett M, et al. Human Epidermal Growth Factor Receptor 2 Testing in Breast Cancer: ASCO-College of American Pathologists Guideline Update. J Clin Oncol. 2023;41(22):3867-3872. [CrossRef]
- Dieci MV, Guarneri V, Tosi A, et al. Neoadjuvant chemotherapy and immunotherapy in luminal B-like breast cancer: results of the phase II GIADA trial. Clin Cancer Res 2022; 28: 308–317.
- Ibrahim A, Toss MS, Makhlouf S, Miligy IM, Minhas F, Rakha EA. Improving mitotic cell counting accuracy and efficiency using phosphohistone-H3 (PHH3) antibody counterstained with haematoxylin and eosin as part of breast cancer grading. Histopathology. 2023;82(3):393-406. [CrossRef]
- Liu F, Wu Y, Mi Y, Gu L, Sang M, Geng C. Identification of core genes and potential molecular mechanisms in breast cancer using bioinformatics analysis. Pathol Res Pract. 2019;215(7):152436. [CrossRef]
- Li T-F, Zeng H-J, Shan Z, Ye R-Y, Cheang T-Y, Zhang Y-J, et al. Overexpression of kinesin superfamily members as prognostic biomarkers of breast cancer. Cancer cell international 2020;20: 123.
- Lu Y, Yang G, Xiao Y, et al. Upregulated cyclins may be novel genes for triple-negative breast cancer based on bioinformatic analysis. Breast Cancer. 2020;27(5):903-911. [CrossRef]
- Sotiriou, C., & Piccart, M. J. (2007). Taking gene-expression profiling to the clinic: when will molecular signatures become relevant to patient care?. Nature reviews. Cancer, 7(7), 545–553. [CrossRef]
| Target | Positive/negative Cut off values |
|---|---|
| ESR1 | -1.98 |
| PGR | -0.63 |
| ERBB2 | 2.00 |
| MKI67 | -0.64 |
| Immunohistochemistry | APIS Breast Cancer Subtyping Kit Analysis | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Serial No | Type/Met | ER | PR | HER-2 | MIB1 IHC | ESR1 | PGR | ERBB2 | MK167 | Proliferation | Molecula Subtype |
| 1 | IDC G2 | 8/8 | 8/8 | Low 2+ ISH -neg | Low | +ve | +ve | -ve | Low | Low | Luminal A |
| 2 | IDC G2 | 8/8 | 8/8 | Low 2+ ISH -ve | Low | +ve | +ve | -ve | high | high | Luminal B Her-2 -ve |
| 3 | IDC G3 | 8/8 | 8/8 | Low 1+ | high | +ve | +ve | -ve | high | high | Luminal B Her2 -ve |
| 4 | IDC G3 | 0/8 | 6/8 | -ve | high | -ve | -ve | -ve | high | high | TNBC |
| 5 | IDC G3 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | high | high | Luminal B her-2 -ve |
| 6 | IDC G3 | 7/8 | 7/8 | -ve | high | -ve | -ve | -ve | high | high | TNBC |
| 7 | IDC G2 | 8/8 | 8/8 | Low 1+ | high | +ve | +ve | -ve | high | high | Luminal B her-2 -ve |
| 8 | IDC G3 | 8/8 | 8/8 | Low 1+ | high | +ve | +ve | -ve | High | High | Luminal B her-2 -ve |
| 9 | IDC G2 | 8/8 | 0/8 | -ve | low | +ve | -ve | -ve | low | high | Luminal A |
| 10 | IDC G2 | 8/8 | 0/8 | Low 1+ | low | +ve | -ve | -ve | low | Low | Luminal A |
| 11 | IDC G3 | 0/8 | 0/8 | +ve 3+ | low | -ve | -ve | +ve | Low | High | HER-2 Enriched |
| 12 | IDC G2 | 8/8 | 0/8 | -ve | low | +ve | -ve | -ve | Low | High | Luminal A |
| 13 | IDC G2 | 8/8 | 0/8 | -ve | low | NA | NA | NA | NA | NA | NA |
| 14 | IDC G2 | 8/8 | 4/8 | -ve | high | +ve | -ve | -ve | high | High | Luminal B her-2 -ve |
| 15 | IDC G1 | 7/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | Low | Luminal A |
| 16 | IDC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | low | Luminal A |
| 17 | IDC G3 | 0/8 | 0/8 | -ve | high | -ve | -ve | -ve | high | high | Triple negative |
| 18 | IDC G3 | 8/8 | 7/8 | -ve | high | +ve | +ve | -ve | High | High | Luminal B Her-2 -ve |
| 19 | IDC G2 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | high | Luminal B |
| 20 | IDC G3 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | High | Luminal B Her 2 -ve |
| 21 | IDC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | low | Luminal A |
| 22 | IDC G2 | 8/8 | 0/8 | -ve | low | +ve | -ve | -ve | low | High | Luminal A |
| 23 | IDC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | High | Luminal A |
| 24 | IDC G3 | 8/8 | 4/8 | -ve | high | +ve | -ve | -ve | High | High | Luminal B Her-2 -ve |
| 25 | IDC G2 | 8/8 | 6/8 | Low 1+ | low | -ve | +ve | -ve | low | High | Luminal A |
| 26 | IDC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | Low | High | Luminal A |
| 27 | IDC G3 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | low | Luminal A |
| 28 | IDC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | High | Luminal A |
| 29 | IDC G3 | 0/8 | 0/8 | +ve 3+ | low | -ve | -ve | -ve | low | High | HER-2 Enriched |
| 30 | IDC G2 | 8/8 | 7/8 | -ve | low | +ve | +ve | -ve | low | low | Luminal A |
| 31 | IDC G3 | 8/8 | 8/8 | +ve 3+ | high | +ve | +ve | +ve | High | high | Luminal B her-2 +ve |
| 32 | IDC G3 | 0/8 | 2/8 | -ve | high | -ve | -ve | -ve | low | high | TNBC |
| 33 | IDC G2 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | High | Luminal B Her-2 -ve |
| 34 | IDC G2 | 8/8 | 8/8 | Low 2+ ISH -ve | high | +ve | +ve | -ve | High | High | Luminal B Her 2 -ve |
| 35 | IDC G2 | 8/8 | 0/8 | -ve | low | +ve | -ve | -ve | low | High | Luminal A |
| 36 | IDC G3 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | High | Luminal B Her-2 -ve |
| 37 | IDC G2 | 0/8 | 0/8 | -ve | low | -ve | -ve | -ve | low | low | Triple negative |
| 38 | IDC G2 | 7/8 | 8/8 | Low 1+ | low | +ve | +ve | neg | low | low | Luminal A |
| 39 | IDC G3 | 7/8 | 8/8 | Low 1+ | high | positive | positive | neg | high | high | luminal B her-2 -ve |
| 40 | IDC G3 | 0/8 | 0/8 | +ve 3+ | high | -ve | +ve | High | High | luminal B Her2 +ve | |
| 41 | IDC G2 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | High | Luminal B Her2 -ve |
| 42 | IDC G3 | 8/8 | 8/8 | +ve 3+ | high | +ve | +ve | +ve | High | High | luminal B Her-2 +ve |
| 43 | IDC G3 | 0/8 | 3/8 | Low 1+ | high | -ve | -ve | -ve | High | High | TNBC |
| 44 | IDC G2 | 8/8 | 6/8 | 2+ISH -ve | low | +ve | -ve | -ve | low | low | Luminal A |
| 45 | ILC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | negative | low | high | luminal A |
| 46 | ILC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | High | Luminal A |
| 47 | IDC G2 | 8/8 | 8/8 | Low +1 | high | +ve | +ve | -ve | High | High | Luminal B Her-2 -ve |
| 48 | IDC G3 | 0/8 | 3/8 | -ve | high | -ve | -ve | -ve | High | High | TNBC |
| 49 | ILC G2 | 8/8 | 8/8 | Low 1+ | low | +ve | +ve | -ve | low | low | Luminal A |
| 50 | IDC G3 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | High | Luminal B Her 2 -ve |
| 51 | IDC G3 | 8/8 | 6/8 | +ve 3+ | high | +ve | -ve | +ve | High | High | Luminal B Her-2 +ve |
| 52 | IDC G3 | 8/8 | 8/8 | Low 1+ | high | +ve | +ve | -ve | High | High | Luminal B Her 2 -ve |
| 53 | IDC G3 | 0/8 | 2/8 | +ve 3+ | high | -ve | +ve | +ve | High | High | Luminal B Her-2 +ve |
| 54 | IDC G2 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | High | Luminal B Her-2 -ve |
| 55 | IDC G3 | 8/8 | 8/8 | Low 1+ | high | +ve | +ve | -ve | High | High | Luminal B Her-2 -ve |
| 56 | IDC G2 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | High | Luminal B Her-2 -ve |
| 57 | IDC G2 | 0/8 | 3/8 | +ve 3+ | high | -ve | +ve | +ve | High | High | Luminal B Her 2 +ve |
| 58 | IDC G3 | 8/8 | 8/8 | +ve 3+ | high | +ve | +ve | +ve | High | High | Lum B Her-2 +ve |
| 59 | mucinous ca G2 | 8/8 | 8/8 | Low 1+ | low | +ve | +ve | -ve | low | High | Luminal A |
| 60 | IDC G2 | 8/8 | 8/8 | low 2+, ISH -ve | low | -ve | -ve | -ve | low | High | Triple negative |
| 61 | IDC G2 | 8/8 | 8/8 | +ve 3+ | low | +ve | +ve | +ve | low | High | Lum B Her-2 +ve |
| 62 | IDC G2 | 8/8 | 8/8 | Low 1+ | low | +ve | +ve | -ve | low | low | Luminal A |
| 63 | IDC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | High | Luminal A |
| 64 | IDC G3 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | high | high | luminal B |
| 65 | IDC G2 | 8/8 | 2/8 | -ve | low | +ve | -ve | -ve | low | High | Luminal A |
| 66 | IDC G1 | 8/8 | 5/8 | -ve | low | +ve | +ve | -ve | low | low | Luminal A |
| 67 | IDC G3 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | High | Luminal B her-2 -ve |
| 68 | IDC G2 | 8/8 | 8/8 | Low 1+ | high | +ve | +ve | -ve | High | High | Luminal B Her-2 -ve |
| 69 | IDC G3 | 0/8 | 0/8 | -ve | High | -ve | -ve | -ve | High | High | TNBC |
| 70 | ILC G2 | 8/8 | 8/8 | Low 1+ | low | +ve | +ve | -ve | low | low | Luminal A |
| 71 | IDC G3 | 7/8 | 7/8 | -ve | high | -ve | -ve | -ve | High | High | TNBC |
| 72 | ILC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | High | Luminal A |
| 73 | ILC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | high | luminal B her-2 -ve |
| 74 | Nodal met IDC G2 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | High | Luminal B her-2 -ve |
| 75 | IDC G3 | 8/8 | 5/8 | Low 1+ | high | +ve | +ve | -ve | High | High | Luminal B Her2 -ve |
| 76 | IDC G3 | 8/8 | 3/8 | -ve | high | +ve | -ve | -ve | High | High | Luminal B Her2 -ve |
| 77 | IDC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | High | Luminal A |
| 78 | ILC G2 | 8/8 | 7/8 | -ve | low | +ve | +ve | -ve | low | high | Luminal A |
| 79 | IDC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | -ve | low | low | Luminal A |
| 80 | IDC G2 | 8/8 | 3/8 | -ve | low | +ve | +ve | -ve | low | low | Luminal A |
| 81 | ILC G2 | 8/8 | 3/8 | -ve | low | +ve | -ve | -ve | low | high | Luminal A |
| 82 | IDC G3 | 7/8 | 7/8 | -ve | high | +ve | +ve | +ve | High | High | Luminal B Her-2 -ve |
| 83 | IDC G3 | 8/8 | 8/8 | +ve 3+ | High | +ve | +ve | +ve | High | High | Luminal B Her-2 +ve |
| 84 | IDC G2 | 8/8 | 0/8 | -ve | low | +ve | -ve | -ve | low | low | Luminal A |
| 85 | IDC G2 | 8/8 | 8/8 | Low 1+ | low | -ve | +ve | -ve | low | high | Luminal A |
| 86 | IDC G3 | 8/8 | 8/8 | -ve | Low | +ve | +ve | -ve | low | High | Luminal A |
| 87 | IDC N met | 0/8 | 0/8 | 2+ISH -ve | High | -ve | -ve | -ve | High | High | TNBC |
| 88 | IDC G1 | 8/8 | 4/8 | -ve | Low | +ve | +ve | -ve | low | low | Luminal A |
| 89 | IDC G3 | 8/8 | 5/8 | +ve 3+ | high | +ve | -ve | +ve | High | high | Luminal B Her-2 +ve |
| 90 | IDC G2 | 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | High | High | luminal B her-2 -ve |
| 91 | Muc G2 | 8/8 | 8/8 | -ve | Low | +ve | +ve | -ve | low | low | luminal A |
| 92 | IDC G2 | 8/8 | 8/8 | -ve | High | +ve | +ve | -ve | High | High | Luminal B her-2 -ve |
| 93 | Pleom ILC | 8/8 | 3/8 | +ve 3+ | low | -ve | -ve | +ve | low | High | HER-2 Enriched |
| 94 | IDC G3 | 7/8 | 6/8 | low 1+ | high | +ve | +ve | -ve | high | high | luminal B her-2 -ve |
| 95 | IDC G3 | 7/8 | 8/8 | low 1+ | high | +ve | +ve | -ve | high | High | Luminal B her-2 -ve |
| 96 | IDC G2 | 8/8 | 4/8 | -ve | low | +ve | -ve | -ve | high | High | luminal B her-2 -ve |
| 97 | IDC G2 8/8 | 8/8 | -ve | high | +ve | +ve | -ve | igh hig | High | Lum B Her-2 -ve | |
| 98 | IDC G2 | 8/8 | 8/8 | Low 1+ | low | +ve | +ve | -ve | low | high | luminal A |
| 99 | IDC G3 | 0/8 | 3/8 | -ve | high | -ve | -ve | +ve | high | high | HER-2 enrcihed |
| 100 | ILC G2 | 8/8 | 8/8 | -ve | low | +ve | +ve | neg | low | low | luminal A |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




